KEGG and GO analyses identified crucial DEGs and DAPs involved in the osmotic stress response in maize leaves. We further analyzed these DEGs and DAPs with respect to the enriched pathways. Similar to previous studies (Shi et al. 2013; Kanojia et al. 2020; Singh et al. 2022), we mainly focused on upregulated DEGs and more abundant DAPs, highlighting their important roles in abiotic stress resistance in plants (Zhang et al. 2022).
Primary and energy metabolism
Photosynthesis and glycolysis provide essential compounds and energy for maintaining cellular homeostasis. Under osmotic stress, almost all DEGs linked to photosynthesis were downregulated, including Chl a/b binding protein, PsbQ, PsaN, OHP, cytb6f, ATP synthase, and RuBisCO (Supplementary Fig. 4A, B; Supplementary Dataset 5). These genes were suppressed the most in L1 and the least in L3, consistent with decreased photosynthesis. Only PsbP, ferredoxin, and triose-phosphate isomerase were upregulated in stressed leaves. Osmotic stress stimulated genes related to Chl degradation, such as pheophorbide-a oxygenase and senescence-inducible chloroplast stay-green proteins, but inhibited genes related to Chl synthesis, especially in L1 (Supplementary Fig. 4C). At the proteomic level, two Chl a/b-binding proteins and PsbP showed inconsistent changes in their corresponding gene levels, with PsbP being more abundant only in L3 (Supplementary Fig. 4A; Supplementary Dataset 5).
In photorespiration, the gene glycolate oxidase linked to peroxisomes was significantly downregulated in L1, but not in L3, whereas two genes encoding mitochondrial glycine cleavage system H protein2 (GDH2) were over-represented (Supplementary Fig. 4D), especially in mature leaves. Thus, the upregulated GDH2 enhances the release of CO2 that can enter the Calvin cycle again, thereby protecting the photosynthetic apparatus under water deficit conditions.
Transitory starch is the main storage form of photosynthetic products; it accumulates in starch granules (SG) in chloroplasts during the day, and is hydrolyzed to soluble sugars at night (AbdElgawad et al. 2020). Microscopy observation indicated that SG existed only in vascular bundle sheath cells but not in mesophyll cells (Fig. 5A, B). Upon 4 h stress, SG degraded during light significantly more in L3 than in mature leaves, and the isolated SG (1.4 to 2.5 μm) from L3 were reduced in size compared with those from mature leaves (Fig. 5C). Meanwhile, maltose, glucose, and sucrose content increased in stressed leaves (Fig. 5D). After removing mature leaves, L3 accumulated adequate SG, comparable to the control (Fig. 5E).
Most genes linked to starch and sugar metabolism were upregulated in the stressed leaves (Supplementary Fig. 5A; Supplementary Dataset 6). Particularly, four β-amylases involved in starch degradation were upregulated, of which ZmBAM8 had unusually high log2FC of 8.96, 8.63 and 6.84 in L1, L2 and L3, respectively. Most genes linked to sugar conversion and transport were also over-represented in stressed leaves, e.g., sucrose-phosphate synthase, three UDP-glycosyltransferase genes and β-frucotofuranosidase (Supplementary Fig. 5B; Supplementary Dataset 6). Interestingly, sucrose transporter 3 (Suc3) was only upregulated in L1. In Arabidopsis, sucrose export from the source tissues is mainly controlled by SUC2 activity (Tong et al. 2022). Hence, the upregulation of Suc3 enhanced sucrose export from L1 to young leaves under osmotic stress. These results imply that SG degradation during the day plays a vital role in leaf stress adaptation, especially in L3, which differs from starch degradation at night to sustain plant growth and metabolism.
The glycolytic tricarboxylic acid (TCA) cycle plays a central role in energy metabolism. Key genes linked to glycolytic TCA were over-presented in stressed leaves, especially in L1, e.g., pyruvate dehydrogenase kinase, NADH dehydrogenase, alternative oxidase AOX3 precursor, cyclic oxidative cytochrome C-2, and isocitrate lyase (Supplementary Fig. 5C; Supplementary Dataset 6). Therefore, when chloroplast energy production declines under osmotic stress, mitochondrial and cytosolic energy pathways are enhanced to supply energy for plant stress adaptation.
Differential defense response to osmotic stress via specific DEGs/DAPs
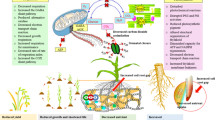
Acute osmotic stress caused a reduction in leaf RWC (Fig. 1C), leading to dynamic alterations in gene expression and protein synthesis (Figs. 3 and 4). The heat map displayed significant changes in the selected DEGs and DAPs in individual leaves (Fig. 6), which were mainly involved in stress signaling, TF regulation, stress tolerance, ROS scavenging, and the synthesis and degradation of proteins and lipids.
A large number of MAPK cascade components were over-represented under osmotic stress, especially in L1 (Fig. 6A; Supplementary Dataset 7), including MAPK, MAPK2, MAPK8, phospholipase C, cysteine-rich receptor-like kinase, leucine-rich repeat receptors, lectin receptor-like kinase, G-proteins, 14-3-3 proteins, and phosphoinositides, which are known to encode key signal molecules that regulate plant development, metabolism, and stress response (Zhu 2016). Many genes associated with calcium sensing, binding, transport, and responsiveness were upregulated under osmotic stress, especially in L1 (Fig. 6A; Supplementary Dataset 7); encoding for plasma membrane calcium-transporting ATPase, Ca-binding protein, calmodulin, and chloroplastic GTP diphosphokinase. In particular, calcium-transporting ATPase 5 was upregulated with high log2FC values of 5.05, 5.10 and 3.67 in L1, L2 and L3, respectively.
The upregulated MAPK and Ca2+ signaling pathways would trigger the activity of TFs, heat shock proteins (HSPs) and other stress proteins. The stress-responsive TFs genes include many MYB, WRKY, APETALA2/ethylene-responsive factor (AP2/ERF), bHLH, dehydration-responsive element-binding proteins (DREB), and heat shock factor (HSF) (Supplementary Dataset 7), which are implicated in plant responses to multiple abiotic stresses (Zhu 2016; Zhang et al. 2022). Overall, more upregulated WRKY, AP2/ERF, MYB, bHLH and HSF genes were observed in L1 than in L2 and L3. The upregulated bHLH genes were eight, five, and four in L1, L2, and L3, respectively. The bHLH93 is upregulated in mature leaves, and its homolog is involved in the differentiation of stomatal guard cells (Ohashi-Ito and Bergmann 2006).
Stress proteins such as HSPs and dehydrin play a vital role in conferring abiotic stress tolerance (Szlachtowska and Rurek 2023). Many genes encoding HSPs, chaperones, and dehydrin were upregulated more in L1 than in L2 and L3 (Fig. 6A). Among the 27 identified HSPs, 20 exhibited higher levels in L1. The chloroplast chaperone ClpD was upregulated in all stressed leaves; however, ClpD only increased in L1. Five different dehydrin genes were highly induced by osmotic stress, with the average log2FC of 5.48, 4.69, 3.17 in L1, L2, and L3, respectively. Immunoblotting revealed increased dehydrin levels in mature leaves under osmotic stress. Similarly, aquaporin TIP1-1 (Fig. 6B) increased in mature leaves, consistent with TIP1-1 expression levels, likely permitting the rapid influx of water from old leaves into young leaves under water-deficient conditions.
A common strategy of plant responses to intracellular osmotic fluctuations is to accumulate osmolytes, such as proline. Delta-1-pyrroline-5-carboxylate synthase (P5CS) controlling the rate-limiting step in glutamate-derived proline biosynthesis. Three P5CS were upregulated in all stressed leaves (Fig. 6A; Supplementary Dataset 7). Meanwhile, a MYB that positively regulates proline synthesis and transport, was only upregulated in L1 (Supplementary Dataset 7).
Protein and lipid degradation pathways were enriched more evidently in L1 under osmotic stress (Fig. 3). Many genes involved in ubiquitin-mediated protein degradation, such as cysteine protease, aspartate protease, ubiquitin-protein ligase, and RING-type E3 ubiquitin transferase were upregulated in stressed leaves, with higher levels in L1 (Fig. 6C). Protease do-like 5, which is involved in chloroplast protein degradation, increased only in L1 (Fig. 6D). Genes linked to glycerolipid degradation (α/β-hydrolases) and fatty acid degradation (acetyl-CoA acetyltransferase) were upregulated in mature stressed leaves (Fig. 6C). Similarly, acetyl-CoA acetyltransferase was significantly upregulated in mature stressed leaves. Sphingolipids are fundamental components of the plant membrane system and are involved in multiple cellular and stress-response processes (Huby et al. 2020). The expression of sphingolipid biosynthesis-related genes, such as ASC1-like protein 1 decreased in all stressed leaves, particularly in L1 (Fig. 6C; Supplementary Dataset 7).
Hormones signal transduction
Hormone signal transduction was enriched in all leaves under osmotic stress (Fig. 3C). Numerous DEGs have been linked to the biosynthesis and signaling of auxins, ABA, zeatin, jasmonic acid (JA), and ethylene. Most of the DEGs (8 of 11) related to the ABA biosynthesis and response, were upregulated in all stressed leaves. The 9-cis-Epoxycarotenoid dioxygenase (NCED) is a key enzyme in ABA biosynthesis. HAV22 and ABF4 (ABRE binding factor 4) are involved in ABA-activated signaling. Four NCED and two HAV22 genes, as well as ABF4, were upregulated by osmotic stress, with higher levels observed in mature leaves (Supplementary Fig. 6). Likewise, MYB139 was upregulated in mature leaves (Supplementary Dataset 7), and its homolog AtMYB44 repressed genes encoding phosphatase 2C in response to ABA (Jung et al. 2008). The expression levels of ABA-related genes positively correlated with leaf age, which was consistent with the actual ABA levels (Fig. 2C).
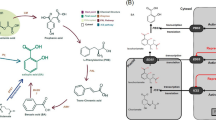
One SA-related gene was downregulated in all leaves, which was consistent with the reduced SA levels in all leaves (Fig 2; Supplementary Fig. 6). Most genes regulating JA biosynthesis, including LOX1 (3), LOX3 (1), AOS (1), and OPR1 (2), were upregulated in L1, followed by L2, but were downregulated in L3, except for one LOX1, which was inconsistent with decreased JA levels in stressed leaves.
Ethylene is a well-known senescence-inducing hormone in plants (Rankenberg et al. 2021). Many TFs genes AP2/ERF were overrepresented under osmotic stress in L1 (17), followed by L2 (9), and L3 (7) (Supplementary Fig. 6). Three ACO genes (involved in ethylene biosynthesis) and two ethylene-responsive protein (ERP) genes were upregulated to comparable levels in all leaves (Supplementary Fig. 6). Without assaying ethylene levels, we could not compare the expression levels of ethylene-related genes.
Remarkably, genes related to hormones that regulate cell division and growth showed decreased expression in all stressed leaves, especially in L1 (Supplementary Fig. 6). Of the 15 auxin-related genes, the upregulated genes were more abundant in L2 (5) and L3 (5), whereas the downregulated genes were more abundant in L1 (6). Of the 11 zeatin-related genes, seven, two, and one were under-represented in L1, L2, and L3, respectively, consistent with zeatin changes (Fig. 2). GA-related genes were downregulated in all leaves, whereas GA20 oxidase 1 was upregulated.
Secondary metabolism
Secondary metabolites such as terpenoids, phenolics, and flavonoids play critical roles in plant adaptation to abiotic stress (Winkel-Shirley 2002). Multiple phenylpropanoid synthesis-related genes were upregulated in all the stressed leaves (Supplementary Fig. 7). In particular, many genes linked to the biosynthesis of terpenoids, betaine, lignin, and wax, were upregulated, especially in L1 (Supplementary Dataset 7). Genes encoding UDP-glucose 4-epimerase, UDP-glucose 6-dehydrogenase, UDP-glucuronate decarboxylase, and cellulose synthase which are associated with cell wall synthesis, were upregulated in mature leaves. In particular, MYB20 which is involved in the regulation of secondary cell wall biogenesis, showed a high log2FC of 7.10, 7.54, 5.40 in L1, L2 and L3, respectively. By contrast, the genes encoding glycosyl hydrolase family 10 protein and mannan endo-1,4-β-mannosidase that are related to cell wall degradation were upregulated in mature leaves (Supplementary Fig. 7; Supplementary Dataset 7). Of the 15 genes linked to flavonoid biosynthesis, five were upregulated in L1. Five MYBs that regulate flavonoid and riboflavin biosynthesis were upregulated in all the stressed leaves. Moreover, the ZF-HD homeobox protein gene ZHD15 which controls flavonoid biosynthesis, was highly expressed in L1 (log2FC 6.11) and L2 (log2FC 3.05), but slightly increased in L3 (log2FC 0.90).