Abstract
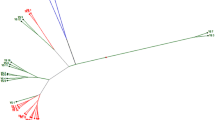
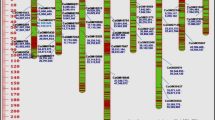
Eleven Chinese and twelve Swedish rapeseed (Brassica napus) genotypes were analysed by PCR with 41 microsatellite primers, generating a total of 50 loci. For these 50 loci, the number of alleles ranged from 1 to 14, and the average number of alleles per loci was 2.7. As an example of simple sequence repeat (SSR) scoring in Metaphor agarose gel, a single marker could distinguish 14 different DNA profiles. Based on cluster analysis (UPGMA), the dendrogram clearly distinguished three clusters, a cluster with exclusively Swedish genotypes, and two clusters with Chinese genotypes. The genetic diversity within the Chinese genotypes was broad compared to the genetic diversity within the Swedish material. The genetic similarity within the Swedish breeding lines ranged from 69.5 to 95.6%, while that of Chinese genotypes ranged from 57.1 to 81.6%. The results in this report will permit to establish a set of microsatellite primers that can be used for selecting appropriate parents for Brassica napus hybrids and for monitoring hybridity level.
Similar content being viewed by others

References
Y.M. Charters A. Robertson M.J. Wilkinson G. Ramsey (1996) ArticleTitlePCR analysis of oilseed rape cultivars (Brassica napus L. ssp. oleifera) using 5′-anchored simple sequence repeat (SSR) primers Theor. Appl. Genet. 92 442–447 Occurrence Handle1:CAS:528:DyaK28XltFOjtLs%3D
Cheung W.Y., Hubert N. and Landry B.S. 1993. A simple and rapid DNA microextraction method for plant, animal, and insect suitable for RAPD and other PCR analysis. Technical tips. In: PCR Methods and Applications, Vol. 3. Cold Spring Harbor Laboratory Press, USA, pp. 69–70.
B.W. Diers P.B.E. McVetty T.C. Osborn (1996) ArticleTitleRelationship between heterosis and genetic distance based on restriction fragment length polymorphism markers in oilseed rape (Brassica napus L.) Crop Sci. 36 79–83
H.H. Gu P. Hagberg W.J. Zhou (2004) ArticleTitleCold pretreatment enhances microspore embryogenesis in oilseed rape (Brassica napus L.) Plant Growth Regul. 42 137–143 Occurrence Handle10.1023/B:GROW.0000017488.29181.fa Occurrence Handle1:CAS:528:DC%2BD2cXhsFOhtLc%3D
S. Kresovich A.K. Szewc-McFadden S.M. Bliek J.R. McFerson (1995) ArticleTitleAbundance and characterization of simple-sequence repeats (SSRs) isolated from a size-fractionated genomic library of Brassica napus L. (rapeseed) Theor. Appl. Genet. 91 206–211 Occurrence Handle10.1007/BF00220879 Occurrence Handle1:CAS:528:DyaK2MXotl2qt7Y%3D
C.Z. Ma T.D. Fu S. Tuvesson B. Gertsson (2003) ArticleTitleGenetic diversity of Chinese and Swedish rapeseed (Brassica napus L.) analysed by Inter-Simple Sequence Repeats (ISSR) Agric. Sci. China 2 137–143
R.J. Mailer R. Scarth B. Fristensky (1994) ArticleTitleDiscrimination among cultivars of rapeseed (Brassica napus L.) using DNA polymorphisms amplified from arbitrary primers Theor. Appl. Genet. 87 697–704 Occurrence Handle10.1007/BF00222895 Occurrence Handle1:CAS:528:DyaK2MXkt1Cqtw%3D%3D
E.J.J. Momoh W.J. Zhou B. Kristiansson (2002) ArticleTitleVariation in the development of secondary dormancy in oilseed rape genotypes under conditions of stress Weed Res. 42 446–455 Occurrence Handle10.1046/j.1365-3180.2002.00308.x
M. Nei (1978) ArticleTitleEstimation of average heterozygosity and genetic distance from a small number of individuals Genetics 89 583–590
J. Plieske D. Struss (2001) ArticleTitleMicrosatellite markers for genome analysis in Brassica. I. Development in Brassica napusabundance in Brassicaceae species Theor. Appl. Genet. 102 689–694 Occurrence Handle1:CAS:528:DC%2BD3MXjvVerurg%3D
A.K. Szewc-McFadden S. Kresovich S.M. Bliek S.E. Mitchell J.R. McFerson (1996) ArticleTitleIdentification of polymorphic, conserved simple sequence repeats (SSRs) in cultivated Brassica species Theor. Appl. Genet. 93 534–538 Occurrence Handle1:CAS:528:DyaK28XmtF2nsLk%3D
G.X. Tang W.J. Zhou H.Z. Li B.Z. Mao Z.H. He K. Yoneyama (2003) ArticleTitleMediumexplant and genotype factors influencing shoot regeneration in oilseed Brassica spp J. Agron. Crop Sci. 189 351–358 Occurrence Handle10.1046/j.1439-037X.2003.00060.x
C.E. Thormann M.E. Ferreira L.E.A. Camargo J.G. Tivang T.C. Osborn (1994) ArticleTitleComparison of genetic relationship estimates within and among cruciferous species based on RFLP and RAPD markers Theor. Appl. Genet. 88 973–980 Occurrence Handle10.1007/BF00220804
M.I. Uzunova W. Ecke (1999) ArticleTitleAbundancepolymorphism and genetic map** of microsatellites in oilseed rape (Brassica napus L.) Plant Breed. 118 323–326 Occurrence Handle10.1046/j.1439-0523.1999.00371.x Occurrence Handle1:CAS:528:DyaK1MXmvVSns7k%3D
F.C. Yeh R.C. Yang T. Boyle (1999) Popgene Version 1.31, Microsoft Window-Based Freeware for Population Genetic Analysis. Quick User Guide University of Alberta Edmonton, Canada 29
G.Q. Zhang W.J. Zhou H.H. Gu W.J. Song E.J.J. Momoh (2003) ArticleTitlePlant regeneration from the hybridization of Brassica napus and B. juncea through embryo culture J. Agron. Crop Sci. 189 347–350 Occurrence Handle10.1046/j.1439-037X.2003.00059.x
W.J. Zhou (2001) Oilseed rape G.P. Zhang W.J. Zhou (Eds) Crop Cultivation Zhejiang University Press Hangzhou, China 153–178
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, W.J., Zhang, G.Q., Tuvesson, S. et al. Genetic Survey of Chinese and Swedish Oilseed Rape (Brassica napus L.) by Simple Sequence Repeats (SSRs). Genet Resour Crop Evol 53, 443–447 (2006). https://doi.org/10.1007/s10722-004-7862-6
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s10722-004-7862-6