Abstract
Honey, a multifaceted blend of sugars, amino acids, vitamins, proteins, and minerals, exhibits compositional variability dependent upon the floral source. While previous studies have attempted to categorize honey, the use of glycomic profiles for honey classification remains an unexplored avenue. This investigation seeks to establish a methodology for distinguishing honey types, specifically multi-floral and pine honey, employing mass spectrometry-based glycomic analysis in tandem with machine learning. In this search, seven samples of pine honey and eight samples of multi-floral honey were obtained from diverse regions of Turkey. Subsequently, the proteins within these honey samples were extracted, and glycans were enzymatically released. The released glycans were labeled with 2-aminobenzoic acid (2-AA) and subjected to analysis via matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS). The glycan profiles of pine and multi-floral honey were determined through these analytical procedures, revealing a total of 76 distinct N-glycan structures. Among these, 13 N-glycan profiles consistently established at high levels across experimental replicates and were incorporated in subsequent analyses. Following the quantification of individual glycan abundances, statistically significant differences in glycan profiles were determined. Notably, N-glycans Hex5HexNAc2, Hex4HexNAc3, and Hex5HexNAc3 displayed considerable differences. Using the 13 N-glycan profiles, an accuracy rate of 93.5% was obtained from machine learning analysis, which increased to 100% when incorporating the identified significantly changed glycans. The most productive models were identified as “subspace and fine k-nearest neighbors (KNN).” The findings underscore the potential of mass spectrometry-based glycomics in conjunction with machine learning as a robust tool for precise honey type classification and its prospective utility in quality control and honey product authentication.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Honey, a natural sweetener formed by bees from the nectar of various types of flowers, has a composition that varies depending on factors such as botanical and geographical origin, as well as processing and storage conditions [1, 2]. Therefore, it is essential to develop reliable analytical methods for evaluating the quality and authenticity of honey. The components found in honey, including carbohydrates, amino acids, vitamins, and minerals, are crucial factors that determine its nutritional and therapeutic properties [2]. Honey sourced from different types of flowers possesses unique chemical constituents, including antimicrobial, anti-inflammatory, and antioxidant properties [2,3,4,5,6,7,8]. However, concerns have arisen about the quality and authenticity of honey due to common issues in the market, such as impurity and mislabeling [9].
In honey authentication research, conventional approaches are used to determine its botanical origin. Sensory and physicochemical assessments are employed to learn the provenance of monofloral honey [10]. At the same time, the customary practice involves the utilization of melissopalynological analysis for the microscopic scrutiny and identification of floral pollen grains within honey [11]. Nevertheless, it should be noted that melissopalynological methodologies may not be as pertinent when applied to other honey varieties due to the inherent variability in pollen content, which typically exhibits lower levels [12]. In addition, the melissopalynological analysis is an intricate process that requires a limited number of highly trained experts, possibly owing to the rigorous training needed for this specialized field [13].
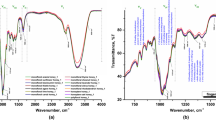
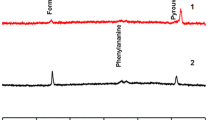
The limitations of conventional methods for verifying the botanical and geographical origins of honey emphasize the necessity for more dependable and contemporary analytical approaches. Cutting-edge analytical instruments and sensor arrays, including chromatography [14], mass spectrometry (MS) [ In this study, samples were obtained from two different honey sources (pine honey and multi-floral honey) from various regions. Initially, honey proteins were extracted from the samples, and then glycan release was performed, followed by labeling with 2-AA. Subsequently, purification of 2-AA labeled glycans was carried out. Glycans were analyzed using MALDI-MS. The peak areas for the identified glycans were extracted, and the relative abundance of each glycan was calculated. Finally, statistical and machine learning analyses were conducted. As a result of the analyses, a total of 76 different N-glycan compositions were identified (Table 1). In Fig. 1, MALDI mass spectra for multi-floral and pine kinds of honey are presented. Out of these, 13 N-glycan profiles that were consistently present in experimental replicates were utilized in the statistical analyses. After determining the relative abundance of each glycan, statistically significant variations in glycan profiles were identified. A classical t-test was applied to detect statistically significant changes in glycan abundance between the two honey types. Significantly regulated N-glycans that contributed to the differentiation between pine and multi-floral honey are shown in Fig. 2. The volcano plot seen in Fig. 2A reveals that the glycans Hex5HexNAc3, Hex4HexNAc3, and Hex5HexNAc2 exhibited statistically significant differences between multi-floral and pine honeys. These glycans displayed notable differences in abundance and were found to be significantly higher in one honey type compared to the other. These specific glycans emerge as key discriminators facilitating the differentiation between pine and multi-floral honey. Specifically, the glycan Hex5HexNAc3 exhibited substantial up-regulation in multi-floral honey compared to pine honey. Additionally, Hex5HexNAc2 demonstrated the most pronounced variation between multi-floral and pine honey types. The figure elucidates the graphical representations employed in the comparative analysis of N-glycans between pine honey and multi-floral honey. Firstly, A encompasses a volcano plot, visually depicting the distinctively regulated N-glycans between the two honey types. And, B displays a diagram derived from principal component analysis, providing a visual representation of the multivariate variations among the N-glycan profiles of the honey varieties (Color figure online) The principal component analysis in Fig. 2B demonstrates the ability to distinguish between pine and multi-floral honey using the significant change of three N-glycans in the honey samples. Red dots represent glycans from pine honey samples, while blue dots represent multi-floral honey. In the two-dimensional analysis, component 1 explains 85.4% of the variance, and component 2 explains 14.1% of it. This analysis indicates a high differentiation between pine and multi-floral honeys. The differentiation between pine and multi-floral kinds of honey can be attributed to significant quantitative differences in N-glycan profiles for significantly changed profiles. The clustering of the samples observed in the chart demonstrates that the glycan compositions of the two honey types are pretty different. Component 1, capturing the most significant variance in the data, likely reflects the fundamental differences in glycan abundance between the two honey types. Similarly, Component 2 highlights additional differences contributing to the differentiation between pine and multi-floral honey. In the machine learning analysis, the relative areas of the glycans were determined based on the mass spectrometric data. Subsequently, machine learning models were developed using MATLAB, employing a 5% cross-validation. The performance of all available models within the MATLAB classification application was comprehensively assessed. To evaluate the machine learning models’ effectiveness, various criteria were considered, including accuracy and the calculation of the area under the curve (AUC). Figure 3A in the study provides a detailed insight into the machine learning classification results based on 13 N-glycans obtained from pine and multi-floral glycoproteins. The findings from this figure indicate that the true positive score for predicting pine honey is an impressive 100%, signifying that all pine honey samples were accurately classified. On the other hand, there is a false positive rate of 12.5% in the classification of multi-floral honey. This indicates that while the model excels at classifying pine honey, there is room for improvement in distinguishing multi-floral honey samples. Overall, the classification accuracy of the model stands at 93.5%. Figure 3B presents the ROC curve analysis for this model, displaying the area under the curve (AUC). The obtained AUC value is 0.93, indicating a high level of classification performance (Fig. 3B). The fact that the ROC curve predominantly lies above the diagonal line signifies the model’s success in distinguishing between pine and multi-floral honey. The high AUC value further substantiates the effectiveness of accurately classifying samples based on the selected glycans. The confusion matrix presented herein pertains to a machine-learning model employed in the discriminatory classification of pine honey versus multi-floral honey. The model’s training involved two distinct feature sets: A comprised 13 N-glycans, and C restricted to three N-glycans demonstrating significant alterations. Additionally, the receiver operating characteristic (ROC) curves, delineating the models’ performance characteristics, are illustrated in (B) for the 13 N-glycans dataset and D specifically for the subset of three significantly changed N-glycans In Fig. 3C, a confusion matrix is presented for the machine learning classification results obtained using three statistically significant glycans (Hex5HexNAc3, Hex4HexNAc3, and Hex5HexNAc2). Remarkably, in this scenario, all pine honey and multi-floral honey samples were perfectly and precisely distinguished from one another, resulting in a 100% accuracy. This remarkable achievement in accurately discerning the two honey types underlines the discriminative capability of the three selected glycans. Figure 3D showcases the ROC curve analysis for the classification model employing these three statistically significant glycans. The ROC curve consistently remains positioned above the diagonal line, with an AUC value of 1.00 reported. This AUC value signifies an exceptionally high level of classification performance. This analysis unequivocally demonstrates the algorithm and model’s exceptional effectiveness in the precise classification of samples based on the selected glycans. The literature extensively covers the proteomic characterization of various honey types. A recent study thoroughly examined the protein composition of 13 honeys, identifying a total of 130 proteins [25]. Utilizing a proteomic approach, honey from distinct geographical and botanical sources was clearly differentiated [26]. It has been understood that the proteomic profiles of honey types differ among honey types. However, there is currently a lack of literature on the N-glycans of honey glycoproteins. This study is the pioneer in utilizing MALDI-MS-based N-glycomics and machine learning for the classification of honey types. The variation in glycosylation machinery across organisms is known. Unlike proteomes, there is no genetic template for the glycosylation machinery within cells [27]. It is emphasized that multiple genes play a role in the formation of glycans attached to proteins, rendering glycans highly intricate and site-, tissue- and species-specific [28]. In contrast to comprehensive proteomics experiments, N-glycan profiling can be accomplished through MALDI-MS. In this method, N-glycans extracted from pine and multi-floral honey were swiftly profiled and utilized for classification purposes. The categorization of different types of honey is crucial to protect consumers from deceptive practices [29]. The existing literature has explored various methodologies for classifying different kinds of honey, with a predominant focus on categorizing them according to botanical and regional origins. The primary objective is to establish a bioanalytical method that exhibits high accuracy and specificity in discerning between various honey types. Mass spectrometry stands out as a highly suitable approach owing to its rapidity, accuracy, and precision. An illustrative instance is the utilization of gas chromatography-mass spectrometry (GC–MS) analysis to assess volatile organic compounds in honey, serving as an effective means for classification [30]. Applying this approach alongside Kohonen’s self-organizing map has enabled the discrimination between Turkish pine honey and Greek pine honey [31]. In conclusion, a bioanalytical technique integrating MALDI-MS-based N-glycomics and machine learning was created for the classification of Turkish pine honey in comparison to multi-floral honey. This method demonstrated high accuracy and specificity in distinguishing pine honey from multi-floral honey. Utilizing statistically significant glycans for discrimination resulted in a 100% accuracy rate. This approach enables swift differentiation between Turkish pine honey and multi-floral honey. The application of this approach could extend to distinguishing between pine and multi-floral honey originating from diverse botanical sources. Additionally, this method has the potential to differentiate between various types of multi-floral honey and mono-flower honey. This suggests that MALDI-MS-based glycomics coupled with machine learning hold promise as a reliable candidate for identifying honey-type adulteration and mitigating variations.Results and discussion
Conclusion
Data Availability
Data will be made available on request.
References
P.M. da Silva, C. Gauche, L.V. Gonzaga, A.C.O. Costa, R. Fett, Honey: chemical composition, stability and authenticity. Food Chem. 196, 309–323 (2016). https://doi.org/10.1016/j.foodchem.2015.09.051
T. Eteraf-Oskouei, M. Najafi, Traditional and modern uses of natural honey in human diseases: a review. Iran. J. Basic Med. Sci. 16(6), 731–742 (2013)
V.R. Pasupuleti, L. Sammugam, N. Ramesh, S.H. Gan, Honey, propolis, and royal jelly: a comprehensive review of their biological actions and health benefits. Oxid. Med. Cell. Longev. (2017). https://doi.org/10.1155/2017/1259510
S. Samarghandian, T. Farkhondeh, F. Samini, Honey and health: a review of recent clinical research. Pharmacogn. Res. 9(2), 121–127 (2017). https://doi.org/10.4103/0974-8490.204647
J.M. Alvarez-Suarez, S. Tulipani, S. Romandini, E. Bertoli, M. Battino, Contribution of honey in nutrition and human health: a review. Mediterr. J. Nutr. Metab. 3(1), 15–23 (2010). https://doi.org/10.1007/s12349-009-0051-6
S. Bogdanov, T. Jurendic, R. Sieber, P. Gallmann, Honey for nutrition and health: a review. J. Am. Coll. Nutr. 27(6), 677–689 (2008). https://doi.org/10.1080/07315724.2008.10719745
M. Küçük, S. Kolayli, S. Karaoglu, E. Ulusoy, C. Baltaci, F. Candan, Biological activities and chemical composition of three honeys of different types from Anatolia. Food Chem. 100(2), 526–534 (2007). https://doi.org/10.1016/j.foodchem.2005.10.010
G. Beretta, P. Granata, M. Ferrero, M. Orioli, R.M. Facino, Standardization of antioxidant properties of honey by a combination of spectrophotometric/fluorimetric assays and chemometrics. Anal. Chim. Acta 533(2), 185–191 (2005). https://doi.org/10.1016/j.aca.2004.11.010
R. Fakhlaei, J. Selamat, A. Khatib, A.F.A. Razis, R. Sukor, S. Ahmad, A.A. Babadi, The toxic impact of honey adulteration: a review. Foods (2020). https://doi.org/10.3390/foods9111538
J.M.B. de Sousa, E.L. de Souza, G. Marques, M. de Toledo Benassi, B. Gullón, M.M. Pintado, M. Magnani, Sugar profile, physicochemical and sensory aspects of monofloral honeys produced by different stingless bee species in Brazilian semi-arid region. LWT 65, 645–651 (2016). https://doi.org/10.1016/j.lwt.2015.08.058
M.A. Rodopoulou, C. Tananaki, M. Dimou, V. Liolios, D. Kanelis, G. Goras, A. Thrasyvoulou, The determination of the botanical origin in honeys with over-represented pollen: combination of melissopalynological, sensory and physicochemical analysis. J. Sci. Food Agric. 98(7), 2705–2712 (2018). https://doi.org/10.1002/jsfa.8764
P. Molan, The limitations of the methods of identifying the floral source of honeys. Bee World 79(2), 59–68 (1998). https://doi.org/10.1080/0005772x.1998.11099381
I. Escriche, M. Juan-Borrás, M. Visquert, J.M. Valiente, An overview of the challenges when analysing pollen for monofloral honey classification. Food Control 143, 109305 (2023). https://doi.org/10.1016/j.foodcont.2022.109305
E. Guzelmeric, I. Ciftci, P.I. Yuksel, E. Yesilada, Importance of chromatographic and spectrophotometric methods in determining authenticity, classification and bioactivity of honey. LWT 132, 109921 (2020). https://doi.org/10.1016/j.lwt.2020.109921
H.E. Tahir, Z. **aobo, H. **aowei, S. Jiyong, A.A. Mariod, Discrimination of honeys using colorimetric sensor arrays, sensory analysis and gas chromatography techniques. Food Chem. 206, 37–43 (2016). https://doi.org/10.1016/j.foodchem.2016.03.032
A. Noviyanto, W.H. Abdulla, Honey botanical origin classification using hyperspectral imaging and machine learning. J. Food Eng. 265, 109684 (2020). https://doi.org/10.1016/j.jfoodeng.2019.109684
D.A. Magdas, F. Guyon, C. Berghian-Grosan, M.C. Muller, Challenges and a step forward in honey classification based on Raman spectroscopy. Food Control 123, 107769 (2021). https://doi.org/10.1016/j.foodcont.2020.107769
K. Rachineni, V.M. Rao Kakita, N.P. Awasthi, V.S. Shirke, R.V. Hosur, S.S. Chandra, Identifying type of sugar adulterants in honey: combined application of NMR spectroscopy and supervised machine learning classification. Curr. Res. Food Sci. 5, 272–277 (2022). https://doi.org/10.1016/j.crfs.2022.01.008
S.P. Kek, N.L. Chin, S.W. Tan, Y.A. Yusof, L.S. Chua, Classification of honey from its bee origin via chemical profiles and mineral content. Food Anal. Methods 10(1), 19–30 (2017). https://doi.org/10.1007/s12161-016-0544-0
H.M. Kayili, M. Atakay, A. Hayatu, B. Salih, Sample preparation methods for N-glycomics. Adv. Sample Prep. 4, 100042 (2022). https://doi.org/10.1016/j.sampre.2022.100042
H.M. Kayili, R. Sakhta, B. Salih, Comparison of denaturing agent effects in enzymatic N-glycan release for human plasma N-glycan analysis. Turk. J. Chem. 46(5), 1524 (2022). https://doi.org/10.55730/1300-0527.3457
D.B. Demirhan, H. Yılmaz, H. Erol, H.M. Kayili, B. Salih, Prediction of gastric cancer by machine learning integrated with mass spectrometry-based N-glycomics. Analyst 148(9), 2073–2080 (2023). https://doi.org/10.1039/D2AN02057B
H.M. Kayili, B. Salih, Site-specific N-glycosylation analysis of human thyroid thyroglobulin by mass spectrometry-based glyco-analytical strategies. J. Proteomics (2022). https://doi.org/10.1016/j.jprot.2022.104700
P.S. Reel, S. Reel, E. Pearson, E. Trucco, E. Jefferson, Using machine learning approaches for multi-omics data analysis: a review. Biotechnol. Adv. 49, 107739 (2021). https://doi.org/10.1016/j.biotechadv.2021.107739
T. Erban, E. Shcherbachenko, P. Talacko, K. Harant, The unique protein composition of honey revealed by comprehensive proteomic analysis: allergens, venom-like proteins, antibacterial properties, royal jelly proteins, serine proteases, and their inhibitors. J. Nat. Prod. 82(5), 1217–1226 (2019). https://doi.org/10.1021/acs.jnatprod.8b00968
Y.-F. Zheng, M.-C. Wu, H.-J. Chien, W.-C. Wang, C.-Y. Kuo, C.-C. Lai, Honey proteomic signatures for the identification of honey adulterated with syrup, producing country, and nectar source using SWATH-MS approach. Food Chem. 354, 129590 (2021). https://doi.org/10.1016/j.foodchem.2021.129590
H.M. Kayili, M. Atakay, A. Hayatu, B. Salih, Sample preparation methods for N-glycomics. Adva. Sample Prep. 4, 100042 (2022)
C.M. West, D. Malzl, A. Hykollari, I.B.H. Wilson, Glycomics, glycoproteomics, and glycogenomics: an inter-taxa evolutionary perspective. Mol. Cell. Proteomics 20, 100024 (2021). https://doi.org/10.1074/mcp.R120.002263
A. Noviyanto, W.H. Abdulla, Honey botanical origin classification using hyperspectral imaging and machine learning. J. Food Eng. (2020). https://doi.org/10.1016/j.jfoodeng.2019.109684
A.C. Soria, I. Martínez-Castro, J. Sanz, Study of the precision in the purge-and-trap–gas chromatography–mass spectrometry analysis of volatile compounds in honey. J. Chromatogr. A 1216(15), 3300–3304 (2009). https://doi.org/10.1016/j.chroma.2009.01.065
C. Tananaki, A. Thrasyvoulou, J.L. Giraudel, M. Montury, Determination of volatile characteristics of Greek and Turkish pine honey samples and their classification by using Kohonen self organising maps. Food Chem. 101(4), 1687–1693 (2007). https://doi.org/10.1016/j.foodchem.2006.04.042
Acknowledgements
Partial funding for the research was received from the Ministry of Development-Republic of Türkiye, with Project Number 2016 K121230. Bekir Salih extends his gratitude to the Turkish Academy of Science (TUBA) for their financial support.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Masri, S., Aksoy, S., Duman, H. et al. Distinguishing Turkish pine honey from multi-floral honey through MALDI-MS-based N-glycomics and machine learning. Food Measure (2024). https://doi.org/10.1007/s11694-024-02597-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11694-024-02597-5