Abstract
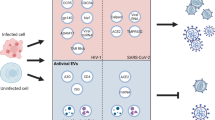
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has caused a global pandemic. Antibody resistance dampens neutralizing antibody therapy and threatens current global Coronavirus (COVID-19) vaccine campaigns. In addition to the emergence of resistant SARS-CoV-2 variants, little is known about how SARS-CoV-2 evades antibodies. Here, we report a novel mechanism of extracellular vesicle (EV)-mediated cell-to-cell transmission of SARS-CoV-2, which facilitates SARS-CoV-2 to escape from neutralizing antibodies. These EVs, initially observed in SARS-CoV-2 envelope protein-expressing cells, are secreted by various SARS-CoV-2-infected cells, including Vero E6, Calu-3, and HPAEpiC cells, undergoing infection-induced pyroptosis. Various SARS-CoV-2-infected cells produce similar EVs characterized by extra-large sizes (1.6–9.5 μm in diameter, average diameter > 4.2 μm) much larger than previously reported virus-generated vesicles. Transmission electron microscopy analysis and plaque assay reveal that these SARS-CoV-2-induced EVs contain large amounts of live virus particles. In particular, the vesicle-cloaked SARS-CoV-2 virus is resistant to neutralizing antibodies and able to reinfect naïve cells independent of the reported receptors and cofactors. Consistently, the constructed 3D images show that intact EVs could be taken up by recipient cells directly, supporting vesicle-mediated cell-to-cell transmission of SARS-CoV-2. Our findings reveal a novel mechanism of receptor-independent SARS-CoV-2 infection via cell-to-cell transmission, provide new insights into antibody resistance of SARS-CoV-2 and suggest potential targets for future antiviral therapeutics.
Similar content being viewed by others
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) explosively spreads and has clinical manifestations ranging from asymptomatic infection to respiratory failure and even death1,2,3. SARS-CoV-2 is a novel virus belonging to the Beta coronavirus genus and exhibits high similarities to another two coronaviruses, severe acute respiratory syndrome coronavirus (SARS-CoV) and Middle East respiratory syndrome coronavirus (MERS-CoV), which have caused large-scale outbreaks over the past two decades4,5. Previous studies found that the SARS-CoV-2 lifecycle commences with classic binding of the spike (S) protein to its cognate receptor on the surface of the host cell, human angiotensin-converting enzyme 2 (hACE-2). The cleavage of the S1/S2 site by the membrane protease serine 2 (TMPRSS2) and virus–cell membrane fusion mediated by lysosomal cathepsin L determine the efficiency of virus entry6,7,8,9. Single-cell sequencing data exhibit that the hACE-2 receptor is expressed in less than 1% cells in certain tissues at low expression levels, including heart, liver, brain, lung, and trachea, yet SARS-CoV-2 RNA may still be detected in these organs10,11,12,20. Current antibody therapies are divided into anti-viral and anti-inflammatory treatments. Among antibody options, convalescent plasma (CP) treatment is receiving significant attention, which may provide patients with immediate passive immunity21,22. However, CP therapy is suboptimal and fails to reverse respiratory failure and reduce mortality23,24. Another promising treatment option was monoclonal antibodies designed to mainly target the S protein of the virus membrane or the hACE-2 receptor of the host cell plasma membrane, thereby preventing viral binding with its receptor. To date, at least eight antibody candidates targeting the S protein have entered different stages of clinical studies20. The LY-CoV555 antibody from Lilly was the first neutralizing antibody to receive FDA emergency use authorization for the treatment of COVID-19. In a phase II trial, LY-CoV555 appeared to accelerate the natural decline in viral load at day 2 in outpatients diagnosed with mild or moderate COVID-19, but did not have the same effect in patients with severe COVID-19 or those with prolonged illness25,26. However, LY-CoV555 exhibited an unsatisfactory therapeutic effect on SARS-CoV-2 variant B.1.1.727. In particular, the “cocktail antibodies” BRII-196 plus BRII-198, developed against variants, were stopped early due to lack of utility28. Appearance of spontaneous mutations in SARS-CoV-2 is the main reason for the unsatisfactory effect of neutralizing antibodies targeting the virus. SARS-CoV-2 variants may dampen the efficacy and specificity of antibodies and further lead to new viral strains that may gradually develop resistance to existing antibodies29,30. Thus, the problem of SARS-CoV-2 esca** from antibodies needs much effort to be solved.
Results
The SARS-CoV-2 envelope protein induces extracellular vesicles containing virus particles
Our previous research has demonstrated that the SARS-CoV-2 structural envelope (2-E) protein forms a type of pH-sensitive cation channel, and that heterogeneous expression of 2-E channels causes host cell death44,45. Multiple characteristic visual fields for CoV-2-EVs were captured. First, there were numerous dense virus particles and mitochondria in a shedding CoV-2-EV. Second, a large number of SARS-CoV-2 virions were encapsulated in the shed CoV-2-EVs (Fig. 3a, yellow dotted circles). These virions displayed an average diameter of 75 ± 10 nm, consistent with previous reports43. To further verify the presence of SARS-CoV-2 virions in CoV-2-EVs, we performed immunohistochemistry analyses using SARS-CoV-2 nucleocapsid immunogold labeling. SARS-CoV-2-infected (MOI = 1) Vero E6 cells showed strong labeling for the nucleocapsid in the cytosol and in viral particles that accumulated intracellularly (Fig. 3b). In shed CoV-2-EVs, in addition to cellular contents, viral particles were visible and marked by gold particles (Fig. 3b, yellow arrows). Plaque-reduction assay and qRT-PCR also supported that these CoV-2-EVs contained a large number of infectious viruses, as high as 2.3 × 107 PFU/mL and 2 × 109 viral copies/mL (Fig. 3c–e). We noticed that the virial copies in the 2-E-EVs were higher than those in the virus-induced EVs, implying that redundant 2-E proteins may facilitate virus production, packaging, or EV secretion.
a TEM images of Vero E6 cells infected with SARS-CoV-2. Blue triangles, double-membrane vesicles (DMVs); red triangles, SMVs; yellow dotted circles, virus particles; red arrow, damaged mitochondria; blue dotted circle, lysosomes (MOI = 1). b Immunoelectron microscopy analyses of infected Vero E6 cells using SARS-CoV-2 nucleocapsid immunogold labeling (MOI = 1). Yellow arrows, virus particles. c Schematic representation of infection experiments. Vero E6 cells were infected with SARS-CoV-2. On the next day, the cells, supernatant, and CoV-2-EVs were collected. d qRT-PCR analysis of SARS-CoV-2 RNA in supernatant and isolated EVs from SARS-CoV-2-infected Vero E6 cells. e Plaque reduction assay of CoV-2-EVs. Data are the means ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; unpaired Student’s t-test.
SARS-CoV-2-induced vesicles help viruses escape from neutralizing antibodies and establish a productive infection
References
WHO Coronavirus Disease (COVID-19) Dashboard. https://covid19.who.int/ (2022).
Wu, J. et al. Clinical features of maintenance hemodialysis patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. Clin. J. Am. Soc. Nephrol. 15, 1139–1145 (2020).
Chu, H. et al. Comparative tropism, replication kinetics, and cell damage profiling of SARS-CoV-2 and SARS-CoV with implications for clinical manifestations, transmissibility, and laboratory studies of COVID-19: an observational study. Lancet Microbe 1, e14–e23 (2020).
Kim, D. et al. The architecture of SARS-CoV-2 transcriptome. Cell 181, 914–921.e10 (2020).
Piroth, L. et al. Comparison of the characteristics, morbidity, and mortality of COVID-19 and seasonal influenza: a nationwide, population-based retrospective cohort study. Lancet Respir. Med. 9, 251–259 (2021).
Harrison, A. G., Lin, T. & Wang, P. Mechanisms of SARS-CoV-2 transmission and pathogenesis. Trends Immunol. 41, 1100–1115 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e8 (2020).
Shang, J. et al. Cell entry mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 117, 11727–11734 (2020).
Matheson, N. J. & Lehner, P. J. How does SARS-CoV-2 cause COVID-19? Science 369, 510–511 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Wang, S. et al. AXL is a candidate receptor for SARS-CoV-2 that promotes infection of pulmonary and bronchial epithelial cells. Cell Res. 31, 126–140 (2021).
Puelles, V. G. et al. Multiorgan and renal tropism of SARS-CoV-2. N. Engl. J. Med. 383, 590–592 (2020).
Chen, L., Li, X., Chen, M., Feng, Y. & **ong, C. The ACE2 expression in human heart indicates new potential mechanism of heart injury among patients infected with SARS-CoV-2. Cardiovasc. Res. 116, 1097–1100 (2020).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 26, 681–687 (2020).
Li, H. et al. SARS-CoV-2 and viral sepsis: observations and hypotheses. Lancet 395, 1517–1520 (2020).
Hikmet, F. et al. The protein expression profile of ACE2 in human tissues. Mol. Syst. Biol. 16, e9610 (2020).
Nie, X. et al. Multi-organ proteomic landscape of COVID-19 autopsies. Cell 184, 775–791 (2021).
Daly, J. L. et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science 370, 861–865 (2020).
Cantuti-Castelvetri, L. et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science 370, 856–860 (2020).
Renn, A., Fu, Y., Hu, X., Hall, M. D. & Simeonov, A. Fruitful neutralizing antibody pipeline brings hope to defeat SARS-Cov-2. Trends Pharm. Sci. 41, 815–829 (2020).
Rojas, M. et al. Convalescent plasma in Covid-19: possible mechanisms of action. Autoimmun. Rev. 19, 102554 (2020).
Focosi, D., Anderson, A. O., Tang, J. W. & Tuccori, M. Convalescent plasma therapy for COVID-19: state of the art. Clin. Microbiol. Rev. 33, e00072–20 (2020).
Simonovich, V. A. et al. A randomized trial of convalescent plasma in Covid-19 severe pneumonia. N. Engl. J. Med. 384, 619–629 (2021).
Rubin, R. Testing an old therapy against a new disease: convalescent plasma for COVID-19. JAMA 323, 2114–2117 (2020).
Chen, P. et al. SARS-CoV-2 neutralizing antibody LY-CoV555 in outpatients with Covid-19. N. Engl. J. Med. 384, 229–237 (2020).
Group, A.-T. L.-C. S. et al. A neutralizing monoclonal antibody for hospitalized patients with Covid-19. N. Engl. J. Med. 384, 905–914 (2021).
Terry, M. Citing Variants, Eli Lilly Asks FDA To Revoke EUA for Single Antibody Therapy. https://www.biospace.com/article/eli-lilly-requests-fda-revoke-eua-for-bam-monotherapy/ (2021).
Group, A. C.-T. F. I. W. C.-S. Efficacy and safety of two neutralising monoclonal antibody therapies, sotrovimab and BRII-196 plus BRII-198, for adults hospitalised with COVID-19 (TICO): a randomised controlled trial. Lancet Infect. Dis. 22, 622–635 (2021).
Baum, A. et al. Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies. Science 369, 1014–1018 (2020).
Weisblum, Y. et al. Escape from neutralizing antibodies by SARS-CoV-2 spike protein variants. Elife 9, e61312 (2020).
**a, B. et al. SARS-CoV-2 envelope protein causes acute respiratory distress syndrome (ARDS)-like pathological damages and constitutes an antiviral target. Cell Res. 31, 847–860 (2021).
Shi, J., Gao, W. & Shao, F. Pyroptosis: gasdermin-mediated programmed necrotic cell death. Trends Biochem. Sci. 42, 245–254 (2017).
Wang, Y. et al. Chemotherapy drugs induce pyroptosis through caspase-3 cleavage of a gasdermin. Nature 547, 99–103 (2017).
Samson, A. L. et al. MLKL trafficking and accumulation at the plasma membrane control the kinetics and threshold for necroptosis. Nat. Commun. 11, 3151 (2020).
Chen, X. et al. Pyroptosis is driven by non-selective gasdermin-D pore and its morphology is different from MLKL channel-mediated necroptosis. Cell Res. 26, 1007–1020 (2016).
Sims, N. R. & Anderson, M. F. Isolation of mitochondria from rat brain using Percoll density gradient centrifugation. Nat. Protoc. 3, 1228–1239 (2008).
Liao, P. C., Bergamini, C., Fato, R., Pon, L. A. & Pallotti, F. Isolation of mitochondria from cells and tissues. Methods Cell Biol. 155, 3–31 (2020).
Ghosh, S. et al. beta-Coronaviruses use lysosomes for egress instead of the biosynthetic secretory pathway. Cell 183, 1520–1535 (2020).
Amara, A. & Mercer, J. Viral apoptotic mimicry. Nat. Rev. Microbiol. 13, 461–469 (2015).
Altan-Bonnet, N. Extracellular vesicles are the Trojan horses of viral infection. Curr. Opin. Microbiol. 32, 77–81 (2016).
Alenquer, M. & Amorim, M. J. Exosome biogenesis, regulation, and function in viral infection. Viruses 7, 5066–5083 (2015).
Lam, S. M. et al. A multi-omics investigation of the composition and function of extracellular vesicles along the temporal trajectory of COVID-19. Nat. Metab. 3, 909–922 (2021).
Wang, P. et al. A cross-talk between epithelium and endothelium mediates human alveolar-capillary injury during SARS-CoV-2 infection. Cell Death Dis. 11, 1042 (2020).
Cortese, M. et al. Integrative imaging reveals SARS-CoV-2-induced resha** of subcellular morphologies. Cell Host Microbe 28, 853–866.e5 (2020).
Klein, S. et al. SARS-CoV-2 structure and replication characterized by in situ cryo-electron tomography. Nat. Commun. 11, 5885 (2020).
Wang, T., Fang, L., Zhao, F., Wang, D. & **ao, S. Exosomes mediate intercellular transmission of porcine reproductive and respiratory syndrome virus. J. Virol. 92, e01734–17 (2018).
Ramakrishnaiah, V. et al. Exosome-mediated transmission of hepatitis C virus between human hepatoma Huh7.5 cells. Proc. Natl. Acad. Sci. USA 110, 13109–13113 (2013).
SARS-CoV-2 (2019-nCoV) Spike Neutralizing Antibody, Rabbit Mab. https://cn.sinobiological.com/antibodies/cov-spike-40592-r001.
Pan, X. et al. Immunoglobulin fragment F(ab’)2 against RBD potently neutralizes SARS-CoV-2 in vitro. Antivir. Res. 182, 104868 (2020).
Zeng, C. et al. SARS-CoV-2 spreads through cell-to-cell transmission. Proc. Natl. Acad. Sci. USA 119, e2111400119 (2022).
Rogers, C. et al. Cleavage of DFNA5 by caspase-3 during apoptosis mediates progression to secondary necrotic/pyroptotic cell death. Nat. Commun. 8, 14128 (2017).
Vanden Berghe, T. et al. Necroptosis, necrosis, and secondary necrosis converge on similar cellular disintegration features. Cell Death Differ. 17, 922–930 (2010).
Freundt, E. C. et al. The open reading frame 3a protein of severe acute respiratory syndrome-associated coronavirus promotes membrane rearrangement and cell death. J. Virol. 84, 1097–1109 (2010).
Yorimitsu, T. & Klionsky, D. J. Eating the endoplasmic reticulum: quality control by autophagy. Trends Cell Biol. 17, 279–285 (2007).
Chen, Y. H. et al. Phosphatidylserine vesicles enable efficient en bloc transmission of enteroviruses. Cell 160, 619–630 (2015).
Plescia, C. B. et al. SARS-CoV-2 viral budding and entry can be modeled using BSL-2 level virus-like particles. J. Biol. Chem. 296, 100103 (2021).
Caobi, A., Nair, M. & Raymond, A. D. Extracellular vesicles in the pathogenesis of viral infections in humans. Viruses 12, 1200 (2020).
Zhu, N. et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 382, 727–733 (2020).
Arantes, T. S. et al. The large marseillevirus explores different entry pathways by forming giant infectious vesicles. J. Virol. 90, 5246–5255 (2016).
Andino, R. & Domingo, E. Viral quasispecies. Virology 479-480, 46–51 (2015).
Muslin, C., Joffret, M. L., Pelletier, I., Blondel, B. & Delpeyroux, F. Evolution and emergence of enteroviruses through intra- and inter-species recombination: plasticity and phenotypic impact of modular genetic exchanges in the 5’ untranslated region. PLoS Pathog. 11, e1005266 (2015).
Pelchen-Matthews, A., Raposo, G. & Marsh, M. Endosomes, exosomes, and Trojan viruses. Trends Microbiol. 12, 310–316 (2004).
Borderia, A. V. et al. Group selection and contribution of minority variants during virus adaptation determines virus fitness and phenotype. PLoS Pathog. 11, e1004838 (2015).
Duchene, S. et al. Temporal signal and the phylodynamic threshold of SARS-CoV-2. Virus Evol. 6, veaa061 (2020).
Leung, K., Shum, M. H., Leung, G. M., Lam, T. T. & Wu, J. T. Early transmissibility assessment of the N501Y mutant strains of SARS-CoV-2 in the United Kingdom, October to November 2020. Eur. Surveill. 26, 2002106 (2021).
Guo, Y. Q. et al. SARS-CoV-2 induced intestinal responses with a biomimetic human gut-on-chip. Sci. Bull. 66, 783–793 (2021).
Yao, X. H. et al. A cohort autopsy study defines COVID-19 systemic pathogenesis. Cell Res. 31, 836–846 (2021).
Mathieu, M., Martin-Jaular, L., Lavieu, G. & Théry, C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 21, 9–17 (2019).
Mulcahy, L. A., Pink, R. C. & Carter, D. R. Routes and mechanisms of extracellular vesicle uptake. J. Extracell. Vesicles https://doi.org/10.3402/jev.v3.24641 (2014).
Colombo, M., Raposo, G. & Thery, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 30, 255–289 (2014).
Acknowledgements
We thank electron microscope platform and Chemical Biology Core Facility in Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences. We thank the staff at the Core Facilities at the School of Life Sciences, Peking University for their professional technical assistance in EM sample preparation and image analysis. We are grateful to the National Science Fund of Distinguished Young Scholars (81825021), Fund of Youth Innovation Promotion Association (2019285, 2021333), the National Natural Science Foundation of China (81773707, 31700732), Project supported by Shanghai Municipal Science and Technology Major Project, Fund of Shanghai Science and Technology Innovation Action Plan (20ZR1474200), Shanghai Rising-Star Program (22QA1411000), the National Key Laboratory Program of China (LG202101-01-04), the National Key Research and Development Program of China (2020YFC0842000), the Strategic Leading Science and Technology Projects of Chinese Academy of Sciences (XDA12050308), Fund of National Science and Technology Major Project (2018ZX09711002-002-006), the Hubei Science and Technology Project (2020FCA003) and Yunnan Key Research and Development Program (202103AC100001) for financial support.
Author information
Authors and Affiliations
Contributions
Z.G., B.X., and J.L. conceived the project. Z.G., B.X., L.-K.Z., and J.L. designed the experiments; X.S., Y. Wang, X.Z., S.L., and Q.L. carried out the cell-based assays and SEM of transfected or virus-infected cells; G.X., X.P., and Y. Wu carried out the virus assays in vitro; Y.-T.Z., R.-H.L., and Y.G. carried out TEM of virus-infected cells; X.-Y.L., X.-Y.H., H.-Y.Z., Y.L., and W.P. carried out golden hamster infection assays. All authors analyzed and discussed the data. Z.G., B.X., and L.-K.Z. wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
**a, B., Pan, X., Luo, RH. et al. Extracellular vesicles mediate antibody-resistant transmission of SARS-CoV-2. Cell Discov 9, 2 (2023). https://doi.org/10.1038/s41421-022-00510-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41421-022-00510-2
- Springer Nature Singapore Pte Ltd.