Abstract
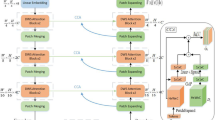
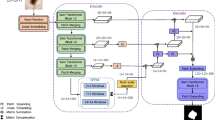
The combination of convolution and Transformer applied to medical image segmentation has achieved great success. However, it still cannot reach extremely accurate segmentation on complex and low-contrast anatomical structures under lower calculation. To solve this problem, we propose a lite Transformer based medical image segmentation framework called LiteTrans, which deeply integrates Transformer and CNN in an Encoder-Decoder-Skip-Connection U-shaped architecture. Inspired by Transformer, a novel multi-branch module with convolution operation and Local-Global Self-Attention (LGSA) is incorporated into LiteTrans to unify local and non-local feature interactions. In particular, LGSA is a global self-attention approximation scheme with lower computational complexity. We evaluate LiteTrans by conducting extensive experiments on synapse multi-organ and ACDC datasets, showing that this approach achieves state-of-the-art performance over other segmentation methods, with fewer parameters and lower FLOPs.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bernard, O., et al.: Deep learning techniques for automatic MRI cardiac multi-structures segmentation and diagnosis: is the problem solved? IEEE Trans. Med. Imaging 37, 2514–2525 (2018)
Cao, H., et al.: Swin-Unet: Unet-like pure transformer for medical image segmentation. ar**v preprint ar**v:2105.05537 (2021)
Chen, J., et al.: TransUNet: transformers make strong encoders for medical image segmentation. ar**v preprint ar**v:2102.04306 (2021)
Chen, X., Wang, H., Ni, B.: X-volution: on the unification of convolution and self-attention. ar**v preprint ar**v:2106.02253 (2021)
Chollet, F.: Xception: deep learning with depthwise separable convolutions. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 1251–1258 (2017)
Dai, Z., Liu, H., Le, Q.V., Tan, M.: CoAtNet: marrying convolution and attention for all data sizes. ar**v preprint ar**v:2106.04803 (2021)
Dosovitskiy, A., et al.: An image is worth 16x16 words: transformers for image recognition at scale. In: International Conference on Learning Representations (2020)
Fu, S., et al.: Domain adaptive relational reasoning for 3D multi-organ segmentation. In: Martel, A.L. (ed.) MICCAI 2020. LNCS, vol. 12261, pp. 656–666. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-59710-8_64
Gibson, E., et al.: Multi-organ abdominal CT reference standard segmentations. This data set was developed as part of independent research supported by Cancer Research UK (Multidisciplinary C28070/A19985) and the National Institute for Health Research UCL/UCL Hospitals Biomedical Research Centre (2018)
Han, K., et al.: A survey on visual transformer (2020)
Hu, J., Shen, L., Sun, G.: Squeeze-and-excitation networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 7132–7141 (2018)
Huang, G., Liu, Z., Laurens, V., Weinberger, K.Q.: Densely connected convolutional networks. IEEE Computer Society (2016)
Huang, L., Yuan, Y., Guo, J., Zhang, C., Chen, X., Wang, J.: Interlaced sparse self-attention for semantic segmentation (2019)
Huang, Z., Ben, Y., Luo, G., Cheng, P., Yu, G., Fu, B.: Shuffle transformer: rethinking spatial shuffle for vision transformer. ar**v preprint ar**v:2106.03650 (2021)
Huang, Z., Wang, X., Huang, L., Huang, C., Wei, Y., Liu, W.: CCNET: criss-cross attention for semantic segmentation. In: Proceedings of the IEEE/CVF International Conference on Computer Vision, pp. 603–612 (2019)
Jha, D., Riegler, M.A., Johansen, D., Halvorsen, P., Johansen, H.D.: DoubleU-Net: a deep convolutional neural network for medical image segmentation. In: 2020 IEEE 33rd International Symposium on Computer-Based Medical Systems (CBMS), pp. 558–564. IEEE (2020)
Kaul, C., Manandhar, S., Pears, N.: FocusNet: an attention-based fully convolutional network for medical image segmentation. In: 2019 IEEE 16th International Symposium on Biomedical Imaging, ISBI 2019, pp. 455–458. IEEE (2019)
Lei, T., Wang, R., Wan, Y., Du, X., Meng, H., Nandi, A.: Medical image segmentation using deep learning: a survey (2020)
Li, D., et al.: Involution: inverting the inherence of convolution for visual recognition. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 12321–12330 (2021)
Li, J., Yan, Y., Liao, S., Yang, X., Shao, L.: Local-to-global self-attention in vision transformers. ar**v preprint ar**v:2107.04735 (2021)
Liu, Z., et al.: Swin transformer: hierarchical vision transformer using shifted windows (2021)
Milletari, F., Navab, N., Ahmadi, S.A.: V-Net: fully convolutional neural networks for volumetric medical image segmentation. In: 2016 4th International Conference on 3D Vision (3DV), pp. 565–571. IEEE (2016)
Oktay, O., et al.: Attention U-Net: learning where to look for the pancreas (2018)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Srinivas, A., Lin, T.Y., Parmar, N., Shlens, J., Abbeel, P., Vaswani, A.: Bottleneck transformers for visual recognition. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 16519–16529 (2021)
Taghanaki, S.A., Abhishek, K., Cohen, J.P., Cohen-Adad, J., Hamarneh, G.: Deep semantic segmentation of natural and medical images: a review. Artif. Intell. Rev. 54(1), 137–178 (2021)
Vaswani, A., et al.: Attention is all you need. ar**v (2017)
Vaswani, A., Ramachandran, P., Srinivas, A., Parmar, N., Hechtman, B., Shlens, J.: Scaling local self-attention for parameter efficient visual backbones. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 12894–12904 (2021)
Wang, H., Zhu, Y., Green, B., Adam, H., Yuille, A., Chen, L.-C.: Axial-DeepLab: stand-alone axial-attention for panoptic segmentation. In: Vedaldi, A., Bischof, H., Brox, T., Frahm, J.-M. (eds.) ECCV 2020. LNCS, vol. 12349, pp. 108–126. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-58548-8_7
Wang, W., **e, E., Li, X., Fan, D.P., Shao, L.: Pyramid vision transformer: a versatile backbone for dense prediction without convolutions (2021)
Wang, Z., Zou, N., Shen, D., Ji, S.: Non-local U-Nets for biomedical image segmentation. In: Proceedings of the AAAI Conference on Artificial Intelligence, vol. 34, pp. 6315–6322 (2020)
Wu, H., et al.: CvT: introducing convolutions to vision transformers. ar**v preprint ar**v:2103.15808 (2021)
Wu, Z., Liu, Z., Lin, J., Lin, Y., Han, S.: Lite transformer with long-short range attention. In: International Conference on Learning Representations (2019)
Xu, W., Xu, Y., Chang, T., Tu, Z.: Co-scale conv-attentional image transformers (2021)
Yu, Q., **a, Y., Bai, Y., Lu, Y., Yuille, A., Shen, W.: Glance-and-gaze vision transformer. ar**v preprint ar**v:2106.02277 (2021)
Zhang, Y., Liu, H., Hu, Q.: Transfuse: fusing transformers and CNNs for medical image segmentation. ar**v preprint ar**v:2102.08005 (2021)
Zheng, S., et al.: Rethinking semantic segmentation from a sequence-to-sequence perspective with transformers. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 6881–6890 (2021)
Zhou, Z., Siddiquee, M.M.R., Tajbakhsh, N., Liang, J.: Unet++: redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans. Med. Imaging 39(6), 1856–1867 (2019)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Xu, S., Quan, H. (2021). LiteTrans: Reconstruct Transformer with Convolution for Medical Image Segmentation. In: Wei, Y., Li, M., Skums, P., Cai, Z. (eds) Bioinformatics Research and Applications. ISBRA 2021. Lecture Notes in Computer Science(), vol 13064. Springer, Cham. https://doi.org/10.1007/978-3-030-91415-8_26
Download citation
DOI: https://doi.org/10.1007/978-3-030-91415-8_26
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-91414-1
Online ISBN: 978-3-030-91415-8
eBook Packages: Computer ScienceComputer Science (R0)