Abstract
The relative contribution of intrinsic genetic factors and extrinsic environmental ones to cancer aetiology and natural history is a lengthy and debated issue. Gene–environment interactions (G x E) arise when the combined presence of both a germline genetic variant and a known environmental factor modulates the risk of disease more than either one alone. A panel of experts discussed our current understanding of cancer aetiology, known examples of G × E interactions in cancer, and the expanded concept of G × E interactions to include somatic cancer mutations and iatrogenic environmental factors such as anti-cancer treatment. Specific genetic polymorphisms and genetic mutations increase susceptibility to certain carcinogens and may be targeted in the near future for prevention and treatment of cancer patients with novel molecularly based therapies. There was general consensus that a better understanding of the complexity and numerosity of G × E interactions, supported by adequate technological, epidemiological, modelling and statistical resources, will further promote our understanding of cancer and lead to novel preventive and therapeutic approaches.
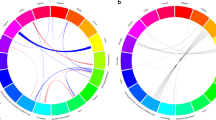
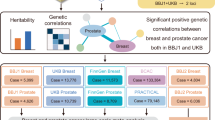
Similar content being viewed by others
Facts
-
Gene–environment interaction indicates that combination of a genetic and an environmental factor modulates the risk of cancer more than either one alone.
-
Lifetime risk of cancers of many different types was shown to strongly correlate with the total number of divisions of the normal self-renewing stem cells.
-
Specific genetic polymorphisms and mutations increase susceptibility to certain carcinogens.
-
p53 is a prototype genetic factor for cancer, it can be somatically or inheritably mutated, as well as subjected to polymorphism.
-
All carriers of inherited BAP1 mutations have developed one or more cancers during their lifetime. Exposure to asbestos and UV-light further increases their risk of develo** mesothelioma, melanoma and skin cancers: GXE interaction.
Questions
-
Why environmental carcinogens cause cancer only in a fraction of exposed individuals?
-
Alterations in TP53 and BAP1 genes both predispose to cancer, but with a different spectrum. What are the underlining molecular determinants of this specificity?
-
Will single cell analysis resolve the difficulties in diagnosis and study associated to tumour heterogeneity?
-
Can we design prophylactic or therapeutic anti-cancer approaches based on genetic of polymorphisms and mutations?
The origin of cancer: studies clash over ‘intrinsic’ versus ‘extrinsic’ risk
Hypotheses on the origin of cancer are integral part of the history of medicine. The first three evidences supporting the presence of extrinsic chemical risk factors date back to the XVIII century: John Hill’s record on tobacco snuff and nasal cancer; Percivall Pott’s association between exposure to soot and scrotal cancer; Samuel von Sömmerring’s observation of a relationship between lip carcinoma and clay-pipe smoking [1]. Recognition of the existence of intrinsic genetic risk factors is more recent, and originated from analyses of familial clustering of cancer cases and is experimentally supported by numerous efforts to generate murine models of hereditary cancers [2]. Epidemiological studies have demonstrated that environmental carcinogens cause cancer only in a fraction of exposed individuals; the biological reasons are often unknown.
Taking advantage of the extremely large amount of data generated in the last 20 years, bioinformatics represents a powerful tool to investigate both the interactions between genes and environment in determining cancer risk and the origin of cancer itself. In the last few years, a great interest has been raised on this topic by a series of articles. Tomasetti & Vogelstein’s in 2015 reported that lifetime risk of cancers of many different types is strongly correlated with the total number of divisions of the normal self-renewing stem cells maintaining tissues’ homoeostasis [3], thus supporting a prevalent ‘bad-luck’ intrinsic origin of cancer. Wu et al. instead proposed that the overwhelming majority of cancers develop following exogenous damage, with only ~ 10–30% of the risk attributable to intrinsic factors, thus supporting the ‘toxic insults’ theory [4]. These studies provide correlations, without entering into causal molecular mechanisms, but the message of these papers could not be in more strident contradiction, with the effect of igniting a public debate on this very hot topic.
Unique occurrences of cancer epidemics in remote regions of the world have provided definitive proof for the gene X environment hypothesis. In particular, studies of an epidemic of mesothelioma in Cappadocia, Turkey, where over 50% of the population exposed to carcinogenic erionite fibers dies of mesothelioma, the most devastating cancer epidemic ever recorded in medicine, revealed that susceptibility to mesothelioma was transmitted in a Mendelian fashion and that the cause of the epidemic was gene–environment interaction [5,6,7].
Expanding the concept of gene–environment interaction (G × E)
In the complex picture of cancer pathogenesis and progression, it is now clear that both external environmental and genetic factors affect the so-called microenvironment, i.e. the local cellular milieu where cancer develops, which has a critical role in determining cell fate (Fig. 1).
Gene–environment interactions arise when the combined presence of both a genetic and an environmental factor modulates the risk of cancer more than either one alone. The traditional definition of G × E interaction most often implies ‘intrinsic’ genetic mutations (i.e. polymorphisms and rare germline mutations) and ‘non-iatrogenic’ environmental factors. Moreover, in cancer patients two additional levels of complexity arise: ‘acquired’ genetic mutations and ‘acquired iatrogenic’ environmental factors (e.g. drugs, radiotherapy) contribute to the cancer phenotype (Figs 2 and 3). Understanding the mutual relationships between all these factors is crucial both from a scientific and from a clinical perspective.
Environmental pressure and cancer susceptibility. A number of extrinsic factors associated to lifestyle, early-life influence or pre-existing chronic conditions exerts pressure on cancer insurgence. The genetic background, including somatic mutations, germline mutations and single nucleotide polymorphism contributes to the outcome of the impact of these environmental factors on the organism. Some extrinsic factors can also promote alteration of the genetic information producing mutagenic events that trigger cancer
The International Weinman Conferences, held at the University of Hawaii Cancer Center, Honolulu, HI, in 2017 (January 26, 27 and November 30–December 1) were conceived to stimulate discussions among experts from different fields of cancer research on complex inter-relations between genetics, external environment and microenvironment relevant to cancer aetiology—thus trying to find common ground in the lengthy conflict between ‘bad-luck’ and ‘toxic insults’, and to cancer progression and therapy. Here, we describe exemplificative topics on G × E interactions in both solid and haematologic malignancies presented by the authors, and the conclusions they reached following the discussions.
Genome-wide association studies, environment and molecular biology
Despite the unclear contribution of each single factor, it is commonly accepted that the risk of develo** cancer is determined by a complex interplay of both genetic and environmental factors [8,9,10]. There are many well-established environmental and lifestyle risk factors for various cancers, including smoking, alcohol, ionizing and solar radiations, exposure to carcinogenic fibres present in the environment, such as asbestos and erionite, as well as occupational exposures to these same fibres, infectious agents, obesity, and physical inactivity, which together are thought to account for up to 60% of cancer deaths in the United States [11]. However, only a small fraction of individuals exposed to these risk factors develop cancer, suggesting the co-existence of intrinsic genetic susceptibility.
Recent genome-wide association studies (GWAS) and next-generation sequencing studies have identified many common and rare germline genetic variations that contribute to increased cancer risks. Many of these cancer susceptibility loci are located in or near genes involved in DNA repair, or epigenetic regulators of gene transcription; however several examples exist of variants in genes involved in carcinogen metabolism. A paradigmatic example is the interaction between N-acetyltransferase 2 (NAT2) gene polymorphisms and smoking in elevating bladder cancer risk. NAT2 is a phase II detoxification enzyme that catalyses the metabolic inactivation of aromatic amines and heterocyclic amines, constituents of cigarette smoking and other environmental exposure. NAT2 polymorphisms allow classification of the human population into rapid, intermediate and slow ‘acetylator’ phenotypes. Numerous population studies and a meta-analysis [12] confirmed that NAT2 slow acetylators, resulting in slow detoxification of tobacco carcinogens, have a 1.4-fold increased risk of bladder cancer, and identified a significant synergistic interaction between NAT2 genotype and smoking: individuals with the NAT2 slow acetylating genotype who are also heavy smokers exhibited the highest risk of develo** bladder cancer [12]. Another eminent example is represented by the interaction between aldehyde dehydrogenase (ALDH2) gene variants, alcohol metabolism and oesophageal cancer [13].
Based on these and other data, there is a clear need for so-called ‘gene–environment-wide association studies’ which take into account environmental factors together with genetic variations to explain complex traits and phenomena [14]. Moreover, while in selected cases such as those represented by NAT2 and ALDH2, it is possible to infer the mechanistic link between the genetic variants and the consequent molecular alterations, a further step is needed to improve our understanding of G × E interactions from a molecular perspective, as shown by recent efforts in this direction [15].
Taking advantage of unique epidemiological resource
The study of G × E interaction requires the capability of discerning as much as possible the contribution of each of the two elements to the observed phenotype. Compared to small families and close relatives, which often share environment and lifestyle besides genes, such interactions can be more clearly elucidated studying distant relatives and high-risk pedigrees.
A unique resource that combines extensive genealogy with cancer and exposure data, the Utah Population Data Base (UPDB), provides the opportunity to better understand the contribution of genes, of the environment, and their combined effects to cancer. The UPDB represents the genealogy of the Utah Mormon pioneers and their descendants. The original Utah genealogy included 1.6 million individuals linked in genealogies 6–7 generations deep [16], and has been expanded from the 1970s with relationship data from Utah Vital Statistics records. State-wide SEER cancer data since 1966 for over 300,000 individuals have been linked to this database. The UPDB today represents 7 million Utah residents, of which 3 million have 3–16 generations of data. The power of the UPDB for G × E interaction studies lies in the simultaneous record of exposure data (such as tobacco use or radon exposure, both available in the UPDB), data for cancer incidence, and data on genetic relationships.
Health-related predisposition genes can be identified, and G × E interactions can be elucidated in high-risk pedigrees using these powerful genealogical resources. Analysis of UPDB high-risk pedigrees identified BRCA1, BRCA2, and CDKN2A as cancer predisposition genes [17,18,19], and studies of environmental exposures in individuals sharing genetic predisposition variants have clarified G × E interactions in cancer [20,21,22].
Similar efforts in creating large databases including both environmental and genetics data are steadily increasing in number, as they provide invaluable opportunities to study the complexity of cancer. Among the most recent examples, are the Iceland cohort [23], the Swedish Family-Cancer Database [24], and the US Veterans Genealogy Project [25].
The case of malignant mesothelioma and BAP1
Malignant mesothelioma (MM) is often used as the “standard” example of a cancer caused by G × E interaction. Epidemiologically, MM is an aggressive polyclonal cancer [26] associated with occupational and/or environmental exposure to tumorigenic mineral fibres, such as asbestos, erionite, or other non-regulated asbestos-like fibres [27,28,29,30]. However, “only” ~ 5% of asbestos workers exposed to asbestos continuously for over 10 years developed MM, suggesting that monogenic or polygenic genetic predisposition also contributes to MM risk [27]. Therefore, inherited genetic variants in genes encoding for proteins involved in the DNA damage response and/or in the inflammatory response may likely modulate the risk of asbestos-induced MM. Once they reach the mesothelium, asbestos fibres cause mesothelial cell death and chronic inflammation, associated with the release of damage-associated molecular patterns (DAMPs) molecules and cytokines [27]. Among these DAMPs, the active and passive release of different isoforms of high-mobility group box 1 (HMGB1) protein [31] sustains the chronic inflammatory response, with secretion of TNF-α and reactive oxygen species (ROS) [32, 33] (Fig. 4). Moreover, asbestos fibres can also directly induce ROS production because of the iron they contain, which behaves as a catalyst for free-radical generation [34, 35]. These molecules in turn activate NF-κB, leading to the survival of mesothelial cells that have accumulated genetic damage [32]. Accordingly, anti-inflammatory drugs and HMGB1 inhibitors impair MM growth in vitro and in experimental animal models and could represent prophylactic and therapeutic approaches [36,37,38].
Asbestos pathogenesis leads to mesothelioma. Asbestos fibers travel to pleura and cause human mesothelial cells (HM) to die of necrosis, leading to the release of HMGB1 into the extracellular space. HMGB1 induces macrophages and other inflammatory cells to accumulate and triggers the inflammatory response and leads to the secretion of cytokines such as TNF-α and IL-1β, which stimulate survival signalling pathways that increase survival of asbestos-damaged HM. This allows key genetic alterations to accumulate within HM that sustain asbestos-induced DNA damage that over time may lead to HM transformation and mesothelioma development
BRCA1-associated protein 1, BAP1 is the most frequently mutated gene in MM, as about 2/3 of sporadic cases—i.e., MM not develo** in carriers of germline BAP1 mutations—carry somatic BAP1 mutations [39, 40]. Besides single nucleotide variations and other intragenic alterations, frequently MM have large copy number variations and micro-deletions reminiscent of chromothripsis. Most common are minute deletions (ranging between 100 bp and 3000 bp) in chromosome band 3p21 in genes encoding for epigenetic modifiers such as BAP1 and others, e.g. SETD2, SCAP, SMARCC1, PBRM1 [39]. Because BAP1 mutations are very rare in lung cancer, the presence or absence of wild-type BAP1, easily detected as presence or absence of nuclear BAP1 immunostaining helps pathologists differentiate MM from lung cancer [41]. A better characterization of the most common molecular modifications in MM should pave the way to novel and more specific therapies [42].
In this respect, germline BAP1 mutations significantly increase the risk of MM, even among individuals not exposed to asbestos [43, 44]. Intriguingly, patients with MM who also carry germline BAP1 mutations have prolonged survival [45].
How to reconcile the environmental and the genetic factor in MM pathogenesis? The incidence of germline BAP1 mutations is low among sporadic MM patients, suggesting that other yet unknown genetic mutations might contribute to MM risk [46]. Based on the experimental evidence in mice, germline BAP1 mutations significantly increase the risk of asbestos-induced MM after exposure to low levels of asbestos, levels that rarely cause MM in wild-type mice [47, 48]. Germline BAP1 mutations are not only associated with MM, but also with a wider recently identified cancer predisposition syndrome which includes cutaneous and uveal melanoma, squamous and basal cell carcinoma (all associated to sunlight exposure) as well as to clear cell renal cell carcinoma (ccRCC) a malignancy not yet associated with a specific carcinogen. Moreover, although less frequent, carriers of germline BAP1 mutations develop also several other malignancies, such as breast cancer, cholangiocarcinoma, sarcomas, brain tumors, etc., suggesting that BAP1 mutations increase the overall cancer risk although those associated to environmental carcinogens predominate [49].
The case of clear cell renal cell carcinoma and BAP1
BAP1 is commonly mutated at the somatic level in ccRCC, and is one of two genes that underlie the first molecular genetic classification of ccRCC [50]. Mutations in BAP1 (found in ~ 15% of ccRCC) tend to anti-correlate with mutations in PBRM1 (found in ~ 50% of ccRCC). Both BAP1 and PBRM1 are two-hit tumour suppressor genes and they reside on chromosome 3p. Chromosome 3p harbours the VHL tumour suppressor gene, which is mutated (or silenced) in the majority of ccRCC. VHL mutation is an initiating event and this mutation is followed by loss of 3p, which eliminates the second copy of VHL as well as one copy of BAP1 and PBRM1 [217]. A mutation analysis shows that only 18% of mutations occurred in oncogene/tumour suppressors, whilst 82% of mutations were in other genes, including genes regulating immune-surveillance and epigenetic modifiers consistent with random mutation theory [217].
A recent paper by Tomasetti et al. tried to clarify the contribution of random replication errors, environmental factors and heredity, emphasising the difference between aetiology of cancer mutations and cancer prevention [218]. These authors proposed that most cancers are preventable, as mutations dependent on environmental factors –even though numerically less prevalent than those dependent on random replication errors or hereditary—still contribute to cancer development, and their absence would result in prevention of a large number of cancer cases.
The current hypothesis is that while intrinsic random DNA errors are the most frequent type of mutations in all tissues and cells, including stem-cells, development of cancer is often facilitated and accelerated by the combinatory effect of these “inevitable” errors—as cells accumulate about 3 new mutations/cell division, together with those caused by exposure to mutagenic environmental carcinogens and hereditary mutations.
References
Olson JS. The history of cancer: an annotated bibliography. New York: Greenwood Press;1989.
Cardiff RD, Kenney N. A compendium of the mouse mammary tumor biologist: from the initial observations in the house mouse to the development of genetically engineered mice. Cold Spring Harb Perspect Biol 2011;3.
Tomasetti C, Vogelstein B. Cancer etiology. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science. 2015;347:78–81.
Wu S, Powers S, Zhu W, Hannun YA. Substantial contribution of extrinsic risk factors to cancer development. Nature. 2016;529:43–47.
Roushdy-Hammady I, Siegel J, Emri S, Testa JR, Carbone M. Genetic-susceptibility factor and malignant mesothelioma in the Cappadocian region of Turkey. Lancet. 2001;357:444–5.
Carbone M, Emri S, Dogan AU, Steele I, Tuncer M, Pass HI, et al. A mesothelioma epidemic in Cappadocia: scientific developments and unexpected social outcomes. Nat Rev Cancer. 2007;7:147–54.
Emri SA. The Cappadocia mesothelioma epidemic: its influence in Turkey and abroad. Ann Transl Med. 2017;5:239.
Simonds NI, Ghazarian AA, Pimentel CB, Schully SD, Ellison GL, Gillanders EM, et al. Review of the Gene-Environment Interaction Literature in Cancer: What Do We Know? Genet Epidemiol. 2016;40:356–65.
Rudolph A, Chang-Claude J, Schmidt MK. Gene-environment interaction and risk of breast cancer. Br J Cancer. 2016;114:125–33.
Carr SR, Akerley W, Hashibe M, Cannon-Albright LA. Evidence for a genetical contribution to non-smoking-related lung cancer. Thorax. 2015;70:1033–9.
Schottenfeld D, Beebe-Dimmer JL, Buffler PA, Omenn GS. Current perspective on the global and United States cancer burden attributable to lifestyle and environmental risk factors. Annu Rev Public Health. 2013;34:97–117.
Garcia-Closas M, Malats N, Silverman D, Dosemeci M, Kogevinas M, Hein DW, et al. NAT2 slow acetylation, GSTM1 null genotype, and risk of bladder cancer: results from the Spanish Bladder Cancer Study and meta-analyses. Lancet. 2005;366:649–59.
Lewis SJ, Smith GD. Alcohol, ALDH2, and esophageal cancer: a meta-analysis which illustrates the potentials and limitations of a Mendelian randomization approach. Cancer Epidemiol Biomark Prev. 2005;14:1967–71.
Thomas D. Gene--environment-wide association studies: emerging approaches. Nat Rev Genet. 2010;11:259–72.
Moyerbrailean GA, Richards AL, Kurtz D, Kalita CA, Davis GO, Harvey CT, et al. High-throughput allele-specific expression across 250 environmental conditions. Genome Res. 2016;26:1627–38.
Cairns J, Lyon JL, Skolnick M. Cancer incidence in defined populations. Cold Spring Harbor, N.Y.: Cold Spring Harbor Laboratory; 1980.
Miki Y, Swensen J, Shattuck-Eidens D, Futreal PA, Harshman K, Tavtigian S, et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science. 1994;266:66–71.
Wooster R, Neuhausen SL, Mangion J, Quirk Y, Ford D, Collins N, et al. Localization of a breast cancer susceptibility gene, BRCA2, to chromosome 13q12-13. Science. 1994;265:2088–90.
Kamb A, Shattuck-Eidens D, Eeles R, Liu Q, Gruis NA, Ding W, et al. Analysis of the p16 gene (CDKN2) as a candidate for the chromosome 9p melanoma susceptibility locus. Nat Genet. 1994;8:23–26.
Churpek JE, Marquez R, Neistadt B, Claussen K, Lee MK, Churpek MM, et al. Inherited mutations in cancer susceptibility genes are common among survivors of breast cancer who develop therapy-related leukemia. Cancer. 2016;122:304–11.
Pijpe A, Andrieu N, Easton DF, Kesminiene A, Cardis E, Nogues C, et al. Exposure to diagnostic radiation and risk of breast cancer among carriers of BRCA1/2 mutations: retrospective cohort study (GENE-RAD-RISK). BMJ. 2012;345:e5660.
Cannon-Albright LA, Thomas A, Goldgar DE, Gholami K, Rowe K, Jacobsen M, et al. Familiality of cancer in Utah. Cancer Res. 1994;54:2378–85.
Amundadottir LT, Thorvaldsson S, Gudbjartsson DF, Sulem P, Kristjansson K, Arnason S, et al. Cancer as a complex phenotype: pattern of cancer distribution within and beyond the nuclear family. PLoS Med. 2004;1:e65.
Hemminki K, Rawal R, Chen B, Bermejo JL. Genetic epidemiology of cancer: from families to heritable genes. Int J Cancer. 2004;111:944–50.
Cannon-Albright LA, Dintelman S, Maness T, Backus S, Thomas A, Meyer LJ. Creation of a national resource with linked genealogy and phenotypic data: the Veterans Genealogy Project. Genet Med. 2013;15:541–7.
Comertpay S, Pastorino S, Tanji M, Mezzapelle R, Strianese O, Napolitano A, et al. Evaluation of clonal origin of malignant mesothelioma. J Transl Med. 2014;12:301.
Carbone M, Ly BH, Dodson RF, Pagano I, Morris PT, Dogan UA, et al. Malignant mesothelioma: facts, myths, and hypotheses. J Cell Physiol. 2012;227:44–58.
Berry G, Reid A, Aboagye-Sarfo P, de Klerk NH, Olsen NJ, Merler E, et al. Malignant mesotheliomas in former miners and millers of crocidolite at Wittenoom (Western Australia) after more than 50 years follow-up. Br J Cancer. 2012;106:1016–20.
Baumann F, Buck BJ, Metcalf RV, McLaurin BT, Merkler DJ, Carbone M. The Presence of Asbestos in the Natural Environment is Likely Related to Mesothelioma in Young Individuals and Women from Southern Nevada. J Thorac Oncol. 2015;10:731–7.
Baumann F, Ambrosi JP, Carbone M. Asbestos is not just asbestos: an unrecognised health hazard. Lancet Oncol. 2013;14:576–8.
Napolitano A, Antoine DJ, Pellegrini L, Baumann F, Pagano I, Pastorino S, et al. HMGB1 and Its Hyperacetylated Isoform are Sensitive and Specific Serum Biomarkers to Detect Asbestos Exposure and to Identify Mesothelioma Patients. Clin Cancer Res. 2016;22:3087–96.
Yang H, Rivera Z, Jube S, Nasu M, Bertino P, Goparaju C, et al. Programmed necrosis induced by asbestos in human mesothelial cells causes high-mobility group box 1 protein release and resultant inflammation. Proc Natl Acad Sci USA. 2010;107:12611–6.
Caputa G, Zhao S, Criado AE, Ory DS, Duncan JG, Schaffer JE. RNASET2 is required for ROS propagation during oxidative stress-mediated cell death. Cell Death Differ. 2016;23:347–57.
Simeonova PP, Luster MI. Iron and reactive oxygen species in the asbestos-induced tumor necrosis factor-alpha response from alveolar macrophages. Am J Respir Cell Mol Biol. 1995;12:676–83.
Croce A, Allegrina M, Rinaudo C, Gaudino G, Yang H, Carbone M. Numerous Iron-Rich Particles Lie on the Surface of Erionite Fibers from Rome (Oregon, USA) and Karlik (Cappadocia, Turkey). Microsc Microanal. 2015;21:1341–7.
Yang H, Pellegrini L, Napolitano A, Giorgi C, Jube S, Preti A, et al. Aspirin delays mesothelioma growth by inhibiting HMGB1-mediated tumor progression. Cell Death Dis. 2015;6:e1786.
Pellegrini L, Xue J, Larson D, Pastorino S, Jube S, Forest KH, et al. HMGB1 targeting by ethyl pyruvate suppresses malignant phenotype of human mesothelioma. Oncotarget. 2017;8:22649–61.
Bononi A, Napolitano A, Pass HI, Yang H, Carbone M. Latest developments in our understanding of the pathogenesis of mesothelioma and the design of targeted therapies. Expert Rev Respir Med. 2015;9:633–54.
Yoshikawa Y, Emi M, Hashimoto-Tamaoki T, Ohmuraya M, Sato A, Tsujimura T, et al. High-density array-CGH with targeted NGS unmask multiple noncontiguous minute deletions on chromosome 3p21 in mesothelioma. Proc Natl Acad Sci USA. 2016;113:13432–7.
Nasu M, Emi M, Pastorino S, Tanji M, Powers A, Luk H, et al. High Incidence of Somatic BAP1 alterations in sporadic malignant mesothelioma. J Thorac Oncol. 2015;10:565–76.
Carbone M, Shimizu D, Napolitano A, Tanji M, Pass HI, Yang H, et al. Positive nuclear BAP1 immunostaining helps differentiate non-small cell lung carcinomas from malignant mesothelioma. Oncotarget. 2016;7:59314–21.
Napolitano A, Carbone M. Malignant Mesothelioma: Time toTranslate? Trends Cancer. 2016;2:467–74.
Testa JR, Cheung M, Pei J, Below JE, Tan Y, Sementino E, et al. Germline BAP1 mutations predispose to malignant mesothelioma. Nat Genet. 2011;43:1022–5.
Carbone M, Flores EG, Emi M, Johnson TA, Tsunoda T, Behner D, et al. Combined Genetic and Genealogic Studies Uncover a Large BAP1 Cancer Syndrome Kindred Tracing Back Nine Generations to a Common Ancestor from the 1700s. PLoS Genet. 2015;11:e1005633.
Baumann F, Flores E, Napolitano A, Kanodia S, Taioli E, Pass H, et al. Mesothelioma patients with germline BAP1 mutations have 7-fold improved long-term survival. Carcinogenesis. 2015;36:76–81.
Rusch A, Ziltener G, Nackaerts K, Weder W, Stahel RA, Felley-Bosco E. Prevalence of BRCA-1 associated protein 1 germline mutation in sporadic malignant pleural mesothelioma cases. Lung Cancer. 2015;87:77–79.
Napolitano A, Pellegrini L, Dey A, Larson D, Tanji M, Flores EG, et al. Minimal asbestos exposure in germline BAP1 heterozygous mice is associated with deregulated inflammatory response and increased risk of mesothelioma. Oncogene. 2016;35:1996–2002.
Betti M, Casalone E, Ferrante D, Romanelli A, Grosso F, Guarrera S, et al. Inference on germline BAP1 mutations and asbestos exposure from the analysis of familial and sporadic mesothelioma in a high-risk area. Genes Chromosomes Cancer. 2015;54:51–62.
Carbone M, Yang H, Pass HI, Krausz T, Testa JR, Gaudino G. BAP1 and cancer. Nat Rev Cancer. 2013;13:153–9.
Pena-Llopis S, Vega-Rubin-de-Celis S, Liao A, Leng N, Pavia-Jimenez A, Wang S, et al. BAP1 loss defines a new class of renal cell carcinoma. Nat Genet. 2012;44:751–9.
Pena-Llopis S, Christie A, **e XJ, Brugarolas J. Cooperation and antagonism among cancer genes: the renal cancer paradigm. Cancer Res. 2013;73:4173–9.
Gu YF, Cohn S, Christie A, McKenzie T, Wolff N, Do QN, et al. Modeling renal cell carcinoma in mice: Bap1 and Pbrm1 inactivation drive tumor grade. Cancer Discov. 2017;7:900–17.
Popova T, Hebert L, Jacquemin V, Gad S, Caux-Moncoutier V, Dubois-d’Enghien C, et al. Germline BAP1 mutations predispose to renal cell carcinomas. Am J Hum Genet. 2013;92:974–80.
Schmidt LS, Linehan WM. Genetic predisposition to kidney cancer. Semin Oncol. 2016;43:566–74.
Farley MN, Schmidt LS, Mester JL, Pena-Llopis S, Pavia-Jimenez A, Christie A, et al. A novel germline mutation in BAP1 predisposes to familial clear-cell renal cell carcinoma. Mol Cancer Res: MCR. 2013;11:1061–71.
Melkonian SC, Daniel CR, Ye Y, Tannir NM, Karam JA, Matin SF, et al. Gene-environment interaction of genome-wide association study-identified susceptibility loci and meat-cooking mutagens in the etiology of renal cell carcinoma. Cancer. 2016;122:108–15.
Scelo G, Riazalhosseini Y, Greger L, Letourneau L, Gonzalez-Porta M, Wozniak MB, et al. Variation in genomic landscape of clear cell renal cell carcinoma across Europe. Nat Commun. 2014;5:5135.
Beckerman R, Prives C. Transcriptional regulation byp53. Cold Spring Harb Perspect Biol. 2010;2:a000935.
Charni M, Molchadsky A, Goldstein I, Solomon H, Tal P, Goldfinger N, et al. Novel p53 target genes secreted by the liver are involved in non-cell-autonomous regulation. Cell Death Differ. 2016;23:509–20.
Prives C, Bargonetti J, Farmer G, Ferrari E, Friedlander P, Wang Y, et al. DNA-binding properties of the p53 tumor suppressor protein. Cold Spring Harb Symp Quant Biol. 1994;59:207–13.
Kruiswijk F, Labuschagne CF, Vousden KH. p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol. 2015;16:393–405.
Charni M, Aloni-Grinstein R, Molchadsky A, Rotter V. p53 on the crossroad between regeneration and cancer. Cell Death Differ. 2017;24:8–14.
Lowe JM, Nguyen TA, Grimm SA, Gabor KA, Peddada SD, Li L, et al. The novel p53 target TNFAIP8 variant 2 is increased in cancer and offsets p53-dependent tumor suppression. Cell Death Differ. 2017;24:181–91.
Lopez I, Tournillon AS, Prado Martins R, Karakostis K, Malbert-Colas L, Nylander K, et al. p53-mediated suppression of BiP triggers BIK-induced apoptosis during prolonged endoplasmic reticulum stress. Cell Death Differ. 2017;24:1717–29.
Muller PA, Vousden KH. p53 mutations in cancer. Nat Cell Biol. 2013;15:2–8.
Muller PA, Vousden KH. Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell. 2014;25:304–17.
Aggarwal M, Saxena R, Sinclair E, Fu Y, Jacobs A, Dyba M, et al. Reactivation of mutant p53 by a dietary-related compound phenethyl isothiocyanate inhibits tumor growth. Cell Death Differ. 2016;23:1615–27.
Kastenhuber ER, Lowe SW. Putting p53 in Context. Cell. 2017;170:1062–78.
Liu Y, Chen C, Xu Z, Scuoppo C, Rillahan CD, Gao J, et al. Deletions linked to TP53 loss drive cancer through p53-independent mechanisms. Nature. 2016;531:471–5.
Weissmueller S, Manchado E, Saborowski M, Morris JPt, Wagenblast E, Davis CA, et al. Mutant p53 drives pancreatic cancer metastasis through cell-autonomous PDGF receptor beta signaling. Cell. 2014;157:382–94.
Adorno M, Cordenonsi M, Montagner M, Dupont S, Wong C, Hann B, et al. A Mutant-p53/Smad complex opposes p63 to empower TGFbeta-induced metastasis. Cell. 2009;137:87–98.
Muller PA, Caswell PT, Doyle B, Iwanicki MP, Tan EH, Karim S, et al. Mutant p53 drives invasion by promoting integrin recycling. Cell. 2009;139:1327–41.
Kehrloesser S, Osterburg C, Tuppi M, Schafer B, Vousden KH, Dotsch V. Intrinsic aggregation propensity of the p63 and p73 TI domains correlates with p53R175H interaction and suggests further significance of aggregation events in the p53 family. Cell Death Differ. 2016;23:1952–60.
Freed-Pastor WA, Mizuno H, Zhao X, Langerod A, Moon SH, Rodriguez-Barrueco R, et al. Mutant p53 disrupts mammary tissue architecture via the mevalonate pathway. Cell. 2012;148:244–58.
Alexandrova EM, Moll UM. Depleting stabilized GOF mutant p53 proteins by inhibiting molecular folding chaperones: a new promise in cancer therapy. Cell Death Differ. 2017;24:3–5.
Stracquadanio G, Wang X, Wallace MD, Grawenda AM, Zhang P, Hewitt J, et al. The importance of p53 pathway genetics in inherited and somatic cancer genomes. Nat Rev Cancer. 2016;16:251–65.
Sarkar J, Dominguez E, Li G, Kusewitt DF, Johnson DG. Modeling gene-environment interactions in oral cavity and esophageal cancers demonstrates a role for the p53 R72P polymorphism in modulating susceptibility. Mol Carcinog. 2014;53:648–58.
Costanzo A, Pediconi N, Narcisi A, Guerrieri F, Belloni L, Fausti F, et al. TP63 and TP73 in cancer, an unresolved “family” puzzle of complexity, redundancy and hierarchy. FEBS Lett. 2014;588:2590–9.
Gebel J, Luh LM, Coutandin D, Osterburg C, Lohr F, Schafer B, et al. Mechanism of TAp73 inhibition by DeltaNp63 and structural basis of p63/p73 hetero-tetramerization. Cell Death Differ. 2016;23:1930–40.
Van Nostrand JL, Bowen ME, Vogel H, Barna M, Attardi LD. The p53 family members have distinct roles during mammalian embryonic development. Cell Death Differ. 2017;24:575–9.
Amelio I, Melino G. The p53 family and the hypoxia-inducible factors (HIFs): determinants of cancer progression. Trends Biochem Sci. 2015;40:425–34.
Wu Q, Shi Y, Ge L, Ma D, Zhang H, Wang J. Relationship of p73 gene polymorphism and additional gene-smoking and gene-obesity interaction with non-small cell lung cancer risk. Oncotarget. 2017;8:34423–34428.
Levine AJ, Tomasini R, McKeon FD, Mak TW, Melino G. The p53 family: guardians of maternal reproduction. Nat Rev Mol Cell Biol. 2011;12:259–65.
Memmi EM, Sanarico AG, Giacobbe A, Peschiaroli A, Frezza V, Cicalese A, et al. p63 Sustains self-renewal of mammary cancer stem cells through regulation of Sonic Hedgehog signaling. Proc Natl Acad Sci USA. 2015;112:3499–504.
Viticchie G, Agostini M, Lena AM, Mancini M, Zhou H, Zolla L, et al. p63 supports aerobic respiration through hexokinase II. Proc Natl Acad Sci USA. 2015;112:11577–82.
Tomasini R, Tsuchihara K, Wilhelm M, Fujitani M, Rufini A, Cheung CC, et al. TAp73 knockout shows genomic instability with infertility and tumor suppressor functions. Genes & Dev. 2008;22:2677–91.
Tomasini R, Tsuchihara K, Tsuda C, Lau SK, Wilhelm M, Ruffini A, et al. TAp73 regulates the spindle assembly checkpoint by modulating BubR1 activity. Proc Natl Acad Sci USA. 2009;106:797–802.
Amelio I, Inoue S, Markert EK, Levine AJ, Knight RA, Mak TW, et al. TAp73 opposes tumor angiogenesis by promoting hypoxia-inducible factor 1alpha degradation. Proc Natl Acad Sci USA. 2015;112:226–31.
Amelio I, Markert EK, Rufini A, Antonov AV, Sayan BS, Tucci P, et al. p73 regulates serine biosynthesis in cancer. Oncogene. 2014;33:5039–46.
Rufini A, Niklison-Chirou MV, Inoue S, Tomasini R, Harris IS, Marino A, et al. TAp73 depletion accelerates aging through metabolic dysregulation. Genes & Dev. 2012;26:2009–14.
Velletri T, Romeo F, Tucci P, Peschiaroli A, Annicchiarico-Petruzzelli M, Niklison-Chirou MV, et al. GLS2 is transcriptionally regulated by p73 and contributes to neuronal differentiation. Cell Cycle. 2013;12:3564–73.
D’Alessandro A, Amelio I, Berkers CR, Antonov A, Vousden KH, Melino G, et al. Metabolic effect of TAp63alpha: enhanced glycolysis and pentose phosphate pathway, resulting in increased antioxidant defense. Oncotarget. 2014;5:7722–33.
Hu W, Zhang C, Wu R, Sun Y, Levine A, Feng Z. Glutaminase 2, a novel p53 target gene regulating energy metabolism and antioxidant function. Proc Natl Acad Sci USA. 2010;107:7455–60.
Amelio I, Antonov AA, Catani MV, Massoud R, Bernassola F, Knight RA, et al. TAp73 promotes anabolism. Oncotarget. 2014;5:12820–934.
Solomon H, Brauning B, Fainer I, Ben-Nissan G, Rabani S, Goldfinger N, et al. Post-translational regulation of p53 function through 20S proteasome-mediated cleavage. Cell Death Differ. 2017;24:2187–98.
Nemajerova A, Amelio I, Gebel J, Dotsch V, Melino G, Moll UM. Non-oncogenic roles of TAp73: from multiciliogenesis to metabolism. Cell death Differ. 2017;25:144–153.
Amelio I, Cutruzzola F, Antonov A, Agostini M, Melino G. Serine and glycine metabolism in cancer. Trends Biochem Sci. 2014;39:191–8.
Antonov A, Agostini M, Morello M, Minieri M, Melino G, Amelio I. Bioinformatics analysis of the serine and glycine pathway in cancer cells. Oncotarget. 2014;5:11004–13.
Sharif T, Ahn DG, Liu RZ, Pringle E, Martell E, Dai C, et al. The NAD(+) salvage pathway modulates cancer cell viability via p73. Cell Death Differ. 2016;23:669–80.
Clendening JW, Pandyra A, Boutros PC, El Ghamrasni S, Khosravi F, Trentin GA, et al. Dysregulation of the mevalonate pathway promotes transformation. Proc Natl Acad Sci USA. 2010;107:15051–6.
Ingallina E, Sorrentino G, Bertolio R, Lisek K, Zannini A, Azzolin L, et al. Mechanical cues control mutant p53 stability through a mevalonate-RhoA axis. Nat Cell Biol. 2018;20:28–35.
Parrales A, Ranjan A, Iyer SV, Padhye S, Weir SJ, Roy A, et al. DNAJA1 controls the fate of misfolded mutant p53 through the mevalonate pathway. Nat Cell Biol. 2016;18:1233–43.
Ahern TP, Pedersen L, Tarp M, Cronin-Fenton DP, Garne JP, Silliman RA, et al. Statin prescriptions and breast cancer recurrence risk: a Danish nationwide prospective cohort study. J Natl Cancer Inst. 2011;103:1461–8.
Clendening JW, Pandyra A, Li Z, Boutros PC, Martirosyan A, Lehner R, et al. Exploiting the mevalonate pathway to distinguish statin-sensitive multiple myeloma. Blood. 2010;115:4787–97.
Mikoshiba K. IP3 receptor/Ca2+channel: from discovery to new signaling concepts. J Neurochem. 2007;102:1426–46.
Mikoshiba K. Role of IP3 receptor signaling in cell functions and diseases. Adv Biol Regul. 2015;57(Supplement C):217–27.
Berridge MJ. The inositol trisphosphate/calcium signaling pathway in health and disease. Physiol Rev. 2016;96:1261–96.
Furuichi T, Yoshikawa S, Miyawaki A, Wada K, Maeda N, Mikoshiba K. Primary structure and functional expression of the inositol 1,4,5-trisphosphate-binding protein P400. Nature. 1989;342:32–38.
Mignery GA, Newton CL, Archer BT 3rd, Sudhof TC. Structure and expression of the rat inositol 1,4,5-trisphosphate receptor. J Biol Chem. 1990;265:12679–85.
Sudhof TC, Newton CL, Archer BT 3rd, Ushkaryov YA, Mignery GA. Structure of a novel InsP3 receptor. EMBO J. 1991;10:3199–206.
Blondel O, Takeda J, Janssen H, Seino S, Bell GI. Sequence and functional characterization of a third inositol trisphosphate receptor subtype, IP3R-3, expressed in pancreatic islets, kidney, gastrointestinal tract, and other tissues. J Biol Chem. 1993;268:11356–63.
Yamamoto-Hino M, Sugiyama T, Hikichi K, Mattei MG, Hasegawa K, Sekine S, et al. Cloning and characterization of human type 2 and type 3 inositol 1,4,5-trisphosphate receptors. Recept & Channels. 1994;2:9–22.
Yamada N, Makino Y, Clark RA, Pearson DW, Mattei MG, Guénet JL, et al. Human inositol 1,4,5-trisphosphate type-1 receptor, Ins<em>P</em>3R1: structure, function, regulation of expression and chromosomal localization. Biochem J. 1994;302:781–90.
White C, Li C, Yang J, Petrenko NB, Madesh M, Thompson CB, et al. The endoplasmic reticulum gateway to apoptosis by Bcl-X(L) modulation of the InsP3R. Nat Cell Biol. 2005;7:1021–8.
Eckenrode EF, Yang J, Velmurugan GV, Foskett JK, White C. Apoptosis protection by Mcl-1 and Bcl-2 modulation of inositol 1,4,5-trisphosphate receptor-dependent Ca2+signaling. J Biol Chem. 2010;285:13678–84.
Szado T, Vanderheyden V, Parys JB, De Smedt H, Rietdorf K, Kotelevets L, et al. Phosphorylation of inositol 1,4,5-trisphosphate receptors by protein kinase B/Akt inhibits Ca2+release and apoptosis. Proc Natl Acad Sci. 2008;105:2427–32.
Boehning D, Patterson RL, Sedaghat L, Glebova NO, Kurosaki T, Snyder SH. Cytochrome c binds to inositol (1,4,5) trisphosphate receptors, amplifying calcium-dependent apoptosis. Nat Cell Biol. 2003;5:1051–61.
Zhang S, Mizutani A, Hisatsune C, Higo T, Bannai H, Nakayama T, et al. Protein 4.1N is required for translocation of inositol 1,4,5-trisphosphate receptor type 1 to the basolateral membrane domain in polarized Madin-Darby canine kidney cells. J Biol Chem. 2003;278:4048–56.
Kawaai K, Hisatsune C, Kuroda Y, Mizutani A, Tashiro T, Mikoshiba K. 80K-H interacts with inositol 1,4,5-trisphosphate (IP3) receptors and regulates IP3-induced calcium release activity. J Biol Chem. 2009;284:372–80.
Tang T-S, Tu H, Chan EYW, Maximov A, Wang Z, Wellington CL, et al. Huntingtin and Huntingtin-Associated Protein 1 Influence Neuronal Calcium Signaling Mediated by Inositol-(1,4,5) Triphosphate Receptor Type 1. Neuron. 2003;39:227–39.
Sung PJ, Tsai FD, Vais H, Court H, Yang J, Fehrenbacher N, et al. Phosphorylated K-Ras limits cell survival by blocking Bcl-xL sensitization of inositol trisphosphate receptors. Proc Natl Acad Sci. 2013;110:20593–8.
Zhang S, Hisatsune C, Matsu-Ura T, Mikoshiba K. G-protein-coupled receptor kinase-interacting proteins inhibit apoptosis by inositol 1,4,5-triphosphate receptor-mediated Ca2+signal regulation. J Biol Chem. 2009;284:29158–69.
Hedgepeth SC, Garcia MI, Wagner LE, Rodriguez AM, Chintapalli SV, Snyder RR, et al. The BRCA1 Tumor Suppressor Binds to Inositol 1,4,5-Trisphosphate Receptors to Stimulate Apoptotic Calcium Release. J Biol Chem. 2015;290:7304–13.
Kuchay S, Giorgi C, Simoneschi D, Pagan J, Missiroli S, Saraf A, et al. PTEN counteracts FBXL2 to promote IP3R3- and Ca2+-mediated apoptosis limiting tumour growth. Nature. 2017;546:554–8.
Bononi A, Giorgi C, Patergnani S, Larson D, Verbruggen K, Tanji M, et al. BAP1 regulates IP3R3-mediated Ca2+flux to mitochondria suppressing cell transformation. Nature. 2017;546:549–53.
Hamada K, Miyatake H, Terauchi A, Mikoshiba K. IP3-mediated gating mechanism of the IP3 receptor revealed by mutagenesis and X-ray crystallography. Proc Natl Acad Sci USA. 2017;114:4661–6.
Rizzuto R, Brini M, Murgia M, Pozzan T. Microdomains with high Ca2+close to IP3-sensitive channels that are sensed by neighboring mitochondria. Sci (New Y, NY). 1993;262:744–7.
Rizzuto R, De Stefani D, Raffaello A, Mammucari C. Mitochondria as sensors and regulators of calcium signalling. Nat Rev Mol Cell Biol. 2012;13:566–78.
Gomez L, Thiebaut PA, Paillard M, Ducreux S, Abrial M, Crola Da Silva C, et al. The SR/ER-mitochondria calcium crosstalk is regulated by GSK3beta during reperfusion injury. Cell Death Differ. 2016;23:313–22.
Rowland AA, Voeltz GK. Endoplasmic reticulum-mitochondria contacts: function of the junction. Nat Rev Mol Cell Biol. 2012;13:607–25.
Vance JE. Phospholipid synthesis in a membrane fraction associated with mitochondria. J Biol Chem. 1990;265:7248–56.
Rizzuto R, Pinton P, Carrington W, Fay FS, Fogarty KE, Lifshitz LM, et al. Close Contacts with the Endoplasmic Reticulum as Determinants of Mitochondrial Ca2+2+Responses. Sci (New Y, NY). 1998;280:1763–6.
Csordás G, Renken C, Várnai P, Walter L, Weaver D, Buttle KF, et al. Structural and functional features and significance of the physical linkage between ER and mitochondria. J Cell Biol. 2006;174:915–21.
Szabadkai G, Bianchi K, Várnai P, De Stefani D, Wieckowski MR, Cavagna D, et al. Chaperone-mediated coupling of endoplasmic reticulum and mitochondrial Ca2+2+channels. J Cell Biol. 2006;175:901–11.
De Stefani D, Bononi A, Romagnoli A, Messina A, De Pinto V, Pinton P, et al. VDAC1 selectively transfers apoptotic Ca2+signals to mitochondria. Cell Death Differ. 2012;19:267–73.
De Stefani D, Raffaello A, Teardo E, Szabo I, Rizzuto R. A forty-kilodalton protein of the inner membrane is the mitochondrial calcium uniporter. Nature. 2011;476:336–40.
Baughman JM, Perocchi F, Girgis HS, Plovanich M, Belcher-Timme CA, Sancak Y, et al. Integrative genomics identifies MCU as an essential component of the mitochondrial calcium uniporter. Nature. 2011;476:341–5.
Khan AA, Soloski MJ, Sharp AH, Schilling G, Sabatini DM, Li SH, et al. Lymphocyte apoptosis: mediation by increased type 3 inositol 1,4,5-trisphosphate receptor. Sci (New Y, NY). 1996;273:503–7.
Jayaraman T, Marks AR. T cells deficient in inositol 1,4,5-trisphosphate receptor are resistant to apoptosis. Mol Cell Biol. 1997;17:3005–12.
Sugawara H, Kurosaki M, Takata M, Kurosaki T. Genetic evidence for involvement of type 1, type 2 and type 3 inositol 1,4,5-trisphosphate receptors in signal transduction through the B-cell antigen receptor. EMBO J. 1997;16:3078–88.
Joseph SK, Hajnoczky G. IP3 receptors in cell survival and apoptosis: Ca2+release and beyond. Apoptosis: Int J Program Cell death. 2007;12:951–68.
Rasola A, Bernardi P. Mitochondrial permeability transition in Ca(2+)-dependent apoptosis and necrosis. Cell Calcium. 2011;50:222–33.
Boehning D, van Rossum DB, Patterson RL, Snyder SH. A peptide inhibitor of cytochrome c/inositol 1,4,5-trisphosphate receptor binding blocks intrinsic and extrinsic cell death pathways. Proc Natl Acad Sci USA. 2005;102:1466–71.
Greenberg EF, Lavik AR, Distelhorst CW. Bcl-2 regulation of the inositol 1,4,5-trisphosphate receptor and calcium signaling in normal and malignant lymphocytes: potential new target for cancer treatment. Biochim Biophys Acta. 2014;1843:2205–10.
Vervloessem T, Kerkhofs M, La Rovere RM, Sneyers F, Parys JB, Bultynck G Bcl-2 inhibitors as anti-cancer therapeutics: the impact of and on calcium signaling. Cell Calcium 2018;70:102–116.
Ando H, Mizutani A, Matsu-ura T, Mikoshiba K. IRBIT, a novel inositol 1,4,5-trisphosphate (IP3) receptor-binding protein, is released from the IP3 receptor upon IP3 binding to the receptor. J Biol Chem. 2003;278:10602–12.
Ando H, Mizutani A, Kiefer H, Tsuzurugi D, Michikawa T, Mikoshiba K. IRBIT suppresses IP3 receptor activity by competing with IP3 for the common binding site on the IP3 receptor. Mol Cell. 2006;22:795–806.
Ando H, Mizutani A, Mikoshiba K. An IRBIT homologue lacks binding activity to inositol 1,4,5-trisphosphate receptor due to the unique N-terminal appendage. J Neurochem. 2009;109:539–50.
Kawaai K, Ando H, Satoh N, Yamada H, Ogawa N, Hirose M, et al. Splicing variation of Long-IRBIT determines the target selectivity of IRBIT family proteins. Proc Natl Acad Sci USA. 2017;114:3921–6.
Ando H, Kawaai K, Mikoshiba K. Biochimica et biophysica acta. 1843. IRBIT: a regulator of ion channels and ion transporters; 2014. p. 2195–204.
Kawaai K, Mizutani A, Shoji H, Ogawa N, Ebisui E, Kuroda Y, et al. IRBIT regulates CaMKIIalpha activity and contributes to catecholamine homeostasis through tyrosine hydroxylase phosphorylation. Proc Natl Acad Sci USA. 2015;112:5515–20.
Ando H, Hirose M, Gainche L, Kawaai K, Bonneau B, Ijuin T, et al. IRBIT Interacts with the Catalytic Core of Phosphatidylinositol Phosphate Kinase Type Ialpha and IIalpha through Conserved Catalytic Aspartate Residues. PLoS One. 2015;10:e0141569.
Bonneau B, Ando H, Kawaai K, Hirose M, Takahashi-Iwanaga H, Mikoshiba K IRBIT controls apoptosis by interacting with the Bcl-2 homolog, Bcl2l10, and by promoting ER-mitochondria contact. Elife 2016, 5. pii:e19896.
Arnaoutov A, Dasso M. Enzyme regulation. IRBIT Is a Nov Regul ribonucleotide reductase High eukaryotes Sci. 2014;345:1512–5.
Ando H, Kawaai K, Bonneau B, Mikoshiba K Remodeling of Ca(2+) signaling in cancer: Regulation of inositol 1,4,5-trisphosphate receptors through oncogenes and tumor suppressors. Advances in Biological Regulation 2018;6:64–76.
Yu H, Pak H, Hammond-Martel I, Ghram M, Rodrigue A, Daou S, et al. Tumor suppressor and deubiquitinase BAP1 promotes DNA double-strand break repair. Proc Natl Acad Sci USA. 2014;111:285–90.
Ismail IH, Davidson R, Gagne JP, Xu ZZ, Poirier GG, Hendzel MJ. Germline mutations in BAP1 impair its function in DNA double-strand break repair. Cancer Res. 2014;74:4282–94.
Bononi A, Yang H, Giorgi C, Patergnani S, Pellegrini L, Su M, et al. Germline BAP1 mutations induce a Warburg effect. Cell Death Differ. 2017;24:1694–704.
Amelio I. Genes versus Environment: cytoplasmic BAP1 determines the toxic response to environmental stressors in mesothelioma. Cell death & Dis. 2017;8:e2907.
Bennett WP, Hussain SP, Vahakangas KH, Khan MA, Shields PG, Harris CC. Molecular epidemiology of human cancer risk: gene-environment interactions and p53 mutation spectrum in human lung cancer. J Pathol. 1999;187:8–18.
Zhang R, Chu M, Zhao Y, Wu C, Guo H, Shi Y, et al. A genome-wide gene-environment interaction analysis for tobacco smoke and lung cancer susceptibility. Carcinogenesis. 2014;35:1528–35.
Haugen A, Ryberg D, Mollerup S, Zienolddiny S, Skaug V, Svendsrud DH. Gene-environment interactions in human lung cancer. Toxicol Lett. 2000;112-3:233–7.
Wu X, Zhao H, Suk R, Christiani DC. Genetic susceptibility to tobacco-related cancer. Oncogene. 2004;23:6500–623.
Thomas A, Liu SV, Subramaniam DS, Giaccone G. Refining the treatment of NSCLC according to histological and molecular subtypes. Nat Rev Clin Oncol. 2015;12:511–26.
Yu T, Chen X, Zhang W, Liu J, Avdiushko R, Napier DL, et al. KLF4 regulates adult lung tumor-initiating cells and represses K-Ras-mediated lung cancer. Cell Death Differ. 2016;23:207–15.
Jamal-Hanjani M, Wilson GA, McGranahan N, Birkbak NJ, Watkins TBK, Veeriah S, et al. Tracking the Evolution of Non-Small-Cell Lung Cancer. N Engl J Med. 2017;376:2109–2121.
Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H, et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med. 2010;362:2380–8.
Mitsudomi T, Morita S, Yatabe Y, Negoro S, Okamoto I, Tsurutani J, et al. Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): an open label, randomised phase 3 trial. Lancet Oncol. 2010;11:121–8.
Zhou C, Wu YL, Chen G, Feng J, Liu XQ, Wang C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011;12:735–42.
Rosell R, Carcereny E, Gervais R, Vergnenegre A, Massuti B, Felip E, et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2012;13:239–46.
Yang JC, Wu YL, Schuler M, Sebastian M, Popat S, Yamamoto N, et al. Afatinib versus cisplatin-based chemotherapy for EGFR mutation-positive lung adenocarcinoma (LUX-Lung 3 and LUX-Lung 6): analysis of overall survival data from two randomised, phase 3 trials. Lancet Oncol. 2015;16:141–51.
Furnari FB, Cloughesy TF, Cavenee WK, Mischel PS. Heterogeneity of epidermal growth factor receptor signalling networks in glioblastoma. Nat Rev Cancer. 2015;15:302–10.
Piyush T, Chacko AR, Sindrewicz P, Hilkens J, Rhodes JM, Yu LG. Interaction of galectin-3 with MUC1 on cell surface promotes EGFR dimerization and activation in human epithelial cancer cells. Cell Death Differ. 2017;24:1937–47.
Turner KM, Deshpande V, Beyter D, Koga T, Rusert J, Lee C, et al. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature. 2017;543:122–5.
Abbosh C, Birkbak NJ, Wilson GA, Jamal-Hanjani M, Constantin T, Salari R, et al. Phylogenetic ctDNA analysis depicts early stage lung cancer evolution. Nature 2017;545:446–451.
Mok TS, Wu YL, Ahn MJ, Garassino MC, Kim HR, Ramalingam SS, et al. Osimertinib or Platinum-Pemetrexed in EGFR T790M-Positive Lung Cancer. N Engl J Med. 2017;376:629–40.
Brisson GD, Alves LR, Pombo-de-Oliveira MS. Genetic susceptibility in childhood acute leukaemias: a systematic review. Ecancermedicalscience. 2015;9:539.
Ferrara F, Schiffer CA. Acute myeloid leukaemia in adults. Lancet. 2013;381:484–95.
Wang J, Wang H, Wang LY, Cai D, Duan Z, Zhang Y, et al. Silencing the epigenetic silencer KDM4A for TRAIL and DR5 simultaneous induction and antitumor therapy. Cell Death Differ. 2016;23:1886–96.
Losman JA, Looper RE, Koivunen P, Lee S, Schneider RK, McMahon C, et al. (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science. 2013;339:1621–5.
Chen WL, Wang JH, Zhao AH, Xu X, Wang YH, Chen TL, et al. A distinct glucose metabolism signature of acute myeloid leukemia with prognostic value. Blood. 2014;124:1645–54.
Tabe Y, Konopleva M. Advances in understanding the leukaemia microenvironment. Br J Haematol. 2014;164:767–78.
Herst PM, Howman RA, Neeson PJ, Berridge MV, Ritchie DS. The level of glycolytic metabolism in acute myeloid leukemia blasts at diagnosis is prognostic for clinical outcome. J Leukoc Biol. 2011;89:51–55.
Chen WL, Wang YY, Zhao A, **a L, **e G, Su M, et al. Enhanced fructose utilization mediated by SLC2A5 Is a unique metabolic feature of acute myeloid leukemia with therapeutic potential. Cancer Cell. 2016;30:779–91.
Liu H, Huang D, McArthur DL, Boros LG, Nissen N, Heaney AP. Fructose induces transketolase flux to promote pancreatic cancer growth. Cancer Res. 2010;70:6368–76.
Barone S, Fussell SL, Singh AK, Lucas F, Xu J, Kim C, et al. Slc2a5 (Glut5) is essential for the absorption of fructose in the intestine and generation of fructose-induced hypertension. J Biol Chem. 2009;284:5056–66.
Monzavi-Karbassi B, Hine RJ, Stanley JS, Ramani VP, Carcel-Trullols J, Whitehead TL, et al. Fructose as a carbon source induces an aggressive phenotype in MDA-MB-468 breast tumor cells. Int J Oncol. 2010;37:615–22.
Hopkins BD, Goncalves MD, Cantley LC.Obesity and Cancer Mechanisms: Cancer Metabolism. J Clin Oncol: Off J Am Soc Clin Oncol. 2016;34:4277–83.
Hammarsten J, Hogstedt B. Hyperinsulinaemia: a prospective risk factor for lethal clinical prostate cancer. Eur J Cancer. 2005;41:2887–95.
Yoon YS, Keum N, Zhang X, Cho E, Giovannucci EL. Hyperinsulinemia, insulin resistance and colorectal adenomas: A meta-analysis. Metab: Clin Exp. 2015;64:1324–33.
Ferguson RD, Novosyadlyy R, Fierz Y, Alikhani N, Sun H, Yakar S, et al. Hyperinsulinemia enhances c-Myc-mediated mammary tumor development and advances metastatic progression to the lung in a mouse model of type 2 diabetes. Breast Cancer Res: BCR. 2012;14:R8.
Lien EC, Lyssiotis CA, Cantley LC. Metabolic Reprogramming by the PI3K-Akt-mTOR Pathway in Cancer. Recent Results Cancer Res Fortschr der Krebsforsch Progres dans Les Rech sur Le Cancer. 2016;207:39–72.
Huang S, Czech MP. The GLUT4 glucose transporter. Cell Metab. 2007;5:237–52.
Fruman DA, Chiu H, Hopkins BD, Bagrodia S, Cantley LC, Abraham RT. The PI3K Pathway in Human Disease. Cell. 2017;170:605–35.
Kalaany NY, Sabatini DM. Tumours with PI3K activation are resistant to dietary restriction. Nature. 2009;458:725–31.
Matassa DS, Amoroso MR, Lu H, Avolio R, Arzeni D, Procaccini C, et al. Oxidative metabolism drives inflammation-induced platinum resistance in human ovarian cancer. Cell Death Differ. 2016;23:1542–54.
Slager SL, Caporaso NE, de Sanjose S, Goldin LR. Genetic susceptibility to chronic lymphocytic leukemia. Semin Hematol. 2013;50:296–302.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA. 2002;99:15524–9.
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M, et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA. 2005;102:13944–9.
Suzuki HI, Young RA, Sharp PA. Super-enhancer-mediated rna processing revealed by integrative microrna network analysis. Cell. 2017;168:1000–14 e1015.
Kang H, Kim C, Lee H, Rho JG, Seo JW, Nam JW, et al. Downregulation of microRNA-362-3p and microRNA-329 promotes tumor progression in human breast cancer. Cell Death Differ. 2016;23:484–95.
Zhang J, Manley JL. Misregulation of pre-mRNA alternative splicing in cancer. Cancer Discov. 2013;3:1228–37.
Zhang J, Lieu YK, Ali AM, Penson A, Reggio KS, Rabadan R, et al. Disease-associated mutation in SRSF2 misregulates splicing by altering RNA-binding affinities. Proc Natl Acad Sci USA. 2015;112:E4726–4734.
Souers AJ, Leverson JD, Boghaert ER, Ackler SL, Catron ND, Chen J, et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat Med. 2013;19:202–8.
Roberts AW, Davids MS, Pagel JM, Kahl BS, Puvvada SD, Gerecitano JF, et al. Targeting BCL2 with Venetoclax in relapsed chronic lymphocytic leukemia. N Engl J Med. 2016;374:311–22.
Howlett NG, Taniguchi T, Olson S, Cox B, Waisfisz Q, De Die-Smulders C, et al. Biallelic inactivation of BRCA2 in Fanconi anemia. Science. 2002;297:606–9.
Ceccaldi R, Liu JC, Amunugama R, Hajdu I, Primack B, Petalcorin MI, et al. Homologous-recombination-deficient tumours are dependent on Poltheta-mediated repair. Nature. 2015;518:258–62.
Konecny GE, Kristeleit RS. PARP inhibitors for BRCA1/2-mutated and sporadic ovarian cancer: current practice and future directions. Br J Cancer. 2016;115:1157–73.
Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017;357:409–13.
Bai F, Morcos F, Cheng RR, Jiang H, Onuchic JN. Elucidating the druggable interface of protein-protein interactions using fragment docking and coevolutionary analysis. Proc Natl Acad Sci USA. 2016;113:E8051–E8058.
Lipper CH, Karmi O, Sohn YS, Darash-Yahana M, Lammert H, Song L, et al. Structure of the human monomeric NEET protein MiNT and its role in regulating iron and reactive oxygen species in cancer cells. Proc Natl Acad Sci USA. 2018;115:272–7.
Sohn YS, Tamir S, Song L, Michaeli D, Matouk I, Conlan AR, et al. NAF-1 and mitoNEET are central to human breast cancer proliferation by maintaining mitochondrial homeostasis and promoting tumor growth. Proc Natl Acad Sci USA. 2013;110:14676–81.
Bai F, Morcos F, Sohn YS, Darash-Yahana M, Rezende CO, Lipper CH, et al. The Fe-S cluster-containing NEET proteins mitoNEET and NAF-1 as chemotherapeutic targets in breast cancer. Proc Natl Acad Sci USA. 2015;112:3698–703.
Blokzijl F, de Ligt J, Jager M, Sasselli V, Roerink S, Sasaki N, et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature. 2016;538:260–4.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Moresco EM, Li X, Beutler B. Going forward with genetics: recent technological advances and forward genetics in mice. Am J Pathol. 2013;182:1462–73.
Zhu L, Finkelstein D, Gao C, Shi L, Wang Y, Lopez-Terrada D, et al. Multi-organ map** of cancer risk. Cell. 2016;166:1132–46 e1137.
Tomasetti C, Li L, Vogelstein B. Stem cell divisions, somatic mutations, cancer etiology, and cancer prevention. Science. 2017;355:1330–4.
Acknowledgements
The meeting was made possible by a generous donation from the Barry and Virginia Weinman Foundation and by the generous support of the International Association for the Study of Lung Cancer (IASLC). M.C. research is supported by 1R01CA198138-01; DoD Translational Team Award; DoD Idea Award; University of Hawaii Foundation (through the Melohn Endowed Chair in Cancer Biology and Genetics; and through unrestricted donations from Honeywell International and from UNITED-FOR-A-CURE); H.Y research is supported by U01CA214195-01; DoD Translational Team Award; DoD Idea Award; H.I.P. is supported by U01CA214195-01, DoD Translational Team Award; J.B. is supported by P50CA196516, R01CA175754 and RP180192.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
M.C. has pending patent applications on BAP1; H.I.P., has patents pending for use of fibulin-3 for diagnosis of mesothelioma; M.C., H.Y and H.I.P. have patents pending for use of HMGB1 for diagnosis of mesothelioma; M.C. and H.Y. have patents pending for use of HMGB1 and its isoforms for diagnosis of mesothelioma; M.C. provides cost-free consultation for MM expertise and diagnosis to patients and colleagues and paid medical-legal expertise. The remaining authors declare that they have no conflict of interest.
Additional information
Edited by: R.A. Knight.
Rights and permissions
About this article
Cite this article
Carbone, M., Amelio, I., Affar, E.B. et al. Consensus report of the 8 and 9th Weinman Symposia on Gene x Environment Interaction in carcinogenesis: novel opportunities for precision medicine. Cell Death Differ 25, 1885–1904 (2018). https://doi.org/10.1038/s41418-018-0213-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-018-0213-5
- Springer Nature Limited
This article is cited by
-
Effect of regulatory cell death on the occurrence and development of head and neck squamous cell carcinoma
Biomarker Research (2023)
-
Deer antlers: the fastest growing tissue with least cancer occurrence
Cell Death & Differentiation (2023)
-
Gene–environment interactions and their impact on human health
Genes & Immunity (2022)
-
Metabolic rewiring and redox alterations in malignant pleural mesothelioma
British Journal of Cancer (2020)
-
The biochemistry of cell death
Cell Death & Disease (2020)