Abstract
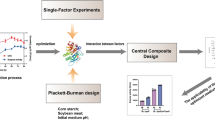
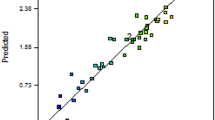
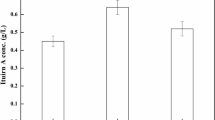
Amidase, a biocatalyst for the biotransformation of nitrile amides into various commercial carboxylic acids, is used in the pharmaceutical industry, in the chemical industry, and in the treatment of wastewater. It has been observed that the synthesis of amidase enzyme relies on several media compositions. The present study focused on increased synthesis of intracellular amidase from a novel thermotolerant bacteria Bacillus tequilensis by response surface methodology (RSM). RSM with a central composite design (CCD) was used to examine the effect of independent variables (starch, peptone, and mineral base) on response variables. From the RSM model, it was confirmed that the experimental results were appropriate in the statistical model of quadratic polynomial with R2 values greater than 0.900 for the two responses. An optimized condition consisting of 5.413 g/l starch, 1.113 g/l peptone, and 150 ml/l mineral base respectively was reported for amidase activity. The experimental values of the enzyme activity and biomass production were 8.453 I.U. and 3.213 g/l at optimized conditions. Using the statistical approach, 1.4 times increment on acrylamidase production was observed over the unoptimized experimental values.






Similar content being viewed by others
Data availability
The authors confirm that the data supporting the findings of this study are provided within the manuscript.
Code availability
Not applicable.
References
Gong JS, Shi JS, Lu ZM et al (2017) Nitrile-converting enzymes as a tool to improve biocatalysis in organic synthesis: recent insights and promises. Crit Rev Biotechnol 37:69–81. https://doi.org/10.3109/07388551.2015.1120704
Agarwal A, Nigam VK (2014) Nitrilase mediated conversion of indole-3-acetonitrile to indole-3-acetic acid. Biocatal Agric Biotechnol 3:351–357. https://doi.org/10.1016/j.bcab.2014.05.005
Mehta PK, Bhatia SK, Bhatia RK, Bhalla TC (2016) Enhanced production of thermostable amidase from Geobacillus subterraneus RL-2a MTCC 11502 via optimization of physicochemical parameters using Taguchi DOE methodology. 3 Biotech 6:66–72. https://doi.org/10.1007/s13205-016-0390-1
Matoso V, Bargi-Souza P, Ivanski F et al (2019) Acrylamide: a review about its toxic effects in the light of Developmental Origin of Health and Disease (DOHaD) concept. Food Chem 283:422–430. https://doi.org/10.1016/j.foodchem.2019.01.054
World Health Organization (2003) Acrylamide in drinking-water: background document for development of WHO guidelines for drinking-water quality
Barouki R, Melen E, Herceg Z et al (2018) Epigenetics as a mechanism linking developmental exposures to long-term toxicity. Environ Int 114:77–86. https://doi.org/10.1016/j.envint.2018.02.014
Ghamdi A, Alenezi F, Algoferi M et al (2020) A review on the new trends of acrylamide toxicity. BJSTR 27:20638–20644. https://doi.org/10.26717/BJSTR.2020.27.004480
Kadry AM, Friedman MA, Abdel-Rahman MS (1999) Pharmacokinetics of acrylamide after oral administration in male rats. Environ Toxicol Pharmacol 7:127–133. https://doi.org/10.1016/S1382-6689(99)00005-8
Friedman M, Mottram DS (2006) Chemistry and safety of acrylamide in food. Springer, New York 561
Bedade DK, Muley AB, Singhal RS (2019) Magnetic cross-linked enzyme aggregates of acrylamidase from Cupriavidus oxalaticus ICTDB921 for biodegradation of acrylamide from industrial waste water. Bioresour Technol 272:137–145. https://doi.org/10.1016/j.biortech.2018.10.015
Zhang Y, Zhang Y (2007) Formation and reduction of acrylamide in maillard reaction: a review based on the current state of knowledge. Crit Rev Food Sci Nutr 47:521–542. https://doi.org/10.1080/10408390600920070
Liang L-Y, Zheng Y-G, Shen Y-C (2008) Optimization of β-alanine production from β-aminopropionitrile by resting cells of Rhodococcus sp. G20 in a bubble column reactor using response surface methodology. Process Biochem 43:758–764. https://doi.org/10.1016/j.procbio.2008.03.002
Ruan L-T, Zheng R-C, Zheng Y-G, Shen Y-C (2016) Purification and characterization of R -stereospecific amidase from Brevibacterium epidermidis ZJB-07021. Int J Biol Macromol 86:893–900. https://doi.org/10.1016/j.ijbiomac.2016.02.020
Zheng Y-G, Chen J, Liu Z-Q et al (2008) Isolation, identification and characterization of Bacillus subtilis ZJB-063, a versatile nitrile-converting bacterium. Appl Microbiol Biotechnol 77:985–993. https://doi.org/10.1007/s00253-007-1236-x
Chacko S, Ramteke PW, Joseph B (2012) A comparative study on the production of amidase using immobilized and dehydrated immobilized cells of Pseudomonas putida MTCC 6809. J Genet Eng Biotechnol 10:121–127. https://doi.org/10.1016/j.jgeb.2012.01.003
Wu Z-M, Zheng R-C, Zheng Y-G (2016) Exploitation and characterization of three versatile amidase super family members from Delftia tsuruhatensis ZJB-05174. Enzyme Microb Technol 86:93–102. https://doi.org/10.1016/j.enzmictec.2016.02.002
Xue Y-P, Yang Y-K, Lv S-Z et al (2016) High-throughput screening methods for nitrilases. Appl Microbiol Biotechnol 100:3421–3432. https://doi.org/10.1007/s00253-016-7381-7393
Prabha R, Nigam VK (2020) Biotransformation of acrylamide to acrylic acid carried through acrylamidase enzyme synthesized from whole cells of Bacillus tequilensis (BITNR004). Biocatal Biotransfor 38:445–456. https://doi.org/10.1080/10242422.2020.1780211
Varshavsky A (2019) N-degron and C-degron pathways of protein degradation. Proc Natl Acad Sci USA 116:358–366. https://doi.org/10.1073/pnas.1816596116
Emmanuel Joshua Jebasingh S, Lakshmikandan M, Rajesh RP, Raja P (2013) Biodegradation of acrylamide and purification of acrylamidase from newly isolated bacterium Moraxella osloensis MSU11. Int Biodeter Biodegr 85:120–125. https://doi.org/10.1016/j.ibiod.2013.06.012
Nojiri M, Taoka N, Yasohara Y (2014) Characterization of an enantioselective amidase from Cupriavidus sp. KNK-J915 (FERM BP-10739) useful for enzymatic resolution of racemic 3-piperidinecarboxamide. J Mol Catal B Enzym 109:136–142. https://doi.org/10.1016/j.molcatb.2014.08.016
Abdelrazek NA, Elkhatib WF, Raafat MM, Aboulwafa MM (2019) Experimental and bioinformatics study for production of l-asparaginase from Bacillus licheniformis: a promising enzyme for medical application. AMB Expr 9:39. https://doi.org/10.1186/s13568-019-0751-3
Agarwal A, Nigam VK (2017) Enhanced production of nitrilase from Streptomyces sp. MTCC 7546 by response surface method. Proc Natl Acad Sci India B - Biol Sci 87:603–609. https://doi.org/10.1007/s40011-015-0638-2
Nigam VK, Arfi T, Kumar V, Shukla P (2017) Bioengineering of nitrilases towards its use as green catalyst: applications and perspectives. Indian J Microbiol 57:131–138. https://doi.org/10.1007/s12088-017-0645-5
Shukla P (2019) Synthetic biology perspectives of microbial enzymes and their innovative applications. Indian J Microbiol 59:401–409. https://doi.org/10.1007/s12088-019-00819-9
Uppuluri KB, V. R. Dasari RK, Sajja V et al (2013) Optimization of l-asparaginase production by isolated Aspergillus niger C4 from sesame (black) oil cake under SSF using Box–Behnken design in column bioreactor. Int J Chem React Eng 11:103–109. https://doi.org/10.1515/ijcre-2012-0064
da Cunha MC, Silva LC, Sato HH, de Castro RJS (2018) Using response surface methodology to improve the L-asparaginase production by Aspergillus niger under solid-state fermentation. Biocatal Agric Biotechnol 16:31–36. https://doi.org/10.1016/j.bcab.2018.07.018
Sun Y, Yang G, Li K et al (2016) CO2 mineralization using basic oxygen furnace slag: process optimization by response surface methodology. Environ Earth Sci 75:1335. https://doi.org/10.1007/s12665-016-6147-7
Lakshmikandan M, Sivaraman K, Elaiya Raja S et al (2014) Biodegradation of acrylamide by acrylamidase from Stenotrophomonas acidaminiphila MSU12 and analysis of degradation products by MALDI-TOF and HPLC. Int Biodeterior Biodegrad 94:214–221. https://doi.org/10.1016/j.ibiod.2014.07.014
Yorseng K, Siengchin S, Ashok B, Rajulu AV (2020) Nanocomposite egg shell powder with in situ generated silver nanoparticles using inherent collagen as reducing agent. J Bioresour Bioproducts 5:101–107. https://doi.org/10.1016/j.jobab.2020.04.003
Varshney AK, Mohan MK, Vidyarthi AS et al (2013) Statistical optimization of medium components to increase the manganese peroxidase productivity by Phanerochaete chrysosporium NCIM 1197. Biotechnol Bioprocess Eng 18:1176–1184. https://doi.org/10.1007/s12257-013-0233-4
Mirizadeh S, Yaghmaei S, Ghobadi Nejad Z (2014) Biodegradation of cyanide by a new isolated strain under alkaline conditions and optimization by response surface methodology (RSM). J Environ Health Sci Eng 12. https://doi.org/10.1186/2052-336X-12-85
Khandelwal AK, Nigam VK, Choudhury B et al (2007) Optimization of nitrilase production from a new thermophilic isolate. J Chem Technol Biotechnol 82:646–651. https://doi.org/10.1002/jctb.1721
Banerjee A, Kaul P, Sharma R, Banerjee UC (2003) A high-throughput amenable colorimetric assay for enantioselective screening of nitrilase-producing microorganisms using pH sensitive indicators. J Biomol Screen 8:559–565. https://doi.org/10.1177/1087057103256910
Mehmood T, Ahmed A, Ahmad A et al (2018) Optimization of mixed surfactants-based β-carotene nanoemulsions using response surface methodology: an ultrasonic homogenization approach. Food Chem 253:179–184. https://doi.org/10.1016/j.foodchem.2018.01.136
Mehmood T, Ahmad A, Ahmed A, Ahmed Z (2017) Optimization of olive oil based O/W nanoemulsions prepared through ultrasonic homogenization: a response surface methodology approach. Food Chem 229:790–796. https://doi.org/10.1016/j.foodchem.2017.03.023
Vaidya BK, Mutalik SR, Joshi RM et al (2009) Enhanced production of amidase from Rhodococcus erythropolis MTCC 1526 by medium optimization using a statistical experimental design. J Ind Microbiol Biotechnol 36:671–678. https://doi.org/10.1007/s10295-009-0536-9
Shan WY, Xue J, Zheng RC (2008) Improvement of amidase production by a newly isolated Delftia tsuruhatensis ZJB-05174 through optimization of culture medium. J Microbiol Biotechnol 18:1932–1937
Ajamani A, Kumar R, Bhargava P, Vats S (2019) Mathematically optimized production, purification and characterization of penicillin G acylase from soil bacterial isolates AA17A and AA17B. Indian J Biotechnol 9
Bhatia SK, Mehta PK, Bhatia RK, Bhalla TC (2014) Optimization of arylacetonitrilase production from Alcaligenes sp. MTCC 10675 and its application in mandelic acid synthesis. Appl Microbiol Biotechnol 98:83–94. https://doi.org/10.1007/s00253-013-5288-9
Muralidhar RV, Chirumamila RR, Marchant R, Nigam P (2001) A response surface approach for the comparison of lipase production by Candida cylindracea using two different carbon sources. Biochem Eng J 9:17–23. https://doi.org/10.1016/S1369-703X(01)00117-6
Funding
The financial assistance received from Seed Money Scheme from Birla Institute of Technology, Mesra, Ranchi (Ref No.: GO/Estb/SMS/2019–2020/2122), for carrying the present work is duly acknowledged.
Author information
Authors and Affiliations
Contributions
Both the authors have equal contribution in this work.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Authorship principles
Both authors agreed with the content and gave consent to submit the manuscript.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Prabha, R., Nigam, V.K. Improved production of acrylamidase from Bacillus tequilensis through response surface methodology. Biomass Conv. Bioref. 13, 10085–10095 (2023). https://doi.org/10.1007/s13399-021-01874-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13399-021-01874-3