Abstract
Metabolic reprogramming is one of the hallmarks of cancer. As nutrients are scarce in the tumor microenvironment (TME), tumor cells adopt multiple metabolic adaptations to meet their growth requirements. Metabolic reprogramming is not only present in tumor cells, but exosomal cargos mediates intercellular communication between tumor cells and non-tumor cells in the TME, inducing metabolic remodeling to create an outpost of microvascular enrichment and immune escape. Here, we highlight the composition and characteristics of TME, meanwhile summarize the components of exosomal cargos and their corresponding sorting mode. Functionally, these exosomal cargos-mediated metabolic reprogramming improves the "soil" for tumor growth and metastasis. Moreover, we discuss the abnormal tumor metabolism targeted by exosomal cargos and its potential antitumor therapy. In conclusion, this review updates the current role of exosomal cargos in TME metabolic reprogramming and enriches the future application scenarios of exosomes.
Similar content being viewed by others
Background
Extracellular vesicles (EVs) are nanoscale cellular secretions that act as key mediators in many pathological/physiological processes [145]. Another method that uses electricity and acoustic forces to manipulate biological particles and submicron particles for deterministic sorting has been applied to the purification of exosomes. The purity of the exosomes purified by this method is more than 95% and the recovery rate is 81% [146].
Cargos in exosomes
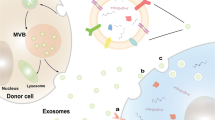
Exosome cargos are the core components that confer biological effects on exosomes. Nearly 100,000 proteins and over 1,000 lipids have been reported to be associated with exosomes [147]. Nucleic acids mainly mRNAs and ncRNAs were enriched in exosomes, including more than 27,000 mRNAs and more than 10,000 ncRNAs were identified in sEV [147]. Genomic DNA (gDNA) and mitochondria DNA are present in exosomes in the form of s single-stranded or double-stranded [148, 149]. Together, these biologically active substances make up the exosomal cargos. (Fig. 2).
Biomarkers of exosomes and main components of exosomal cargos. Several exosome surface proteins are considered to be biomarkers of exosomes, including transmembrane 4 superfamily (CD9, CD63, CD81), lipid raft protein (flotillin-1), and Ceramide. Exosome content proteins HSP70, TSG101 and ALIX are also biomarkers of exosomes. Exosomes carry a variety of cargos, such as nucleic acids, proteins, enzymes and metabolites (mainly lipids)
Nucleic acids
The role of exosomal RNAs in tumors has been widely reported, mainly including mRNAs and ncRNAs. Regarding functional exosomal mRNAs, an early study reported that exosomal mRNAs can complete translation in receptor cells [150]. Recently, exosomal mRNAs CUL9, KMT2D, PBRM1, PREX2 and SETD2 were found to be possible novel potential biomarkers for clear cell renal cell carcinoma (ccRCC) [151]. Studies on exosomal ncRNAs have focused on miRNAs, lncRNAs & circRNAs. Usually exosomal ncRNAs are transported to the recipient cells as molecular sponges.
Exosomal DNA mainly includes gDNA and mtDNA [149]. The TME of "hibernating" cancer cells secretes EVs containing mtDNA, leading to endocrine therapy resistance in breast cancer cells [152]. T cells secrete EVs containing gDNA and mtDNA, which activate the cGAS/STING signaling pathway and induce antiviral responses in DCs [153].
Lipids
The lipid cargos of exosomes mainly include sphingolipids, cholesterol, phosphatidylserine, saturated fatty acids, and ceramides, which are mainly associated with exosome biogenesis [154]. The bilayer lipid membrane structure of exosomes determines the enrichment of membrane lipid components (phosphoglycerolipids, sphingolipids, and sterols) [147]. The fact that neutral sphingomyelinase inhibitors can reduce exosome secretion further illustrates the importance of membrane lipids for exosomes [155]. Cholesterol enrichment in exosomes is associated with MVB. Different subcellular organelles have different cholesterol concentrations. Oxysterol-binding protein-related proteins (ORPs) are involved in cholesterol transport and are able to maintain the proper cholesterol concentration required for MVB biogenesis [156]. In addition, when low-density lipoprotein-cholesterol is low in endosomes, endoplasmic reticulum stress-derived cholesterol can be transferred to MVB [157].
Some biologically active lipids are important cargos for exosomes to perform their biological functions. LTB4 is packaged and released in exosomes [158]. This LTB-enriched exosome biogenesis originates from the nuclear envelope of centrocytes and is an unconventional pathway of exosome production [159]. Granulocyte myeloid-derived suppressor cells can secrete exosomal PGE2 to ameliorate collagen-induced arthritis [160]. Ubiquitination of 15-LO2 in hypoxia promotes 15-LO2 sorting to exosomes, which are involved in the regulation of pulmonary vascular homeostasis [176]. Recent studies have shown that two RNA binding proteins, Alyref and Fus, mediate miRNA sorting into sEVs, enriching the understanding of exosomal miRNA biogenesis [99, 177]. In addition, activation of the NLRP3 inflammasome and cleavage of RILP increase exosome production, and the cleaved form of RILP interacts with FMR1 to regulate exosomal miR-155 content [178].
Some lncRNAs are also found in exosomes, probably by forming lncRNA–RBP complexes [179]. Although the specific mechanism of lncRNA sorting to exosomes is still unclear. Interestingly, lncRNA-encoded microproteins were identified in glioma-derived exosomes, indicating the biological functional diversity of exosomal lncRNAs [188].
CircRNA is a component of exosomal ncRNAs, and different circRNAs were found in exosomes originating from different cells, indicating that the sorting process of exosomal circRNAs is selective [189]. SNF8, a key component of the ESCRT-II complex, sorts circRHOBTB3 into exosomes by binding to specific sequences (141-240nt) on circRHOBTB3 [180]. After ectopic expression of miR-7 in HEK293T and MCF-7 cells, the level of circRNA CDR1as was significantly downregulated in exosomes but slightly increased in cells. This result partially suggests that sorting of circRNAs to exosomes was regulated by changes of associated miRNA levels [181].
The sorting mechanism of exosomal DNA is still unclear. Certain physiological pathways may be involved in this sorting process. MtDNA sorting to exosomes may be related to the endosomal pathway [153], and gDNA sorting to exosomes may involve micronuclei (MN) [182].
Protein sorting to exosomes
Typically, proteins can enter cells together with cell surface proteins by endocytosis and invagination of the plasma membrane [12]. The protein sorting process can be done in organelles such as mitochondria, endoplasmic reticulum and Golgi apparatus [190]. During the budding stage of exosome biogenesis, early-sorting endosome (ESE) may fuse and communicate with mitochondria, endoplasmic reticulum and trans-Golgi network, indicating the reason why the protein can be detected in exosomes of host cells [12]. In addition, some regulatory proteins in the formation, transport and secretion of exosomes may be involved in the sorting process of exosomal proteins, such as Rab GTPase, ESCRT proteins, tetraspanins and SNARE protein complexes [172, 191,192,193].
Some non-canonical mechanisms are also involved in the regulation of exosomal protein sorting. A recent study shows that proteins containing the KFERQ pentapeptide sequence can be sorted to exosomes by a process dependent on the membrane protein LAMP2A, a novel mechanism independent of ESCRT [183, 184]. Epigenetic modifications can be involved in the sorting of exosomal proteins. Cav1 can be sorted to MVB and ILV in a phosphorylation and ubiquitination-dependent manner, regulates exosome biogenesis by regulating MVB cholesterol contents, and delivers specific ECM-associated proteins (Tenascin-C, fibronectin, nidogen, elilin, EDIL3 and heparan sulfate proteoglycans) to exosomes [185]. Some proteins have UBL domains (ubiquitin-like sequences), among which UBL3/MUB proteins, as one of the conserved UBLs, can act as post-translational modification (PTM) factors to regulate the process of protein sorting to sEVs [186]. In addition, endosomal microautophagy is also involved in the sorting of exosomal proteins, and the chaperone HSC70-mediated proteins are sorted into exosomes under the electrostatic interaction between the cationic domain of HSC70 and the MVB membrane [187, 194].
Metabolites sorting to exosomes
Exosomes contain intact metabolites (mainly lipid metabolites) with characteristic of the host cells [195]. For example, exosomes secreted by granulocyte-myeloid derived suppressor cells (G-MDSCs) are enriched in PGE2, exosomes secreted by neutrophils are enriched in LTB4 [158, 160]. Triglycerides (TG) and sphingomyelin were found in T cell-derived exosomes isolated from human plasma [196]. Metabolomics of cancer stem cell-derived exosomes from melanoma revealed that multiple lipid metabolites, such as glycerophosphoglycerol (PG), glycerophosphatidylserine (PS), TG, and glycerophosphorylcholine (PC) in exosomes [197]. Currently, the mechanism by which these metabolites are sorted to exosomes remains unclear, but may be related to the interaction of lyn and fotillin-1 through the lipid domains of exosomal lipid membranes [159].
Biology of tumor metabolic reprogramming
Metabolic abnormalities are one of the hallmarkers of tumor cells, which are metabolically reprogrammed to meet their rapid proliferation requirements [198]. Given the specific physicochemical characteristics of high pressure, high pH and hypoxia within the TME, as well as the heterogeneity of the tumor vasculature, local tumor cells often have limited metabolic resources, which accelerates the digestion of nutrients and the accumulation of metabolites. In this particular environment, tumor cells adjust their metabolism in order to maintain their growth, which not only allows them to re-meet their energy supply needs, but also regulates their gene expression and protein modifications, facilitating the spread of tumor cells [199].
Glucose metabolism
Glucose is the main source of energy for cellular metabolism and biosynthesis. Glucose metabolism includes the glycolytic pathway, the pentose phosphate pathway (PPP), the serine synthesis pathway (SSP) and the tricarboxylic acid (TCA) cycle pathway [200]. Tumor cells are able to re-edit these pathways to obtain ATP with various biological macromolecules. In contrast to normal cells, tumor cells produce large amounts of lactate via the aerobic glycolytic pathway, resulting in an acidic TME that contributes to the proliferation [201]. In addition, increased uptake of glucose leads to the accumulation of intermediate metabolites in the glycolytic pathway, activating the cellular PPP while inhibiting the intracellular TCA[297]. During the release of exosomes, phosphorylated PKM2 acts as a protein kinase to promote the formation of SNARE complexes by enhancing the phosphorylation of SNAP23 [298]. During the release of exosomes, phosphorylated PKM2 acts as a protein kinase to promote the formation of SNARE complexes by enhancing the phosphorylation of SNAP23. PKM2 combines metabolic regulation with non-metabolic regulation of exosome secretion, is an ideal target for exosome and tumor metabolic therapy. Shikonin is the active ingredient of Comfrey, a naphthoquinone compound [290]. Shikonin is a specific PKM2 inhibitor that not only inhibits glucose uptake and lactate production in tumor cells, but also inhibits glycolysis by reducing extracellular secretion of exosomal PKM2 and enhances cisplatin sensitivity in NSCLC cells [290]. A study on bladder cancer found that highly-expression of PKM2 was associated with cisplatin resistance. Shikonin can promote cisplatin sensitivity of bladder cancer cells by reducing the release of exosomes, but the specific mechanism remains to be explored [291].
Exosomal ncRNAs, as common cargos in TME, play an important role in the metabolic reprogramming of TME. It was found that oxaliplatin-resistant CRC cells-derived exosomal circRNA ciRS-122 was delivered to sensitive cells, which enhanced glycolysis and chemoresistance in sensitive cells via miR-122/PKM2 signaling axis [292]. Development of exosome-transported si-ciRS-122 can reverse the ciRS-122/miR-122/PKM2 signaling axis to inhibit glycolysis and enhance chemosensitivity in CRC cells. In addition to tumor cells, targeting exosomal circRNAs derived from CAFs in TME has antitumor effects. It was found that exosomal circCCT3 secreted by CAFs could enhance glucose metabolism by regulating the expression of HK2. It was found that exosomal circCCT3 secreted by CAFs could enhance glucose metabolism by regulating the expression of HK2. Treatment of CAFs with coptisine inhibited the secretion of exosomal circCCT3 and suppressed HCC cell proliferation and invasion [293]. In addition, docosahexaenoic acid (DHA) as an omega 3 free fatty acid has been reported to exert anti-angiogenesis effects. DHA can alter the expression of angiogenesis-related exosomal miRNAs in breast cancer cells, inhibits angiogenesis by up-regulating exosomal miR-101, miR-199, and miR-342, and down-regulating exosomal mir-382 and miR-21 to exert anti-tumor effects [294, 299].
Benefiting from the targeting and biocompatibility of exosomes, exploitation of exosomes as carriers for drug delivery targeting tumor metabolism has a bright future. Although there are currently no engineered exosomes to directly target various metabolic pathways in the TME, exosomes can be combined with classical drugs or modalities as an adjuvant therapy to improve the efficacy of anti-tumor therapy. It has been shown that combination of exosome-mediated cPLA2 siRNA and metformin reduces the growth of glioblastoma xenografts by impairing the energy metabolism of mitochondria [295]. Photodynamic therapy (PDT) is a novel method of treating tumors with photosensitizing drugs and laser activation [300]. Aggregation-induced emission luminogens (AIEgens) are photosensitizers for PDT whose efficacy is limited by GSH. A recent work developed TEX for the co-delivery of AIEgens and proton pump inhibitor (PPI) for tumor combination therapy. TEX can specifically deliver AIEgens and PPI to tumor sites, and PPI inhibits GSH and ATP produced by glutamine metabolism in tumor cells, which contribute to the efficacy of AIEgens [296]. The combination of exosomes, glutamine metabolism and PDT may be a new option for future tumor treatment, but treatments that inhibit glutamine metabolism still need to be approached with caution. Glutamine depletion may stimulate release from Rab11a compartments of exosomes with pro-tumorigenic functions [301]. Therefore, exosome-based tumor metabolic therapy still needs further refinement to find the balance between pro-tumorigenesis and anti-tumorigenesis.
Conclusion
This review highlights the multiple roles and molecular mechanisms of exosome-mediated metabolic reprogramming in TME reprogramming. The field of exosomes (or EVs) has made great progress in recent years benefiting from technological breakthroughs in isolation, purification, in vivo tracking and content analysis [2]. This has led to the identification of other types of EVs besides exosomes and their functions becoming a novel hotspot in the field of EVs. In the future, the understanding of exosomes will be enriched by how to precisely distinguish exosomes from other EVs subtypes and exclude contaminants to further obtain high purity exosomes. This will also help to improve the targeting and biosafety of antitumor therapies developed with exosomes as vectors.
We describe the cell–cell communication mediated by exosomal cargos in TME and how these cargos are sorted to exosomes. Along with technological advances, the way of sorting various types of cargos into exosomes (or specific subtypes of EVs) is the bottleneck for further development in the field of exosomes. The bioactive cargos are the key to the function of exosomes. In addition to the cell-derived exosomal cargos in human TME, milk exosomes have a bright future as an oral drug delivery system, due to the biocompatibility of milk exosomes with exogenous cargos [302].
The heterogeneity of TME promotes tumor proliferation, metastasis, stemness and drug resistance. We summarized the main components and characteristics of TME, and highlighted the role and mechanism of exosomal cargos-mediated metabolic reprogramming in the heterogeneity of TME. Improving TME becomes an emerging strategy for anti-tumor treatment. The plasticity of tumor metabolism is both promising and challenging. Given the complex composition of TME, targeting one component for metabolic remodeling is difficult, and we need to consider more whether altered metabolism has the same therapeutic effects on multiple components of TME. Application of tumor organoid platforms to exosomes may be used to simulate the effect of exosomes on TME.
Exosomal cargos-mediated abnormalities metabolism in TME remains to be extensively studied. Considering the widespread of exosomal cargos, exploring the molecular mechanisms of exosomal cargos-induced metabolic reprogramming is beneficial for tumor precision treatment. As more and more biologic companies are entering the exosome field, the development of exosome-based drug delivery modalities to reshape metabolism in TME is promising. Combining chemotherapy, radiotherapy or targeted therapy with novel metabolic therapies may be the future trend in tumor treatment.
Availability of data and materials
Not applicable.
Abbreviations
- TME:
-
Tumor microenvironment
- EVs:
-
Extracellular vesicles
- MVs:
-
Microvesicles
- ncRNA:
-
Non-coding RNA
- CAFs:
-
Cancer-associated fibroblasts
- ECM:
-
Extracellular matrix
- NFs:
-
Normal fibroblasts
- HGSOC:
-
High-grade serous ovarian cancer
- TAMs:
-
Tumor-associated macrophages
- MDMs:
-
Monocyte-derived macrophages
- RCC:
-
Renal cell carcinoma
- APCs:
-
Antigen-presenting cells
- TCR:
-
T cell receptor
- CTLs:
-
Cytotoxic T lymphocytes
- ECs:
-
Endothelial cells
- ISF:
-
Interstitial fluid
- LECs:
-
Lymphatic endothelial cells
- WAT:
-
White adipose tissue
- CAAs:
-
Cancer-associated adipocytes
- HSCs:
-
Hepatic stellate cells
- PSCs:
-
Pancreatic stellate cells
- LOXs:
-
Lysyl oxidases
- LHs:
-
Lysyl hydroxylases
- HIFs:
-
Hypoxia-inducible factors
- ILV:
-
Intraluminal vescicles
- MVE:
-
Multivesicular endosome
- sEVs:
-
Small extracellular vesicles
- ESCRT:
-
Endosomal sorting complex
- MVB:
-
Multivesicular body
- MMPs:
-
Matrix metalloproteinases
- DGUC:
-
Density gradient differential ultracentrifugatio
- SEC:
-
Size exclusion chromatography
- UF:
-
Ultrafiltration
- AIEX:
-
Anion exchange chromatography
- gDNA:
-
Genomic DNA
- ccRCC:
-
Clear cell renal cell carcinoma
- OPRs:
-
Oxysterol-binding protein-related proteins
- nSMase2:
-
Neural sphingomyelinase 2
- hnRNP:
-
Heterogeneous nuclear ribonucleoprotein
- Ago2:
-
Argonaute 2
- MN:
-
Micronuclei
- ESE:
-
Early-sorting endosome
- PTM:
-
Post-translational modification
- G-MDSCs:
-
Granulocyte-myeloid derived suppressor cells
- TG:
-
Triglycerides
- PG:
-
Glycerophosphoglycerol
- PS:
-
Glycerophosphatidylserine
- PC:
-
Glycerophosphorylcholine
- PPP:
-
Pentose phosphate pathway
- SSP:
-
Serine synthesis pathway
- TCA:
-
Tricarboxylic acid
- PKM2:
-
Pyruvate kinase 2
- FA:
-
Fatty acid
- FATP:
-
Fatty acid transporter protein
- FABPpm:
-
Plasma membrane fatty acid binding protein
- ACLY:
-
ATP-citrate lyase
- FASN:
-
Fatty acid synthase
- EMT:
-
Epithelial-mesenchymal transition
- Gln:
-
Glutamine
- Ser:
-
Serine
- Trp:
-
Tryptophan
- Kyn:
-
Kynurenine
- 5-HT:
-
5-Hydroxytryptamine
- MDSCs:
-
Myeloid-derived suppressor cells
- TDO:
-
Tryptophan 2, 3-dioxygenase
- BC:
-
Breast cancer
- HCC:
-
Hepatocellular carcinoma
- TNBC:
-
Triple-negative breast cancer
- CRC:
-
Colorectal cancer
- AML:
-
Acute myeloid leukemia
- EPC:
-
Endothelial progenitor cell
- HUVECs:
-
Human umbilical vein endothelial cells
- HNSCC:
-
Head and neck squamous cell carcinoma
- TEXs:
-
Tumor cell-derived exosomes
- ADO:
-
Adenosine
- DCs:
-
Dendritic cells
- AM:
-
Adrenomedullin
- CRLM:
-
Metastasis of colorectal cancer
- PDAC:
-
Pancreatic ductal adenocarcinoma
- OXPHOS:
-
Oxidative phosphorylation
- LUAD:
-
Lung adenocarcinoma
- HTR:
-
Hormonal therapy-resistant
- FAO:
-
Fatty acid oxidation
- NSCLC:
-
Non-small cell lung cancer
- DHA:
-
Docosahexaenoic acid
- PDT:
-
Photodynamic therapy
- PPI:
-
Proton pump inhibitor
References
Qian F, Huang Z, Zhong H, Lei Q, Ai Y, **e Z, et al. Analysis and biomedical applications of functional cargo in extracellular vesicles. ACS Nano. 2022;16(12):19980–20001.
Lucotti S, Kenific CM, Zhang H, Lyden D. Extracellular vesicles and particles impact the systemic landscape of cancer. EMBO J. 2022;41(18):e109288.
Thery C, Witwer KW, Aikawa E, Alcaraz MJ, Anderson JD, Andriantsitohaina R, et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the international society for extracellular vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles. 2018;7(1):1535750.
**e F, Zhou X, Fang M, Li H, Su P, Tu Y, et al. Extracellular vesicles in cancer immune microenvironment and cancer immunotherapy. Adv Sci (Weinh). 2019;6(24):1901779.
Zhou M, Li YJ, Tang YC, Hao XY, Xu WJ, **ang DX, et al. Apoptotic bodies for advanced drug delivery and therapy. J Control Release. 2022;351:394–406.
Bian X, **ao YT, Wu T, Yao M, Du L, Ren S, et al. Microvesicles and chemokines in tumor microenvironment: mediators of intercellular communications in tumor progression. Mol Cancer. 2019;18(1):50.
Zhang H, Freitas D, Kim HS, Fabijanic K, Li Z, Chen H, et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat Cell Biol. 2018;20(3):332–43.
Zhang Q, Jeppesen DK, Higginbotham JN, Graves-Deal R, Trinh VQ, Ramirez MA, et al. Supermeres are functional extracellular nanoparticles replete with disease biomarkers and therapeutic targets. Nat Cell Biol. 2021;23(12):1240–54.
Ma L, Li Y, Peng J, Wu D, Zhao X, Cui Y, et al. Discovery of the migrasome, an organelle mediating release of cytoplasmic contents during cell migration. Cell Res. 2015;25(1):24–38.
Zhao X, Lei Y, Zheng J, Peng J, Li Y, Yu L, et al. Identification of markers for migrasome detection. Cell Discov. 2019;5:27.
Jiao H, Jiang D, Hu X, Du W, Ji L, Yang Y, et al. Mitocytosis, a migrasome-mediated mitochondrial quality-control process. Cell. 2021;184(11):2896-910 e13.
Kalluri R, LeBleu V. The biology function and biomedical applications of exosomes. Science (New York, NY). 2020;367(6478):eaau6977.
Shao H, Im H, Castro CM, Breakefield X, Weissleder R, Lee H. New Technologies for Analysis of Extracellular Vesicles. Chem Rev. 2018;118(4):1917–50.
Srivastava A, Rathore S, Munshi A, Ramesh R. Organically derived exosomes as carriers of anticancer drugs and imaging agents for cancer treatment. Semin Cancer Biol. 2022;86:80.
Ma YS, Yang XL, **n R, Liu JB, Fu D. Power and promise of exosomes as clinical biomarkers and therapeutic vectors for liquid biopsy and cancer control. Biochim Biophys Acta Rev Cancer. 2021;1875(1):188497.
Cappello F, Fais S. Extracellular vesicles in cancer pros and cons: The importance of the evidence-based medicine. Semin Cancer Biol. 2022;86:4.
Yang K, Zhou Q, Qiao B, Shao B, Hu S, Wang G, et al. Exosome-derived noncoding RNAs: Function, mechanism, and application in tumor angiogenesis. Mol Ther Nucleic Acids. 2022;27:983–97.
Banik A, Sharma R, Chauhan A, Singh S. Cutting the umbilical cord: Cancer stem cell-targeted therapeutics. Life Sci. 2022;299:120502.
Wu Y, Niu D, Deng S, Lei X, **e Z, Yang X. Tumor-derived or non-tumor-derived exosomal noncodingRNAs and signaling pathways in tumor microenvironment. Int Immunopharmacol. 2022;106:108626.
Lampropoulou D, Pliakou E, Aravantinos G, Filippou D, Gazouli M. The Role of Exosomal Non-Coding RNAs in Colorectal Cancer Drug Resistance. Int J Mol Sci. 2022;23(3):1473.
Arora S, Khan S, Zaki A, Tabassum G, Mohsin M, Bhutto H, et al. Integration of chemokine signaling with non-coding RNAs in tumor microenvironment and heterogeneity in different cancers. Semin Cancer Biol. 2022;86:720.
Aghanejad A, Bonab S, Sepehri M, Haghighi F, Tarighatnia A, Kreiter C, et al. A review on targeting tumor microenvironment: The main paradigm shift in the mAb-based immunotherapy of solid tumors. Int J Biol Macromol. 2022;207:592–610.
Petroni G, Buqué A, Coussens L, Galluzzi L. Targeting oncogene and non-oncogene addiction to inflame the tumour microenvironment. Nat Rev Drug Discovery. 2022;21(6):440–62.
Arora S, Khan S, Zaki A, Tabassum G, Mohsin M, Bhutto HN, et al. Integration of chemokine signaling with non-coding RNAs in tumor microenvironment and heterogeneity in different cancers. Semin Cancer Biol. 2022;86(Pt 2):720–36.
Maman S, Witz IP. A history of exploring cancer in context. Nat Rev Cancer. 2018;18(6):359–76.
**a L, Oyang L, Lin J, Tan S, Han Y, Wu N, et al. The cancer metabolic reprogramming and immune response. Mol Cancer. 2021;20(1):28.
** Q, Yan R, Cheng X, Wang W, Zhong Y, Hou Z, et al. Cancer-associated fibroblasts: overview, progress, challenges, and directions. Cancer Gene Ther. 2021;28(9):984–99.
Czekay R, Cheon D, Samarakoon R, Kutz S, Higgins P. Cancer-associated fibroblasts: mechanisms of tumor progression and novel therapeutic targets. Cancers. 2022;14(5):1231.
Poon S, Ailles L. Modeling the role of cancer-associated fibroblasts in tumor cell invasion. Cancers. 2022;14(4):962.
Jia W, Liang S, Cheng B, Ling C. The role of cancer-associated fibroblasts in hepatocellular carcinoma and the value of traditional Chinese medicine treatment. Front Oncol. 2021;11:763519.
Bejarano L, Jordāo M, Joyce J. Therapeutic targeting of the tumor microenvironment. Cancer Discov. 2021;11(4):933–59.
Costa A, Kieffer Y, Scholer-Dahirel A, Pelon F, Bourachot B, Cardon M, et al. Fibroblast heterogeneity and immunosuppressive environment in human breast cancer. Cancer Cell. 2018;33(3):463-79.e10.
Givel A, Kieffer Y, Scholer-Dahirel A, Sirven P, Cardon M, Pelon F, et al. miR200-regulated CXCL12β promotes fibroblast heterogeneity and immunosuppression in ovarian cancers. Nat Commun. 2018;9(1):1056.
Pelon F, Bourachot B, Kieffer Y, Magagna I, Mermet-Meillon F, Bonnet I, et al. Cancer-associated fibroblast heterogeneity in axillary lymph nodes drives metastases in breast cancer through complementary mechanisms. Nat Commun. 2020;11(1):404.
Han C, Zhang C, Wang H, Zhao L. Exosome-mediated communication between tumor cells and tumor-associated macrophages: implications for tumor microenvironment. Oncoimmunology. 2021;10(1):1887552.
DeNardo D, Ruffell B. Macrophages as regulators of tumour immunity and immunotherapy. Nat Rev Immunol. 2019;19(6):369–82.
Inagaki K, Kunisho S, Takigawa H, Yuge R, Oka S, Tanaka S, et al. Role of tumor-associated macrophages at the invasive front in human colorectal cancer progression. Cancer Sci. 2021;112(7):2692–704.
Tan S, **a L, Yi P, Han Y, Tang L, Pan Q, et al. Exosomal miRNAs in tumor microenvironment. J Exp Clin Cancer Res: CR. 2020;39(1):67.
Sica A, Mantovani A. Macrophage plasticity and polarization: in vivo veritas. J Clin Investig. 2012;122(3):787–95.
Nirschl T, El Asmar M, Ludwig W, Ganguly S, Gorin M, Johnson M, et al. Transcriptional profiling of tumor associated macrophages in human renal cell carcinoma reveals significant heterogeneity and opportunity for immunomodulation. Am J Clin Exp Urol. 2020;8(1):48–58.
Marku M, Verstraete N, Raynal F, Madrid-Mencía M, Domagala M, Fournié J, et al. Insights on TAM formation from a Boolean model of macrophage polarization based on in vitro studies. Cancers. 2020;12(12):3664.
Dang M, Gonzalez M, Gaonkar K, Rathi K, Young P, Arif S, et al. Macrophages in SHH subgroup medulloblastoma display dynamic heterogeneity that varies with treatment modality. Cell Rep. 2021;34(13):108917.
**e F, Zhou X, Fang M, Li H, Su P, Tu Y, et al. Extracellular vesicles in cancer immune microenvironment and cancer immunotherapy. Adv Sci (Weinheim, Baden-Wurttemberg, Germany). 2019;6(24):1901779.
Saillard M, Cenerenti M, Romero P, Jandus C. Impact of immunotherapy on CD4 T cell phenotypes and function in cancer. Vaccines. 2021;9(5):454.
Miggelbrink A, Jackson J, Lorrey S, Srinivasan E, Waibl-Polania J, Wilkinson D, et al. CD4 T-cell exhaustion: does it exist and what are its roles in cancer? Clin Cancer Res J Am Asso Cancer Res. 2021;27(21):5742–52.
Yang P, Peng Y, Feng Y, Xu Z, Feng P, Cao J, et al. Immune cell-derived extracellular vesicles - new strategies in cancer immunotherapy. Front Immunol. 2021;12:771551.
Nanke Y, Kobashigawa T, Yago T, Kawamoto M, Yamanaka H, Kotake S. Detection of IFN-γ+IL-17+ cells in salivary glands of patients with Sjögren’s syndrome and Mikulicz’s disease: Potential role of Th17 Th1 in the pathogenesis of autoimmune diseases Nihon Rinsho Men’eki Gakkai kaishi. Jap J Clin Immunol. 2016;39(5):473–7.
Farhood B, Najafi M, Mortezaee K. CD8 cytotoxic T lymphocytes in cancer immunotherapy: A review. J Cell Physiol. 2019;234(6):8509–21.
Maimela N, Liu S, Zhang Y. Fates of CD8+ T cells in Tumor Microenvironment. Comput Struct Biotechnol J. 2019;17:1–13.
Henning A, Roychoudhuri R, Restifo N. Epigenetic control of CD8 T cell differentiation. Nat Rev Immunol. 2018;18(5):340–56.
Dong D, Fan P, Feng Y, Liu G, Peng Y, Dong T, et al. Association between circulating CD39+CD8+ T cells pre-chemoradiotherapy and prognosis in patients with nasopharyngeal carcinoma. Chin Med J. 2021;134(17):2066–72.
Jørgensen N, Hviid T, Nielsen L, Sønderstrup I, Eriksen J, Ejlertsen B, et al. Tumour-infiltrating CD4-, CD8- and FOXP3-positive immune cells as predictive markers of mortality in BRCA1- and BRCA2-associated breast cancer. Br J Cancer. 2021;125(10):1388–98.
Giatromanolaki A, Anestopoulos I, Panayiotidis M, Mitrakas A, Pappa A, Koukourakis M. Prognostic relevance of the relative presence of CD4, CD8 and CD20 expressing tumor infiltrating lymphocytes in operable non-small cell lung cancer patients. Anticancer Res. 2021;41(8):3989–95.
Toor S, Sasidharan Nair V, Saleh R, Taha R, Murshed K, Al-Dhaheri M, et al. Transcriptome of tumor-infiltrating T cells in colorectal cancer patients uncovered a unique gene signature in CD4 T cells associated with poor disease-specific survival. Vaccines. 2021;9(4):334.
Borsetto D, Tomasoni M, Payne K, Polesel J, Deganello A, Bossi P, et al. Prognostic significance of CD4+ and CD8+ tumor-infiltrating lymphocytes in head and neck squamous cell carcinoma: a meta-analysis. Cancers. 2021;13(4):781.
Gu Y, Chen Y, ** K, Cao Y, Liu X, Lv K, et al. Intratumoral CD103CD4 T cell infiltration defines immunoevasive contexture and poor clinical outcomes in gastric cancer patients. Oncoimmunology. 2020;9(1):1844402.
Wang W, Green M, Choi J, Gijón M, Kennedy P, Johnson J, et al. CD8 T cells regulate tumour ferroptosis during cancer immunotherapy. Nature. 2019;569(7755):270–4.
Ma X, Bi E, Lu Y, Su P, Huang C, Liu L, et al. Cholesterol induces CD8 T cell exhaustion in the tumor microenvironment. Cell Metab. 2019;30(1):143-56.e5.
Knox MC, Ni J, Bece A, Bucci J, Chin Y, Graham PH, et al. A clinician’s guide to cancer-derived exosomes: immune interactions and therapeutic implications. Front Immunol. 2020;11:1612.
Taskaeva I, Bgatova N. Microvasculature in hepatocellular carcinoma: an ultrastructural study. Microvasc Res. 2021;133:104094.
Schito L, Rey S. Hypoxia: turning vessels into vassals of cancer immunotolerance. Cancer Lett. 2020;487:74–84.
Schaaf M, Garg A, Agostinis P. Defining the role of the tumor vasculature in antitumor immunity and immunotherapy. Cell Death Dis. 2018;9(2):115.
Peri S, Biagioni A, Versienti G, Andreucci E, Staderini F, Barbato G, et al. Enhanced vasculogenic capacity induced by 5-fluorouracil chemoresistance in a gastric cancer cell line. Int J Mol Sci. 2021;22(14):7698.
Morales-Guadarrama G, García-Becerra R, Méndez-Pérez E, García-Quiroz J, Avila E, Díaz L. Vasculogenic mimicry in breast cancer: clinical relevance and drivers. Cells. 2021;10(7):1758.
Jia T, Jacquet T, Dalonneau F, Coudert P, Vaganay E, Exbrayat-Héritier C, et al. FGF-2 promotes angiogenesis through a SRSF1/SRSF3/SRPK1-dependent axis that controls VEGFR1 splicing in endothelial cells. BMC Biol. 2021;19(1):173.
Haider T, Sandha K, Soni V, Gupta P. Recent advances in tumor microenvironment associated therapeutic strategies and evaluation models. Mater Sci Eng, C Mater Biol Appl. 2020;116:111229.
Ichikawa K, Watanabe Miyano S, Minoshima Y, Matsui J, Funahashi Y. Activated FGF2 signaling pathway in tumor vasculature is essential for acquired resistance to anti-VEGF therapy. Sci Rep. 2020;10(1):2939.
Tsioumpekou M, Cunha S, Ma H, Åhgren A, Cedervall J, Olsson A, et al. Specific targeting of PDGFRβ in the stroma inhibits growth and angiogenesis in tumors with high PDGF-BB expression. Theranostics. 2020;10(3):1122–35.
Petrova T, Koh G. Biological functions of lymphatic vessels. Sci (New York, NY). 2020;369(6500):eaax4063.
das Neves SP, Delivanoglou N, Da Mesquita S. CNS-draining meningeal lymphatic vasculature: roles conundrums and future challenges. Front Pharmacol. 2021;12:655052.
Garnier L, Gkountidi A, Hugues S. Tumor-associated lymphatic vessel features and immunomodulatory functions. Front Immunol. 2019;10:720.
Brouillard P, Dupont L, Helaers R, Coulie R, Tiller G, Peeden J, et al. Loss of ADAMTS3 activity causes Hennekam lymphangiectasia-lymphedema syndrome 3. Hum Mol Genet. 2017;26(21):4095–104.
Jha S, Rauniyar K, Chronowska E, Mattonet K, Maina E, Koistinen H, et al. KLK3/PSA and cathepsin D activate VEGF-C and VEGF-D. eLife. 2019;8:e44478.
Jha S, Rauniyar K, Jeltsch M. Key molecules in lymphatic development, function, and identification. Ann Anatomy Anatomischer Anzeiger Organ Anatomische Gesellschaft. 2018;219:25–34.
Petrova TV, Koh GY. Biological functions of lymphatic vessels. Science. 2020;369(6500):eaax063.
Yu P, Wu G, Lee H, Simons M. Endothelial metabolic control of lymphangiogenesis. BioEssays. 2018;40(6):e1700245.
Wilczak W, Wittmer C, Clauditz T, Minner S, Steurer S, Büscheck F, et al. marked prognostic impact of minimal lymphatic tumor spread in prostate cancer. Eur Urol. 2018;74(3):376–86.
Chen C, Zheng H, Luo Y, Kong Y, An M, Li Y, et al. SUMOylation promotes extracellular vesicle-mediated transmission of lncRNA ELNAT1 and lymph node metastasis in bladder cancer. J Clin Invest. 2021;131(8):146431.
Qu W, Li S, Zhang M, Qiao Q. Pattern and prognosis of distant metastases in nasopharyngeal carcinoma: A large-population retrospective analysis. Cancer Med. 2020;9(17):6147–58.
Jain R, Martin J, Stylianopoulos T. The role of mechanical forces in tumor growth and therapy. Annu Rev Biomed Eng. 2014;16:321–46.
Li Z, Zhang X, Liu C, Ma J. Non-immune cell components in the gastrointestinal tumor microenvironment influencing tumor immunotherapy. Front Cell Dev Biol. 2021;9:729941.
Leary N, Walser S, He Y, Cousin N, Pereira P, Gallo A, et al. Melanoma-derived extracellular vesicles mediate lymphatic remodelling and impair tumour immunity in draining lymph nodes. J Extracell Vesicles. 2022;11(2):e12197.
Liu C, Wang Y, Li L, He D, Chi J, Li Q, et al. Engineered extracellular vesicles and their mimetics for cancer immunotherapy. J Control Release. 2022;349:679–98.
Tang Y, Zhang W, Sheng T, He X, **ong X. Overview of the molecular mechanisms contributing to the formation of cancer-associated adipocytes (Review). Mol Med Rep. 2021;24(5):768.
Iyengar N, Zhou X, Mendieta H, Giri D, El-Hely O, Winston L, et al. Effects of adiposity and exercise on breast tissue and systemic metabo-inflammatory factors in women at high risk or diagnosed with breast cancer. Cancer Prev Res (Phila). 2021;14(5):541–50.
Saha A, Hamilton-Reeves J, DiGiovanni J. White adipose tissue-derived factors and prostate cancer progression: mechanisms and targets for interventions. Cancer Metastasis Rev. 2022;41:649.
Liu A, Pan W, Zhuang S, Tang Y, Zhang H. Cancer cell-derived exosomal miR-425-3p induces white adipocyte atrophy. Adipocyte. 2022;11(1):487–500.
Wu Y, Li X, Li Q, Cheng C, Zheng L. Adipose tissue-to-breast cancer crosstalk: Comprehensive insights. Biochim Biophys Acta Rev Cancer. 2022;1877(5):188800.
Seki T, Yang Y, Sun X, Lim S, **e S, Guo Z, et al. Brown-fat-mediated tumour suppression by cold-altered global metabolism. Nature. 2022;608(7922):421–8.
Oguri Y, Shinoda K, Kim H, Alba D, Bolus W, Wang Q, et al. CD81 controls beige fat progenitor cell growth and energy balance via FAK signaling. Cell. 2020;182(3):563-77.e20.
Han J, Jeon Y, Oh M, Lee G, Nahmgoong H, Han S, et al. Adipocyte HIF2α functions as a thermostat via PKA Cα regulation in beige adipocytes. Nat Commun. 2022;13(1):3268.
Gantov M, Pagnotta P, Lotufo C, Rindone G, Riera M, Calvo J, et al. Beige adipocytes contribute to breast cancer progression. Oncol Rep. 2021;45(1):317–28.
Suárez-Nájera L, Chanona-Pérez J, Valdivia-Flores A, Marrero-Rodríguez D, Salcedo-Vargas M, García-Ruiz D, et al. Morphometric study of adipocytes on breast cancer by means of photonic microscopy and image analysis. Microsc Res Tech. 2018;81(2):240–9.
Wu Q, Li B, Li Z, Li J, Sun S, Sun S. Cancer-associated adipocytes: key players in breast cancer progression. J Hematol Oncol. 2019;12(1):95.
Munteanu R, Onaciu A, Moldovan C, Zimta A, Gulei D, Paradiso A, et al. Adipocyte-based cell therapy in oncology: the role of cancer-associated adipocytes and their reinterpretation as delivery platforms. Pharmaceutics. 2020;12(5):402.
Zhao C, Wu M, Zeng N, **ong M, Hu W, Lv W, et al. Cancer-associated adipocytes: emerging supporters in breast cancer. J Exp Clin Cancer Res CR. 2020;39(1):156.
Trivedi P, Wang S, Friedman S. The power of plasticity-metabolic regulation of hepatic stellate cells. Cell Metab. 2021;33(2):242–57.
Yin C, Evason K, Asahina K, Stainier D. Hepatic stellate cells in liver development, regeneration, and cancer. J Clin Investig. 2013;123(5):1902–10.
Zhao S, Mi Y, Zheng B, Wei P, Gu Y, Zhang Z, et al. Highly-metastatic colorectal cancer cell released miR-181a-5p-rich extracellular vesicles promote liver metastasis by activating hepatic stellate cells and remodelling the tumour microenvironment. J Extracell Vesicles. 2022;11(1):e12186.
Zhou Y, Ren H, Dai B, Li J, Shang L, Huang J, et al. Hepatocellular carcinoma-derived exosomal miRNA-21 contributes to tumor progression by converting hepatocyte stellate cells to cancer-associated fibroblasts. J Exp Clin Cancer Res CR. 2018;37(1):324.
Masamune A, Yoshida N, Hamada S, Takikawa T, Nabeshima T, Shimosegawa T. Exosomes derived from pancreatic cancer cells induce activation and profibrogenic activities in pancreatic stellate cells. Biochem Biophys Res Commun. 2018;495(1):71–7.
Zhang Y, Zhou Y, Zhang B, Huang S, Li P, He X, et al. Pancreatic cancer-derived exosomes promoted pancreatic stellate cells recruitment by pancreatic cancer. J Cancer. 2019;10(18):4397–407.
Schnittert J, Bansal R, Prakash J. Targeting pancreatic stellate cells in cancer. Trends Cancer. 2019;5(2):128–42.
Abyaneh H, Regenold M, McKee T, Allen C, Gauthier M. Towards extracellular matrix normalization for improved treatment of solid tumors. Theranostics. 2020;10(4):1960–80.
Henke E, Nandigama R, Ergün S. Extracellular matrix in the tumor microenvironment and its impact on cancer therapy. Front Mol Biosci. 2019;6:160.
Ayad N, Weaver V. Tension in tumour cells keeps metabolism high. Nature. 2020;578(7796):517–8.
Chaudhuri O, Cooper-White J, Janmey P, Mooney D, Shenoy V. Effects of extracellular matrix viscoelasticity on cellular behaviour. Nature. 2020;584(7822):535–46.
Fiore V, Krajnc M, Quiroz F, Levorse J, Pasolli H, Shvartsman S, et al. Publisher correction: mechanics of a multilayer epithelium instruct tumour architecture and function. Nature. 2020;586(7827):E9.
Mendonça D, Miguez P, Mendonça G, Yamauchi M, Aragão F, Cooper L. Titanium surface topography affects collagen biosynthesis of adherent cells. Bone. 2011;49(3):463–72.
Smithen D, Leung L, Challinor M, Lawrence R, Tang H, Niculescu-Duvaz D, et al. 2-Aminomethylene-5-sulfonylthiazole Inhibitors of Lysyl Oxidase (LOX) and LOXL2 show significant efficacy in delaying tumor growth. J Med Chem. 2020;63(5):2308–24.
Sullivan W, Mullen P, Schmid E, Flores A, Momcilovic M, Sharpley M, et al. Extracellular matrix remodeling regulates glucose metabolism through TXNIP destabilization. Cell. 2018;175(1):117-32.e21.
Torrino S, Grasset E, Audebert S, Belhadj I, Lacoux C, Haynes M, et al. Mechano-induced cell metabolism promotes microtubule glutamylation to force metastasis. Cell Metab. 2021;33(7):1342-57.e10.
Lequeux A, Noman MZ, **ao M, Van Moer K, Hasmim M, Benoit A, et al. Targeting HIF-1 alpha transcriptional activity drives cytotoxic immune effector cells into melanoma and improves combination immunotherapy. Oncogene. 2021;40(28):4725–35.
Bai R, Li Y, Jian L, Yang Y, Zhao L, Wei M. The hypoxia-driven crosstalk between tumor and tumor-associated macrophages: mechanisms and clinical treatment strategies. Mol Cancer. 2022;21(1):177.
**a X, Wang S, Ni B, **ng S, Cao H, Zhang Z, et al. Hypoxic gastric cancer-derived exosomes promote progression and metastasis via MiR-301a-3p/PHD3/HIF-1alpha positive feedback loop. Oncogene. 2020;39(39):6231–44.
Yan Y, He M, Zhao L, Wu H, Zhao Y, Han L, et al. A novel HIF-2alpha targeted inhibitor suppresses hypoxia-induced breast cancer stemness via SOD2-mtROS-PDI/GPR78-UPR(ER) axis. Cell Death Differ. 2022;29(9):1769–89.
Bui BP, Nguyen PL, Lee K, Cho J. Hypoxia-inducible factor-1: a novel therapeutic target for the management of cancer, drug resistance, and cancer-related pain. Cancers (Basel). 2022;14(24):6054.
Anvari G, Bellas E. Hypoxia induces stress fiber formation in adipocytes in the early stage of obesity. Sci Rep. 2021;11(1):21473.
Rosell-Garcia T, Palomo-Alvarez O, Rodriguez-Pascual F. A hierarchical network of hypoxia-inducible factor and SMAD proteins governs procollagen lysyl hydroxylase 2 induction by hypoxia and transforming growth factor beta1. J Biol Chem. 2019;294(39):14308–18.
Maruyama T, Shimoda M, Sako A, Ueda K, Hakoda H, Sakata A, et al. Predictive effectiveness of the glasgow prognostic score for gastrointestinal stromal tumors. Nutr Cancer. 2021;73(8):1333–9.
Jiang Y, Jiang H, Wang K, Liu C, Man X, Fu Q. Hypoxia enhances the production and antitumor effect of exosomes derived from natural killer cells. Ann Transl Med. 2021;9(6):473.
Qian D, **e Y, Huang M, Gu J. Tumor-derived exosomes in hypoxic microenvironment: release mechanism, biological function and clinical application. J Cancer. 2022;13(5):1685–94.
He G, Peng X, Wei S, Yang S, Li X, Huang M, et al. Exosomes in the hypoxic TME: from release, uptake and biofunctions to clinical applications. Mol Cancer. 2022;21(1):19.
Dorayappan KDP, Wanner R, Wallbillich JJ, Saini U, Zingarelli R, Suarez AA, et al. Hypoxia-induced exosomes contribute to a more aggressive and chemoresistant ovarian cancer phenotype: a novel mechanism linking STAT3/Rab proteins. Oncogene. 2018;37(28):3806–21.
Panigrahi GK, Praharaj PP, Peak TC, Long J, Singh R, Rhim JS, et al. Hypoxia-induced exosome secretion promotes survival of African-American and Caucasian prostate cancer cells. Sci Rep. 2018;8(1):3853.
Li B, Antonyak MA, Zhang J, Cerione RA. RhoA triggers a specific signaling pathway that generates transforming microvesicles in cancer cells. Oncogene. 2012;31(45):4740–9.
Kumar A, Deep G. Hypoxia in tumor microenvironment regulates exosome biogenesis: molecular mechanisms and translational opportunities. Cancer Lett. 2020;479:23–30.
Crewe C, Funcke J, Li S, Joffin N, Gliniak C, Ghaben A, et al. Extracellular vesicle-based interorgan transport of mitochondria from energetically stressed adipocytes. Cell Metab. 2021;33(9):1853-68.e11.
Tang X, Guo T, Gao X, Wu X, **ng X, Ji J, et al. Exosome-derived noncoding RNAs in gastric cancer: functions and clinical applications. Mol Cancer. 2021;20(1):99.
Book A. ISEV2017. J Extracell Vesicles. 2017;6(sup1):1310414.
Fan Q, Yang L, Zhang X, Peng X, Wei S, Su D, et al. The emerging role of exosome-derived non-coding RNAs in cancer biology. Cancer Lett. 2018;414:107–15.
Fukuda M. Rab27 effectors, pleiotropic regulators in secretory pathways. Traffic (Copenhagen, Denmark). 2013;14(9):949–63.
Wang M, Yu F, Li P, Wang K. Emerging function and clinical significance of exosomal circRNAs in cancer. Mol Ther Nucleic Acids. 2020;21:367–83.
Fukuda M. Rab27 effectors, pleiotropic regulators in secretory pathways. Traffic. 2013;14(9):949–63.
Klinkert K, Echard A. Rab35 GTPase: a central regulator of phosphoinositides and F-actin in endocytic recycling and beyond. Traffic (Copenhagen, Denmark). 2016;17(10):1063–77.
Arbo B, Cechinel L, Palazzo R, Siqueira I. Endosomal dysfunction impacts extracellular vesicle release: Central role in Aβ pathology. Ageing Res Rev. 2020;58:101006.
Corbeil D, Santos M, Karbanová J, Kurth T, Rappa G, Lorico A. Uptake and fate of extracellular membrane vesicles: nucleoplasmic reticulum-associated late endosomes as a new gate to intercellular communication. Cells. 2020;9(9):1931.
Li X, Wang Y, Wang Q, Liu Y, Bao W, Wu S. Exosomes in cancer: Small transporters with big functions. Cancer Lett. 2018;435:55–65.
Simonetti B, Cullen P. Actin-dependent endosomal receptor recycling. Curr Opin Cell Biol. 2019;56:22–33.
Nam G, Choi Y, Kim G, Kim S, Kim S, Kim I. Emerging prospects of exosomes for cancer treatment: from conventional therapy to immunotherapy. Adv Materials (Deerfield Beach, Fla). 2020;32(51):e2002440.
Fu M, Gu J, Jiang P, Qian H, Xu W, Zhang X. Exosomes in gastric cancer: roles, mechanisms, and applications. Mol Cancer. 2019;18(1):41.
Qiu Y, Chen T, Hu R, Zhu R, Li C, Ruan Y, et al. Next frontier in tumor immunotherapy: macrophage-mediated immune evasion. Biomark Res. 2021;9(1):72.
Gupta S, Rawat S, Arora V, Kottarath S, Dinda A, Vaishnav P, et al. An improvised one-step sucrose cushion ultracentrifugation method for exosome isolation from culture supernatants of mesenchymal stem cells. Stem Cell Res Ther. 2018;9(1):180.
Tian Y, Gong M, Hu Y, Liu H, Zhang W, Zhang M, et al. Quality and efficiency assessment of six extracellular vesicle isolation methods by nano-flow cytometry. J Extracell Vesicles. 2020;9(1):1697028.
Shire**i S, Inci F. The Yin and Yang of exosome isolation methods: conventional practice, microfluidics, and commercial kits. Biotechnol Adv. 2022;54:107814.
Tayebi M, Yang D, Collins D, Ai Y. Deterministic sorting of submicrometer particles and extracellular vesicles using a combined electric and acoustic field. Nano Lett. 2021;21(16):6835–42.
Tenchov R, Sasso JM, Wang X, Liaw WS, Chen CA, Zhou QA. Exosomes horizontal line nature’s lipid nanoparticles, a rising star in drug delivery and diagnostics. ACS Nano. 2022;16(11):17802–46.
Balaj L, Lessard R, Dai L, Cho YJ, Pomeroy SL, Breakefield XO, et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat Commun. 2011;2:180.
Isaac R, Reis FCG, Ying W, Olefsky JM. Exosomes as mediators of intercellular crosstalk in metabolism. Cell Metab. 2021;33(9):1744–62.
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9(6):654–9.
He X, Tian F, Guo F, Zhang F, Zhang H, Ji J, et al. Circulating exosomal mRNA signatures for the early diagnosis of clear cell renal cell carcinoma. BMC Med. 2022;20(1):270.
Sansone P, Savini C, Kurelac I, Chang Q, Amato LB, Strillacci A, et al. Packaging and transfer of mitochondrial DNA via exosomes regulate escape from dormancy in hormonal therapy-resistant breast cancer. Proc Natl Acad Sci U S A. 2017;114(43):E9066–75.
Torralba D, Baixauli F, Villarroya-Beltri C, Fernandez-Delgado I, Latorre-Pellicer A, Acin-Perez R, et al. Priming of dendritic cells by DNA-containing extracellular vesicles from activated T cells through antigen-driven contacts. Nat Commun. 2018;9(1):2658.
Skotland T, Sagini K, Sandvig K, Llorente A. An emerging focus on lipids in extracellular vesicles. Adv Drug Deliv Rev. 2020;159:308–21.
Menck K, Sonmezer C, Worst TS, Schulz M, Dihazi GH, Streit F, et al. Neutral sphingomyelinases control extracellular vesicles budding from the plasma membrane. J Extracell Vesicles. 2017;6(1):1378056.
Kobuna H, Inoue T, Shibata M, Gengyo-Ando K, Yamamoto A, Mitani S, et al. Multivesicular body formation requires OSBP-related proteins and cholesterol. PLoS Genet. 2010;6(8):e1001055.
Eden ER, Sanchez-Heras E, Tsapara A, Sobota A, Levine TP, Futter CE. Annexin A1 tethers membrane contact sites that mediate ER to endosome cholesterol transport. Dev Cell. 2016;37(5):473–83.
Majumdar R, Tavakoli Tameh A, Arya SB, Parent CA. Exosomes mediate LTB4 release during neutrophil chemotaxis. PLoS Biol. 2021;19(7):e3001271.
Arya SB, Chen S, Jordan-Javed F, Parent CA. Ceramide-rich microdomains facilitate nuclear envelope budding for non-conventional exosome formation. Nat Cell Biol. 2022;24(7):1019–28.
Wu X, Zhu D, Tian J, Tang X, Guo H, Ma J, et al. Granulocytic myeloid-derived suppressor cell exosomal prostaglandin E2 ameliorates collagen-induced arthritis by enhancing IL-10(+) B cells. Front Immunol. 2020;11:588500.
Zhang M, **n W, Ma C, Zhang H, Mao M, Liu Y, et al. Exosomal 15-LO2 mediates hypoxia-induced pulmonary artery hypertension in vivo and in vitro. Cell Death Dis. 2018;9(10):1022.
Eguchi T, Sheta M, Fujii M, Calderwood SK. Cancer extracellular vesicles, tumoroid models, and tumor microenvironment. Semin Cancer Biol. 2022;86(Pt 1):112–26.
Shi F, Deng Z, Zhou Z, Jiang B, Jiang CY, Zhao RZ, et al. Heat injured stromal cells-derived exosomal EGFR enhances prostatic wound healing after thulium laser resection through EMT and NF-kappaB signaling. Prostate. 2019;79(11):1238–55.
Zhang Z, Zhou Y, Jia Y, Wang C, Zhang M, Xu Z. PRR34-AS1 promotes exosome secretion of VEGF and TGF-beta via recruiting DDX3X to stabilize Rab27a mRNA in hepatocellular carcinoma. J Transl Med. 2022;20(1):491.
Patnam S, Samal R, Koyyada R, Joshi P, Singh AD, Nagalla B, et al. Exosomal PTEN as a predictive marker of aggressive gliomas. Neurol India. 2022;70(1):215–22.
Qin Z, Li Y, Li J, Jiang L, Zhang Z, Chang K, et al. Exosomal STAT1 derived from high phosphorus-stimulated vascular endothelial cells induces vascular smooth muscle cell calcification via the Wnt/beta-catenin signaling pathway. Int J Mol Med. 2022;50(6):139.
Clement E, Lazar I, Attane C, Carrie L, Dauvillier S, Ducoux-Petit M, et al. Adipocyte extracellular vesicles carry enzymes and fatty acids that stimulate mitochondrial metabolism and remodeling in tumor cells. EMBO J. 2020;39(3):e102525.
Wang D, Zhao C, Xu F, Zhang A, ** M, Zhang K, et al. Cisplatin-resistant NSCLC cells induced by hypoxia transmit resistance to sensitive cells through exosomal PKM2. Theranostics. 2021;11(6):2860–75.
Zhu L, Xu Y, Kang S, Lin B, Zhang C, You Z, et al. Quantification-promoted discovery of glycosylated exosomal PD-L1 as a potential tumor biomarker. Small Methods. 2022;6(9):e2200549.
Tan Y, Huang Y, Mei R, Mao F, Yang D, Liu J, et al. HucMSC-derived exosomes delivered BECN1 induces ferroptosis of hepatic stellate cells via regulating the xCT/GPX4 axis. Cell Death Dis. 2022;13(4):319.
Mańka R, Janas P, Sapoń K, Janas T, Janas T. Role of RNA motifs in RNA interaction with membrane lipid rafts: implications for therapeutic applications of exosomal RNAs. Int J Mol Sci. 2021;22(17):9416.
Matsui T, Sakamaki Y, Nakashima S, Fukuda M. Rab39 and its effector UACA regulate basolateral exosome release from polarized epithelial cells. Cell Rep. 2022;39(9):110875.
Kosaka N, Iguchi H, Hagiwara K, Yoshioka Y, Takeshita F, Ochiya T. Neutral sphingomyelinase 2 (nSMase2)-dependent exosomal transfer of angiogenic microRNAs regulate cancer cell metastasis. J Biol Chem. 2013;288(15):10849–59.
Villarroya-Beltri C, Gutierrez-Vazquez C, Sanchez-Cabo F, Perez-Hernandez D, Vazquez J, Martin-Cofreces N, et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat Commun. 2013;4:2980.
Koppers-Lalic D, Hackenberg M, Bijnsdorp IV, van Eijndhoven MAJ, Sadek P, Sie D, et al. Nontemplated nucleotide additions distinguish the small RNA composition in cells from exosomes. Cell Rep. 2014;8(6):1649–58.
Guduric-Fuchs J, O’Connor A, Camp B, O’Neill CL, Medina RJ, Simpson DA. Selective extracellular vesicle-mediated export of an overlap** set of microRNAs from multiple cell types. BMC Genomics. 2012;13:357.
Garcia-Martin R, Wang G, Brandão B, Zanotto T, Shah S, Kumar Patel S, et al. MicroRNA sequence codes for small extracellular vesicle release and cellular retention. Nature. 2022;601(7893):446–51.
Willson J. RILP gets cleaved and exosomes leave. Nat Rev Mol Cell Biol. 2020;21(11):658–9.
Statello L, Maugeri M, Garre E, Nawaz M, Wahlgren J, Papadimitriou A, et al. Identification of RNA-binding proteins in exosomes capable of interacting with different types of RNA: RBP-facilitated transport of RNAs into exosomes. PLoS ONE. 2018;13(4):e0195969.
Chen C, Yu H, Han F, Lai X, Ye K, Lei S, et al. Tumor-suppressive circRHOBTB3 is excreted out of cells via exosome to sustain colorectal cancer cell fitness. Mol Cancer. 2022;21(1):46.
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J, et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 2015;25(8):981–4.
Yokoi A, Villar-Prados A, Oliphint PA, Zhang J, Song X, De Hoff P, et al. Mechanisms of nuclear content loading to exosomes. Sci Adv. 2019;5(11):eaax8849.
Ferreira J, da Rosa SA, Pereira P. LAMP2A mediates the loading of proteins into endosomes and selects exosomal cargo. Autophagy. 2022;18(9):2263–5.
Ferreira J, da Rosa Soares A, Ramalho J, Máximo Carvalho C, Cardoso M, Pintado P, et al. LAMP2A regulates the loading of proteins into exosomes. Sci Adv. 2022;8(12):eabm1140.
Albacete-Albacete L, Navarro-Lérida I, López J, Martín-Padura I, Astudillo A, Ferrarini A, et al. ECM deposition is driven by caveolin-1-dependent regulation of exosomal biogenesis and cargo sorting. J Cell Biol. 2020;219(11):e202006178.
Ageta H, Ageta-Ishihara N, Hitachi K, Karayel O, Onouchi T, Yamaguchi H, et al. UBL3 modification influences protein sorting to small extracellular vesicles. Nat Commun. 2018;9(1):3936.
Sahu R, Kaushik S, Clement C, Cannizzo E, Scharf B, Follenzi A, et al. Microautophagy of cytosolic proteins by late endosomes. Dev Cell. 2011;20(1):131–9.
Cai T, Zhang Q, Wu B, Wang J, Li N, Zhang T, et al. LncRNA-encoded microproteins: a new form of cargo in cell culture-derived and circulating extracellular vesicles. J Extracell Vesicles. 2021;10(9):e12123.
Hinger S, Cha D, Franklin J, Higginbotham J, Dou Y, ** J, et al. Diverse long RNAs are differentially sorted into extracellular vesicles secreted by colorectal cancer cells. Cell Rep. 2018;25(3):715-25.e4.
Vats S, Galli T. Role of SNAREs in unconventional secretion-focus on the VAMP7-dependent secretion. Front Cell Dev Biol. 2022;10:884020.
Vietri M, Radulovic M, Stenmark H. The many functions of ESCRTs. Nat Rev Mol Cell Biol. 2020;21(1):25–42.
Guo L, Zhang Y, Wei R, Zhang X, Wang C, Feng M. viaProinflammatory macrophage-derived microvesicles exhibit tumor tropism dependent on CCL2/CCR2 signaling axis and promote drug delivery SNARE-mediated membrane fusion. Theranostics. 2020;10(15):6581–98.
An M, Lohse I, Tan Z, Zhu J, Wu J, Kurapati H, et al. Quantitative proteomic analysis of serum exosomes from patients with locally advanced pancreatic cancer undergoing chemoradiotherapy. J Proteome Res. 2017;16(4):1763–72.
De Lellis L, Florio R, Di Bella MC, Brocco D, Guidotti F, Tinari N, et al. Exosomes as pleiotropic players in pancreatic cancer. Biomedicines. 2021;9(3):275.
Zhao H, Yang L, Baddour J, Achreja A, Bernard V, Moss T, et al. Tumor microenvironment derived exosomes pleiotropically modulate cancer cell metabolism. Elife. 2016;5:e10250.
Zebrowska A, Jelonek K, Mondal S, Gawin M, Mrowiec K, Widłak P, et al. Proteomic and metabolomic profiles of T cell-derived exosomes isolated from human plasma. Cells. 2022;11(12):1965.
Palacios-Ferrer J, García-Ortega M, Gallardo-Gómez M, García M, Díaz C, Boulaiz H, et al. Metabolomic profile of cancer stem cell-derived exosomes from patients with malignant melanoma. Mol Oncol. 2021;15(2):407–28.
Pavlova N, Zhu J, Thompson C. The hallmarks of cancer metabolism: Still emerging. Cell Metab. 2022;34(3):355–77.
Stine Z, Schug Z, Salvino J, Dang C. Targeting cancer metabolism in the era of precision oncology. Nat Rev Drug Discovery. 2022;21(2):141–62.
Brunner J, Finley L. SnapShot: Cancer metabolism. Mol cell. 2021;81(18):3878-e1.
Noe J, Rendon B, Geller A, Conroy L, Morrissey S, Young L, et al. Lactate supports a metabolic-epigenetic link in macrophage polarization. Sci Adv. 2021;7(46):eabi8602.
Du D, Liu C, Qin M, Zhang X, ** T, Yuan S, et al. Metabolic dysregulation and emerging therapeutical targets for hepatocellular carcinoma. Acta pharmaceutica Sinica B. 2022;12(2):558–80.
Xu I, Lai R, Lin S, Tse A, Chiu D, Koh H, et al. Transketolase counteracts oxidative stress to drive cancer development. Proc Natl Acad Sci USA. 2016;113(6):E725–34.
Tseng H, Zeng Y, Lin Y, Huang J, Lin C, Lee M, et al. A novel AMPK activator shows therapeutic potential in hepatocellular carcinoma by suppressing HIF1α-mediated aerobic glycolysis. Mol Oncol. 2022;16(11):2274–94.
Huang Y, Chen Z, Lu T, Bi G, Li M, Liang J, et al. HIF-1α switches the functionality of TGF-β signaling via changing the partners of smads to drive glucose metabolic reprogramming in non-small cell lung cancer. J Exp Clin Cancer Res: CR. 2021;40(1):398.
Li Y, Li Y, Liu X, Zhang L, Chen Y, Zhao Q, et al. Blockage of citrate export prevents TCA cycle fragmentation via Irg1 inactivation. Cell Rep. 2022;38(7):110391.
Qi X, Zhang Y, Ji H, Wu X, Wang F, **e M, et al. Knockdown of prohibitin expression promotes glucose metabolism in eutopic endometrial stromal cells from women with endometriosis. Reprod Biomed Online. 2014;29(6):761–70.
Mittal A, Nenwani M, Sarangi I, Achreja A, Lawrence T, Nagrath D. Radiotherapy-induced metabolic hallmarks in the tumor microenvironment. Trends Cancer. 2022;8:855.
Liu L, Yuan L, Huang D, Han Q, Cai J, Wang S, et al. miR126 regulates the progression of epithelial ovarian cancer in vitro and in vivo by targeting VEGFA. Int J Oncol. 2020;57(3):825–34.
Castillo-Sanchez R, Churruca-Schuind A, Martinez-Ival M, Salazar EP. Cancer-associated fibroblasts communicate with breast tumor cells through extracellular vesicles in tumor development. Technol Cancer Res Treat. 2022;21:15330338221131648.
Kumagai S, Koyama S, Itahashi K, Tanegashima T, Lin Y, Togashi Y, et al. Lactic acid promotes PD-1 expression in regulatory T cells in highly glycolytic tumor microenvironments. Cancer Cell. 2022;40(2):201-18.e9.
Zappasodi R, Serganova I, Cohen I, Maeda M, Shindo M, Senbabaoglu Y, et al. CTLA-4 blockade drives loss of T stability in glycolysis-low tumours. Nature. 2021;591(7851):652–8.
Koundouros N, Poulogiannis G. Reprogramming of fatty acid metabolism in cancer. Br J Cancer. 2020;122(1):4–22.
Chassen S, Ferchaud-Roucher V, Gupta M, Jansson T, Powell T. Alterations in placental long chain polyunsaturated fatty acid metabolism in human intrauterine growth restriction. Clin Sci (London, England : 1979). 2018;132(5):595–607.
Zhao H, Li Y. Cancer metabolism and intervention therapy. Mol Biomed. 2021;2(1):5.
de Araujo JR, Eich C, Jorquera C, Schomann T, Baldazzi F, Chan A, et al. Ceramide and palmitic acid inhibit macrophage-mediated epithelial-mesenchymal transition in colorectal cancer. Mol Cell Biochem. 2020;468:153–68.
Das U. Essential fatty acids and their metabolites in the pathobiology of inflammation and its resolution. Biomolecules. 2021;11(12):1873.
Wang B, Wu L, Chen J, Dong L, Chen C, Wen Z, et al. Metabolism pathways of arachidonic acids: mechanisms and potential therapeutic targets. Signal Transduct Target Ther. 2021;6(1):94.
Kennedy B, Harris R. Cyclooxygenase and lipoxygenase gene expression in the inflammogenesis of breast cancer. Inflammopharmacology. 2018;26:909.
Vecchi L, Araújo T, Azevedo F, Mota S, Ávila V, Ribeiro M, et al. Phospholipase a drives tumorigenesis and cancer aggressiveness through its interaction with annexin A1. Cells. 2021;10(6):1472.
Osman W, Youssef N. Combined use of COX-1 and VEGF immunohistochemistry refines the histopathologic prognosis of renal cell carcinoma. Int J Clin Exp Pathol. 2015;8(7):8165–77.
O’Sullivan D, van der Windt G, Huang S, Curtis J, Chang C, Buck M, et al. Memory CD8(+) T cells use cell-intrinsic lipolysis to support the metabolic programming necessary for development. Immunity. 2014;41(1):75–88.
Liu QP, Lin JY, An P, Chen YY, Luan X, Zhang H. LncRNAs in tumor microenvironment: The potential target for cancer treatment with natural compounds and chemical drugs. Biochem Pharmacol. 2021;193:114802.
Machado M, Patente T, Rouillé Y, Heumel S, Melo E, Deruyter L, et al. Streptococcus pneumoniaeAcetate improves the killing of by alveolar macrophages NLRP3 inflammasome and glycolysis-HIF-1α Axis. Front Immunol. 2022;13:773261.
Pranzini E, Pardella E, Paoli P, Fendt S, Taddei M. Metabolic reprogramming in anticancer drug resistance: a focus on amino acids. Trends Cancer. 2021;7(8):682–99.
Reinfeld B, Madden M, Wolf M, Chytil A, Bader J, Patterson A, et al. Cell-programmed nutrient partitioning in the tumour microenvironment. Nature. 2021;593(7858):282–8.
Durán R, Oppliger W, Robitaille A, Heiserich L, Skendaj R, Gottlieb E, et al. Glutaminolysis activates Rag-mTORC1 signaling. Mol Cell. 2012;47(3):349–58.
Mattaini K, Sullivan M, Vander HM. The importance of serine metabolism in cancer. J Cell Biol. 2016;214(3):249–57.
**e M, Pei D. Serine hydroxymethyltransferase 2: a novel target for human cancer therapy. Invest New Drugs. 2021;39(6):1671–81.
Peyraud F, Guegan J, Bodet D, Cousin S, Bessede A, Italiano A. Targeting tryptophan catabolism in cancer immunotherapy era: challenges and perspectives. Front Immunol. 2022;13:807271.
Chen Y, Chen J, Guo D, Yang P, Chen S, Zhao C, et al. Tryptophan metabolites as biomarkers for esophageal cancer susceptibility, metastasis, and prognosis. Front Oncol. 2022;12:800291.
Wu D, Zhu Y. Role of kynurenine in promoting the generation of exhausted CD8 T cells in colorectal cancer. Am J Transl Res. 2021;13(3):1535–47.
Luo P, Yin P, Hua R, Tan Y, Li Z, Qiu G, et al. A Large-scale, multicenter serum metabolite biomarker identification study for the early detection of hepatocellular carcinoma. Hepatology (Baltimore, MD). 2018;67(2):662–75.
Udumula M, Sakr S, Dar S, Alvero A, Ali-Fehmi R, Abdulfatah E, et al. Ovarian cancer modulates the immunosuppressive function of CD11bGr1 myeloid cells via glutamine metabolism. Mol Metab. 2021;53:101272.
Oh M, Sun I, Zhao L, Leone R, Sun I, Xu W, et al. Targeting glutamine metabolism enhances tumor-specific immunity by modulating suppressive myeloid cells. J Clin Investig. 2020;130(7):3865–84.
Oh MH, Sun IH, Zhao L, Leone RD, Sun IM, Xu W, et al. Targeting glutamine metabolism enhances tumor-specific immunity by modulating suppressive myeloid cells. J Clin Invest. 2020;130(7):3865–84.
Chen L, Zhu S, Liu T, Zhao H, Chen P, Duan Y, et al. Cancer associated fibroblasts promote renal cancer progression through a TDO/Kyn/AhR dependent signaling pathway. Front Oncol. 2021;11:628821.
Yan W, Wu X, Zhou W, Fong M, Cao M, Liu J, et al. Cancer-cell-secreted exosomal miR-105 promotes tumour growth through the MYC-dependent metabolic reprogramming of stromal cells. Nat Cell Biol. 2018;20(5):597–609.
Sung J, Kang C, Kang S, Jang Y, Chae Y, Kim B, et al. ITGB4-mediated metabolic reprogramming of cancer-associated fibroblasts. Oncogene. 2020;39(3):664–76.
Rai A, Greening D, Chen M, Xu R, Ji H, Simpson R. Exosomes derived from human primary and metastatic colorectal cancer cells contribute to functional heterogeneity of activated fibroblasts by reprogramming their proteome. Proteomics. 2019;19(8):e1800148.
Zhang C, Wang XY, Zhang P, He TC, Han JH, Zhang R, et al. Cancer-derived exosomal HSPC111 promotes colorectal cancer liver metastasis by reprogramming lipid metabolism in cancer-associated fibroblasts. Cell Death Dis. 2022;13(1):57.
Ma X, Wang J, Li J, Ma C, Chen S, Lei W, et al. Loading MiR-210 in endothelial progenitor cells derived exosomes boosts their beneficial effects on hypoxia/Reoxygeneation-injured human endothelial cells via protecting mitochondrial function. Cell Physiol Biochem: Int J Exp Cell Physiol, Biochem, Pharmacol. 2018;46(2):664–75.
Wang B, Wang X, Hou D, Huang Q, Zhan W, Chen C, et al. Exosomes derived from acute myeloid leukemia cells promote chemoresistance by enhancing glycolysis-mediated vascular remodeling. J Cell Physiol. 2019;234(7):10602–14.
Ludwig N, Yerneni S, Azambuja J, Gillespie D, Menshikova E, Jackson E, et al. Tumor-derived exosomes promote angiogenesis via adenosine a receptor signaling. Angiogenesis. 2020;23(4):599–610.
Liu F, Bu Z, Zhao F, **ao D. Increased T-helper 17 cell differentiation mediated by exosome-mediated microRNA-451 redistribution in gastric cancer infiltrated T cells. Cancer Sci. 2018;109(1):65–73.
Zhou C, Zhang Y, Yan R, Huang L, Mellor A, Yang Y, et al. Exosome-derived miR-142-5p remodels lymphatic vessels and induces IDO to promote immune privilege in the tumour microenvironment. Cell Death Differ. 2021;28(2):715–29.
Muller L, Simms P, Hong C, Nishimura M, Jackson E, Watkins S, et al. Human tumor-derived exosomes (TEX) regulate Treg functions via cell surface signaling rather than uptake mechanisms. Oncoimmunology. 2017;6(8):e1261243.
Salimu J, Webber J, Gurney M, Al-Taei S, Clayton A, Tabi Z. Dominant immunosuppression of dendritic cell function by prostate-cancer-derived exosomes. J Extracell Vesicles. 2017;6(1):1368823.
Park J, Dutta B, Tse S, Gupta N, Tan C, Low J, et al. Hypoxia-induced tumor exosomes promote M2-like macrophage polarization of infiltrating myeloid cells and microRNA-mediated metabolic shift. Oncogene. 2019;38(26):5158–73.
Zhou S, Lan Y, Li Y, Li Z, Pu J, Wei L. Hypoxic tumor-derived exosomes induce M2 macrophage polarization via PKM2/AMPK to promote lung cancer progression. Cell Transplant. 2022;31:9636897221106998.
Wang S, Xu M, **ao X, Wang L, Sun Z, Guan M, et al. Pancreatic cancer cell exosomes induce lipidomics changes in adipocytes. Adipocyte. 2022;11(1):346–55.
Wu Q, Sun S, Li Z, Yang Q, Li B, Zhu S, et al. Tumour-originated exosomal miR-155 triggers cancer-associated cachexia to promote tumour progression. Mol Cancer. 2018;17(1):155.
Wu Q, Li J, Li Z, Sun S, Zhu S, Wang L, et al. Exosomes from the tumour-adipocyte interplay stimulate beige/brown differentiation and reprogram metabolism in stromal adipocytes to promote tumour progression. J Exp Clin Cancer Res. 2019;38(1):223.
Wu Q, Li J, Li Z, Sun S, Zhu S, Wang L, et al. Exosomes from the tumour-adipocyte interplay stimulate beige/brown differentiation and reprogram metabolism in stromal adipocytes to promote tumour progression. J Exp Clin Cancer Res : CR. 2019;38(1):223.
Sagar G, Sah R, Javeed N, Dutta S, Smyrk T, Lau J, et al. Pathogenesis of pancreatic cancer exosome-induced lipolysis in adipose tissue. Gut. 2016;65(7):1165–74.
Li J, Wu X, Schiffmann L, MacVicar T, Zhou C, Wang Z, et al. IL-17B/RB activation in pancreatic stellate cells promotes pancreatic cancer metabolism and growth. Cancers. 2021;13(21):5338.
Shu S, Yang Y, Allen C, Maguire O, Minderman H, Sen A, et al. Publisher Correction: Metabolic reprogramming of stromal fibroblasts by melanoma exosome microRNA favours a pre-metastatic microenvironment. Sci Rep. 2019;9(1):4959.
Zhang H, Deng T, Liu R, Ning T, Yang H, Liu D, et al. CAF secreted miR-522 suppresses ferroptosis and promotes acquired chemo-resistance in gastric cancer. Mol Cancer. 2020;19(1):43.
Li Y, Zhao Z, Liu W, Li X. SNHG3 functions as miRNA sponge to promote breast cancer cells growth through the metabolic reprogramming. Appl Biochem Biotechnol. 2020;191(3):1084–99.
Lu L, Huang J, Mo J, Da X, Li Q, Fan M, et al. Exosomal lncRNA TUG1 from cancer-associated fibroblasts promotes liver cancer cell migration, invasion, and glycolysis by regulating the miR-524-5p/SIX1 axis. Cell Mol Biol Lett. 2022;27(1):17.
Liu T, Han C, Fang P, Ma Z, Wang X, Chen H, et al. Cancer-associated fibroblast-specific lncRNA LINC01614 enhances glutamine uptake in lung adenocarcinoma. J Hematol Oncol. 2022;15(1):141.
Sansone P, Savini C, Kurelac I, Chang Q, Amato L, Strillacci A, et al. Packaging and transfer of mitochondrial DNA via exosomes regulate escape from dormancy in hormonal therapy-resistant breast cancer. Proc Natl Acad Sci USA. 2017;114(43):E9066–75.
Huang S, Fan P, Zhang C, **e J, Gu X, Lei S, et al. Exosomal microRNA-503-3p derived from macrophages represses glycolysis and promotes mitochondrial oxidative phosphorylation in breast cancer cells by elevating DACT2. Cell Death Discov. 2021;7(1):119.
Wang P, Li GY, Zhou L, Jiang HL, Yang Y, Wu HT. Exosomes from M2 macrophages promoted glycolysis in FaDu cells by inhibiting PDLIM2 expression to stabilize PFKL. Neoplasma. 2022;69(5):1041–53.
Chen F, Chen J, Yang L, Liu J, Zhang X, Zhang Y, et al. Extracellular vesicle-packaged HIF-1α-stabilizing lncRNA from tumour-associated macrophages regulates aerobic glycolysis of breast cancer cells. Nat Cell Biol. 2019;21(4):498–510.
Lazar I, Clement E, Dauvillier S, Milhas D, Ducoux-Petit M, LeGonidec S, et al. Adipocyte exosomes promote melanoma aggressiveness through fatty acid oxidation: a novel mechanism linking obesity and cancer. Can Res. 2016;76(14):4051–7.
Liu Y, Tan J, Ou S, Chen J, Chen L. Adipose-derived exosomes deliver miR-23a/b to regulate tumor growth in hepatocellular cancer by targeting the VHL/HIF axis. J Physiol Biochem. 2019;75(3):391–401.
Yin H, Qiu X, Shan Y, You B, **e L, Zhang P, et al. HIF-1α downregulation of miR-433-3p in adipocyte-derived exosomes contributes to NPC progression via targeting SCD1. Cancer Sci. 2021;112(4):1457–70.
Alamoudi A, Alnoury A, Gad H. miRNA in tumour metabolism and why could it be the preferred pathway for energy reprograming. Brief Funct Genomics. 2018;17(3):157–69.
Sung JY, Cheong JH. New immunometabolic strategy based on cell type-specific metabolic reprogramming in the tumor immune microenvironment. Cells. 2022;11(5):768.
Martínez-Reyes I, Chandel N. Cancer metabolism: looking forward. Nat Rev Cancer. 2021;21(10):669–80.
Huang W, Fan L, Tang Y, Chi Y, Li J. A pan-cancer analysis of the oncogenic role of integrin Beta4 (ITGB4) in human tumors. Int J Gen Med. 2021;14:9629–45.
Wang WT, ** WL, Li X. Intercellular communication in the tumour microecosystem: Mediators and therapeutic approaches for hepatocellular carcinoma. Biochim Biophys Acta Mol Basis Dis. 2022;1868(12):166528.
Bai S, Wei Y, Liu R, Xu R, **ang L, Du J. Role of tumour-derived exosomes in metastasis. Biomed Pharmacother. 2022;147:112657.
Ludwig N, Gillespie D, Reichert T, Jackson E, Whiteside T. Purine metabolites in tumor-derived exosomes may facilitate immune escape of head and neck squamous cell carcinoma. Cancers. 2020;12(6):1602.
Guo W, Qiao T, Dong B, Li T, Liu Q, Xu X. The effect of hypoxia-induced exosomes on anti-tumor immunity and its implication for immunotherapy. Front Immunol. 2022;13:915985.
Zhang Y, Mao Q, **a Q, Cheng J, Huang Z, Li Y, et al. Noncoding RNAs link metabolic reprogramming to immune microenvironment in cancers. J Hematol Oncol. 2021;14(1):169.
Jhunjhunwala S, Hammer C, Delamarre L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat Rev Cancer. 2021;21(5):298–312.
Nazari N, Jafari F, Ghalamfarsa G, Hadinia A, Atapour A, Ahmadi M, et al. The emerging role of microRNA in regulating the mTOR signaling pathway in immune and inflammatory responses. Immunol Cell Biol. 2021;99(8):814–32.
Sun L, Wang Y, Wang X, Navarro-Corcuera A, Ilyas S, Jalan-Sakrikar N, et al. PD-L1 promotes myofibroblastic activation of hepatic stellate cells by distinct mechanisms selective for TGF-β receptor I versus II. Cell Rep. 2022;38(6):110349.
Zhao L, Ma X, Yu J. Exosomes and organ-specific metastasis. Mol Ther Methods Clin Dev. 2021;22:133–47.
Chen Y, McAndrews K, Kalluri R. Clinical and therapeutic relevance of cancer-associated fibroblasts. Nat Rev Clin Oncol. 2021;18(12):792–804.
Miki Y, Yashiro M, Moyano-Galceran L, Sugimoto A, Ohira M, Lehti K. Crosstalk between cancer associated fibroblasts and cancer cells in scirrhous type gastric cancer. Front Oncol. 2020;10:568557.
Chen L, Huang L, Gu Y, Cang W, Sun P, **ang Y. Lactate-lactylation hands between metabolic reprogramming and immunosuppression. Int J Mol Sci. 2022;23(19):11943.
Zhang M, Di Martino JS, Bowman RL, Campbell NR, Baksh SC, Simon-Vermot T, et al. Adipocyte-derived lipids mediate melanoma progression via FATP proteins. Cancer Discov. 2018;8(8):1006–25.
Dogra S, Hannafon BN. Breast cancer microenvironment cross talk through extracellular vesicle RNAs. Am J Pathol. 2021;191(8):1330–41.
Li X, Shen Y, Zhang L, Guo X, Wu J. Understanding initiation and progression of hepatocellular carcinoma through single cell sequencing. Biochim Biophys Acta. 2022;1877(3):188720.
Han W, Wang S, Qi Y, Wu F, Tian N, Qiang B, et al. Targeting HOTAIRM1 ameliorates glioblastoma by disrupting mitochondrial oxidative phosphorylation and serine metabolism. iScience. 2022;25(8):104823.
Yuan Y, Li H, Pu W, Chen L, Guo D, Jiang H, et al. Cancer metabolism and tumor microenvironment: fostering each other? Sci China Life Sci. 2022;65(2):236–79.
Dai Y, Liu Y, Li J, ** M, Yang H, Huang G. Shikonin inhibited glycolysis and sensitized cisplatin treatment in non-small cell lung cancer cells via the exosomal pyruvate kinase M2 pathway. Bioengineered. 2022;13(5):13906–18.
Wang Y, Hao F, Nan Y, Qu L, Na W, Jia C, et al. PKM2 inhibitor Shikonin overcomes the cisplatin resistance in bladder cancer by inducing necroptosis. Int J Biol Sci. 2018;14(13):1883–91.
Wang X, Zhang H, Yang H, Bai M, Ning T, Deng T, et al. Exosome-delivered circRNA promotes glycolysis to induce chemoresistance through the miR-122-PKM2 axis in colorectal cancer. Mol Oncol. 2020;14(3):539–55.
Lv B, Zhu W, Feng C. circCCT3Coptisine blocks secretion of exosomal from cancer-associated fibroblasts to reprogram glucose metabolism in hepatocellular carcinoma. DNA and Cell Biol. 2020;39:2281.
Aslan C, Maralbashi S, Kahroba H, Asadi M, Soltani-Zangbar M, Javadian M, et al. Docosahexaenoic acid (DHA) inhibits pro-angiogenic effects of breast cancer cells via down-regulating cellular and exosomal expression of angiogenic genes and microRNAs. Life Sci. 2020;258:118094.
Zhan Q, Yi K, Cui X, Li X, Yang S, Wang Q, et al. Blood exosomes-based targeted delivery of cPLA2 siRNA and metformin to modulate glioblastoma energy metabolism for tailoring personalized therapy. Neuro-Oncology. 1871;2022:24.
Zhu D, Zhang T, Li Y, Huang C, Suo M, **a L, et al. Tumor-derived exosomes co-delivering aggregation-induced emission luminogens and proton pump inhibitors for tumor glutamine starvation therapy and enhanced type-I photodynamic therapy. Biomaterials. 2022;283:121462.
Chen M, Liu H, Li Z, Ming A, Chen H. Mechanism of PKM2 affecting cancer immunity and metabolism in tumor microenvironment. J Cancer. 2021;12(12):3566–74.
Wei Y, Wang D, ** F, Bian Z, Li L, Liang H, et al. Pyruvate kinase type M2 promotes tumour cell exosome release via phosphorylating synaptosome-associated protein 23. Nat Commun. 2017;8:14041.
Tan Y, Luo X, Lv W, Hu W, Zhao C, **ong M, et al. Tumor-derived exosomal components: the multifaceted roles and mechanisms in breast cancer metastasis. Cell Death Dis. 2021;12(6):547.
Yang Z, Wang D, Zhang C, Liu H, Hao M, Kan S, et al. The applications of gold nanoparticles in the diagnosis and treatment of gastrointestinal cancer. Front Oncol. 2021;11:819329.
Fan S, Kroeger B, Marie P, Bridges E, Mason J, McCormick K, et al. Glutamine deprivation alters the origin and function of cancer cell exosomes. EMBO J. 2020;39(16):e103009.
Zhong J, **a B, Shan S, Zheng A, Zhang S, Chen J, et al. High-quality milk exosomes as oral drug delivery system. Biomaterials. 2021;277:121126.
Acknowledgements
We wish to thank lab members for their valuable and enthusiastic work and scientific discussion.
Funding
This work was supported in part by grants from the following sources: the National Natural Science Foundation of China (82,203,233, 82,202,966, 82,173,142, 81,972,636, 81,872,281), the Natural Science Foundation of Hunan Province (2022JJ80078, 2020JJ5336), the Research Project of Health Commission of Hunan Province (202,203,034,978, 202,202,055,318, 202,109,031,837, 202,109,032,010, 20,201,020), Key Research and Development Program of Hunan Province (2022SK2051), Hunan Provincial Science and Technology Department (2020TP1018), the Changsha Science and Technology Board (kh2201054), Ascend Foundation of National cancer center (NCC201909B06), and by Hunan Cancer Hospital Climb Plan (ZX2020001-3, YF2020002).
Author information
Authors and Affiliations
Contributions
ST, YY and YW drafted the manuscript and prepared the figures. YH, LH, RY, ZH, YT, LL, YL, LO, JL, QP, XJ, XX, LX, MP, NW and YT helped in collecting the related literatures and participated in discussion. DC, QL and YZ designed the review and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Tan, S., Yang, Y., Yang, W. et al. Exosomal cargos-mediated metabolic reprogramming in tumor microenvironment. J Exp Clin Cancer Res 42, 59 (2023). https://doi.org/10.1186/s13046-023-02634-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13046-023-02634-z