Search
Search Results
-
Chromatin lncRNA Platr10 controls stem cell pluripotency by coordinating an intrachromosomal regulatory network
BackgroundA specific 3-dimensional intrachromosomal architecture of core stem cell factor genes is required to reprogram a somatic cell into...

-
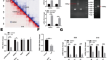
JUN-induced super-enhancer RNA forms R-loop to promote nasopharyngeal carcinoma metastasis
Oncogenic super-enhancers (SEs) generate noncoding enhancer/SE RNAs (eRNAs/seRNAs) that exert a critical function in malignancy through powerful...
-
MoDLE: high-performance stochastic modeling of DNA loop extrusion interactions
DNA loop extrusion emerges as a key process establishing genome structure and function. We introduce MoDLE, a computational tool for fast, stochastic...

-
Diverse silent chromatin states modulate genome compartmentalization and loop extrusion barriers
The relationships between chromosomal compartmentalization, chromatin state and function are poorly understood. Here by profiling long-range contact...

-
Fold-back mechanism originating inv-dup-del rearrangements in chromosomes 13 and 15
Intrachromosomal rearrangements involve a single chromosome and can be formed by several proposed mechanisms. We reported two patients with...

-
Interruption of aberrant chromatin loo** is required for regenerating RB1 function and suppressing tumorigenesis
RB transcriptional corepressor 1 ( RB1 ) is a critical regulatory gene in physiological and pathological processes. Genetic mutation is considered to...

-
Smc3 acetylation, Pds5 and Scc2 control the translocase activity that establishes cohesin-dependent chromatin loops
Cohesin is a DNA translocase that is instrumental in the folding of the genome into chromatin loops, with functional consequences on DNA-related...

-
Chromosomal Interaction in Chromatin Organization
Genes are arranged linearly in long DNA strands. Genomic DNA must be packaged tightly to be contained in the nucleus. Studies have shown that...
-
Hi-C Analysis to Identify Genome-Wide Chromatin Structural Aberration in Cancer
Hi-C is a method that analyzes genome-wide chromatin structureGenome-wide chromatin structure using next-generation sequencerNext-generation...
-
Multiscale reorganization of the genome following DNA damage facilitates chromosome translocations via nuclear actin polymerization
Nuclear actin-based movements have been shown to orchestrate clustering of DNA double-strand breaks (DSBs) into homology-directed repair domains....

-
The roles of long noncoding RNAs in the regulation of OCT4 expression
OCT4 is a major transcription factor that maintains the pluripotency of stem cells, including embryonic stem cells, induced pluripotent stem cells...

-
Analysis of HiChIP Data
HiChIP is a novel method for the analysis of chromatin interactions based on in situ Hi-C that adds an immuno-precipitation (ChIP) step for the...
-
Open chromatin interaction maps reveal functional regulatory elements and chromatin architecture variations during wheat evolution
BackgroundBread wheat ( Triticum aestivum ) is an allohexaploid that is generated by two subsequent allopolyploidization events. The large genome size...

-
Non-Mendelian Genetics: Finding of Transposable Elements in Corns and Flies
The essence of Mendelian genetics is the rule of chromosome segregation and the independent assortment of different chromosomes. However, genes on...
-
A graph neural network-based interpretable framework reveals a novel DNA fragility–associated chromatin structural unit
BackgroundDNA double-strand breaks (DSBs) are among the most deleterious DNA lesions, and they can cause cancer if improperly repaired. Recent...

-
Lineage-specific rearrangement of chromatin loops and epigenomic features during adipocytes and osteoblasts commitment
Human mesenchymal stem cells (hMSCs) can be differentiated into adipocytes and osteoblasts. The processes are driven by the rewiring of chromatin...

-
SnapHiC: a computational pipeline to identify chromatin loops from single-cell Hi-C data
Single-cell Hi-C (scHi-C) analysis has been increasingly used to map chromatin architecture in diverse tissue contexts, but computational tools to...

-
Chromatin structure in cancer
In the past decade, we have seen the emergence of sequence-based methods to understand chromosome organization. With the confluence of in situ ...

-
Diurnal RNAPII-tethered chromatin interactions are associated with rhythmic gene expression in rice
BackgroundThe daily cycling of plant physiological processes is speculated to arise from the coordinated rhythms of gene expression. However, the...

-
3D organization of regulatory elements for transcriptional regulation in Arabidopsis
BackgroundAlthough spatial organization of compartments and topologically associating domains at large scale is relatively well studied, the spatial...

