Abstract
Haematopoietic stem cell transplantation is used clinically to treat many haematological conditions. Isolation of a patient’s own stem cells is a feasible approach for gene therapy and autologous transplantation, which removes the requirement for an immunologically matched donor and the restriction this causes. Manipulating the genome of blood progenitor cells is possible and represents a promising approach to correct genetic and acquired disorders using the patient’s own material. However, the efficiency and safety of gene correction in these stem cells is not yet at the level required for translation into large-scale clinical application. Current approaches, challenges and possible solutions will be discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction: The Haematopoietic System
Haematopoietic stem cells (HSCs) are capable of self-renewal and differentiation into all cell lineages found in the blood, providing the lifelong basis for many vital processes including immunological defence, transport of materials, blood clotting following injury, and oxygenation of the body’s tissues. Under homeostatic conditions, the blood system is required to generate in the region of 1011 neutrophils and red blood cells daily [1]. Although HSCs are predominantly quiescent in vivo, and more rapidly cycling transit amplifying or progenitor cells are responsible for generating this large number of mature blood cells in the short term, HSCs are ultimately responsible for ensuring the long-term production of these progenitor cells and are therefore essential for the lifelong maintenance of haematopoiesis. Under conditions of stress, such as following infection, blood loss or the ablation of mature blood cells via chemotherapy or irradiation, HSCs demonstrate enormous regenerative potential.
This property makes HSCs ideal for treatment of acquired and inherited diseases as they can engraft into a recipient patient’s bone marrow space and repopulate the entire haematopoietic system. HSCs from bone marrow, peripheral blood and cord blood have been developed and applied clinically over the last 60 years, and transplantation is the gold standard treatment for many haematological disorders including lymphoma, leukaemia, thalassemia and anaemias [2]. This treatment is not without drawbacks, which include difficulties associated with finding an HLA-matched donor, graft rejection and the donor cells reacting against the recipient’s tissues (graft versus host disease) [3]. All of these can affect the morbidity and mortality of treated patients.
Correction of Autologous Stem Cells
Since HSCs can be harvested from their bone marrow niche and then re-transplanted back into a new recipient (allogeneic transplant) or the original donor (autologous transplant), they are particularly amenable to ex vivo manipulation using gene therapy, to attenuate, augment or correct mutated disease-causing genes. In an autologous setting, this means that HSCs from a patient can be phenotypically corrected and returned to treat blood diseases, without the need of finding a matched donor or the risk of graft rejection or graft versus host disease. To date, integrating viral vector systems such as attenuated replication-defective lentivirus or gammaretrovirus have been predominantly used to deliver a correcting gene copy into the HSCs of patients with inherited haematologic disorders. These vectors integrate their payload into the genome of target cells, a necessity for longevity of transgene expression in cells that have the potential to self-renew for the lifespan of the patient. The majority of current gene therapy approaches relies on the delivery of a functional copy of a gene’s complementary DNA (cDNA) to express the functioning version of a protein, the lack of which is the cause of disease. This is generally performed ex vivo where patient HSCs are removed from the bone marrow or collected from cells mobilized into the peripheral blood. These cells are then enriched using the CD34 cell surface marker which is prevalent, but not unique, in stem cells and cultured to enable gene delivery using a viral vector. Cells are then returned to the patient by intravenous injection, and the corrected cells home back to the bone marrow, engraft in the correct niche and can produce all mature blood cell lineages expressing the correct protein. While this approach is adequate for treating many disorders, it does have limitations including the size of gene that can be delivered, production of non-physiological levels of RNA from a heterologous promoter and the associated lack of response to relevant transcription factors that control endogenous gene expression. Trials to treat disorders including combined immunodeficiencies, metabolic diseases and globinopathies have shown remarkable benefits in early clinical trials [3–8] where treatment options can be limited if matched HSC donors cannot be found.
The use of integrating vectors is required for long-term therapy, but this approach has additional associated disadvantages. Lack of control over the genomic integration site of the viral vector means that expression in cell populations is heterogeneous and affected by ‘position effects’ where the chromatin state of surrounding DNA can exert control and restrict gene expression. A more serious consequence of integrating cassettes into genomic DNA in an undirected manner is insertional mutagenesis, whereby the foreign vector sequence can unintentionally interfere with genes flanking the integration site, up-regulating or inhibiting them. This can then have severe consequences if oncogenes or tumour suppressor genes are involved. Early gene therapy trials highlighted these pitfalls, where the gene therapy vehicle contributed to initiation of blood cell clonal expansions and leukaemia [9–11]. These serious adverse effects directed increased research efforts to produce safety modifications in the vectors, and the newest generation of viruses are considered less mutagenic [8, 12, 13], but the unknown effects of non-targeted integration remain.
Genome Engineering
One solution to the lack of control caused by random vector integration would be to place therapeutic sequences in a precise location within the genome or alternatively repair an endogenous gene locus. This would enable a mutation to be engineered out of cells and replaced with the correct sequence or the faulty dominant negative version of a gene to be simply deleted. Alternatively, exogenous coding regions could be placed in ‘safe harbours’ in intrageneic regions [14] or inserted upstream of an endogenous promoter to provide equivalent physiological levels of expression [15, 16]. This genome engineering approach would avoid the random insertion of foreign DNA and exogenous controlling elements while allowing for the seamless correction of mutations or the exact placement of other sequences.
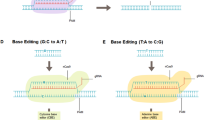
Gene editing can be achieved by using targeting endonucleases, which create double-strand breaks in host cell DNA. Such breaks can be created using an increasing array of tools, including zinc-finger nucleases, homing endonucleases, transcription activator-like effector nucleases (TALENs) and the clustered regularly interspaced short palindromic repeats/CRISPR-associated protein 9 (CRISPR/Cas9) system, which have been reviewed elsewhere [17, 18] and within this edition. This approach creates a genetic lesion which is recognized by the cell and repaired by either the non-homologous end joining (NHEJ) pathway or homologous recombination (HR). NHEJ is error-prone, often introducing deletions, while HR can repair the break specifically using a DNA template which, in the setting of genome engineering, consists of an exogenous synthetic DNA donor molecule. The synthetic donor DNA spans the genomic locus that is to be modified, with the perfect sequence homology between the arms of the synthetic donor and the endogenous locus directing the two DNA molecules to align. The cells’ HR DNA repair machinery then mediates the replacement of the endogenous genomic locus with the DNA template molecule. Normally, this is a very inefficient process and is dependent on the size of homologous region, the cell type targeted and the cell-cycle status of the cell, as HR is up-regulated at progression into mitosis [19–21]. Dramatic improvements in efficiency have been made by creating a double strand break in the target DNA which activates the cellular DNA repair machinery and increases the rate of homologous recombination, but the efficiency of HR even with the use of site-specific nucleases is still limiting for most clinical applications using HSC.
If a double-strand break is engineered and the cell uses NHEJ to repair the lesion, it often leads to a loss of function if the break is within a gene coding region due to the low fidelity of ligating the DNA ends, resulting in varying sizes of genomic insertions or deletions. Creating double-strand breaks without providing any donor sequence can help increase the level of NHEJ and is useful for disrupting a disease-causing allele or removing an entire region of DNA.
Gene Editing in Haematopoietic Stem Cells
Experience of ex vivo manipulation and nucleic acid delivery to HSCs made them an obvious early choice for the application for gene editing. Some haematologic disease backgrounds, such as the immunodeficiency SCID-X1, provide an environment where there is a strong selective advantage for cells that express the wild-type protein. This was observed in a SCID-X1 patient who fortuitously developed a point mutation, resulting in attenuation of the disease-causing mutation in the IL2RG gene, thus allowing the progenitor cells to receive survival, growth and differentiation signals sufficient to reconstitute the patient’s immune system and correct the disease phenotype [22••]. Such a clinical outcome proves that correction of minimal numbers of cells could have a major impact in the treatment of some diseases providing principal efficacy for gene editing in small numbers of primitive HSCs. Building upon this, the IL2RG cDNA has been inserted upstream of the endogenous promoter in CD34+ HSC cell populations, resulting in the expression of the transgene with the correct controlling elements [15], providing an approach for correcting any given mutation within the gene. This paradigm exists in other immunodeficiencies [23] and haematologic disease models where the selective proliferative advantage conferred to corrected cells is elicited at the level of the HSC, such as in the most frequently inherited bone marrow failure syndrome, Fanconi anaemia. In Fanconi anaemia, the phenomenon of reverse mosaicism has been observed for a number of patients, where a single somatic HSC has undergone a spontaneous correcting mutation that has conferred a huge selective advantage on the resulting clone and its progeny [24–26] suggesting that even genome editing at a very low efficiency may have clinical impact.
A second approach was directed by observations in people who are resistant to HIV infection. These individuals harbour a homozygous mutation, delta 32, in the CCR5 gene which encodes a co-receptor for HIV cell entry [27, 28]. The importance of this was emphasized by reports of the HIV-positive ‘Berlin patient’ who received an HSC transplantation for leukaemia. The bone marrow donor had a homozygous delta 32 CCR5 mutation. As donor T cells developed from the engrafted stem cells, the HIV could no longer interact with cells, effectively curing him of HIV as well as treating the leukaemia [29]. This has led to efforts using gene editing to disrupt the CCR5 locus as a treatment for HIV using zinc-finger nuclease technology in T cells [30, 31] or in HSC [32••]. An alternative approach has been to use this location as a ‘safe harbour’ to integrate gene expression cassettes with minimal genotoxic consequences [14] due to the fact that cells lacking CCR5 do not appear to have diminished functionality, at least in in vivo experiments [32••].
Building on these approaches, modifications are now being attempted in HSC to treat other diseases. Haemoglobinopathies are caused by mutations within genes that produce different globin chains, with mutations within β globin resulting in β-thalassaemia or sickle cell anaemia. Hoban et al. were the first group to demonstrate that gene editing could be used to correct the sickle mutation in β globin in clinically relevant patient bone marrow CD34+ cells [33•].
The discovery of further elements that control globin switching has revealed new targets for gene editing to treat β-thalassaemia and sickle cell anaemia. Persistence of fetal haemoglobin in adults can reduce or ameliorate symptoms in these diseases, as the reliance on β-globin to form functional haemoglobin molecules is reduced [34]. Bcl11A represses expression of fetal haemoglobin after birth, representing a novel target for gene therapy [35]. Application of CRISPR-Cas9 libraries has assisted the map** of erythroid-specific enhancer regions that control Bcl11A [36] revealing potential clinical targets for DNA disruption in non-coding sequences. Modification of these enhancers could down-regulate fetal haemoglobin repression and help alleviate symptoms of β-globinopathies. As a proof of concept, bcl11a expression has been disrupted in mouse erythroleukaemia cells using TALENs, increasing the level of fetal haemoglobin [37].
Challenges Facing Gene Editing in HSC
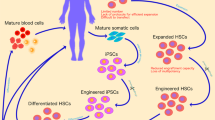
Despite promising experiments and trials, major challenges remain for the widespread clinical application of gene editing in HSCs. For the majority of diseases, the efficiency of gene correction or disruption is insufficient. More primitive, long-term repopulating stem cells are thought to be more difficult to correct than more differentiated cells [15] probably largely due to the fact that HSCs are extremely difficult to transfect or electroporate. In addition, a number of groups have identified that certain DNA repair pathways, such as HR-mediated repair, may be differentially regulated in HSCs compared to their more mature progeny [38, 39]. Such defects in DNA repair may also negatively impact on the efficiency of genome editing in HSCs. An associated problem relates to the risks inherent to the creation of double-strand breaks in genomic DNA. Designer nucleases greatly improve the specificity of cutting sequences, but off-target cleavage of DNA does occur, especially where sequences are similar. For instance, when targeting the β globin locus, cleavage was also observed in the δ-globin gene [33•]. In this example, δ-globin is functionally dispensable but highlights the challenge of targeting specific sequences. Measuring off-target cleavage effects also poses a significant problem, as non-specific cleavage can be rapidly repaired, often resulting in small insertions or deletions that would require clonal genome-wide sequencing to identify [17, 40]. Bioinformatic analysis can be used to reveal similar genomic sequences that have an increased chance of being affected, so they can be checked in detail, but this approach is limiting. Gene editing in polyclonal populations is effective to a degree [15, 33•, 41] but from a safety perspective, being able to correct and expand a thoroughly characterized cell or limited population of clones would be preferable. While this is possible in induced pluripotent stem cells which can be differentiated into some lineages of blood cell [42], the technology is not yet available to recreate all blood lineages from pluripotent stem cells. Equally, expansion of HSC in vitro is not yet possible as the media used to achieve cell cycling also leads to differentiation, reducing the engraftment and self-renewal capacity of the cells [43]. The lack of suitable culture conditions to effectively maintain or even expand HSCs in vitro is one of the most significant barriers to effective gene editing of these cells.
Expansion of Haematopoietic Stem Cells
Much effort has been put into discovering pathways that control HSC self-renewal and the maintenance of the stem cell phenotype as in vitro approaches to induce HSC replication cause differentiation and loss of self-renewal capacity. Such information could be practically applied to the expansion of HSCs from sources where the number of cells is limited, such as umbilical cord blood. However, the development of such technology would also potentially create a paradigm shift in the setting of HSC gene editing, as it would facilitate the clonal expansion of individual HSCs that had been subject to ex vivo manipulation.
A number of experimental strategies have been explored to facilitate the expansion of murine and/or human HSCs in vitro. Perhaps the best characterised is the expansion and enhanced engraftment of murine and human HSCs via the ectopic overexpression of HOXB4. HOXB4 is a member of the homeobox transcription factor family which is involved in developmental cell-fate determination and embryonic patterning during development. Overexpression of HOXB4 can expand murine HSCs in vivo and in vitro [44–46], and it is also able to mediate expansion of human HSCs, although to a much lesser degree than has been shown for mouse. However, constitutive overexpression of HOXB4 has also been associated with some serious deleterious effects, including pronounced dosage-dependent lineage skewing [47], delays to differentiation [48] and, in large animal models, contributing towards leukaemic transformation by blocking differentiation [49]. Although it has been reported that HSC expansion can also be facilitated by the delivery of HOXB4 protein directly to cells in culture [50, 51], this is a much more modest effect than that achieved by retroviral mediated constitutive overexpression. Clearly, for HOXB4 to become clinically relevant, a methodology must be devised for transient or regulated delivery of HOXB4 itself, or activation of its downstream effectors that are important for HSC expansion. In addition to HOXB4, a significant degree of interest has been shown the use of angiopoietin-like proteins for the maintenance and expansion of human and murine HSCs in vitro. Originally, these proteins were implicated in the expansion of HSCs by the laboratory of Harvey Lodish, who recognized that they were a highly expressed component of the cellular secretome from a fetal liver cell population that could facilitate HSC expansion when co-cultured with HSCs [52]. It was subsequently shown that angiopoietin-like proteins (1, 2, 3, 4, 6 and 7) may assist HSC survival and expansion in vivo in media supplemented with additional growth factors [53, 54].
Another promising candidate for facilitating the expansion of human, but not murine, HSCs in vitro is the inhibition of the aryl-hydrocarbon receptor, which has previously been identified as an important mediator of the cellular response to environmental toxins [55]. A novel small molecule antagonist of the aryl-hydrocarbon receptor, StemRegenin (SR1), was identified during a chemical library screen for factors that could improve the frequency of immunophenotypically defined human HSCs in vitro. It has subsequently been shown that SR1 treatment allows functional HSCs to propagate and even expand in culture [55].
Currently, SR1 probably represents the most likely candidate for translation towards clinical expansion of HSCs. However, a number of other potential experimental strategies for HSC expansion have been documented (Table 1), and with further development, these may prove to be even more effective than SR1.
Conclusions
Gene editing in HSCs holds a great deal of promise for the treatment of both inherited and acquired defects within the blood in terms of targeting a cell population that can easily be reintroduced back into patients without the need for complicated surgery and which can subsequently regenerate the whole of the haematopoietic system with modified cells. If the efficiency of gene editing within HSCs can be improved, then it will potentially allow the field to completely overcome toxicities associated with viral vector integration effects that have been documented in several gene therapy clinical trials. Advances in our ability to maintain functional stem cells during in vitro culture/manipulation will be an essential step towards realising the promise of HSC gene editing.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Gordon MY, Lewis JL, Marley SB. Of mice and men…and elephants. Blood. 2002;100(13):4679–80. doi:10.1182/blood-2002-08-2517.
Appelbaum FR. Hematopoietic-cell transplantation at 50. N Engl J Med. 2007;357(15):1472–5. doi:10.1056/NEJMp078166.
Aiuti A, Cassani B, Andolfi G, Mirolo M, Biasco L, Recchia A, et al. Multilineage hematopoietic reconstitution without clonal selection in ADA-SCID patients treated with stem cell gene therapy. J Clin Invest. 2007;117(8):2233–40. doi:10.1172/JCI31666.
Biffi A, Montini E, Lorioli L, Cesani M, Fumagalli F, Plati T, et al. Lentiviral hematopoietic stem cell gene therapy benefits metachromatic leukodystrophy. Science. 2013;341(6148):1233158. doi:10.1126/science.1233158.
Cartier N, Hacein-Bey-Abina S, Bartholomae CC, Veres G, Schmidt M, Kutschera I, et al. Hematopoietic stem cell gene therapy with a lentiviral vector in X-linked adrenoleukodystrophy. Science. 2009;326(5954):818–23. doi:10.1126/science.1171242.
Cavazzana-Calvo M, Payen E, Negre O, Wang G, Hehir K, Fusil F, et al. Transfusion independence and HMGA2 activation after gene therapy of human beta-thalassaemia. Nature. 2010;467(7313):318–22. doi:10.1038/nature09328.
Gaspar HB, Cooray S, Gilmour KC, Parsley KL, Adams S, Howe SJ, et al. Long-term persistence of a polyclonal T cell repertoire after gene therapy for X-linked severe combined immunodeficiency. Sci Transl Med. 2011;3(97):97ra79. doi:10.1126/scitranslmed.3002715.
Hacein-Bey-Abina S, Pai SY, Gaspar HB, Armant M, Berry CC, Blanche S, et al. A modified gamma-retrovirus vector for X-linked severe combined immunodeficiency. N Engl J Med. 2014;371(15):1407–17. doi:10.1056/NEJMoa1404588.
Braun CJ, Boztug K, Paruzynski A, Witzel M, Schwarzer A, Rothe M, et al. Gene therapy for Wiskott-Aldrich syndrome--long-term efficacy and genotoxicity. Sci Transl Med. 2014;6(227):227ra33. doi:10.1126/scitranslmed.3007280.
Hacein-Bey-Abina S, Garrigue A, Wang GP, Soulier J, Lim A, Morillon E, et al. Insertional oncogenesis in 4 patients after retrovirus-mediated gene therapy of SCID-X1. J Clin Invest. 2008;118(9):3132–42. doi:10.1172/JCI35700.
Howe SJ, Mansour MR, Schwarzwaelder K, Bartholomae C, Hubank M, Kempski H, et al. Insertional mutagenesis combined with acquired somatic mutations causes leukemogenesis following gene therapy of SCID-X1 patients. J Clin Invest. 2008;118(9):3143–50. doi:10.1172/JCI35798.
Thornhill SI, Schambach A, Howe SJ, Ulaganathan M, Grassman E, Williams D, et al. Self-inactivating gammaretroviral vectors for gene therapy of X-linked severe combined immunodeficiency. Mol Ther. 2008;16(3):590–8. doi:10.1038/sj.mt.6300393.
Zhou S, Mody D, DeRavin SS, Hauer J, Lu T, Ma Z, et al. A self-inactivating lentiviral vector for SCID-X1 gene therapy that does not activate LMO2 expression in human T cells. Blood. 2010;116(6):900–8. doi:10.1182/blood-2009-10-250209.
Lombardo A, Cesana D, Genovese P, Di Stefano B, Provasi E, Colombo DF, et al. Site-specific integration and tailoring of cassette design for sustainable gene transfer. Nat Methods. 2011;8(10):861–9. doi:10.1038/nmeth.1674.
Genovese P, Schiroli G, Escobar G, Di Tomaso T, Firrito C, Calabria A, et al. Targeted genome editing in human repopulating haematopoietic stem cells. Nature. 2014;510(7504):235–40. doi:10.1038/nature13420.
Lombardo A, Genovese P, Beausejour CM, Colleoni S, Lee YL, Kim KA, et al. Gene editing in human stem cells using zinc finger nucleases and integrase-defective lentiviral vector delivery. Nat Biotechnol. 2007;25(11):1298–306. doi:10.1038/nbt1353.
Cox DB, Platt RJ, Zhang F. Therapeutic genome editing: prospects and challenges. Nat Med. 2015;21(2):121–31. doi:10.1038/nm.3793.
Gaj T, Gersbach CA, Barbas 3rd CF. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013;31(7):397–405. doi:10.1016/j.tibtech.2013.04.004.
Ciccia A, Elledge SJ. The DNA damage response: making it safe to play with knives. Mol Cell. 2010;40(2):179–204. doi:10.1016/j.molcel.2010.09.019.
Hei**k AM, Krajewska M, van Vugt MA. The DNA damage response during mitosis. Mutat Res. 2013;750(1–2):45–55. doi:10.1016/j.mrfmmm.2013.07.003.
Maruyama T, Dougan SK, Truttmann MC, Bilate AM, Ingram JR, Ploegh HL. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat Biotechnol. 2015;33(5):538–42. doi:10.1038/nbt.3190.
Stephan V, Wahn V, Le Deist F, Dirksen U, Broker B, Muller-Fleckenstein I, et al. Atypical X-linked severe combined immunodeficiency due to possible spontaneous reversion of the genetic defect in T cells. N Engl J Med. 1996;335(21):1563–7. doi:10.1056/NEJM199611213352104. This study describes the occurance of a spontaneous correcting mutation with a HSC/progenitor cell in a patient suffering from an inherited form of severe combined imunodeficiency. The corrected clone expanded sufficiently to ameliorate the symptoms of the disease, thereby providing a proof of concept for effective cell therapy from a single gene-edited HSC clone.
Wada T, Candotti F. Somatic mosaicism in primary immune deficiencies. Curr Opin Allergy Clin Immunol. 2008;8(6):510–4. doi:10.1097/ACI.0b013e328314b651.
Gross M, Hanenberg H, Lobitz S, Friedl R, Herterich S, Dietrich R, et al. Reverse mosaicism in Fanconi anemia: natural gene therapy via molecular self-correction. Cytogenet Genome Res. 2002;98(2–3):126–35. 69805.
Mankad A, Taniguchi T, Cox B, Akkari Y, Rathbun RK, Lucas L, et al. Natural gene therapy in monozygotic twins with Fanconi anemia. Blood. 2006;107(8):3084–90. doi:10.1182/blood-2005-07-2638.
Waisfisz Q, Morgan NV, Savino M, de Winter JP, van Berkel CG, Hoatlin ME, et al. Spontaneous functional correction of homozygous fanconi anaemia alleles reveals novel mechanistic basis for reverse mosaicism. Nat Genet. 1999;22(4):379–83. doi:10.1038/11956.
Liu R, Paxton WA, Choe S, Ceradini D, Martin SR, Horuk R, et al. Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell. 1996;86(3):367–77.
Samson M, Libert F, Doranz BJ, Rucker J, Liesnard C, Farber CM, et al. Resistance to HIV-1 infection in Caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature. 1996;382(6593):722–5. doi:10.1038/382722a0.
Hutter G, Nowak D, Mossner M, Ganepola S, Mussig A, Allers K, et al. Long-term control of HIV by CCR5 Delta32/Delta32 stem-cell transplantation. N Engl J Med. 2009;360(7):692–8. doi:10.1056/NEJMoa0802905.
Perez EE, Wang J, Miller JC, Jouvenot Y, Kim KA, Liu O, et al. Establishment of HIV-1 resistance in CD4+ T cells by genome editing using zinc-finger nucleases. Nat Biotechnol. 2008;26(7):808–16. doi:10.1038/nbt1410.
Tebas P, Stein D, Tang WW, Frank I, Wang SQ, Lee G, et al. Gene editing of CCR5 in autologous CD4 T cells of persons infected with HIV. N Engl J Med. 2014;370(10):901–10. doi:10.1056/NEJMoa1300662.
Holt N, Wang J, Kim K, Friedman G, Wang X, Taupin V, et al. Human hematopoietic stem/progenitor cells modified by zinc-finger nucleases targeted to CCR5 control HIV-1 in vivo. Nat Biotechnol. 2010;28(8):839–47. doi:10.1038/nbt.1663. This study describes the approach of using genome editing to disrupt the expression of the CCR5 HIV-1 co-receptor in HSCs so that, following transplantation, gene modified donor derived T cells will be resistant to HIV-1 infection.
Hoban MD, Cost GJ, Mendel MC, Romero Z, Kaufman ML, Joglekar AV, et al. Correction of the sickle cell disease mutation in human hematopoietic stem/progenitor cells. Blood. 2015;125(17):2597–604. doi:10.1182/blood-2014-12-615948. This work demonstrates that the combined delivery of zinc finger nuclease and donor template DNA into HSCs can be used to correct the β-globin locus in cells from sickle cell patients.
Shaeffer JR, Moake JL. Sickle cell—betao thalassemia variant with high hemoglobin F and mild clinical course. Am J Med. 1976;61(3):437–8.
Uda M, Galanello R, Sanna S, Lettre G, Sankaran VG, Chen W, et al. Genome-wide association study shows BCL11A associated with persistent fetal hemoglobin and amelioration of the phenotype of beta-thalassemia. Proc Natl Acad Sci U S A. 2008;105(5):1620–5. doi:10.1073/pnas.0711566105.
Canver MC, Smith EC, Sher F, Pinello L, Sanjana NE, Shalem O, et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature. 2015;527(7577):192–7. doi:10.1038/nature15521.
Bauer DE, Kamran SC, Lessard S, Xu J, Fujiwara Y, Lin C, et al. An erythroid enhancer of BCL11A subject to genetic variation determines fetal hemoglobin level. Science. 2013;342(6155):253–7. doi:10.1126/science.1242088.
Milyavsky M, Gan OI, Trottier M, Komosa M, Tabach O, Notta F, et al. A distinctive DNA damage response in human hematopoietic stem cells reveals an apoptosis-independent role for p53 in self-renewal. Cell Stem Cell. 2010;7(2):186–97. doi:10.1016/j.stem.2010.05.016.
Mohrin M, Bourke E, Alexander D, Warr MR, Barry-Holson K, Le Beau MM, et al. Hematopoietic stem cell quiescence promotes error-prone DNA repair and mutagenesis. Cell Stem Cell. 2010;7(2):174–85. doi:10.1016/j.stem.2010.06.014.
Frock RL, Hu J, Meyers RM, Ho YJ, Kii E, Alt FW. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat Biotechnol. 2015;33(2):179–86. doi:10.1038/nbt.3101.
Heckl D, Kowalczyk MS, Yudovich D, Belizaire R, Puram RV, McConkey ME, et al. Generation of mouse models of myeloid malignancy with combinatorial genetic lesions using CRISPR-Cas9 genome editing. Nat Biotechnol. 2014;32(9):941–6. doi:10.1038/nbt.2951.
Dreyer AK, Hoffmann D, Lachmann N, Ackermann M, Steinemann D, Timm B, et al. TALEN-mediated functional correction of X-linked chronic granulomatous disease in patient-derived induced pluripotent stem cells. Biomaterials. 2015;69:191–200. doi:10.1016/j.biomaterials.2015.07.057.
Ema H, Takano H, Sudo K, Nakauchi H. In vitro self-renewal division of hematopoietic stem cells. J Exp Med. 2000;192(9):1281–8.
Antonchuk J, Sauvageau G, Humphries RK. HOXB4 overexpression mediates very rapid stem cell regeneration and competitive hematopoietic repopulation. Exp Hematol. 2001;29(9):1125–34.
Antonchuk J, Sauvageau G, Humphries RK. HOXB4-induced expansion of adult hematopoietic stem cells ex vivo. Cell. 2002;109(1):39–45.
Sauvageau G, Thorsteinsdottir U, Eaves CJ, Lawrence HJ, Largman C, Lansdorp PM, et al. Overexpression of HOXB4 in hematopoietic cells causes the selective expansion of more primitive populations in vitro and in vivo. Genes Dev. 1995;9(14):1753–65.
Schiedlmeier B, Klump H, Will E, Arman-Kalcek G, Li Z, Wang Z, et al. High-level ectopic HOXB4 expression confers a profound in vivo competitive growth advantage on human cord blood CD34+ cells, but impairs lymphomyeloid differentiation. Blood. 2003;101(5):1759–68. doi:10.1182/blood-2002-03-0767.
Milsom MD, Woolford LB, Margison GP, Humphries RK, Fairbairn LJ. Enhanced in vivo selection of bone marrow cells by retroviral-mediated coexpression of mutant O6-methylguanine-DNA-methyltransferase and HOXB4. Mol Ther. 2004;10(5):862–73. doi:10.1016/j.ymthe.2004.07.019.
Zhang XB, Beard BC, Trobridge GD, Wood BL, Sale GE, Sud R, et al. High incidence of leukemia in large animals after stem cell gene therapy with a HOXB4-expressing retroviral vector. J Clin Invest. 2008;118(4):1502–10. doi:10.1172/JCI34371.
Amsellem S, Pflumio F, Bardinet D, Izac B, Charneau P, Romeo PH, et al. Ex vivo expansion of human hematopoietic stem cells by direct delivery of the HOXB4 homeoprotein. Nat Med. 2003;9(11):1423–7. doi:10.1038/nm953.
Krosl J, Austin P, Beslu N, Kroon E, Humphries RK, Sauvageau G. In vitro expansion of hematopoietic stem cells by recombinant TAT-HOXB4 protein. Nat Med. 2003;9(11):1428–32. doi:10.1038/nm951.
Zhang CC, Kaba M, Ge G, **e K, Tong W, Hug C, et al. Angiopoietin-like proteins stimulate ex vivo expansion of hematopoietic stem cells. Nat Med. 2006;12(2):240–5. doi:10.1038/nm1342.
Akhter S, Rahman MM, Lee HS, Kim HJ, Hong ST. Dynamic roles of angiopoietin-like proteins 1, 2, 3, 4, 6 and 7 in the survival and enhancement of ex vivo expansion of bone-marrow hematopoietic stem cells. Protein Cell. 2013;4(3):220–30. doi:10.1007/s13238-013-2066-5.
Zheng J, Huynh H, Umikawa M, Silvany R, Zhang CC. Angiopoietin-like protein 3 supports the activity of hematopoietic stem cells in the bone marrow niche. Blood. 2011;117(2):470–9. doi:10.1182/blood-2010-06-291716.
Boitano AE, Wang J, Romeo R, Bouchez LC, Parker AE, Sutton SE, et al. Aryl hydrocarbon receptor antagonists promote the expansion of human hematopoietic stem cells. Science. 2010;329(5997):1345–8. doi:10.1126/science.1191536.
Dolnikov A, Xu N, Shen S, Song E, Holmes T, Klamer G, et al. GSK-3beta inhibition promotes early engraftment of ex vivo-expanded haematopoietic stem cells. Cell Prolif. 2014;47(2):113–23. doi:10.1111/cpr.12092.
Kim JA, Kang YJ, Park G, Kim M, Park YO, Kim H, et al. Identification of a stroma-mediated Wnt/beta-catenin signal promoting self-renewal of hematopoietic stem cells in the stem cell niche. Stem Cells. 2009;27(6):1318–29. doi:10.1002/stem.52.
Ko KH, Holmes T, Palladinetti P, Song E, Nordon R, O’Brien TA, et al. GSK-3beta inhibition promotes engraftment of ex vivo-expanded hematopoietic stem cells and modulates gene expression. Stem Cells. 2011;29(1):108–18. doi:10.1002/stem.551.
Delaney C, Heimfeld S, Brashem-Stein C, Voorhies H, Manger RL, Bernstein ID. Notch-mediated expansion of human cord blood progenitor cells capable of rapid myeloid reconstitution. Nat Med. 2010;16(2):232–6. doi:10.1038/nm.2080.
Delaney C, Varnum-Finney B, Aoyama K, Brashem-Stein C, Bernstein ID. Dose-dependent effects of the Notch ligand Delta1 on ex vivo differentiation and in vivo marrow repopulating ability of cord blood cells. Blood. 2005;106(8):2693–9. doi:10.1182/blood-2005-03-1131.
North TE, Goessling W, Walkley CR, Lengerke C, Kopani KR, Lord AM, et al. Prostaglandin E2 regulates vertebrate haematopoietic stem cell homeostasis. Nature. 2007;447(7147):1007–11. doi:10.1038/nature05883.
de Lima M, McMannis J, Gee A, Komanduri K, Couriel D, Andersson BS, et al. Transplantation of ex vivo expanded cord blood cells using the copper chelator tetraethylenepentamine: a phase I/II clinical trial. Bone Marrow Transplant. 2008;41(9):771–8. doi:10.1038/sj.bmt.1705979.
Peled T, Glukhman E, Hasson N, Adi S, Assor H, Yudin D, et al. Chelatable cellular copper modulates differentiation and self-renewal of cord blood-derived hematopoietic progenitor cells. Exp Hematol. 2005;33(10):1092–100. doi:10.1016/j.exphem.2005.06.015.
Peled T, Landau E, Mandel J, Glukhman E, Goudsmid NR, Nagler A, et al. Linear polyamine copper chelator tetraethylenepentamine augments long-term ex vivo expansion of cord blood-derived CD34+ cells and increases their engraftment potential in NOD/SCID mice. Exp Hematol. 2004;32(6):547–55. doi:10.1016/j.exphem.2004.03.002.
Peled T, Landau E, Prus E, Treves AJ, Nagler A, Fibach E. Cellular copper content modulates differentiation and self-renewal in cultures of cord blood-derived CD34+ cells. Br J Haematol. 2002;116(3):655–61.
Aguila JR, Liao W, Yang J, Avila C, Hagag N, Senzel L, et al. SALL4 is a robust stimulator for the expansion of hematopoietic stem cells. Blood. 2011;118(3):576–85. doi:10.1182/blood-2011-01-333641.
Milanovich S, Peterson J, Allred J, Stelloh C, Rajasekaran K, Fisher J, et al. Sall4 overexpression blocks murine hematopoiesis in a dose-dependent manner. Exp Hematol. 2015;43(1):53–64. doi:10.1016/j.exphem.2014.09.004. e1-8.
Yang J, Liao W, Ma Y. Role of SALL4 in hematopoiesis. Curr Opin Hematol. 2012;19(4):287–91. doi:10.1097/MOH.0b013e328353c684.
Himburg HA, Muramoto GG, Daher P, Meadows SK, Russell JL, Doan P, et al. Pleiotrophin regulates the expansion and regeneration of hematopoietic stem cells. Nat Med. 2010;16(4):475–82. doi:10.1038/nm.2119.
Isern J, Martin-Antonio B, Ghazanfari R, Martin AM, Lopez JA, del Toro R, et al. Self-renewing human bone marrow mesenspheres promote hematopoietic stem cell expansion. Cell Rep. 2013;3(5):1714–24. doi:10.1016/j.celrep.2013.03.041.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Steven J. Howe and Michael D. Milsom declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Genome Editing
Rights and permissions
About this article
Cite this article
Howe, S.J., Milsom, M.D. Barriers to Effective Genome Editing of Haematopoietic Stem Cells. Curr Stem Cell Rep 2, 2–8 (2016). https://doi.org/10.1007/s40778-016-0032-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40778-016-0032-x