Abstract
Background
Root hair, a special type of tubular-shaped cell, outgrows from root epidermal cell and plays important roles in the acquisition of nutrients and water, as well as interactions with biotic and abiotic stress. Although many genes involved in root hair development have been identified, genetic basis of natural variation in root hair growth has never been explored.
Results
Here, we utilized a maize association panel including 281 inbred lines with tropical, subtropical, and temperate origins to decipher the phenotypic diversity and genetic basis of root hair length. We demonstrated significant associations of root hair length with many metabolic pathways and other agronomic traits. Combining root hair phenotypes with 1.25 million single nucleotide polymorphisms (SNPs) via genome-wide association study (GWAS) revealed several candidate genes implicated in cellular signaling, polar growth, disease resistance and various metabolic pathways.
Conclusions
These results illustrate the genetic basis of root hair length in maize, offering a list of candidate genes predictably contributing to root hair growth, which are invaluable resource for the future functional investigation.
Similar content being viewed by others
Background
Root hairs are special tubular-shaped outgrowth from root epidermal cells, which vastly enlarge the root surface area and assist in water and nutrient absorption such as NO3−, Cl−, Ca2+, K+, Zn2+, and Mn2+, as well as the interactions with biotic and abiotic stresses [1, 2]. On the other hand, the length, density and morphology of root hair are influenced by various endogenous and environmental factors, including phytohormones and mineral nutrients, especially under phosphate (Pi) limiting condition [3,4,5,6,7]. The development of root hair can be separated into three basic stages: specification of the epidermal cell fate, initiation of the root hair outgrowth, and elongation of the hair via tip growth [2, 8].
As a unique single cell type in plant biology, the development, physiology, and cell biology of root hair have been intensively studied in Arabidopsis [2, 6, 9,27,28,29,30]. In maize, LD decay is rapid due to its extensive genetic diversity. Therefore, maize is recognized as an ideal model plant for conducting association studies [31,32,33,34]. To date, GWAS has successfully used to identify numerous candidate loci/genes controlling sever-al morphological or metabolic traits in maize, such as shoot apical meristem size, husk trait, plant height, kernel weight, drought tolerance, grain drying rate and grain moisture [35,36,37,38,39,40,41]. In this study, we used a maize association panel including 281 inbred lines with tropical, subtropical, and temperate backgrounds to interpret the phenotypic diversity and the genetic basis of root hair development. Several candidate genes putatively involved in root hair development were identified, providing a useful resource for further functional studies to elucidate molecular pathways involved in maize root hair growth and development.
Results
Phenotypic variation among root hair length
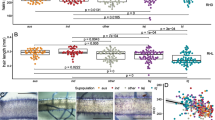
The association population in this study consists of a global collection of 281 diverse maize inbred lines [42, 43]. The root hair length of primary roots was measured from 3-day-old plants (Table S1). The noticeable variation in root hair length was represented by B73 and Mo17, two inbred lines commonly used in maize biology (Fig. 1a). The measured root hair trait followed a normal distribution with a slight right skew (Fig. 1b). The trait maximum, minimum, mean, standard deviation and coefficient of variation were listed in Table 1. The ratio of root hair length of inbred lines to B73 control ranged from 0.2 to 1.42 with the mean of 0.95, indicating that the root hair length exhibits broad variations in the association population.
Phenotypic variation of root hair length. a Diagram of B73 (left) and Mo17 (right) root hair length. Bars = 1 mm. b Frequency distributions of root hair length. Classes of trait values are shown on X axis and counts of inbred lines with the phenotypic values for these bins are shown on Y axis. c Boxplot of root hair length distribution in different subpopulations. Kruskal-Wallis test was applied to examine the difference of traits among subpopulations. Different letters indicate significance levels at P ≤ 0.05. No. of inbred lines included in each subpopulation are 117, 30, 22 and 112 for MIXED, NSS, SS and TST, respectively
All 281 lines used in this study include three subpopulations and one mixed group, which are referred to TST, SS, NSS, and MIXED [42]. TST subpopulation is of tropical or subtropical origin, consisting of 112 lines. SS and NSS subpopulations are of temperate origin, consisting of 22 and 30 lines, respectively. MIXED subpopulation is inbred lines which were not accurately assigned into the above three subpopulations based on the phylogenic analysis [42, 43]. To investigate the effect of population structure on root hair phenotypes, the root hair length was compared between different subpopulations. Compared with SS, NSS and MIXED subpopulations, the mean value of TST subpopulation was significantly less, suggesting that maize inbred lines from tropical/subtropical origin tend to have shorter root hairs (Fig. 1c).
Associations of root hair phenotype with agronomic traits and metabolic pathways
As root hairs play a crucial role in the plant acquisition of nutrients and water, we postulated that root hair morphology is likely coordinated with other agronomic traits and amino acid metabolism. To verify our hypothesis, the Pearson-correlations were calculated after comparing root hair length with 17 agronomic traits and 18 amino acid contents in maize kernel, which were previously measured in the same association panel [44, 45]. The 17 agronomic traits include seven morphological traits, i.e. plant height (PH), ear height (EH), ear leaf width (ELW), ear leaf length (ELL), tassel maximum axis length (TMAL), tassel branch number (TBN), leaf number above ear (LNAE); seven yield-related traits, i.e. ear length (EL), ear diameter (ED), cob diameter (CD), kernel number per row (KNPR), cob grain weight (GW), cob weight (CW), kernel width (KW); three flowering-related traits, i.e. days to anthesis (DTA), days to silking (DTS) and days to heading (DTH) [44]. The amino acids measured in dry maize kernel include Ala, Arg, Asx, Glx, Gly, Lle, Leu, Lys, Met, Pro, Phe, Val, Tyr, His, Cys, Thr and Ser, and the total concentration of amino acids [45].
Of the 17 agronomic traits examined, 7 were correlated with root hair length (P ≤ 0.05), which were EH, ELW, TBN, LNAE, and all three flowering-related traits (Fig. 2a). Meanwhile, of the 18 amino acids, 3 were correlated with root hair length (P ≤ 0.05), which were Gly, Lys and Arg (Fig. 2b). The Pathway Association Study Tool (PAST) was further performed to elucidate the biochemical pathways likely contributing to root hair elongation [46,47,48]. Under the statistical threshold P < 0.01, a total of 8 and 1 metabolic pathways exhibited significant associations with increased or decreased root hair length, respectively (Table S2 and S3). Overall, the close correlations of root hair length with other agronomic or metabolic traits above-mentioned suggest that the root hair occurrence is correlated to other aspects of plant development and growth.
Correlation coefficients of root hair length with other agronomic and amino acidmetabolic traits. a Correlation coefficients between root hair length with 17 agronomic traits. b Correlation coefficients between root hair length with 18 metabolic traits. *Significant at P ≤ 0.05; **significant at P ≤ 0.01. The correlation level is color-coded according to the color key plotted in the middle
GWAS on maize root hair length
Using 1,253,814 SNPs covering the whole maize genome with a minor allele frequency (MAF) ≥ 0.05, we performed GWAS to explore the genetic loci underlying the root hair length. Under the mixed linear model (MLM) [49, 50], which accounts for false positives arising from the population structure (Q matrix) and kinship relationship (K matrix) of the natural variation in the population, a stringent threshold of –log10P ≥ 5.2 was designated as the threshold for calling significantly associated SNPs. The Manhattan plots for the SNPs associated with root hair length were shown in Fig. 3. In total, 18 significant SNPs were detected, which are located on chromosomes 1, 2, 4, 5, 6, and 10, explaining the phenotypic variations ranged from 6.3 to 10.1%. Moreover, GWAS was also performed using the general linear model (GLM). Under the stringent threshold of –log10P ≥ 6.5, a total of 14 SNPs were identified, 12 out of which overlapped with MLM-derived SNPs (Fig. S1). As there were 8 MLM-derived SNPs on chromosome 6 present within the same LD region (r2 ≥ 0.2), and the leading SNP_ 75630277 was selected as the representative of this locus (Table 2), yielding a total of 11 SNPs used for the further analysis.
GWAS showing significant P-values associated with root hair length based on the mixed linear model (MLM). a Manhattan plots of MLM for root hair length. The horizontal dashed line represents the significance threshold -log10(P) = 5.2. b QQ plot of MLM showing the ratio of the observed and the expected P-values for root hair length. The solid diagonal lines represent agreement between observed and expected probability distributions assuming null SNP–trait association
Genes co-localized with significant SNPs
Protein-coding genes harboring or nearest to the significant SNPs were nominated as the candidates associated with root hair (Table 2). The allelic effect of haplotype coordinated with significant SNPs on root hair phenotypes was assessed (Fig. 4 and Figure S2). The most significant SNP (chr6.S_ 34221678, P-value = 1.70E-07, R2 = 9.0%) locates in the fourth exon of GRMZM2G403003 (Fig. 4a), encoding a TON1 recruiting motif (TRM) protein. The average root hair length for A allele was substantially shorter than T allele (P ≤ 0.01, Fig. 4d). The second significant SNP (chr5.S_198197360, P-value = 1.80E-07, R2 = 9.8%) locates in the third intron of GRMZM2G044851 (Fig. 4b), encoding the nitrate transporter 1.5. The average root hair length for allele with A was greatly longer than allele with C (P ≤ 0.01, Fig. 4e). The third significant SNP (chr6.S_ 75630277, P-value = 3.97E-07, R2 = 9.7%), which is the one with other 7 SNPs within the same LD region, locates at the promoter region of GRMZM5G825276 (Fig. 4c), encoding a GDSL-like lipase. The average root hair length for allele with A was greatly longer than allele with G (P ≤ 0.01, Fig. 4f). It is noted that the haplotype analysis for the other eight genes was depicted in Fig. S2.
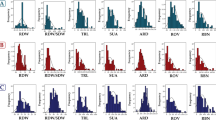
The allele effects of significant SNPs located around representative genes for root hair length. a–c Regional plots showing association map** results for SNPs located around GRMZM2G403003 (a), GRMZM2G044851 (b), GRMZM5G0825276 (c). d–f Allele effects of the most significant SNPs for root hair traits. *Significant at P ≤ 0.05; **significant at P ≤ 0.01. (d) GRMZM2G403003, (e) GRMZM2G044851, (f) GRMZM2G825276. Each dot represents an SNP. The horizontal dashed black line represents the significant threshold –log10(P) = 5.2
Discussion
Root hairs enlarge the root surface area and thereby play vital roles in plant absorbing water and nutrient, as well as co** with biotic and abiotic stress. In recent years, extensive mutation-based studies have been carried out to dissect the genetic regulatory network determining root hair development, but we lack fundamental knowledge about whether and how root hair is controlled in a plant natural variation population. In this work, we interpreted the natural variation and the associated genetic architecture of root hair length in a maize association panel, and a set of putative candidate genes controlling root hair development were revealed by performing GWAS analysis.
Phenotypic variability and coordination of root hair length with other processes of plant development and growth
Maize was domesticated from its wild ancestor, teosinte (Zea mays ssp. parviglumis) about 8000–9000 years ago. Nowadays, modern maize displays a large geographic distribution from tropical to temperate climate. Therefore, population structure is associated with significant differences in maize morphology [51]. In this study, we observed that root hair exhibits wide length variation with normal distribution. Interestingly, tropical lines displayed shorter root hairs relative to temperate lines, implying that the adaption of maize from tropical to temperate regions was probably accompanied by the character of lessening root hair growth.
In our investigation, the root hair at the length dimension was negatively rather than positively correlated with several morphological and metabolic traits. This is somewhat counter-intuitive. However, it is known that the maize plant is a monocot and has a fibrous root system consisting of massive adventitious and lateral roots, the anatomy of which is markedly different from dicotyledonous model Arabidopsis [52,53,54]. In this context, it is possible that the other aspects of root system may enable maize to have adequate larger surface area to support plant growth under normal soil condition. Meanwhile, it is worthy to be mention that to facilitate high throughput phenoty**, the root hair from the emerging primary roots after 3-days post germination was assessed in this study, thereby whether the growth rate will alter distinctly in different inbred lines is uncertain at this point.
Putative genes involved in root hair morphogenesis
We identified a total of 11 candidate genes associated with root hair elongation, and the homologs of three genes have been reported to influence root hair development in Arabidopsis. GRMZM2G100288 encodes a receptor-like protein kinase, and its closest homologs in Arabidopsis is FERONIA (FER), which is well known as a key hub of cell signaling networks mediating various hormone, stress, and immune responses [55,56,57,58,59,60,61,62,63,http://maizego.org/. The maize lines in this association panel were clustered into three subpopulations, including 22 SS lines, 30 NSS lines and 112 TST lines, and the remaining 117 lines fall into a mixed subpopulation. Detailed information about the origins of these lines was described in previous studies [42, 43]. The 1.25 million high-quality SNPs (MAF > 5%) and the estimated population structure and kinship were assessed using the method previously described [99, 100]. The 165,248 markers in approximate linkage equilibrium with each other were found by PLINK (window size 50, step size 50, r2 ≥ 0.2) [101]. Then, we used the uniform Bonferroni-corrected thresholds at α = 1 for MLM and α = 0.05 for GLM as the significance cutoffs as reported in the previous studies [37, 44, 102]. Finally, the suggestive P value was computed by 1/n and 0.05/n (n = 165,248), and we obtained the suggestive threshold 6.05 × 10− 6 for MLM and 3.03 × 10− 7 for GLM, respectively.
To estimate the phenotypic variance explained by each significant SNP, we used ANOVA to construct linear models of Y = αX + βP + ε (1) and Y = βP + ε (2). In this model, Y is the phenotype, X is the SNP genotype, P is the matrix of three subpopulations (NSS, SS and TST), α is the SNP effect, β is the subpopulation effects, and ε is random effects. Thus, the R2 of each significant SNP after adjusting for the population structure effects were reported as previously described [37].
Pathway analysis
The pathway analysis was performed in https://maizegdb.org/past. The resulting SNP-trait association data and effects data generated by TASSEL were implemented in the pathway analysis. During the process of loading data, the LD data is filtered to drop rows where the loci are not the same, and then unneeded columns from the TASSEL output are dropped [103]. Only pathways with five or more mapped genes were considered in the analysis. Significance of the enrichment score was determined by permutation analysis (1000 random permutations of the effect values).
Prediction of candidate genes
To search the candidate genes underlying associated SNPs, we selected the most significant/leading SNPs within the same LD block (R2 < 0.2) to represent the locus, and candidate genes were nominated by the leading SNP positioned. The physical locations of the SNPs were recorded according to the B73 RefGen_v2 (www.maizesequence.org). The corresponding genes were annotated based on the literatures describing the function of their homologs in other species or the information retrieved from conserved domain database (CDD).
Availability of data and materials
All supporting data can be found within the manuscript and its additional files.
Abbreviations
- CW:
-
Cob weight
- DTA:
-
Days to anthesis
- DTH:
-
Days to heading
- DTS:
-
Days to silking
- ED:
-
Ear diameter
- EH:
-
Ear height
- EL:
-
Ear length
- ELL:
-
Ear leaf length
- ELW:
-
Ear leaf width
- GLM:
-
General linear model
- GW:
-
Cob grain weight
- GWAS:
-
Genome-wide association study
- K matrix:
-
Kinship relationship
- KNPR:
-
Kernel number per row
- KW:
-
Kernel width
- LD:
-
Linkage disequilibrium
- LNAE:
-
Leaf number above ear
- MAF:
-
Minor allele of frequency
- MLM:
-
Mixed linear model
- PH:
-
Plant height
- Q matrix:
-
Population structure
- R2 :
-
Percentage of phenotypic variation explained by the identified SNPs
- SNPs:
-
Single nucleotide polymorphisms
- TBN:
-
Tassel branch number
- TMAL:
-
Tassel maximum axis length
References
Petersen LN, Ingle RA, Knight MR, Denby KJ. OXI1 protein kinase is required for plant immunity against Pseudomonas syringae in Arabidopsis. J Exp Bot. 2009;60(13):3727–35. https://doi.org/10.1093/jxb/erp219.
Grierson C, Nielsen E, Ketelaarc T, Schiefelbein J. Root hairs. Arabidopsis Book. 2014;12:210–48. https://doi.org/10.1199/tab.0172.
Gilroy S, Jones DL. Through form to function: root hair development and nutrient uptake. Trends Plant Sci. 2000;5(2):56–60. https://doi.org/10.1016/S1360-1385(99)01551-4.
Lee RDW, Cho HT. Auxin, the organizer of the hormonal/environmental signals for root hair growth. Front Plant Sci. 2013;4. https://doi.org/10.3389/fpls.2013.00448.
Nestler J, Keyes SD, Wissuwa M. Root hair formation in rice (Oryza sativa L.) differs between root types and is altered in artificial growth conditions. J Exp Bot. 2016;67(12):3699–708. https://doi.org/10.1093/jxb/erw115.
Salazar-Henao JE, Vélez-Bermúdez IC, Schmidt W. The regulation and plasticity of root hair patterning and morphogenesis. Development. 2016;143(11):1848–58. https://doi.org/10.1242/dev.132845.
Cui SK, Suzaki T, Tominaga-Wada R, Yoshida S. Regulation and functional diversification of root hairs. Semin Cell Dev Biol. 2018;83:115–22. https://doi.org/10.1016/j.semcdb.2017.10.003.
Berger F, Hung CY, Dolan L, Schiefelbein J. Control of cell division in the root epidermis of Arabidopsis thaliana. Dev Biol. 1998;194(2):235–45. https://doi.org/10.1006/dbio.1997.8813.
Balcerowicz D, Schoenaers S, Vissenberg K. Cell fate determination and the switch from diffuse growth to planar polarity in Arabidopsis root epidermal cells. Front Plant Sci. 2015;6:1163–75. https://doi.org/10.3389/fpls.2015.01163.
Bruex A, Kainkaryam RM, Wieckowski Y, Kang YH, Bernhardt C, **a Y, et al. A gene regulatory network for root epidermis cell differentiation in Arabidopsis. PLoS Genet. 2012;8(1):2446–63. https://doi.org/10.1371/journal.pgen.1002446.
Gu F, Nielsen E. Targeting and regulation of cell wall synthesis during tip growth in plants. J Integr Plant Biol. 2013;55(9):835–46. https://doi.org/10.1111/jipb.12077.
Marzec M, Melzer M, Szarejko I. Root hair development in the grasses: what we already know and what we still need to know. Plant Physiol. 2015;168(2):407–14. https://doi.org/10.1104/pp.15.00158.
Kim CM, Han CD, Dolan L. RSL class I genes positively regulate root hair development in Oryza sativa. New Phytol. 2017;213(1):314–23. https://doi.org/10.1111/nph.14160.
Moon S, Cho LH, Kim YJ, Gho YS, Jeong HY, Hong WJ, et al. RSL class II transcription factors guide the nuclear localization of RHL1 to regulate root hair development. Plant Physiol. 2019;179(2):558–68. https://doi.org/10.1104/pp.18.01002.
Ma N, Wang Y, Qiu S, Kang Z, Che S, Wang G, et al. Overexpression of OsEXPA8, a root-specific gene, improves rice growth and root system architecture by facilitating cell extension. PloS One. 2013;8(10):e75997.
Yu Z, Kang B, He X, Lv S, Bai Y, Ding W, et al. Root hair-specific expansins modulate root hairelongation in rice. Plant J. 2011;66(5):725–34. https://doi.org/10.1111/j.1365-313X.2011.04533.x.
Kim CM, Park SH, Il Je B, Park SH, Park SJ, Piao HL, et al. OsCSLD1, a cellulose synthase-like D1 gene, is required for root hair morphogenesis in rice. Plant Physiol. 2007;143(3):1220–30. https://doi.org/10.1104/pp.106.091546.
Huang J, Kim CM, Xuan YH, Liu J, Kim TH, Kim BK, et al. Formin homology 1 (OsFH1) regulates root-hair elongation in rice (Oryza sativa). Planta. 2013;237(5):1227–39. https://doi.org/10.1007/s00425-013-1838-8.
Huang J, Kim CM, Xuan YH, Park SJ, Piao HL, Je BI, et al. OsSNDP1, a Sec14-nodulin domain-containing protein, plays a critical role in root hair elongation in rice. Plant Mol Biol. 2013;82(1-2):39–50. https://doi.org/10.1007/s11103-013-0033-4.
Guo M, Ruan W, Li C, Huang F, Zeng M, Liu Y, et al. Integrative comparison of the role of the PHOSPHATE RESPONSE1 subfamily in phosphate signaling and homeostasis in Rice. Plant Physiol. 2015;168(4):1762–76. https://doi.org/10.1104/pp.15.00736.
Wen TJ, Hochholdinger F, Sauer M, Bruce W, Schnable PS. The roothairless1 gene of maize encodes a homolog of sec3, which is involved in polar exocytosis. Plant Physiol. 2005;138(3):1637–43. https://doi.org/10.1104/pp.105.062174.
Hochholdinger F, Wen TJ, Zimmermann R, Chimot-Marolle P, da Costa e Silva O, Bruce W, et al. The maize (Zea mays L.) roothairless3 gene encodes a putative GPI-anchored, monocot-specific, COBRA-like protein that significantly affects grain yield. Plant J. 2008;54(5):888–98. https://doi.org/10.1111/j.1365-313X.2008.03459.x.
Nestler J, Liu S, Wen TJ, Paschold A, Marcon C, Tang HM, et al. Roothairless5, which functions in maize (Zea mays L.) root hair initiation and elongation encodes a monocot-specific NADPH oxidase. Plant J. 2014;79(5):729–40. https://doi.org/10.1111/tpj.12578.
Li L, Hey S, Liu S, Liu Q, McNinch C, Hu HC, et al. Characterization of maize roothairless6 which encodes a D-type cellulose synthase and controls the switch from bulge formation to tip growth. Sci Rep. 2016;6:34395.
Wang C, Qi C, Luo J, Liu L, He Y, Chen L. Characterization of LRL5 as a key regulator of root hair growth in maize. Plant J. 2019;98(1):71–82. https://doi.org/10.1111/tpj.14200.
Zhang X, Mi Y, Mao H, Liu S, Chen L, Qin F. Genetic variation in ZmTIP1 contributes to root hair elongation and drought tolerance in maize. Plant Biotechnol J. 2020;18(5):1271–83. https://doi.org/10.1111/pbi.13290.
Yu J, Buckler ES. Genetic association map** and genome organization of maize. Curr Opin Biotechnol. 2006;17(2):155–60. https://doi.org/10.1016/j.copbio.2006.02.003.
Atwell S, Huang YS, Vilhjálmsson BJ, Willems G, Horton M, Li Y, et al. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature. 2010;465(7298):627–31. https://doi.org/10.1038/nature08800.
Huang X, Zhao Y, Wei X, Li C, Wang A, Zhao Q, et al. Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat Genet. 2011;44(1):32–9. https://doi.org/10.1038/ng.1018.
Riedelsheimer C, Lisec J, Czedik-Eysenberg A, Sulpice R, Flis A, Grieder C, et al. Genome-wide association map** of leaf metabolic profiles for dissecting complex traits in maize. Proc Natl Acad Sci U S A. 2012;109(23):8872–7. https://doi.org/10.1073/pnas.1120813109.
Gore MA, Chia JM, Elshire RJ, Sun Q, Ersoz ES, Hurwitz BL, et al. A first-generation haplotype map of maize. Science. 2009;326(5956):1115–7. https://doi.org/10.1126/science.1177837.
Yan J, Shah T, Warburton ML, Buckler ES, McMullen MD, Crouch J. Genetic characterization and linkage disequilibrium estimation of a global maize collection using SNP markers. PLoS ONE. 2009;4(12):e8451.
Kump KL, Bradbury PJ, Wisser RJ, Buckler ES, Belcher AR, Oropeza-Rosas MA, et al. Genome-wide association study of quantitative resistance to southern leaf blight in the maize nested association map** population. Nat Genet. 2011;43(2):163–8. https://doi.org/10.1038/ng.747.
Guo Z, Tucker DM, Wang D, Basten CJ, Ersoz E, Briggs WH, et al. Accuracy of across-environment genome-wide prediction in maize nested association map** populations. G3 (Bethesda). 2013;3(2):263–72.
Leiboff S, Li X, Hu HC, Todt N, Yang J, Li X, et al. Genetic control of morphometric diversity in the maize shoot apical meristem. Nat Commun. 2015;6:8974.
Cui Z, Luo J, Qi C, Ruan Y, Li J, Zhang A, et al. Genome-wide association study (GWAS) reveals the genetic architecture of four husk traits in maize. BMC Genomics. 2016;17(1):946.
Wang X, Wang H, Liu S, Ferjani A, Li J, Yan J, et al. Genetic variation in ZmVPP1 contributes to drought tolerance in maize seedlings. Nat Genet. 2016;48(10):1233–41. https://doi.org/10.1038/ng.3636.
Li X, Zhou Z, Ding J, Wu Y, Zhou B, Wang R, et al. Combined linkage and association map** reveals QTL and candidate genes for plant and ear height in maize. Front Plant Sci. 2016;7. https://doi.org/10.3389/fpls.2016.00833.
Zhang X, Guan Z, Wang L, Fu J, Zhang Y, Li Z, et al. Combined GWAS and QTL analysis for dissecting the genetic architecture of kernel test weight in maize. Mol Gen Genomics. 2019;295(2):409–20. https://doi.org/10.1007/s00438-019-01631-2.
Jia T, Wang L, Li J, Ma J, Cao Y, Lübberstedt T, et al. Integrating a genome-wide association study with transcriptomic analysis to detect genes controlling grain drying rate in maize (Zea may, L.). Theor Appl Genet. 2019;133(2):623–34. https://doi.org/10.1007/s00122-019-03492-0.
Li W, Yu Y, Wang L, Luo Y, Peng Y, Xu Y, et al. The genetic architecture of the dynamic changes in grain moisture in maize. Plant Biotechnol J. 2021;19(6):1195–205. https://doi.org/10.1111/pbi.13541.
Yang X, Gao S, Xu S, Zhang Z, Prasanna BM, Li L, et al. Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize. Mol Breeding. 2010;28(4):511–26. https://doi.org/10.1007/s11032-010-9500-7.
Li H, Peng Z, Yang X, Wang W, Fu J, Wang J, et al. Genome-wide association study dissects the genetic architecture of oil biosynthesis in maize kernels. Nat Genet. 2012;45(1):43–50. https://doi.org/10.1038/ng.2484.
Yang N, Lu Y, Yang X, Huang J, Zhou Y, Ali F, et al. Genome wide association studies using a new nonparametric model reveal the genetic architecture of 17 agronomic traits in an enlarged maize association panel. PLoS Genet. 2014;10(9):e1004573.
Deng M, Li D, Luo J, **ao Y, Liu H, Pan Q, et al. The genetic architecture of amino acids dissection by association and linkage analysis in maize. Plant Biotechnol J. 2017;15(10):1250–63. https://doi.org/10.1111/pbi.12712.
Tang JD, Perkins A, Williams WP, Warburton ML. Using genome-wide associations to identify metabolic pathways involved in maize aflatoxin accumulation resistance. BMC Genomics. 2015;16(1):673. https://doi.org/10.1186/s12864-015-1874-9.
Warburton ML, Womack ED, Tang JD, Thrash A, Smith JS, Xu WW, et al. Genome-wide association and metabolic pathway analysis of corn earworm resistance in maize. Plant Genome. 2018;11(1):170069.
Li H, Thrash A, Tang JD, He L, Yan J, Warburton ML. Leveraging GWAS data to identify metabolic pathways and networks involved in maize lipid biosynthesis. Plant J. 2019;98(5):853–63. https://doi.org/10.1111/tpj.14282.
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, et al. A unified mixed-model method for association map** that accounts for multiple levels of relatedness. Nat Genet. 2005;38(2):203–8. https://doi.org/10.1038/ng1702.
Zhang Z, Ersoz E, Lai CQ, Todhunter RJ, Tiwari HK, Gore MA, et al. Mixed linear model approach adapted for genome-wide association studies. Nat Genet. 2010;42(4):355–60. https://doi.org/10.1038/ng.546.
Castelletti S, Coupel-Ledru A, Granato I, Palaffre C, Cabrera-Bosquet L, Tonelli C, et al. Maize adaptation across temperate climates was obtained via expression of two florigen genes. PloS Genet. 2020;16(7):e1008882.
Hochholdinger F. Genetic dissection of root formation in maize (Zea mays) reveals root-type specific developmental programmes. An Bot. 2004;93(4):359–68. https://doi.org/10.1093/aob/mch056.
Hochholdinger F, Park WJ, Sauer M, Woll K. From weeds to crops: genetic analysis of root development in cereals. Trends Plant Sci. 2004;9(1):42–8. https://doi.org/10.1016/j.tplants.2003.11.003.
Petricka JJ, Winter CM, Benfey PN. Control of Arabidopsis root development. Annu Rev Plant Biol. 2012;63(1):563–90. https://doi.org/10.1146/annurev-arplant-042811-105501.
Guo H, Li L, Ye H, Yu X, Algreen A, Yin Y. Three related receptor-like kinases are required for optimal cell elongation in Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2009;106(18):7648–53. https://doi.org/10.1073/pnas.0812346106.
Deslauriers SD, Larsen PB. FERONIA is a key modulator of brassinosteroid and ethylene responsiveness in Arabidopsis hypocotyls. Mol Plant. 2010;3(3):626–40. https://doi.org/10.1093/mp/ssq015.
Kessler SA, Shimosato-Asano H, Keinath NF, Wuest SE, Ingram G, Panstruga R, et al. Conserved molecular components for pollen tube reception and ungal invasion. Science. 2010;330(6006):968–71. https://doi.org/10.1126/science.1195211.
Stegmann M, Monaghan J, Smakowska-Luzan E, Rovenich H, Lehner A, Holton N, et al. The receptor kinase FER is a RALF-regulated scaffold controlling plant immune signaling. Science. 2017;355(6322):287–9.
Gommers CMM, Keuskamp DH, Buti S, van Veen H, Koevoets IT, Reinen E, et al. Molecular profiles of contrasting shade response strategies in wild plants: differential control of immunity and shoot elongation. Plant Cell. 2017;29(2):331–44. https://doi.org/10.1105/tpc.16.00790.
Barbez E, Dünser K, Gaidora A, Lendl T, Busch W. Auxin steers root cell expansion via apoplastic pH regulation in Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2017;114(24):4884–93. https://doi.org/10.1073/pnas.1613499114.
Feng W, Kita D, Peaucelle A, Cartwright HN, Doan V, Duan Q, et al. The FERONIA receptor kinase maintains cell-wall integrity during salt stress through Ca2+ signaling. Curr Biol. 2018;28(5):666–75. https://doi.org/10.1016/j.cub.2018.01.023.
Zhao C, Zayed O, Yu Z, Jiang W, Zhu P, Hsu CC, et al. Leucine-rich repeat extensin proteins regulate plant salt tolerance in Arabidopsis. Proc Natl Acad Sci U S A. 2018;115(51):13123–8. https://doi.org/10.1073/pnas.1816991115.
Dong Q, Zhang Z, Liu Y, Tao LZ, Liu H. FERONIA regulates auxin-mediated lateral root development and primary root gravitropism. FEBS Lett. 2018;593(1):97–106. https://doi.org/10.1002/1873-3468.13292.
Guo H, Nolan TM, Song G, Liu S, **e Z, Chen J, et al. FERONIA receptor kinase contributes to plant immunity by suppressing Jasmonic acid signaling in Arabidopsis thaliana. Curr Biol. 2018;28(20):3316–24. https://doi.org/10.1016/j.cub.2018.07.078.
Wang L, Yang T, Wang B, Lin Q, Zhu S, Li C, et al. RALF1-FERONIA complex affects splicing dynamics to modulate stress responses and growth in plants. Sci Adv. 2020;6(21):eaaz1622.
Duan Q, Kita D, Li C, Cheung AY, Wu HM. FERONIA receptor-like kinase regulates RHO GTPase signaling of root hair development. Proc Natl Acad Sci U S A. 2010;107(41):17821–6. https://doi.org/10.1073/pnas.1005366107.
Zhu S, Estévez JM, Liao H, Zhu Y, Yang T, Li C, et al. The RALF1–FERONIA complex phosphorylates eIF4E1 to promote protein synthesis and polar root hair growth. Mol Plant. 2020;13(5):698–716. https://doi.org/10.1016/j.molp.2019.12.014.
Zhu S, Martínez Pacheco J, Estevez JM, Yu F. Autocrine regulation of root hair size by the RALF-FERONIA-RSL4 signaling pathway. New Phytol. 2020;227(1):45–9. https://doi.org/10.1111/nph.16497.
Molendijk AJ, Bischoff F, Rajendrakumar CSV, Friml J, Braun M, Gilroy S, et al. Arabidopsis thaliana Rop GTPases are localized to tips of root hairs and control polar growth. EMBO J. 2001;20(11):2779–88. https://doi.org/10.1093/emboj/20.11.2779.
Vernoud V, Horton AC, Yang Z, Nielsen E. Analysis of the small GTPase gene superfamily of Arabidopsis. Plant Physiol. 2003;131(3):1191–208. https://doi.org/10.1104/pp.013052.
Denninger P, Reichelt A, Schmidt VAF, Mehlhorn DG, Asseck LY, Stanley CE, et al. Distinct RopGEFs successively drive polarization and outgrowth of root hairs. Curr Biol. 2019;29(11):1854–65. https://doi.org/10.1016/j.cub.2019.04.059.
Drevensek S, Goussot M, Duroc Y, Christodoulidou A, Steyaert S, Schaefer E, et al. The Arabidopsis TRM1-TON1 interaction reveals a recruitment network common to plant cortical microtubule arrays and eukaryotic centrosomes. Plant Cell. 2012;24(1):178–91. https://doi.org/10.1105/tpc.111.089748.
Takahashi H, Hirota K, Kawahara A, Hayakawa E, Inoue Y. Randomization of cortical microtubules in root epidermal cells induces root hair initiation in lettuce (Lactuca sativa L.) seedlings. Plant Cell Physiol. 2003;44(3):350–9. https://doi.org/10.1093/pcp/pcg043.
Takahashi H, Kawahara A, Inoue Y. Ethylene promotes the induction by auxin of the cortical microtubule randomization required for low-pH-induced root hair initiation in lettuce (Lactuca sativa L.) seedlings. Plant Cell Physiol. 2003;44(9):932–40. https://doi.org/10.1093/pcp/pcg119.
Zhao B, Lin X, Poland J, Trick H, Leach J, Hulbert S. A maize resistance gene functions against bacterial streak disease in rice. Proc Natl Acad Sci U S A. 2005;102(43):15383–8. https://doi.org/10.1073/pnas.0503023102.
Zhang C, Chen H, Cai T, Deng Y, Zhuang R, Zhang N, et al. Overexpression of a novel peanut NBS-LRR gene AhRRS5 enhances disease resistance to Ralstonia solanacearum in tobacco. Plant Biotechnol J. 2017;15(1):39–55. https://doi.org/10.1111/pbi.12589.
McHale L, Tan X, Koehl P, Michelmore RW. Plant NBS-LRR proteins: adaptable guards. Genome Biol. 2006;7(4):212.
Sarazin V, Duclercq J, Mendou B, Aubanelle L, Nicolas V, Aono M, et al. Arabidopsis BNT1, an atypical TIR–NBS–LRR gene, acting as a regulator of the hormonal response to stress. Plant Sci. 2015;239:216–29. https://doi.org/10.1016/j.plantsci.2015.07.017.
Tanaka N, Kato M, Tomioka R, Kurata R, Fukao Y, Aoyama T, et al. Characteristics of a root hair-less line of Arabidopsis thaliana under physiological stresses. J Exp Bot. 2014;65(6):1497–512. https://doi.org/10.1093/jxb/eru014.
Bates TR, Lynch JP. Stimulation of root hair elongation in Arabidopsis thaliana by low phosphorus availability. Plant Cell Environ. 1996;19(5):529–38. https://doi.org/10.1111/j.1365-3040.1996.tb00386.x.
Schikora A, Schmidt W. Iron stress-induced changes in root epidermal cell fate are regulated independently from physiological responses to low iron availability. Plant Physiol. 2001;125(4):1679–87. https://doi.org/10.1104/pp.125.4.1679.
Yang TJW, Perry PJ, Ciani S, Pandian S, Schmidt W. Manganese deficiency alters the patterning and development of root hairs in Arabidopsis. J Exp Bot. 2008;59(12):3453–64. https://doi.org/10.1093/jxb/ern195.
Jung JY, Shin R, Schachtman DP. Ethylene mediates response and tolerance to potassium deprivation in Arabidopsis. Plant Cell. 2009;21(2):607–21. https://doi.org/10.1105/tpc.108.063099.
Niu Y, Chai R, Liu L, ** G, Liu M, Tang C, et al. Magnesium availability regulates the development of root hairs in Arabidopsis thaliana (L.) Heynh. Plant Cell Environ. 2014;37(12):2795–813. https://doi.org/10.1111/pce.12362.
He X, Zeng J, Cao F, Ahmed IM, Zhang G, Vincze E, et al. HvEXPB7, a novel β-expansin gene revealed by the root hair transcriptome of Tibetan wild barley, improves root hair growth under drought stress. J Exp Bot. 2015;66(22):7405–19. https://doi.org/10.1093/jxb/erv436.
Cheng S, Zhou D, Zhao Y. WUSCHEL-related homeobox gene WOX11 increases rice drought resistance by controlling root hair formation and root system development. Plant Signal Behav. 2016;11(2):e1130198.
Zheng Y, Drechsler N, Rausch C, Kunze R. The Arabidopsis nitrate transporter NPF7.3/NRT1.5 is involved in lateral root development under potassium deprivation. Plant Signal Behav. 2016;11(5):e1176819.
Diet A, Link B, Seifert GJ, Schellenberg B, Wagner U, Pauly M, et al. The Arabidopsis root hair cell wall formation mutant lrx1 is suppressed by mutations in the RHM1 gene encoding a UDP-L-rhamnose synthase. Plant Cell. 2006;18(7):1630–41. https://doi.org/10.1105/tpc.105.038653.
Wang X. Regulatory functions of phospholipase D and phosphatidic acid in plant growth, development, and stress responses. Plant Physiol. 2005;139(2):566–73. https://doi.org/10.1104/pp.105.068809.
Kf de Campos M, Schaaf G. The regulation of cell polarity by lipid transfer proteins of the SEC14 family. Curr Opin Plant Biol. 2017;40:158–68.
Reyes F, Orellana A. Golgi transporters: opening the gate to cell wall polysaccharide biosynthesis. Curr Opin Plant Biol. 2008;11(3):244–51. https://doi.org/10.1016/j.pbi.2008.03.008.
Rautengarten C, Ebert B, Moreno I, Temple H, Herter T, Link B, et al. The Golgi localized bifunctional UDP-rhamnose/UDP-galactose transporter family of Arabidopsis. Proc Natl Acad Sci U S A. 2014;111(31):11563–8. https://doi.org/10.1073/pnas.1406073111.
Orellana A, Moraga C, Araya M, Moreno A. Overview of nucleotide sugar transporter gene family functions across multiple species. J Mol Biol. 2016;428(16):3150–65. https://doi.org/10.1016/j.jmb.2016.05.021.
Temple H, Saez-Aguayo S, Reyes FC, Orellana A. The inside and outside: topological issues in plant cell wall biosynthesis and the roles of nucleotide sugar transporters. Glycobiology. 2016;26(9):913–25. https://doi.org/10.1093/glycob/cww054.
Akoh CC, Lee GC, Liaw YC, Huang TH, Shaw JF. GDSL family of serine esterases/lipases. Prog Lipid Res. 2004;43(6):534–52. https://doi.org/10.1016/j.plipres.2004.09.002.
Clauß K, Baumert A, Nimtz M, Milkowski C, Strack D. Role of a GDSL lipase-like protein as sinapine esterase in Brassicaceae. Plant J. 2008;53(5):802–13. https://doi.org/10.1111/j.1365-313X.2007.03374.x.
Liu H, Luo X, Niu L, **ao Y, Chen L, Liu J, et al. Distant eQTLs and non-coding sequences play critical roles in regulating gene expression and quantitative trait variation in maize. Mol Plant. 2017;10(3):414–26. https://doi.org/10.1016/j.molp.2016.06.016.
Gui S, Yang L, Li J, Luo J, Xu X, Yuan J, et al. ZEAMAP, a comprehensive database adapted to the maize Multi-Omics era. iScience. 2020;23(6):101241.
Li MX, Yeung JMY, Cherny SS, Sham PC. Evaluating the effective numbers of independent tests and significant p-value thresholds in commercial genoty** arrays and public imputation reference datasets. Hum Genet. 2011;131(5):747–56. https://doi.org/10.1007/s00439-011-1118-2.
Yang W, Guo Z, Huang C, Duan L, Chen G, Jiang N, et al. Combining high-throughput phenoty** and genome-wide association studies to reveal natural genetic variation in rice. Nat Commun. 2014;5:5087.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Human Genet. 2007;81(3):559–75. https://doi.org/10.1086/519795.
Mao H, Wang H, Liu S, Li Z, Yang X, Yan J, et al. A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat Commun. 2015;6:8326.
Thrash A, Tang JD, DeOrnellis M, Peterson DG, Warburton ML. PAST: The Pathway Association Studies Tool to infer biological meaning from GWAS datasets. Plants (Basel). 2020;9(1):58.
Acknowledgments
We thank all members of our laboratories for the helpful assistances and discussions during the research.
Funding
This research was funded by grants from the National Key Research and Development Program of China (2017YFD0101104) to Y.H. and the National Natural Science Foundation of China (31872668, 31470279) to L.Q.C.
Author information
Authors and Affiliations
Contributions
L.Q.C. and Y.H. conceived and supervised the project; L.L. conducted experiments and performed bioinformatics and statistical analyses; Manuscript was prepared by L. L, L.Q.C. and Y.H. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All the experiments on plant seeds were in accordance with guidelines of China Agricultural University. The source from the maize seeds were publicly available at Maizego.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional files 1:
Figure S1. GWAS showing significant P-values associated with root hair length based on the general linear model (GLM). (a) Manhattan plots of GLM for root hair length. The horizontal dashed line represents the significance threshold -log10(P) = 6.5. (b) QQ plot of GLM showing the ratio of the observed and the expected P-values for root hair length. The solid diagonal lines represent agreement between observed and expected probability distributions assuming null SNP–trait association.
Additional files 2: 0
Figure S2. The allele effects of significant SNPs located around representative genes for root hair length. (a–d; i-l) Regional plots showing association map** results for SNPs located around GRMZM2G100288 (a), GRMZM2G147446 (b), GRMZM2G078013 (c), GRMZM2G000471 (d), GRMZM2G064644 (i), AC193598.3_FG002 (j), GRMZM2G180244 (k), GRMZM2G091579 (l). (e–h; m-p) Allele effects of the most significant SNPs for root hair traits. (e) GRMZM2G100288, (f) GRMZM2G147446, (g) GRMZM2G078013, (h) GRMZM2G000471, (m) GRMZM2G064644, (n) AC193598.3_FG002, (o) GRMZM2G180244, (p) GRMZM2G091579. Each dot represents an SNP. The horizontal dashed black line represents the significant threshold –log10(P) = 5.2. *Significant at P ≤ 0.05; **significant at P ≤ 0.01.
Additional files 3:
Table S1. Root hair length ratio (Inbreds/B73) of the 281 inbred lines from different subpopulations.
Additional files 4:
Supplemental Table 2. Pathways associated with increased root hair length at statistical significance P < 0.05.
Additional files 5:
Supplemental Table 3. Pathways associated with decreased hair length at statistical significance P < 0.05.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Liu, L., Jiang, LG., Luo, JH. et al. Genome-wide association study reveals the genetic architecture of root hair length in maize. BMC Genomics 22, 664 (2021). https://doi.org/10.1186/s12864-021-07961-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-021-07961-z