Abstract
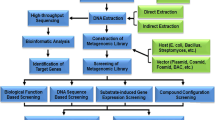
The ability to produce high-value products using bacteria will increasingly rely on continued research to make large-scale bacterial fermentation cost-efficient. Engineering bacteria to use alternate carbon sources as feedstock provides an opportunity to reduce production costs. Using inexpensive carbon sources from various forms of waste provides an opportunity to substantially reduce feedstock costs. Functional carbon metabolism pathways can be identified by the introduction of metagenomic libraries into the organism of interest followed by screening for the desired phenotype. We present here a method to transfer metagenomic libraries from E. coli to Pseudomonas alloputida, followed by screening for use of galactose as a sole carbon source.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Cheng J, Pinnell L, Engel K et al (2014) Versatile broad-host-range cosmids for construction of high quality metagenomic libraries. J Microbiol Methods 99:27–34. https://doi.org/10.1016/j.mimet.2014.01.015
Lam KN, Cheng J, Engel K et al (2015) Current and future resources for functional metagenomics. Front Microbiol 6:1196. https://doi.org/10.3389/fmicb.2015.01196
Cheng J, Romantsov T, Engel K et al (2017) Functional metagenomics reveals novel β-galactosidases not predictable from gene sequences. PLoS One 12:e0172545. https://doi.org/10.1371/journal.pone.0172545
Cheng J, Charles TC (2016) Novel polyhydroxyalkanoate copolymers produced in Pseudomonas putida by metagenomic polyhydroxyalkanoate synthases. Appl Microbiol Biotechnol 100:7611–7627. https://doi.org/10.1007/s00253-016-7666-6
Cheng J, Nordeste R, Trainer MA, Charles TC (2017) Methods for the isolation of genes encoding novel PHA metabolism enzymes from complex microbial communities. In: Streit W, Daniel R (eds) Metagenomics. Methods in molecular biology, vol 1539. Humana Press, New York, pp 237–248. https://doi.org/10.1007/978-1-4939-6691-2_15
Nikel PI, Lorenzo V de (2018) Pseudomonas putida as a functional chassis for industrial biocatalysis: from native biochemistry to trans-metabolism. Metab Eng 50:142–155. https://doi.org/10.1016/j.ymben.2018.05.005
Martínez-García E, Nikel PI, Aparicio T, Lorenzo V de (2014) Pseudomonas 2.0: genetic upgrading of P. putida KT2440 as an enhanced host for heterologous gene expression. Microb Cell Factories 13:1–15. https://doi.org/10.1186/s12934-014-0159-3
Tsang YF, Kumar V, Samadar P et al (2019) Production of bioplastic through food waste valorization. Environ Int 127:625–644. https://doi.org/10.1016/j.envint.2019.03.076
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Press, Cold Spring Harbor
Escapa IF, Morales V, Martino VP et al (2011) Disruption of β-oxidation pathway in Pseudomonas putida KT2442 to produce new functionalized PHAs with thioester groups. Appl Microbiol Biotechnol 89:1583–1598. https://doi.org/10.1007/s00253-011-3099-4
Jones JDG, Gutterson N (1987) An efficient mobilizable cosmid vector, pRK 7813, and its use in a rapid method for marker exchange in Pseudomonas fluorescens strain HV37a. Gene 61:299–306. https://doi.org/10.1016/0378-1119(87)90193-4
Finan TM, Kunkel B, Vos GFD, Signer ER (1986) Second symbiotic megaplasmid in Rhizobium meliloti carrying exopolysaccharide and thiamine synthesis genes. J Bacteriol 167:66–72. https://doi.org/10.1128/jb.167.1.66-72.1986
Okonechnikov K, Golosova O, Fursov M, team U (2012) Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 28:1166–1167. https://doi.org/10.1093/bioinformatics/bts091
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinform Oxf Engl 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Aziz RK, Bartels D, Best AA et al (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Kovach ME, Elzer PH, Hill DS et al (1995) Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166:175–176. https://doi.org/10.1016/0378-1119(95)00584-1
Meade MJ, Waddell RL, Callahan TM (2001) Soil bacteria Pseudomonas putida and Alcaligenes xylosoxidans subsp. denitrificans inactivate triclosan in liquid and solid substrates. FEMS Microbiol Lett 204:45–48. https://doi.org/10.1111/j.1574-6968.2001.tb10860.x
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Thakor, A., Cheng, J., Charles, T.C. (2023). Isolation of Genes Encoding Carbon Metabolism Pathways from Complex Microbial Communities. In: Streit, W.R., Daniel, R. (eds) Metagenomics. Methods in Molecular Biology, vol 2555. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2795-2_8
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2795-2_8
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2794-5
Online ISBN: 978-1-0716-2795-2
eBook Packages: Springer Protocols