Abstract
Dynamic miRNA detection using the qRT-PCR technique requires appropriate reference genes to ensure data reliability. Previous studies have screened internal reference genes in plants during embryonic development and various stress treatment, involving relatively few tissues and organs. There is no relevant miRNA study in Lilium henryi Baker and limited research on the optimal miRNA reference genes in lilies, such as 5S, 18S, U6 and Actin. Twelve genes were selected as candidate reference genes whose expression stability was analyzed in petals at different developmental stages and other tissues using various algorithms, such as geNorm, NormFinder, BestKeeper, and Delta CT. The results revealed that the optimal combination of reference genes for Lilium henryi Baker petals at different developmental stages was osa-miR166m and osa-miR166a-3p, while that for different tissues of Lilium henryi Baker was osa-miR166g-3p and osa-miR166a-3p.Four important genes related to growth and development regulation, namely, osa-miR156a, osa-miR395b, osa-miR396a-3p, and osa-miR396a-5p, were selected for validation. The findings of the present study could contribute to future investigations onmiRNA expression and the related functions in Lilium henryi Baker while providing important references for the normalization of the miRNA expression in other varieties of lily.
Similar content being viewed by others
Introduction
Lilies (Lilium spp.) are considered among the most important floral crops worldwide1,2. Lilies are recognized for their diverse flower colors and forms and are, therefore, widely used in garden beautification and cut flower markets3,4,5. The growth and development of lilies involve a series of biochemical processes in which miRNAs (microRNAs) play important roles, for instance, in cell proliferation, cell differentiation, growth, development, and response to external environmental stimuli6,7,8. Plant miRNAs were identified in Arabidopsis and Oryza sativa in 20029, and since then, hundreds of similar small RNA molecules, which are collectively referred to as miRNAs, have been identified in several model organisms and cells. The above studies have revealed the complexity of gene regulatory networks in plants, due to which miRNAs have become a research hotspot in recent years.
An unstable expression of a reference gene could lead to biased outcomes10. Among the various detection methods available, qRT-PCR is the preferred method for gene detection owing to its high sensitivity, strong specificity, and convenient operation11,12,13. Currently, qRT-PCR based on stem-loop and poly(A) methods, which use primers designed specifically for this purpose, is commonly used for miRNA detection. The stem-loop method14 offers greater specificity and sensitivity compared to the poly(A) method15,16,17. The selection of miRNA-related reference genes varies among different cultivars, and lilies encompass multiple varieties. For instance, in a study on Lilium pumilum DC. Fisch. For somatic embryos, the reference genes selected were lpu-miR159a and the F-box family protein (FP)18. In Lilium × formolongi, 5S RNA and EF were selected as reference genes19. In the OT (Oriental × Trumpet) hybrid ‘Robina’, U4 was reported as the reference gene20. Among the bicolor cultivars of Asiatic hybrid lily, namely, Lollypop, Cancún, Vermeer, Sugar Love, and Enjoys, as well asnthree full-color cultivars of the species, namely, Côte d’Azur, Vivaldi, and Montreux, U6 was identified as the reference gene21. In Lilium leichtlinii var. maximowiczii, 18S rRNA was reported as the reference gene5. Lilium henryi Baker is a species endemic to China and located primarily in regions such as Hubei and Guizhou. This species has characteristic orange flowers, which confer high ornamental value. The bulbs of Lilium henryi Baker are used for human consumption and also in traditional medicine22. The species exhibits strong adaptability23 and excellent overall resistance. It is the only wild species in the Lilium genus with a vining growth habit, owing to which it is considered an important breeding material in horticulture. The species is at the state of NT (Near Threatened) according the IUCN standard (http://www.iplant.cn/rep/prot/Lilium%20henryi). For the above reasons, a detailed investigation of Lilium henryi Baker is of paramount importance. In the same context, the selection of miRNA reference genes and the establishment of a qRT-PCR detection system for Lilium henryi Baker are imperative.
Our research group pioneered miRNA omics studies related to flower color traits in Lilium henryi Baker (yet to be published). A small RNA sequencing of Lilium henryi Baker was conducted and revealed numerous miRNA sequences. Among those sequences, ten miRNAs with relatively stable expression were screened out and used as candidate reference genes. In addition, two commonly used reference genes were also selected. The stability of the candidate reference genes was analyzed using different software tools, such as geNorm24, NormFinder25, and BestKeeper26. The objective of this study was to identify suitable miRNA reference genes for different growth and development stages and various tissues to provide appropriate internal reference choices for accurate quantification of miRNA gene expression levels during the growth and development of Lilium henryi Baker.
Materials and methods
Experimental materials
Lilium henryi Baker was used as the experimental material in the present study (Fig. 1), Fig. 1 generate using Illustrator 2021 software. The plants of this species were cultivated in a greenhouse at the Bei**g Academy of Agriculture and Forestry Sciences. Samples were collected at different stages of flower development (S1: green bud stage; S2: bud stage; S3: coloring stage; S4: early blooming stage; and S5: full blooming stage) and also from the various tissues (petals at the full blooming stage, stem-root, stem, and leaves). Three biological replicates were performed for each group of samples. After collection, the samples were rapidly frozen in liquid nitrogen and stored at − 80 °C until use.
Illustration of the five stages of petal growth and development (first five images on the left) in Lilium henryi Baker. S1: green bud stage, S2: bud stage, S3: coloring stage, S4: early blooming stage, S5: full blooming stage. Illustration of the stem-root, stem, and leaf tissues (three images on the right) of Lilium henryi Baker.
Experimental methods
Extraction and reverse transcription of the miRNAs
The required miRNAs (from the petals of Lilium henryi Baker at the five stages of development and also from the stem-root, stem, and leaf tissues of the species) were extracted using the miRcute miRNA Isolation Kit (Tiangen Biotech Co.). The integrity of the extracted miRNA was assessed using agarose gel electrophoresis, and the purity and concentration of the miRNA were determined using a NanoDrop 2000. After quality control analysis, the miRNAs were reverse transcribed using the miRNA 1st Strand cDNA Synthesis Kit (using the stem-loop method) from Nan**g Vazyme Biotech Co. Ltd., following the manufacturer’s instructions. First-strand cDNA synthesis was performed in a reaction volume of 20 µL, which contained 0.5 µg of the miRNA to be reverse transcribed in an RNase-free centrifuge tube. In this process, the genomic DNA was first removed from the miRNA sample by heating at 42 °C for 2 min. Afterward, the first-strand cDNA synthesis reaction was conducted using the following temperature program: 25 °C for 5 min, 50 °C for 15 min, and finally, 85 °C for 5 min.
Screening of the candidate reference genes and primer design
Using our research group’s published small RNA sequencing data for Lilium henryi Baker (PRJNA1014586), ten miRNAs with relatively stable expression were selected as candidate reference genes. In addition, commonly used reference genes for plant miRNA qPCR were also selected, along with four miRNAs for validation. The sequences of primers used for all of the above analyses are provided in Table 1. These sequences were synthesized at Bei**g Tsingke Biotech Co. Ltd.
qRT-PCR analysis of the candidate reference genes
A 50 µL reaction mixture, which contained 5 µL of the cDNA template, 2 µL each (10 µM) of forward and reverse primers, 25 µL of 2 × miRNA Universal SYBR qPCR Master Mix, and 16 µL of ddH2O, was prepared according to the instructions of the Vazyme miRNA Universal SYBR qPCR Master Mix Kit. Each reaction was performed in triplicate. The PCR was conducted using the following program: initial denaturation at 95 °C for 5 min, followed by 40 cycles of denaturation at 95 °C for 10 s, followed by annealing at 60 °C for 30 s, and a final extension step at 72 °C for 15 s. A melting curve analysis was conducted using the following temperature program: 95 °C for 15 s, 60 °C for 60 s, and 95 °C for 15 s. Each reaction system was run in triplicate.
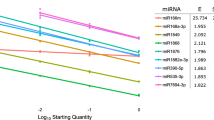
The amplification efficiency of the primers was determined by generating a standard curve using a fivefold dilution series of the cDNA template in water (v:v) –at 1:4, 1:24, 1:124, 1:624, and 1:3124. After each dilution, qRT-PCR was performed for each primer pair to obtain the Cq values and generate the standard curve. The amplification efficiency (E) and the R2 value were calculated, and the correction equation was determined to be: E = (5 −1/slope − 1) × 100%27,28.
Data processing and analysis
The data obtained was processed and analyzed using the geNorm, NormFinder, and BestKeeper software programs, the Delta CT method26. A lower mean SDobtained for an internal reference gene indicates a higher expression stability.
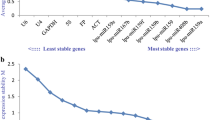
Delta CT analysis of petals at different developmental stages. The mean SD values of osa-miR166a-3p, osa-miR166m, and novel_1 were relatively low, indicating their good expression stability across all the samples. The most stable internal reference gene was osa-miR166a-3p, while U6 presented the largest mean SD value, indicating that U6 has the poorest stability (Fig. 8a). Among the different tissues of Lilium henryi Baker, 18S, novel_20, and osa-miR167d-5p presented relatively small mean SD values, indicating good expression stability across all the samples. U6 presented the largest mean SD, indicating that it had the poorest stability (Fig. 8b).
Expression stability analysis of candidate internal reference genes using the Delta CT method. (a) The results of the expression stability analysis of candidate internal reference genes in the petals at different developmental stages of the species using the Delta CT method. (b) The results of the expression stability analysis of candidate internal reference genes in the different tissues of the species using the Delta CT method.
RefFinder analysis
According to the ranks assigned to the genes based on each of the four programs, RefFinder assigns an appropriate weight to each gene and calculates the geometric mean of these weights. A resultant lower value indicates higher stability of the candidate internal reference gene under the given experimental conditions. Figure 9 shows the Venn diagram of the 5 most stable internal reference genes identified based on the geNorm, NormFinder, BestKeeper, and Delta CT analyses.
The five most stable reference genes selected via geNorm, NormFinder, BestKeeper, and Delta CT analyses. (a) Venn diagram of the expression stability of candidate reference genes in petals at different developmental stages. (b) Venn diagram of the expression stability of candidate reference genes in the different tissues of the species.
The results of the comprehensive analysis conducted using RefFinder revealed that in Lilium henryi Baker petals at different developmental stages (Fig. 10a). According to the results of the analysis based on the five evaluation methods, osa-miR166m, and osa-miR166a-3p exhibited relatively stable expression, with corresponding stability values of 1.19 and 1.41, respectively. U6 exhibited the least stable expression. According to these results of the geNorm analysis, the optimal number of internal reference genes to be introduced in the experiment was 2: osa-miR166m and osa-miR166a-3p. The different tissues of Lilium henryi Baker were used (Fig. 10b). According to the results of the analysis based on the five evaluation methods, osa-miR166g-3p and osa-miR166a-3p exhibited relatively stable expression, with corresponding stability values of 1.41 and 2.74, respectively. U6 exhibited the least stable expression. According to the results of geNorm analysis, the optimal number of internal reference genes to be introduced in the experiment was again 2: osa-miR166g-3p and osa-miR166a-3p. According to all the algorithms used in the present study, in Lilium henryi Baker, the miR166 family members, including osa-miR166a-3p, osa-miR166g-3p, and osa-miR166m, consistently ranked high in stability. Therefore, it was inferred that the miR166 family genes could serve as suitable candidates for use as internal reference genes in both petals at the different developmental stages of Lilium henryi Baker and in the different tissues of the species.
Results of RefFinder analysis of the expression stability of candidate reference genes. (a) The results of RefFinder analysis of the expression stability of candidate reference genes in petals at different developmental stages. (b) The results of the RefFinder analysis of the expression stability of candidate reference genes in the different tissues of the species.
Validation of genes
According to previous studies, miR156 plays an important role in regulating flowering time, lateral root development, and anthocyanidin synthesis33,32,35. miR396 is also widely involved in the regulation of growth and development in plant petals, leaves, and other parts36,37.Therefore, the expression levels of miR156 and miR396 were verified. Among the petals at the different developmental stages of the species, when the most stable reference genes were used, osa-miR166m and osa-miR166a-3p, the expression patterns of the target gene osa-miR156a were consistent (Fig. 11a). This gene exhibited the lowest expression level during the green bud stage, reached its maximum expression level during the coloring stage as the petals developed, then again exhibited a decreased expression level during the initial opening stage, and finally, a slight increase. However, for the least stable reference gene, U6, the expression level of osa-miR156a was quite low during the coloring, initial opening, and full bloom stages.
The expression profiles of the three validated genes in petals at different developmental stages and in the various tissues of the species. (a–c) The expression patterns of osa-miR156a, osa-miR396a-3p, and osa-miR396a-5p, in petals at different developmental stages. (d, e) The expression patterns of osa-miR395b, at different developmental stages and in the lower and upper parts of petals. (f–h) The expression patterns of osa-miR156a, osa-miR396a-3p, and osa-miR396a-5p in different tissues.
Similarly, the expression pattern of the target gene osa-miR396a-3p was consistent (Fig. 11b) in the different developmental stages of petals, with slight decreases during the bud stage, increases during the coloring stage, sharp decreases during the initial opening stage, and a maximum expression level during the full bloom stage. However, when U6, the least stable gene, was used as the reference gene, the expression pattern of osa-miR396a-3p varied and was opposite to that observed when osa-miR166m and osa-miR166a-3p were used as reference genes. The expression pattern of the target gene osa-miR396a-5p was consistent (Fig. 11c) among the different developmental stages of petals, with the maximum expression occuring during the full bloom stage. Therefore, osa-miR156a and osa-miR396a may have regulatory effects on the growth and development of petals, especially during the blooming stage.
Due to the gradual formation of spots in the lower part of petals during development and because the main component of the spots is anthocyanins, some studies suggest that osa-miR395b is a key gene related to anthocyanin synthesis38. Therefore, osa-miR166m and osa-miR166a-3p should be used as reference genes during the five stages of petal development to determine the expression of osa-miR395b in the lower and upper parts of petals o further verify the reliability of osa-miR166m and osa-miR166a-3p as internal reference genes.The expression trend of osa-miR395b was consistent when osa-miR166m and osa-miR166a-3p were used as internal reference genes. The expression level of osa-miR395b was extremely low in the lower part of the bud stage (Fig. 11d) and then significantly increased during the coloring stage. However, in the upper part of the petals (Fig. 11e), the expression level of osa-miR395b gradually increased during the green bud stage, bud stage, coloring stage, and early bloom stage, while the expression level was extremely low in the full blooming stage. This further proves the accuracy of osa-miR166m and osa-miR166a-3p as internal reference genes, whether for the entire petal or for different parts of the petal.
Among the different tissues, when the most stable reference genes, osa-miR166g-3p and osa-miR166a-3p, were used, the expression pattern of the target gene osa-miR156a remained consistent (Fig. 11f), with the lowest expression level in the petals observed during the full bloom stage and the highest expression level observed in the stem. Therefore, osa-miR156a may be involved in the regulation of stem growth.However, when the least stable reference gene U6 was used, the expression pattern was different, with the highest expression level observed in the stem and the lowest expression level observed in the petals at the full blooming stage. Similarly, the expression patterns of the target gene osa-miR396a-3p (Fig. 11g) in the petals and the stem were consistent, with relatively higher expression levels observed, while lower expression levels of this gene were observed in the stem and leaves. However, when the least stable reference gene U6 was used, the expression level of osa-miR396a-3p was extremely high in the stem. The expression pattern of the target gene osa-miR396a-5p (Fig. 11h) also remained consistent, with the highest expression level observed in the stem, while relatively lower expression levels were observed in the stem, leaves, and petals at the full bloom stage. Therefore, osa-miR396a-3p and osa-miR396a-5p may be involved in the growth and development of stem rooting. However, when the least stable reference gene U6 was used, significant differences in the expression of osa-miR396a-5p were observed among the different tissues, with the expression level in the stem approaching 0.
Discussion
The application of qRT-PCR for the quantitative analysis of the expression profiles of key genes is a fundamental and widely employed strategy for deciphering the mechanisms of plant growth and development. The selection of reference genes, particularly the internal control genes, has a crucial influence on the accuracy of gene expression profiles39,40. The screening and validation of miRNA reference genes have been reported for several plants, including Brassica napus41, grapevine (Vitis vinifera L. ‘Muscat Hamburg’)42, and rice (Oryza sativa)43. In poplar, U6 was identified as the most suitable reference gene for miRNA qRT-PCR experiments44. In sugarcane buds, miR171 and 18S rRNA were revealed as the most suitable reference genes for individual use16. In Chinese cedar (Cryptomeria fortunei), the most stable reference genes for miRNA expression were identified as novel16, cln-miR6725, novel1, and U645.
The expression levels of reference genes vary significantly among different plant varieties, tissues, organs, or physiological states. For instance, in a study by Yu et al.46, the reference gene used in the salt stress response experiment in the soybean variety ‘Williams 82’ was gma-miR1520d. However, Liu et al.47 suggested that in Soybean seeds (‘Williams 82’) under different abiotic stress conditions, the most suitable combination of reference miRNAs was miR166a and miR167d in the leaf samples and miR171a and miR156a or miR167a and miR171a in the root samples.
Related research has also been conducted in the field of lilies. Zhang et al.48 used 5S as an internal reference gene in Lilium × formolongi. Gao et al.49 used 18S as an internal reference gene in Lilium regale. Moreover, U4 and U6 have also been used as internal reference genes in lilies50,51. Therefore, it was considered necessary to conduct a study on the screening of miRNA reference genes in Lilium henryi Baker. In this context, the present study was conducted to determine the suitable reference genes for Lilium henryi Baker. A total of 12 candidate genes were selected, and their performance was evaluated in different samples (flower petals at different development stages, stems, stem-roots, and leaves) of Lilium henryi Baker. Four commonly used algorithms (geNorm, NormFinder, BestKeeper, and Delta CT) were applied to analyze the expression stability of these 12 candidate reference genes in the petals at different developmental stages and in various tissues of the species. The objective was to evaluate and determine stable reference genes. While the top five genes selected based on the results of the different algorithms were generally similar, slight differences were observed in their stability levels, which was attributable to the different calculation methods employed by each software package. However, despite these differences, the results obtained using these different algorithms were significantly consistent in terms of the selected best reference genes. According the NormFinder analysis, the stability values of osa miR166m and osa miR66a-3p were both 0.14, indicating that osa-miR166m and osa-miR166a-3p exhibited the highest stability (Fig. 6a). To further analyze the results obtained using the four algorithms, an online analysis tool referred to as RefFinder was used. The results obtained using the geNorm, NormFinder, BestKeeper, and Delta CT methods were consistent with the results obtained using RefFinder, indicating the accuracy of the software analysis. In the case of different tissues, the results obtained using geNorm and NormFinder were generally consistent, while those obtained using BestKeeper and Delta CT presented certain differences. Similar findings were reported in previous studies and therefore, these differences are acceptable from a comprehensive perspective52,51,54.
Further analysis using RefFinder revealed results consistent with those of geNorm and NormFinder analyses. The best combination of reference genes was determined based on the comprehensive analysis conducted using RefFinder. The results indicated that suitable reference genes should be selected for different experimental samples, as not all commonly used reference genes are applicable to different tissue samples or experimental conditions. This finding was similar to that reported for Euscaphis konishii Hayata10. To further validate the accuracy of the selected reference genes, osa-miR156a, osa-miR396a-3p, and osa-miR396a-5p were selected as validation genes, owing to their important regulatory roles in plant growth and development. Moreover, during the development of lily petals, spots gradually form in the lower tissue. To investigate whether the selected internal reference genes involved in petal spot formation are suitable for miRNA transcription level analysis, the expression level of the osa-miR395b gene, which is related to anthocyanin synthesis was detected. The results confirmed the accuracy of the screened reference genes, indicating that the optimal internal reference genes osa-miR166m and osa-miR166a-3p can be used for analyzing miRNA transcription levels in different tissues of petals at different developmental stages.
Previous studies have demonstrated that U6 is highly conserved across different plant species, implying that its sequence is relatively stable among different plants. Consequently, the small nuclear RNA of U6 is widely used as an internal reference gene for miRNA studies on various plants, including tea plants55, rice (Oryza sativa)56, and tomato57. However, while U6 is commonly used as a reference gene in several plant species, differences in the plant genomic structures and regulatory mechanisms may lead to variations in the expression of U6 under certain conditions. For instance, in Lilium species (Lilium pumilum DC. Fisch. and Lilium davidii var. unicolor)51, U6 reportedly exhibited relatively lower stability ranks, while in sweet potato (Ipomoea batatas. L.)58, U6 expression was relatively unstable. The present study revealed that the commonly used reference gene U6 exhibited unstable expression during the growth and development of Lilium henryi Baker. On the other hand, certain miRNAs from the miR166 family exhibited relatively stable expression and were more suitable as reference genes. When miR166 is used as a reference gene, it typically includes members of its subfamily, such as miR166a, miR166b, miR166c, and miR166m. The miR166 family is a group of similar miRNAs that share the same miRNA precursor sequence or have highly conserved mature miRNA sequences. These subfamily members frequently exhibit similar expression patterns and functions. Therefore, when selecting reference genes, the entire miR166 family or its multiple subfamilies may be considered for detection. This approach allows for a further comprehensive and accurate assessment of the expression levels of the miR166 family of genes used as reference genes. Similarly, in Tamarillo (Solanum betaceum) callus tissue samples, miR166a was reportedly the most stable miRNA59.
Previous studies have screened plant internal reference genes such as EF48, 18S49, U650 and Actin 51 during embryonic development and various stress treatment, involving relatively few tissues and organs in lily. Therefore, this study screened and validated internal reference genes involved in various flower development stages and tissue parts of Lilium henryi Baker. A more precise validation was conducted on the selected internal reference genes in the petals, avoiding interference from specific parts of the petals on the results of selecting the best internal reference genes. Therefore,the present study provides important information for studying the role of miRNAs in the growth and development of lilies and offers a strategy and method for screening miRNA reference genes in other plants. However, further research is warranted to validate and optimize the results obtained in the present study. Future studies could consider expanding the sample size and including samples from different developmental stages to further validate the stability and applicability of the miR166 family. Further investigations based on reference gene screening could reveal the differential expression and functions of miRNAs during the growth and development of lilies, providing a solid foundation and theoretical support for unraveling the mechanisms underlying the formation of important traits and the process of molecular breeding in lilies.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Alappat, B. & Alappat, J. Anthocyanin pigments: Beyond aesthetics. Molecules. 25, 5500 (2020).
Yamagishi, M. How genes paint lily flowers: Regulation of colouration and pigmentation patterning. Sci. Hortic-Amsterdam. 163, 27–36 (2013).
Al-Ajlouni, M. G., Othman, Y. A., A’Saf, T. S. & Ayad, J. Y. Lilium morphology, physiology, anatomy and postharvest flower quality in response to plant growth regulators. S. Afr. J. Bot. 156, 43–53 (2023).
Bakhshaie, M., Khosravi, S., Azadi, P., Bagheri, H. & van Tuyl, J. M. Biotechnological advances in lilium. Plant. Cell. Rep. 35, 1799–1826 (2016).
Wang, Z., Li, X., Chen, M. M., Yang, L. Y. & Zhang, Y. C. Molecular and metabolic insights into anthocyanin biosynthesis for spot formation on lilium leichtlinii var. maximowiczii flower petals. Int. J. Mol. Sci. 24, 1844 (2023).
Bajczyk, M. et al. Recent insights into plant mirna biogenesis: multiple layers of mirna level regulation. Plants-Basel. 12, 342 (2023).
Wang, J. W., Czech, B. & Weigel, D. Mir156-regulated spl transcription factors define an endogenous flowering pathway in arabidopsis thaliana. Cell. 138, 738–749 (2009).
Willmann, M. R. & Poethig, R. S. Conservation and evolution of mirna regulatory programs in plant development. Curr. Opin. Plant. Biol. 10, 503–511 (2007).
Reinhart, B. J., Weinstein, E. G., Rhoades, M. W., Bartel, B. & Bartel, D. P. Micrornas in plants. Gene. Dev. 16, 1616–1626 (2002).
Liang, W. X. et al. Selection and evaluation of reference genes for qrt-pcr analysis in euscaphis konishii hayata based on transcriptome data. Plant Methods. 14, 1–9 (2018).
Gachon, C., Mingam, A. & Charrier, B. Real-time pcr: what relevance to plant studies?. J. Exp. Bot. 55, 1445–1454 (2004).
Kou, S. et al. Selection and validation of suitable reference genes for mirna expression normalization by quantitative Rt-Pcr in citrus somatic embryogenic and adult tissues. Plant Cell. Rep. 31, 2151–2163 (2012).
Kozera, B. & Rapacz, M. Reference genes in real-time pcr. J. Appl. Genet. 54, 391–406 (2013).
Chen, C. F. et al. Real-time quantification of micrornas by stem-loop Rt-Pcr. Nucl. Acids Res. 33, e179 (2005).
Shi, R. & Chiang, V. L. Facile means for quantifying microrna expression by real-time Pcr. Biotechniques. 39, 519–525 (2005).
Yang, Y. et al. Selection of reference genes for normalization of microrna expression by rt-qpcr in sugarcane buds under cold stress. Front. Plant. Sci. 7, 86 (2016).
Tashiro, R. M., Philips, J. G. & Winefield, C. S. Identification of suitable grapevine reference genes for Qrt-Pcr derived from heterologous species. Mol Genet Genomics. 291, 483–492 (2016).
Zhang, J. et al. Small Rna and transcriptome sequencing reveal a potential mirna-mediated interaction network that functions during somatic embryogenesis in lilium pumilum Dc. Fisch.. Front Plant. Sci. 8, 566 (2017).
Zhang, Q., Zhao, Y., Gao, X. & Jia, G. Analysis of mirna-mediated regulation of flowering induction in lilium × formolongi. BMC Plant. Biol. 21, 1–16 (2021).
Chen, Y. et al. Key MicroRNAs and target genes involved in regulating maturation in lilium. Ornament. Plant. Res. 2, 1–14 (2022).
Yamagishi, M. & Sakai, M. The Microrna828/Myb12 module mediates bicolor pattern development in asiatic hybrid lily (lilium Spp) flowers. Front Plant. Sci. 11, 590791 (2020).
Yang, C., Shi, L. & Lei, X. Fan**gshan research editorial committee, edited by fan**g mountain medicinal plants editorial committee (Guizhou Science and Technology Press, 2016).
Xue, J. King of bulb fresh cut flowers - Lily (Zhongyuan Farmers’ Publishing House, 2006).
Hellemans, J., Mortier, G., De Paepe, A., Speleman, F. & Vandesompele, J. Qbase relative quantification framework and software for management and automated analysis of real-time quantitative pcr data. Genome Biol. 8, 1–14 (2007).
Andersen, C. L., Jensen, J. L. & Orntoft, T. F. Normalization of real-time quantitative reverse transcription-pcr data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250 (2004).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekee** genes, differentially regulated target genes and sample integrity: bestkeeper–excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515 (2004).
Dong, X. M., Zhang, W. & Zhang, S. B. Selection and validation of reference genes for quantitative real-time PCR analysis of development and tissue-dependent flower color formation in Cymbidium lowianum. Int. J. Mol. Sci. 23, 738 (2022).
Tong, Z. G., Gao, Z.H., Wang, F., Zhou, J. & Zhang, Z. Selection of reliable reference genes for gene expression studies in peach using real-time PCR. BMC Mol. Biol.. 10, 71 (2009).
**e, F., **ao, P., Chen, D., Xu, L. & Zhang, B. Mirdeepfinder: A mirna analysis tool for deep sequencing of plant small Rnas. Plant Mol. Biol. 80, 75–84 (2012).
Yao, L. et al. Selection of Q Rt-Pcr reference genes for amomum tsaoko seeds during dormancy release. China J. Chin. Mater. Med. 46, 3832–3837 (2021).
Paolacci, A. R., Tanzarella, O. A., Porceddu, E. & Ciaffi, M. Identification and validation of reference genes for quantitative Rt-Pcr normalization in wheat. BMC Mol. Biol. 10, 1–27 (2009).
Vandesompele, J. et al. Accurate normalization of real-time quantitative rt-pcr data by geometric averaging of multiple internal control genes. Genome Biol. 3, 34 (2002).
Asgari, M., Mirzaie-asl, A., Abdollahi, M. R. & Khodaei, L. Flowering time regulation by the Mirna156 in the beet (beta vulgaris Ssp. maritima). Plant Cell Tiss Org. 150, 361–370 (2022).
Yu, N., Niu, Q. W., Ng, K. H. & Chua, N. H. The role of Mir156/Spls modules in arabidopsis lateral root development. Plant J. 83, 673–685 (2015).
Gou, J., Felippes, F. F., Liu, C., Weigel, D. & Wang, J. Negative regulation of anthocyanin biosynthesis in arabidopsis by a Mir156-targeted spl transcription factor. Plant. Cell. 23, 1512–1522 (2011).
Cao, D. Y. et al. regulations on growth and development in tomato cotyledon, flower and fruit via destruction of mir396 with short tandem target mimic. Plant. Sci. 247, 1–12 (2016).
Yang, F. X. et al. Organ-specific gene expression reveals the role of the cymbidium ensifolium-Mir396/growth-regulating factors module in flower development of the orchid plant cymbidium ensifolium. Front. Plant. Sci. 12, 7997780 (2022).
Li, H. S. et al. Effects of blue light on flavonoid accumulation linked to the expression of Mir393, Mir394 and Mir395 in longan embryogenic calli. PLOS One. 13, 191444 (2018).
Derveaux, S., Vandesompele, J. & Hellemans, J. How to do successful gene expression analysis using real-time pcr. Methods. 50, 227–230 (2010).
Heid, C. A., Stevens, J., Livak, K. J. & Williams, P. M. Real time quantitative Pcr. Genome. Res. 6, 986–994 (1996).
Machado, R. D. et al. Comprehensive selection of reference genes for quantitative gene expression analysis during seed development in Brassica napus. Plant Cell Rep. 34, 1139–1149 (2015).
Luo, M. et al. Selection of reference genes for Mirna Qrt-Pcr under abiotic stress in grapevine. Sci Rep-UK 8, 4444 (2018).
Verstraeten, B., De Smet, L., Kyndt, T. & De Meyer, T. Selection of mirna reference genes for plant defence studies in rice (Oryza Sativa). Planta. 250, 2101–2110 (2019).
Tang, F. et al. Selection and validation of reference genes for quantitative expression analysis of mirnas and mrnas in poplar. Plant Methods. 15, 1–15 (2019).
Zhang, Y. T. et al. Selection and optimization of reference genes for microrna expression normalization by Qrt-Pcr in Chinese Cedar (Cryptomeria fortunei) under multiple stresses. Int. J. Mol. Sci. 22, 7246 (2021).
Yu, Y. H., Tang, J., Liu, C. & Ni, Z. Y. Microrna4359B positively regulates the soybean response to salt stress by targeting the F-Box protein Gmfbx193. Environ. Exp. Bot. 206, 105177 (2023).
Liu, W. C. et al. Normalization for relative quantification of mrna and microrna in soybean exposed to various abiotic stresses. PLOS One. 11, 155606 (2016).
Zhang, Q., Zhao, Y., Gao, X. & Jia, G. Analysis of mirna-mediated regulation of flowering induction in lilium × formolongi. BMC Plant. Biol. 21, 190 (2021).
Gao, X. et al. The Lre-Mir159a-Lrgamyb pathway mediates resistance to grey mould infection in lilium regale. Mol. Plant. Pathol. 21, 749–760 (2020).
Yamagishi, M. & Sakai, M. The Microrna828/Myb12 module mediates bicolor pattern development in asiatic hybrid lily (Lilium Spp.) flowers. Front. Plant. Sci. 11, 590791 (2020).
Zhang, J. et al. The use of mirnas as reference genes for mirna expression normalization during lilium somatic embryogenesis by real-time reverse transcription pcr analysis. Plant. Cell. Tiss. Org. 129, 105–118 (2017).
Barbosa Amorim, L. L. et al. Cowpea and abiotic stresses identification of reference genes for transcriptional profiling by Qpcr. Plant. Methods. 14, 1–17 (2018).
Hong, S., Seo, P. J., Yang, M., **ang, F. & Park, C. Exploring valid reference genes for gene expression studies in brachypodium distachyon by real-time Pcr. BMC Plant. Biol. 8, 1–1 (2008).
Wan, H. et al. Selection of appropriate reference genes for gene expression studies by quantitative real-time polymerase chain reaction in cucumber. Anal Biochem. 399, 257–261 (2010).
Song, H. et al. Selection and verification of candidate reference genes for mature microrna expression by quantitative Rt-Pcr in the tea plant (Camellia Sinensis). Genes-Basel. 7, 25 (2016).
Ding, Y., Chen, Z. & Zhu, C. Microarray-based analysis of cadmium-responsive micrornas in rice (Oryza Sativa). J. Exp. Bot. 62, 3563–3573 (2011).
Koul, A. et al. Carotenoid profiling, in silico analysis and transcript profiling of mirnas targeting carotenoid biosynthetic pathway genes in different developmental tissues of tomato. Plant. Physiol. Bioch. 108, 412–421 (2016).
Liu, X. Y. et al. Optimization of reference genes for qrt-pcr analysis of microrna expression under abiotic stress conditions in sweetpotato. Plant. Physiol. Bioch. 154, 379–386 (2020).
Cordeiro, D., Rito, M., Borges, F., Canhoto, J. & Correia, S. Selection and validation of reference genes for Qpcr analysis of mirnas and their targets during somatic embryogenesis in Tamarillo (Solanum Betaceumcav.). Plant. Cell. Tiss. Org. 143, 109–120 (2020).
Funding
This research was funded by the China National Natural Science Foundation Projects for Youth (Grant number 32201606), Key research and development projects of the National Forestry and Grassland Administration (Grant number GLM [2021] No. 72), Special projects for capacity building in scientific and technological innovation of the Bei**g Academy of Agriculture and Forestry (Grant number KJCX20230106),Bei**g Innovation Consortium of Agriculture Research Sysem (Grant number BAIC09-2023).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation was performed by G.J., S.Y. and Y.D. Data collection and analysis were performed by G.J. and M.Z. The first draft of the manuscript was written by G.J. and M.Z. The reviewing and editing were performed by X.Z, M.W. and C.Z. The funding acquisition was performed by M.Z. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
**, G., Zhang, X., Yu, S. et al. Screening and validation of optimal miRNA reference genes in different develo** stages and tissues of Lilium henryi Baker. Sci Rep 14, 1545 (2024). https://doi.org/10.1038/s41598-024-51562-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-51562-1
- Springer Nature Limited