Abstract
Proteolysis targeting chimeras (PROTACs) technology has emerged as a novel therapeutic paradigm in recent years. PROTACs are heterobifunctional molecules that degrade target proteins by hijacking the ubiquitin–proteasome system. Currently, about 20–25% of all protein targets are being studied, and most works focus on their enzymatic functions. Unlike small molecules, PROTACs inhibit the whole biological function of the target protein by binding to the target protein and inducing subsequent proteasomal degradation. PROTACs compensate for limitations that transcription factors, nuclear proteins, and other scaffolding proteins are difficult to handle with traditional small-molecule inhibitors. Currently, PROTACs have successfully degraded diverse proteins, such as BTK, BRD4, AR, ER, STAT3, IRAK4, tau, etc. And ARV-110 and ARV-471 exhibited excellent efficacy in clinical II trials. However, what targets are appropriate for PROTAC technology to achieve better benefits than small-molecule inhibitors are not fully understood. And how to rationally design an efficient PROTACs and optimize it to be orally effective poses big challenges for researchers. In this review, we summarize the features of PROTAC technology, analyze the detail of general principles for designing efficient PROTACs, and discuss the typical application of PROTACs targeting different protein categories. In addition, we also introduce the progress of relevant clinical trial results of representative PROTACs and assess the challenges and limitations that PROTACs may face. Collectively, our studies provide references for further application of PROTACs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
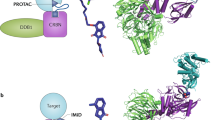
Proteolysis targeting chimeras (PROTACs) were first reported by Sakamoto et al. in 2001 [1]. PROTACs are heterobifunctional molecules that contain three components: the protein-of-interest (POI) binding moiety, a linker, and E3 ubiquitin ligase binding moiety (Fig. 1a) [2, 88]. However, the similar catalytic domain of the same family members prevented researchers from develo** isoform-specific inhibitors. [89]. To address this problem, the highly specific SGK3-PROTAC1 (Fig. 2d) was developed. This PROTAC was designed by Tovell’s group based on the non-SGK3 selective inhibitor 308-R, to degrade SGK3 specifically [90]. At a low micromolar concentration of SGK3-PROTAC1, intracellular SGK3 levels can be significantly reduced without affecting SGK1 and SGK2. It could be assumed that the selectivity and specificity of SGK3-PROTAC1 derives from the selective recognition of SGK3 by VHL during the formation of ternary complexes induced by SGK3-PROTAC1.
Catalytic mode of action (MOA)
Traditional small-molecule inhibitors act in a dose-dependent manner, to achieve clinical effect by maximizing drug-receptor occupancy. Excessive drug concentrations lead to undesirable side effects and off-target effects [91]. PROTACs can initiate the degradation of target protein catalytic and escape from proteasome [92]. Theoretically, PROTACs can be delivered at lower doses, for longer dosing intervals, and with lower toxicity than small molecule inhibitors since their low concentration is sufficient to degrade proteins and is not constrained by equilibrium occupancy. Because of their catalytic nature, low doses of PROTACs may reduce the probability of off-target effects to occur [77].
Eliminate the accumulation of drug targets
The binding of small-molecule inhibitors to target proteins cansues increased protein accumulation even in a relatively short amount of time [93]. It can be attributed to two reasons: 1). drug binding to target proteins can stabilize the protein structure, thereby extending their half-life, and 2). long-term inhibition will cause upregulation of its compensatory expressio. In general, the accumulation of target protein can be detrimental to the efficacy of drugs. Therefore, for these proteins that are insensitive to inhibitors, it’s extremely suitable to take PROTAC-mediated protein degradation. For example, BRD4, as one of the important bromodomain and extraterminal domain (BET) family members [94]. Researchers demonstrated that targeting BRD4 is an effective means of suppressing MYC-driven cancers [95]. However, the small molecule BRD4 inhibitor, JQ1 (Fig. 2e) and OTX015 resulted in robust protein accumulation, and high concentration of inhibitor is required to suppress downstream c-MYC. In 2015, Lu et al. designed a potent BRD4 PROTAC (ARV-825) by hijacking CRBN E3 ligase, which induced a rapid and sustained degradation of BRD4 protein in all BL cell lines [26]. This highlights the advantages of PROTAC over small-molecule inhibitors.
Others
In addition to the points mentioned above, PROTACs also have other advantages. The occurrence of acquired drug resistance is often closely related to point mutations that can decrease the affinity of the inhibitor to the target protein. PROTACs are able to overcome drug resistance issues via the complete elimination of the target mechanism [96]. Besides, the event-driven model of PROTACs do not require high drug exposure to reduce the risk of off-target effects [97]. Unlike other DNA-level protein knockout techniques, PROTACs enable for the rapid degradation of target proteins in vivo at the post-translational level. In the field of targeted protein degradation (TPD), besides UPS based PROTACs, lysosome-targeting chimeras (LYTACs), autophagy-targeting chimeras (AUTACs), and antibody-based PROTACs (AbTACs) degrade target proteins through lysomal. PROTACs cannot degrade extracellular and membrane proteins. Therefore, lysosome induced protein degradation can compensate for the lack of PROTACs. LYTACs were first proposed by Banik et al. and consist of a ligand binds lysosome-targeting receptors (LTRs) and a ligand binds extracellular or membrane protein [98]. Currently, only poly-serine-O-mannose-6-phosphonate (M6Pn) and N-acetyl galactosamine (Tri-GalNAC) were LTRs ligands available [99]. The LYTACs have been used to successfully degrade apolipoprotein E4, epidermal growth factor receptor (EGFR), programmed death protein ligand 1 (PD-L1), and CD71 [99]. However, due to the large molecular weight, poor cell permeability, and the possible emergence of immune response in vivo, further studies are needed [100]. In 2019, Takahashi et al. developed AUTACs based on the autophagic process for the degradation of endogenous proteins [101]. AUTACs are a bifunctional molecule with a linker joints POI ligand and autophagic recruitment tag. However, currently published AUTACs are inefficient due to the lack of efficient autophagy pathway recruiters. The autophagic process is extremely complex and may have an impact on natural autophagy, the mechanism of action of AUTACs remain unclear, so it need to be studied in depth [100]. AbTACs utilize bispecific antibodies, with one arm targeting POIs and the other targeting RNF43 E3 ligases [102]. AbTACs can induce POIs internalization and subsequent lysosomal degradation, but the the exact degradation mechanism remains to be confirmed.
The typical application of PROTACs for targeting diverse proteins
In theory, PROTACs can degrade almost all intracellular proteins if there is an appropriate small molecule that specifically binds with those POI, but not all degraders outperform small-molecule inhibitors. Here, we summarize some typical PROTAC molecules that have demonstrated obvious inhibition activities, several of which have advanced to the clinical trial stage.
PROTACs for targeting protein kinases
The human genome encodes over 500 protein kinases [103], making it the largest protein family. Currently, traditional small-molecule inhibitors are the primary treatment options for protein kinases related diseases. A majority of kinase inhibitors focused on the inhibition of receptor tyrosine kinase (RTK) [104]. However, the emergence of drug resistance impaired the clinical benefit, so it is urgent to apply novel therapeutic strategy to overcome this challenge.
In 2013, Crews’s group reported the earliest kinase PROTACs, which was used to target PI3K to block the human epidermal growth factor receptor 3 (ErbB3)–PI3K-Akt (protein kinase B) signal pathway [105]. This PROTAC is composed of two heterospecific peptide sequences recruiting POI and E3 ligase. An ErbB3-derived sequence that can bind to PI3K after it has been phosphorylated. Another sequence derived from hypoxia-inducible factor-1α (HIF1α) can be identified by VHL [105]. The two moieties were conjugated by a PEG linker, and a cell-penetrating sequence was incorporated to improve cell permeability. However, this PROTAC only display moderate potency because of poor permeability and unstable linker [106].
FAK, a tyrosine kinase, regulates many aspects of tumor progression (e.g., invasion, metastasis, and angiogenesis). The leading FAK kinase inhibitor defactinib, failed in clinical trials to treat malignant pleural mesothelioma stem cancer for the lack of efficacy. FAK also has a scaffolding role other than kinase, but kinase inhibitors cannot inhibit kinase-independent function. Cromm et al. designed PROTAC-3 (Fig. 2f) which could effectively induce the degradation of FAK with the IC50 of 6.5 nM [43]. PROTAC-3 is a bifunctional molecule consisting of defactinib and VHL ligand. It effectively inhibits FAK kinase-independent signaling and kinase-dependent signaling by efficient induction of degradation.
Bruton’s tyrosine kinase (BTK) is a member of the non-receptor cytoplasmic tyrosine kinase of the TEC family and a key regulator of the B cell receptor (BCR) signaling pathway, which plays a critical role in the life activities of B-cells like proliferation, survival, and differentiation [107, 108]. BTK is widely expressed in B cell neoplasms, and the clinical interventions are generally performed by inhibiting the kinase activity of BTK [109]. In 2013, FDA approved the first-in-class covalent inhibitor ibrutinib for the treatment of several B-cell malignancies. Ibrutinib binds covalently to Cysteine481 (C481) of BTK with IC50 of 0.5 nM [110, 111]. However, it has been revealed that a cysteine to serine mutation at position 481 of BTK (C481S) is what causes acquired resistance to ibrutinib [112]. So, induction of BTK protein degradation using PROTAC technology has emerged as a promising alternative approach. To date, four BTK degraders have entered clinical trials. They are NX-2127 (NCT04830137) and, NX-5948 (NCT05131022) from Nurix Therapeutics, HSK-29116 (NCT04861779) and BGB-16673 (NCT05006716) small molecule drugs from Haisco and BeiGene respectively. NX-2127 is an oral dual-target small molecule that possesses the activity of BTK degrader and IMiD neosubstrates degrader. A phase I clinical trial of NX-2127 is currently underway for the treatment of relapsed or refractory B-cell malignancies. Preclinical data have demonstrated that NX-2127 could potently induce the degradation of both ibrutinib-sensitive BTKWT (wild type) and ibrutinib-resistant BTKC481S in multiple cancer cell lines and human peripheral blood mononuclear cells (PBMCs) with the DC50 < 5 nM. Additionally, NX-2127 inhibited cell proliferation of BTKC481S in TMD8 cells more effectively than ibrutinib. NX-2127 exhibits immunomodulatory activity through comprised of thalidomide IMiD [113]. Krönke et al. revealed that lenalidomide causes selective ubiquitination and degradation of CRBN neosubstrates Aiolos (IKZF3) and Ikaros (IKZF1) [35]. Lazarian et al. have shown that the overexpression of IKZF3 is a driver of BTK inhibitor resistance in chronic lymphocytic leukemia (CLL) [114]. Therefore, NX-2127 combines BTK degradation with IKZF degradation is expected to enhance its anti-tumor activity. NX-5948 is another BTK degrader designed by Nurix Therapeutics. Unlike NX-2127, NX-5948 lacks immunomodulatory activity and has the ability to cross the blood brain barrier (BBB) in animal models. NX-5948 displayed similar performance that preclinical data have shown that NX-5948 induced the degradation of BTK (50% degradation efficiency at < 1 n M) in lymphoma cell lines and PBMCs [115].
PROTACs for targeting nuclear receptors
Nuclear receptors (NRs) belong to the family of transcription factors. Unlike other traditional transcription factors, its main function is to convert external the signal to transcriptional output [21]. A typical NR includes three domains: two structural domains that bind DNA and ligand respectively, and an unstructured N-terminal regulatory domain that is highly variable in terms of both sequence and size [116]. Ligand agonist binding confers a conformational change that results in exposure of the nuclear localization signal (NLS), which allows NR to translocate to the nucleus and bind the response elements. Small-molecule inhibitors that bind to ligand binding domain have been designed to activate or block the signal transduction function of nuclear receptors. However, small-molecule inhibitors have several disadvantages. For instance, our understanding of the concept of pure inhibitors is not clear, as continual AR antagonists prove to be agonists when the AR gene is overexpressed or mutated [117, 118]. In addition, some ligands for orphan NRs have not yet been identified, thus making it more complicated to target NRs to treat diseases. The advent of PROTAC technology has made it possible to target a wider range of NRs. NRs such as AR and ER participate in various important physiological progress in the body, and are closely related to prostate cancer and breast cancer. Therefore, a series of PROTACs targeting ER or AR have been developed.
AR signaling is critical in the development and maintenance of the normal function of prostate. AR not only plays a key role in the maintenance of musculoskeletal and male sex-related functions but also in the progression of prostate cancer [119]. Inhibition of AR function with AR antagonists such as enzalutamide and apalutamide is a common strategy in the treatment of prostate cancer [120]. Unfortunately, castration-resistant eventually occurs in patients with antiandrogen therapy [121]. PROTACs emerged as an alternative potential therapeutic approach to compensate for the shortcomings of AR inhibitors. Salami et al. synthesized a potent AR PROTAC ARCC-4 (Fig. 2g), which comprised of enzalutamide derivative and E3 ligand recruiting VHL. Compared with its parent inhibitor enzalutamide, ARCC-4 can effectively degrade AR and AR mutants caused by long-term use of clinical inhibitors, without leading to the presence of drug resistance [118]. It is well-known that ARV-110 (Fig. 2g) is the first AR-targeting PROTAC in clinical trial. The latest clinical trial data indicated that ARV-110 has an acceptable safety profile. The maximum tolerated dose (MTD) has not been established and the determination of the recommended phase 2 dose (RP2D) continues. In addition, ARV-110 has demonstrated antitumor activity in patients with metastatic castrate-resistant prostate cancer (mCRPC) following enzalutamid and/or abiraterone administration [44]. Recently, Wang’s group reporteded two highly potent and orally bioavailable AR PROTACs, ARD-2128 and ARD-2585 (Fig. 2g). ARD-2128 features an optimized AR antagonist linked to thalidomide via a rigid linker, achieving 67% oral bioavailability and better antitumor activity than enzalutamide in mice [151]. Here we introduced a specific and potent STAT3 PROTACs. Bai et al. reported the first STAT3 PROTAC SD-36 (Fig. 2i) that not only could effectively and specifically degraded STAT3 and has the antiproliferative activity of leukemia and lymphoma cell lines [147]. SD-36 consists of a selective STAT3 inhibitor SI-109 and lenalidomide and is a typical successful example of how PROTACs can be applied to target challenging proteins such as transcription factors.
Design and development of PROTACs
The degradation activity of PROTACs not only depends on the affinity of both ends to their respective target, but also relies on the formation of ternary complex that can form stable PPI. Currently, the construction of PROTACs largely relies on empirical analyses and structure–activity relationship (SAR) studies. However, synthetic difficulty presents significant limitations for rapid synthesis of a wealth of PROTAC compound libraries. By analyzing and summarizing published PROTACs structures, we will provide conventional strategies in PROTAC design to accelerate PROTACs discovery. In addition, we have listed some recently reported PROTACs that recruit traditional E3 ligases with corresponding degradation activity (Table 4).
E3 ligase and its ligand
Of the more than 600 ligases identified, only a few with small molecule ligands have been used for PROTAC targeting [163]. We list the commonly used E3 ligases and their ligands (Fig. 3). Cao et al. summarized and analyzed the structures of highly active PROTACs published over 20 years, and they found that CRBN, VHL, and cIAP ligands were used most frequently, of which CRBN accounted for 60.1%, VHL for 30.1%, and cIAP for 5.5% [164]. The main reason is that CRBN is widely expressed in tissues with high abundance and CRBN-based PROTACs have better degradation efficiency. In addition, CRBN ligands have better drug-like properties compared to the VHL ligand. PROTACs recruiting MDM2 and cIAP usually have high molecular weight and poor tissue permeability, indicating that the oral bioavailability may be a potential concern. Some other E3 ligases such as DCAF11 [27], DCAF15 [28], DCAF16 [29], KEAP1 [30], and RNF114 [31] etc., are less used for the following reasons: their ligands are derived from natural products with poor affinity, and are difficult to synthesize, and most of these E3 ligases are recruited by irreversible PROTACs, which have poor degradation activity and some potential toxicity. Of note, different recruited E3 ligases have been shown to induce different degrees of protein degradation [165]. The major reasons are as follow: different expression levels of E3 ligases in different cells may contribute to the different degradation efficiency. And some proteins have different degrees of selectivity for different E3 ligases. Therefore, in the process of designing the PROTACs, ligands targeting CRBN or VHL should be preferentially chosen, as these two E3 ligases have the widest range of applications. As an illustrative example, both ARV-110 and ARV-471 selected CRBN ligase as the E3 ligand. Here, we review the traditional E3 ligases and their ligands used in PROTAC design.
Linker design strategies of PROTACs
Type of linkers
Maple’s group built a database containing more than 400 published PROTACs to find a general principle that has been applied in PROTAC [166]. A summary of the linker structures in the database (Table 5) reveals that the frequently used linkers in PROTACs design are PEG and (un)saturated alkane chains with varying lengths up to now [81]. Due to the facile chemical synthesis feature, alkyl linkers are often used for the synthesis of PROTAC molecules to identify the optimal linker length. However, introduction of alkyl linkers might reduce the cell permeability of PROTACs due to their high hydrophobicity. Alkyl chains containing heteroatoms (oxygen atoms or nitrogen atoms) have improved hydrophilicity over alkyl chains alone. In addition, incorporating PEG chain can enhance the solubility and uptake of PROTACs by cells. More than half of the published PROTACs structure contained alkyl and PEG motifs. Alkyl, PEG, and glycol chains are incorporated into the PROTACs to increase the flexibility. However, their introduction can affect the pharmacokinetics (PK) properties of PROTACs. In recent years, linear linkers are gradually replaced by rigid linkers, such as alkynes and saturated heterocycles (piperazine and piperidine). The incorporation of aromatic rings or alkyne chains imparts some rigidity and promotes stable ternary complex formation. It also facilitates the solubility and cell permeability of PROTAC [206], suggesting that targeting PGES-2 may be a potential approach for INM-based antiviral PROTACs design. Desantis et al. designed four INM-based PROTACs, but the biological evaluation results showed that only two compounds were about 4.5-fold more potent than INM, as well as a wide-spectrum antiviral activity against the β-coronavirus HCoV-OC43 and α-coronavirus HCoV-229E [201].
Other PROTACs
In 2020, Rao et al. reported the first PROTAC of HMG-CoA reductase (HMGCR), which is the rate-limiting enzyme in the cholesterol biosynthetic pathway [207, 208]. They synthesized a series of PROTACs by tethering Atorvastatin and CRBN ligands. After optimization and screening, they ultimately found the most potent degrader P22A (Fig. 2m) with DC50 of 0.1 μM [209]. This PROTAC stressed the potential application for the treatment of hypercholesterolemia and cardiovascular disease. In addition, PROTACs are a promising therapeutic approach in other non-oncoproteins. Li et al. reported the first PROTAC that induced degradation of α1A-adrenergic receptor (α1A-AR) and is also the first PROTAC for G protein-coupled receptors (GPCRs) [210]. They connected α1A-AR inhibitor prazosin with pomalidomide by different linkers and finally found the potent compound 9c (Fig. 2m). 9c could inhibit the proliferation of PC-3 cells and cause tumor growth slowdown, which provided a new strategy for the treatment of prostate cancer. Hu et al. presented the first PROTAC of indoleamine 2,3-dioxygenase 1 (IDO1) [65]. IDO1 has been extensively reported as key immune checkpoint, which overexpressed in multiple cancers [211]. Hu et al. discovered the first PROTAC 2c (Fig. 2m) which induced the pronounced and sustained degradation of IDO1. Si et al. showed that PROTAC of hematopoietic progenitor kinase1 (HPK1) helped to improve CAR-T cell-based immunotherapy [212]. PROTAC technology is so widespread in the field of disease treatment, making it a powerful tool for drug discovery.
Disadvantages and future challenges of PROTAC
As an emerging technology, PROTAC has attracted great attention from academia and the pharmaceutical industry. The development of any new technology comes with various opportunities and challenges, and PROTAC is no exception. The prospect of potential opportunities and challenges for PROTAC will contribute to the research and development of targeted protein-degrading drugs. Although PROTAC has unique advantages over other drug discovery paradigm, it also has some disadvantages, which bring nonnegligible issues and challenges:
-
Pharmaceutical property: PROTAC molecule is more complex than traditional small-molecule drugs and has more potential metabolic sites, which affects the metabolic stability of PROTAC molecules. At the same time, traditional small-molecule inhibitors generally follow the “Rule of Five”, but most of the reported PROTACs tend to have a molecular weight greater than 700, resulting in poor permeability, low solubility and unsatisfactory oral bioavailability [213]. Therefore, how to improve physicochemical properties of PROTAC molecule will be the key to its successful drug formation if “the Rule of Five” are not satisfied.
-
Resistance: First, PROTACs can cause drug resistance through the change in the genome of the core component of the E3 ligase complex. Significantly reduced expression of CRBN gene or CUL2 gene can also cause resistance to PROTACs [214, 215]. Studies have shown that deletion of the CRBN genome is the main reason for myeloma cells to develop resistance to IMiDs. Secondly, the action of PROTAC depends on specific E3 ligase subtype, and the expression of specific E3 ligase limits the application of PROTAC in different cell types. Although the human genome encodes hundreds of E3 ubiquitin ligases, only a few E3 ligases and small molecule ligands have been used for PROTACs. Therefore, finding more kinds of E3 ligases for the research and development of PROTAC drugs might be the way to solve drug resistance [216].
-
“Hook effect” and “Off target”: How to avoid Hook effect and off-target effect is also a major challenge for PROTAC drugs development. The higher the concentration of drugs, the better degradation effect is not necessarily for PROTACs, which is often referred to as the “Hook effect”. In the research of PROTACs, it has been found that significantly higher concentration than DC50 will result in self-inhibition effect to compensate degradation efficiency, called “Hook effect” [217, 218]. In addition, the mechanism of off-target effects of PROTACs molecules have not been fully understood [219]. PROTACs can completely degrade target protein, thus inhibit all functions of target protein. However, in this process, normal protein may be accidentally injured, off-target effect and toxicity are also one of the biggest challenges. For example, studies have shown that thalidomide derivatives can cause degradation of transcription factors such as IKZF1, IKZF3 and GSTP1 [214]. Further studies found that the degradation of thalidomide derivatives on transcription factors such as GSPT1 was due to their “molecular glue” effect.
-
Target selection: To date, what targets are appropriate for PROTAC technology to achieve better benefits than small-molecule inhibitors are not fully understood and most of the target proteins of the PROTACs are part of the “druggable” protein. In fact, one of the greatest advantages of PROTAC technology is its potential to handle “undruggable” target. Because PROTAC technology only needs temporarily mediate the formation of ternary complexes, low affinity POI ligands can be incorporated into PROTAC molecules. Unfortunately, there are only few PROTAC molecules targeting “undruggable” proteins to date. Therefore, another challenge for PROTACs is the need to develop more molecules that target “undruggable” proteins and thus embody the advantages of PROTAC technology.
Discussion and conclusion
As an emerging paradigm for drug discovery, PROTACs have attracted great attention from academia and industry. Although PROTAC technology has many advantages in drug development, there are still many obstacles and challenges in the process of discovery and clinical application, such as off-target, cell permeability, stability, and large molecular weight, etc. In addition, the issues of oral bioavailability and drug integrity are also ongoing challenges for PROTAC drug development. It is worth noting that PROTAC still has many advantages in clinical application compared with other traditional small-molecule inhibitors. First, PROTAC plays a role by inducing the degradation of pathogenic proteins, so it can promote the degradation of multiple rounds of target proteins, assisting to eliminate off-target effects and accumulation of drug targets. PROTAC can also degrade some proteins that are considered “undruggable”, such as transcription factors. Secondly, PROTAC has the advantages of improving selectivity and specificity, overcoming drug resistance. In short, the current status of PROTAC drug development is the coexistence of both advantages and disadvantages, but how to solve these problems will be the key to the success of PROTAC drug development.
The discovery of efficient PROTAC molecules is a time-consuming and challenging process, such as the optimization of linker length and structure. It is urgent to summarize a general method for designing efficient PROTAC molecules. At present, the design and optimization of PROTAC mainly focus on the structure–activity relationships research of POI ligands and linker. Among them, linker is not only critical to the degradation activity of PROTACs, but also greatly affects the membrane permeability, metabolic stability and drug availability. Therefore, how to effectively design and link POI and E3 ligands is the key to the molecular design of PROTACs. Up to now, the principles guiding the design of linker, including length and composition, have not been fully understood. On the other hand, photo-PROTAC designed based on “photo control linkers” also has some advantages over traditional drugs, which is also introduced in this article. It is expected that the newly emerging photo-PRTOAC can become a leading way among PROTAC drugs. In this review, we summarized the general principles in the design of PROTAC, providing a systematic understanding for the research and design of PROTACs. In addition, E3 ligase is also crucial in the composition of the ternary complex. However, among the hundreds of E3 ligases encoded by the human genome, only a few E3 ligases are used in PROTACs, and the progress in discovering new E3 ligases and their ligands is far behind the research of PROTACs. So far, the majority of PROTACs induce target protein degradation by recruiting E3 ligases CRBN, VHL, MDM2 and IAP, and the research on PROTACs by only these E3 ligases is still far from enough. Therefore, it is necessary to explore more novel E3 ligases to accelerate the development of PROTACs. However, it can be predicted that the number of E3 ligands may increase significantly in the future, which will provide more options for the design of PROTACs.
PROTAC technology has been developed for nearly 20 years, and some molecules have entered clinical trials, which reveals the huge therapeutic potential of PROTACs in tumor, immune disease, neurodegenerative disease, cardiovascular disease and viral infection. There are also studies around the world using this technology to treat COVID-19. So far, two PROTAC drugs ARV-110 and ARV-471 have entered the phase II clinical trial, which are used to treat prostate cancer and breast cancer respectively. Although more than ten drugs are in clinical trials, clinical research data are still insufficient, and more clinical studies are needed to prove the prospects of PROTAC technology. With the deepening of research, these obstacles will be basically solved in the near future. Once more drugs enter the clinical application, it will open a new era of drug research and development.
Although there are still many obstacles and challenges to be overcame, PROTACs have great therapeutic potential with its unique advantages. It is believed that in the future, with the development of technology and in-depth research, the design and synthesis of PROTACs will be gradually optimized, which will eventually open up a broad road for the treatment of various diseases, and is expected to provide clinical therapeutic benefits in the near future. In a word, PROTAC technology not only provides a powerful tool for the research in the field of pharmaceutical chemistry, but also brings great hope for the development of clinical drugs in the future.
Availability of data and materials
Not applicable.
Abbreviations
- PROTACs:
-
Proteolysis targeting chimeras
- POI:
-
Protein of interest
- UPS:
-
Ubiquitin-protease system
- MetAP-2:
-
Methionine aminopeptidase-2
- β-TRCP:
-
β-Transducin repeat-containing protein
- ER:
-
Estrogen
- AR:
-
Androgen
- DHT:
-
Dihydrotestosterone
- VHL:
-
Von Hippel-Lindau
- FKBP12:
-
FK506 binding protein 12
- MDM2:
-
Mouse double minute 2
- cIAP:
-
Cell inhibitor of apoptosis protein
- CRBN:
-
Cereblon
- IMiDs:
-
Immunomodulatory drugs
- BL:
-
Burkitt’s lymphoma
- Hyp:
-
Hydroxyproline
- ALK:
-
Anaplastic lymphoma kinase
- FAK:
-
Focal adhesion kinase
- RNAi:
-
RNA interference
- SGK:
-
Serum and glucocorticoid-induced protein kinase
- PI3K:
-
Phosphoinositide 3-kinase
- MOA:
-
Mode of action
- RTK:
-
Receptor tyrosine kinase
- HIF1α:
-
Hypoxia-inducible factor-1α
- PEG:
-
Polyethylene glycol
- BTK:
-
Bruton’s tyrosine kinase
- BCR:
-
B cell receptor
- C481:
-
Cysteine481
- WT:
-
Wild type
- IKZF3:
-
Ikaros family zinc finger 3
- PBMCs:
-
Peripheral blood mononuclear cells
- CLL:
-
Chronic lymphocytic leukemia
- NRs:
-
Nuclear receptors
- NLS:
-
Nuclear localization signal
- MTD:
-
Maximum tolerated dose
- RP2D:
-
Recommended phase 2 dose
- mCRPC:
-
Metastatic castrate-resistant prostate cancer
- PSA:
-
Prostate specific antigen
- ER+:
-
Estrogen receptor-positive
- HER2+:
-
Human epidermal growth factor receptor 2 positive
- SERD:
-
Selective estrogen receptor degraders
- TFs:
-
Transcriptional factors
- STAT3:
-
Signal transducer and activator of transcription 3
- SH2:
-
Src-homology 2
- PPI:
-
Protein–protein interaction
- SAR:
-
Structure–activity relationship
- DMNB:
-
4,5-Dimethoxy-2-nitrobenzyl
- CLIPTACs:
-
In-cell click-formed proteolysis targeting chimeras
- Tz:
-
Tetrazine
- TCO:
-
Trans-cyclo-octene
- ERK1/2:
-
Extracellular regulated protein kinase
- CMGCs:
-
CMGC kinase group
- AD:
-
Alzheimer’s disease
- Aβ:
-
Amyloid-β
- HD:
-
Huntington’s disease
- mHtt:
-
Mutant huntingtin
- IRAK4:
-
Interleukin-1 receptor-associated kinase 4
- TLRs:
-
Toll-like receptors
- 1L-1R:
-
Interleukin1 receptors
- DD:
-
Death domain
- HDACs:
-
Histone deacetylases
- HCV:
-
Hepatitis C virus
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- INM:
-
Indomethacin
- PGES-2:
-
Human prostaglandin E synthase type 2
- HMGCR:
-
HMG-CoA reductase
- RL:
-
Reinforcement Learning
- MWs:
-
Molecular weights
- BRD4:
-
Bromodomain-containing protein 4
- α1A-AR:
-
α1A-adrenergic receptor
- IDO1:
-
Indoleamine 2,3-dioxygenase 1
- HPK1:
-
Hematopoietic progenitor kinase1
- GPCRs:
-
G protein-coupled receptors
- EGFR:
-
Epidermal growth factor receptor
- TPD:
-
Targeted protein degradation
- RAS:
-
RAt Sarcoma
- LYTACs:
-
Lysosome-targeting chimeras
- AUTACs:
-
Autophagy-targeting chimeras
- AbTACs:
-
Antibody-based PROTACs
- LTRs:
-
Lysosome-targeting receptors
- M6Pn:
-
Poly-serine-O-mannose-6-phosphonate
- GalNAC:
-
N-acetyl galactosamine
- PD-L1:
-
Programmed death protein ligand 1
References
Sakamoto KM, Kim KB, Kumagai A, Mercurio F, Crews CM, Deshaies RJ. Protacs: chimeric molecules that target proteins to the Skp1–Cullin–F box complex for ubiquitination and degradation. Proc Natl Acad Sci. 2001;98(15):8554–9. https://doi.org/10.1073/pnas.141230798.
Schneekloth AR, Pucheault M, Tae HS, Crews CM. Targeted intracellular protein degradation induced by a small molecule: En route to chemical proteomics. Bioorg Med Chem Lett. 2008;18(22):5904–8. https://doi.org/10.1016/j.bmcl.2008.07.114.
Yao T, **ao H, Wang H, Xu X. Recent advances in PROTACs for drug targeted protein research. Int J Mol Sci. 2022;23(18):10328. https://doi.org/10.3390/ijms231810328.
Buckley DL, Buckley DL, Crews CM. Small-molecule control of intracellular protein levels through modulation of the ubiquitin proteasome system. Angew Chem. 2014;53(9):2312–30. https://doi.org/10.1002/anie.201307761.
Gadd MS, Testa A, Lucas X, Chan K-H, Chen W, Lamont DJ, et al. Structural basis of PROTAC cooperative recognition for selective protein degradation. Nat Chem Biol. 2017;13(5):514–21. https://doi.org/10.1038/nchembio.2329.
Finley D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu Rev Biochem. 2009;78:477–513. https://doi.org/10.1146/annurev.biochem.78.081507.101607.
Hipp MS, Kasturi P, Hartl FU. The proteostasis network and its decline in ageing. Nat Rev Mol Cell Biol. 2019;20(7):421–35. https://doi.org/10.1038/s41580-019-0101-y.
Kliza K, Husnjak K. Resolving the complexity of ubiquitin networks. Front Mol Biosci. 2020;7:21. https://doi.org/10.3389/fmolb.2020.00021.
Konstantinidou M, Li J, Zhang B, Wang Z, Shaabani S, Brake FT, et al. PROTACs– a game-changing technology. Expert Opin Drug Discov. 2019;14(12):1255–68. https://doi.org/10.1080/17460441.2019.1659242.
Cromm PM, Crews CM. Targeted protein degradation: from chemical biology to drug discovery. Chem Biol. 2017;24(9):1181–90. https://doi.org/10.1016/j.chembiol.2017.05.024.
Crews CM, Hu Z. Recent developments in PROTAC-mediated protein degradation: from bench to clinic. ChemBioChem. 2021. https://doi.org/10.1002/cbic.202100270.
Komander D, Rape M. The ubiquitin code. Annu Rev Biochem. 2012;81:203–29. https://doi.org/10.1146/annurev-biochem-060310-170328.
Chen Y, ** J. The application of ubiquitin ligases in the PROTAC drug design. Acta Biochim Biophys Sin. 2020;52(7):776–90. https://doi.org/10.1093/abbs/gmaa053.
Burslem GM, Crews CM. Proteolysis-targeting chimeras as therapeutics and tools for biological discovery. Cell. 2020;181(1):102–14. https://doi.org/10.1016/j.cell.2019.11.031.
Salami J, Crews CM. Waste disposal-an attractive strategy for cancer therapy. Science. 2017;355(6330):1163–7. https://doi.org/10.1126/science.aam7340.
Eder J, Herrling P. Trends in modern drug discovery. Handb Exp Pharmacol. 2015;232:3–22. https://doi.org/10.1007/164_2015_20.
Valeur E, Jimonet P. New modalities, technologies, and partnerships in probe and lead generation: enabling a mode-of-action centric paradigm. J Med Chem. 2018;61(20):9004–29. https://doi.org/10.1021/acs.jmedchem.8b00378.
Kargbo RB. PROTAC-mediated degradation of KRAS protein for anticancer therapeutics. ACS Med Chem Lett. 2020;11(1):5–6. https://doi.org/10.1021/acsmedchemlett.9b00584.
Hopkins AL, Groom CR. The druggable genome. Nat Rev Drug Discov. 2002;1(9):727–30. https://doi.org/10.1038/nrd892.
Neklesa TK, Snyder L, Willard RR, Vitale N, Pizzano J, Gordon DA, et al. ARV-110: an oral androgen receptor PROTAC degrader for prostate cancer. J Clin Oncol. 2019;37:259–259. https://doi.org/10.1200/jco.2019.37.7_suppl.259.
Flanagan JJ, Neklesa TK. Targeting nuclear receptors with PROTAC degraders. Mol Cell Endocrinol. 2019;493:110452. https://doi.org/10.1016/j.mce.2019.110452.
Sakamoto KM, Kim KB, Verma R, Ransick A, Stein B, Crews CM, et al. Development of Protacs to target cancer-promoting proteins for ubiquitination and degradation. Mol Cell Proteomics. 2003;2(12):1350–8. https://doi.org/10.1074/mcp.T300009-MCP200.
Schneekloth JS, Fonseca FN, Koldobskiy M, Mandal A, Deshaies R, Sakamoto K, et al. Chemical genetic control of protein levels: selective in vivo targeted degradation. J Am Chem Soc. 2004;126(12):3748–54. https://doi.org/10.1021/ja039025z.
Hon W-C, Wilson MI, Harlos K, Claridge TDW, Schofield CJ, Pugh CW, et al. Structural basis for the recognition of hydroxyproline in HIF-1α by pVHL. Nature. 2002;417(6892):975–8. https://doi.org/10.1038/nature00767.
Itoh Y, Ishikawa M, Naito M, Hashimoto Y. Protein knockdown using methyl bestatin−ligand hybrid molecules: design and synthesis of inducers of ubiquitination-mediated degradation of cellular retinoic acid-binding proteins. J Am Chem Soc. 2010;132(16):5820–6. https://doi.org/10.1021/ja100691p.
Lu J, Qian Y, Altieri M, Dong H, Wang J, Raina K, et al. Hijacking the E3 ubiquitin ligase cereblon to efficiently target BRD4. Chem Biol. 2015;22(6):755–63. https://doi.org/10.1016/j.chembiol.2015.05.009.
Zhang X, Luukkonen LM, Eissler CL, Crowley VM, Yamashita Y, Schafroth MA, et al. DCAF11 supports targeted protein degradation by electrophilic proteolysis-targeting chimeras. J Am Chem Soc. 2021;143(13):5141–9. https://doi.org/10.1021/jacs.1c00990.
Li L, Mi D, Pei H, Duan Q, Wang X, Zhou W, et al. In vivo target protein degradation induced by PROTACs based on E3 ligase DCAF15. Signal Transduct Target Ther. 2020;5(1):129. https://doi.org/10.1038/s41392-020-00245-0.
Zhang X, Crowley VM, Wucherpfennig TG, Dix MM, Cravatt BF. Electrophilic PROTACs that degrade nuclear proteins by engaging DCAF16. Nat Chem Biol. 2019;15(7):737–46. https://doi.org/10.1038/s41589-019-0279-5.
Lu M-C, Liu T, Jiao Q, Ji J-A, Tao M, Liu Y, et al. Discovery of a Keap1-dependent peptide PROTAC to knockdown Tau by ubiquitination-proteasome degradation pathway. Eur J Med Chem. 2018;146:251–9. https://doi.org/10.1016/j.ejmech.2018.01.063.
Yang Y, Zhou C, Wang Y, Liu W, Liu C, Wang L, et al. The E3 ubiquitin ligase RNF114 and TAB1 degradation are required for maternal-to-zygotic transition. EMBO Rep. 2017;18(2):205–16. https://doi.org/10.15252/embr.201642573.
Ishida T, Ciulli A. E3 ligase ligands for PROTACs: how they were found and how to discover new ones. SLAS Discov Adv Life Sci R D. 2020;26(4):484–502. https://doi.org/10.1177/2472555220965528.
Nguyen TV. USP15 antagonizes CRL4CRBN-mediated ubiquitylation of glutamine synthetase and neosubstrates. Proc Natl Acad Sci U S A. 2021;118(40):e2111391118. https://doi.org/10.1073/pnas.2111391118.
Lopez-Girona A, Mendy D, Ito T, Miller K, Gandhi A, Kang J, et al. Cereblon is a direct protein target for immunomodulatory and antiproliferative activities of lenalidomide and pomalidomide. Leukemia. 2012;26(11):2326–35. https://doi.org/10.1038/leu.2012.119.
Krönke J, Udeshi ND, Narla A, Grauman P, Hurst SN, McConkey M, et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science. 2014;343(6168):301–5. https://doi.org/10.1126/science.1244851.
Fischer ES, Böhm K, Lydeard JR, Yang H, Stadler MB, Cavadini S, et al. Structure of the DDB1-CRBN E3 ubiquitin ligase in complex with thalidomide. Nature. 2014;512(7512):49–53. https://doi.org/10.1038/nature13527.
Chamberlain PP, Lopez-Girona A, Miller K, Carmel G, Pagarigan B, Chie-Leon B, et al. Structure of the human Cereblon–DDB1–lenalidomide complex reveals basis for responsiveness to thalidomide analogs. Nat Struct Mol Biol. 2014;21(9):803–9. https://doi.org/10.1038/nsmb.2874.
Lu G, Middleton RE, Sun H, Naniong M, Ott CJ, Mitsiades CS, et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science. 2014;343(6168):305–9. https://doi.org/10.1126/science.1244917.
Lee J, Lee Y, Jung YM, Park JH, Yoo HS, Park J. Discovery of E3 ligase ligands for target protein degradation. Molecules. 2022;27(19):6515. https://doi.org/10.3390/molecules27196515.
Buckley DL, Gustafson JL, Van Molle I, Roth AG, Tae HS, Gareiss PC, et al. Small-molecule inhibitors of the interaction between the E3 ligase VHL and HIF1α. Angew Chem Int Ed Engl. 2012;51(46):11463–7. https://doi.org/10.1002/anie.201206231.
Zhao Q, Ren C, Liu L, Chen J, et al. Discovery of SIAIS178 as an effective BCR-ABL degrader by recruiting Von Hippel-Lindau (VHL) E3 ubiquitin ligase. J Med Chem. 2019;62(20):9281–98. https://doi.org/10.1021/acs.jmedchem.9b01264.
Kang CH, Lee DH, Lee CO, Du Ha J, Park CH, Hwang JY. Induced protein degradation of anaplastic lymphoma kinase (ALK) by proteolysis targeting chimera (PROTAC). Biochem Biophys Res Commun. 2018;505(2):542–7. https://doi.org/10.1016/j.bbrc.2018.09.169.
Cromm PM, Samarasinghe KT, Hines J, Crews CM. Addressing kinase-independent functions of Fak via PROTAC-mediated degradation. J Am Chem Soc. 2018;140(49):17019–26. https://doi.org/10.1021/jacs.8b08008.
Petrylak DP, Gao X, Vogelzang NJ, Garfield MH, Taylor IW, Taylor I, et al. First-in-human phase I study of ARV-110, an androgen receptor (AR) PROTAC degrader in patients (pts) with metastatic castrate-resistant prostate cancer (mCRPC) following enzalutamide (ENZ) and/or abiraterone (ABI). J Clin Oncol. 2020;38:3500–3500. https://doi.org/10.1200/jco.2020.38.15_suppl.3500.
**e H, Liu J, Alem Glison DM, Fleming JB. The clinical advances of proteolysis targeting chimeras in oncology. Explor Target Anti-Tumor Ther. 2021;2(6):511–21. https://doi.org/10.37349/etat.2021.00061.
He Y, Koch R, Budamagunta V, Zhang P, Zhang X, Khan S, et al. DT2216-a Bcl-xL-specific degrader is highly active against Bcl-xL-dependent T cell lymphomas. J Hematol OncolJ Hematol Oncol. 2020;13(1):95. https://doi.org/10.1186/s13045-020-00928-9.
You I, Erickson EC, Donovan KA, Eleuteri NA, Fischer ES, Gray NS, et al. Discovery of an AKT degrader with prolonged inhibition of downstream signaling. Cell Chem Biol. 2020;27(1):66-73.e7. https://doi.org/10.1016/j.chembiol.2019.11.014.
Kargbo RB. PROTAC compounds targeting α-synuclein protein for treating neurogenerative disorders: Alzheimer’s and Parkinson’s diseases. ACS Med Chem Lett. 2020;11(6):1086–7. https://doi.org/10.1021/acsmedchemlett.0c00192.
Gasic I, Groendyke BJ, Nowak RP, Yuan JC, Kalabathula J, Fischer ES, et al. Tubulin resists degradation by cereblon-recruiting PROTACs. Cells. 2020;9(5):1083. https://doi.org/10.3390/cells9051083.
Shi W, Feng Z, Chi F, Zhou J, Qiu Q, Jiang Y, et al. Structure-based discovery of receptor tyrosine kinase AXL degraders with excellent anti-tumor activity by selectively degrading AXL and inducing methuosis. Eur J Med Chem. 2022;234:114253. https://doi.org/10.1016/j.ejmech.2022.114253.
Wang Z, He N, Guo Z, Niu C, Song T, Guo Y, et al. Proteolysis targeting chimeras for the selective degradation of Mcl-1/Bcl-2 derived from nonselective target binding ligands. J Med Chem. 2019;62(17):8152–63. https://doi.org/10.1021/acs.jmedchem.9b00919.
Zhang X, Thummuri D, He Y, Liu X, Zhang P, Zhou D, et al. Utilizing PROTAC technology to address the on-target platelet toxicity associated with inhibition of BCL-X L. Chem Commun. 2019;55(98):14765–8. https://doi.org/10.1039/C9CC07217A.
Xue G, Chen J, Liu L, Zhou D, Zuo Y, Fu T, et al. Protein degradation through covalent inhibitor-based PROTACs. Chem Commun. 2020;56(10):1521–4. https://doi.org/10.1039/C9CC08238G.
Nowak RP, DeAngelo SL, Buckley D, He Z, Donovan KA, An J, et al. Plasticity in binding confers selectivity in ligand-induced protein degradation. Nat Chem Biol. 2018;14(7):706–14. https://doi.org/10.1038/s41589-018-0055-y.
Buhimschi AD, Armstrong HA, Toure M, Jaime-Figueroa S, Chen TL, Lehman AM, et al. Targeting the C481S ibrutinib-resistance mutation in Bruton’s tyrosine kinase using PROTAC-mediated degradation. Biochemistry. 2018;57(26):3564–75. https://doi.org/10.1021/acs.biochem.8b00391.
Chi JJ, Li H, Zhou Z, Izquierdo-Ferrer J, Xue Y, Wavelet CM, et al. A novel strategy to block mitotic progression for targeted therapy. EBioMedicine. 2019;49:40–54. https://doi.org/10.1016/j.ebiom.2019.10.013.
Zhou F, Chen L, Cao C, Yu J, Luo X, Zhou P, et al. Development of selective mono or dual PROTAC degrader probe of CDK isoforms. Eur J Med Chem. 2020;187:111952. https://doi.org/10.1016/j.ejmech.2019.111952.
Zhao B, Burgess K. PROTACs suppression of CDK4/6, crucial kinases for cell cycle regulation in cancer. Chem Commun. 2019;55(18):2704–7. https://doi.org/10.1039/C9CC00163H.
Steinebach C, Lindner S, Udeshi ND, Mani DC, Kehm H, Köpff S, et al. Homo-PROTACs for the chemical knockdown of cereblon. ACS Chem Biol. 2018;13(9):2771–82. https://doi.org/10.1021/acschembio.8b00693.
Zhou L, Chen W, Cao C, Shi Y, Ye W, Hu J, et al. Design and synthesis of α-naphthoflavone chimera derivatives able to eliminate cytochrome P450 (CYP)1B1-mediated drug resistance via targeted CYP1B1 degradation. Eur J Med Chem. 2020;189:112028. https://doi.org/10.1016/j.ejmech.2019.112028.
Potjewyd F, Turner A-MW, Beri J, Rectenwald JM, Norris-Drouin JL, Cholensky SH, et al. Degradation of polycomb repressive complex 2 with an EED-targeted bivalent chemical degrader. Cell Chem Biol. 2020;27(1):47-56.e15. https://doi.org/10.1016/j.chembiol.2019.11.006.
Burslem GM, Smith BE, Lai AC, Jaime-Figueroa S, McQuaid DC, et al. The advantages of targeted protein degradation over inhibition: an RTK case study. Chem Biol. 2017;25(1):67. https://doi.org/10.1016/j.chembiol.2017.09.009.
Powell CE, Gao Y, Tan L, Donovan KA, Nowak RP, Loehr A, et al. Chemically induced degradation of anaplastic lymphoma kinase (ALK). J Med Chem. 2018;61(9):4249–55. https://doi.org/10.1021/acs.jmedchem.7b01655.
Smalley JP, Adams GE, Millard CJ, Song Y, Norris JKS, Schwabe JWR, et al. PROTAC-mediated degradation of class I histone deacetylase enzymes in corepressor complexes. Chem Commun. 2020;56(32):4476–9. https://doi.org/10.1039/D0CC01485K.
Hu M, Zhou W, Wang Y, Yao D, Ye T, Yao Y, et al. Discovery of the first potent proteolysis targeting chimera (PROTAC) degrader of indoleamine 2,3-dioxygenase 1. Acta Pharm Sin B. 2020;10(10):1943–53. https://doi.org/10.1016/j.apsb.2020.02.010.
Li Y, Yang J, Aguilar A, McEachern D, Przybranowski S, Liu L, et al. Discovery of MD-224 as a first-in-class, highly potent, and efficacious proteolysis targeting chimera murine double minute 2 degrader capable of achieving complete and durable tumor regression. J Med Chem. 2019;62(2):448–66. https://doi.org/10.1021/acs.jmedchem.8b00909.
Silva MC, Ferguson FM, Cai Q, Donovan KA, Nandi G, Patnaik D, et al. Targeted degradation of aberrant tau in frontotemporal dementia patient-derived neuronal cell models. eLife. 2019;8:e45457. https://doi.org/10.7554/elife.45457.
Maniaci C, Hughes SJ, Testa A, Chen W, Lamont DJ, Rocha S, et al. Homo-PROTACs: bivalent small-molecule dimerizers of the VHL E3 ubiquitin ligase to induce self-degradation. Nat Commun. 2017;8(1):830. https://doi.org/10.1038/s41467-017-00954-1.
Li Z, Pinch BJ, Olson CM, Donovan KA, Nowak RP, Mills CE, et al. Development and characterization of a Wee1 kinase degrader. Cell Chem Biol. 2020;27(1):57-65.e9. https://doi.org/10.1016/j.chembiol.2019.10.013.
Hu M, Li Y, Li J, Zhou H, Liu C, Liu Z, et al. Discovery of potent and selective HER2 PROTAC degrader based Tucatinib with improved efficacy against HER2 positive cancers. Eur J Med Chem. 2022;244:114775. https://doi.org/10.1016/j.ejmech.2022.114775.
Holohan C, Van Schaeybroeck S, Longley DB, Johnston PG. Cancer drug resistance: an evolving paradigm. Nat Rev Cancer. 2013;13(10):714–26. https://doi.org/10.1038/nrc3599.
Zou Y, Ma D, Wang Y. The PROTAC technology in drug development: the PROTAC technology in drug development. Cell Biochem Funct. 2019;37(1):21–30. https://doi.org/10.1002/cbf.3369.
Martín-Acosta P, **ao X. PROTACs to address the challenges facing small molecule inhibitors. Eur J Med Chem. 2021;210: 112993. https://doi.org/10.1016/j.ejmech.2020.112993.
Wang Y, Jiang X, Feng F, Liu W, Sun H. Degradation of proteins by PROTACs and other strategies. Acta Pharm Sin B. 2020;10(2):207–38. https://doi.org/10.1016/j.apsb.2019.08.001.
Bond MJ, Crews CM. Proteolysis targeting chimeras (PROTACs) come of age: entering the third decade of targeted protein degradation. RSC Chem Biol. 2021;2(3):725–42. https://doi.org/10.1039/D1CB00011J.
Lazo JS, Sharlow ER. Drugging undruggable molecular cancer targets. Annu Rev Pharmacol Toxicol. 2016;56:23–40. https://doi.org/10.1146/annurev-pharmtox-010715-103440.
Neklesa TK, Winkler JD, Crews CM. Targeted protein degradation by PROTACs. Pharmacol Ther. 2017;174:138–44. https://doi.org/10.1016/j.pharmthera.2017.02.027.
Moon S, Lee B-H. Chemically induced cellular proteolysis: an emerging therapeutic strategy for undruggable targets. Mol Cells. 2018;41(11):933–42. https://doi.org/10.14348/molcells.2018.0372.
Nero TL, Morton CJ, Holien JK, Wielens J, Parker MW. Oncogenic protein interfaces: small molecules, big challenges. Nat Rev Cancer. 2014;14(4):248–62. https://doi.org/10.1038/nrc3690.
Bondeson DP, Smith BE, Burslem GM, Buhimschi AD, Hines J, et al. Lessons in PROTAC design from selective degradation with a promiscuous warhead. Chem Biol. 2017;25(1):78. https://doi.org/10.1016/j.chembiol.2017.09.010.
Troup RI, Fallan C, Baud MGJ. Current strategies for the design of PROTAC linkers: a critical review. Explor Target Antitumor Ther. 2020;1(5):273–312. https://doi.org/10.37349/etat.2020.00018.
Moore AR, Rosenberg SC, McCormick F, Malek S. RAS-targeted therapies: is the undruggable drugged? Nat Rev Drug Discov. 2020;19(8):533–52. https://doi.org/10.1038/s41573-020-0068-6.
Zeng M, **ong Y, Safaee N, Nowak RP, Donovan KA, et al. Exploring targeted degradation strategy for oncogenic KRASG12C. Chem Biol. 2020;27(1):19. https://doi.org/10.1016/j.chembiol.2019.12.006.
Tanaka N, Lin JJ, Li C, Ryan MB, Zhang J, Kiedrowski LA, et al. Clinical acquired resistance to KRASG12C inhibition through a novel KRAS switch-II pocket mutation and polyclonal alterations converging on RAS-MAPK reactivation. Cancer Discov. 2021;11(8):1913–22. https://doi.org/10.1158/2159-8290.CD-21-0365.
Bond MJ, Chu L, Nalawansha DA, Li K, Crews CM. Targeted degradation of oncogenic KRASG12C by VHL-recruiting PROTACs. ACS Cent Sci. 2020;6(8):1367–75. https://doi.org/10.1021/acscentsci.0c00411.
Bruhn MA, Pearson RB, Hannan RD, Sheppard KE. Second AKT: the rise of SGK in cancer signalling. Growth Factors. 2010;28(6):394–408. https://doi.org/10.3109/08977194.2010.518616.
Halland N, Schmidt F, Weiss T, Saas J, Li Z, Czech J, et al. Discovery of N-[4-(1H-Pyrazolo[3,4-b]pyrazin-6-yl)-phenyl]-sulfonamides as highly active and selective SGK1 inhibitors. ACS Med Chem Lett. 2015;6(1):73–8. https://doi.org/10.1021/ml5003376.
Sherk AB, Frigo DE, Schnackenberg CG, Bray JD, La** NJ, Trizna W, et al. Development of a small-molecule serum- and glucocorticoid-regulated kinase-1 antagonist and its evaluation as a prostate cancer therapeutic. Cancer Res. 2008;68(18):7475–83. https://doi.org/10.1158/0008-5472.CAN-08-1047.
Gong G, Wang K, Dai X, Zhou Y, Basnet R, Chen Y, et al. Identification, structure modification, and characterization of potential small-molecule SGK3 inhibitors with novel scaffolds. Acta Pharmacol Sin. 2018;39:1902–12. https://doi.org/10.1038/s41401-018-0087-6.
Tovell H, Testa A, Zhou H, Shpiro N, Crafter C, Ciulli A, et al. Design and characterization of SGK3-PROTAC1, an isoform specific SGK3 kinase PROTAC degrader. ACS Chem Biol. 2019;14(9):2024–34. https://doi.org/10.1021/acschembio.9b00505.
Adjei AA. What is the right dose? The elusive optimal biologic dose in phase I clinical trials. J Clin Oncol Off J Am Soc Clin Oncol. 2006;24(25):4054–5. https://doi.org/10.1200/JCO.2006.07.4658.
Bondeson DP, Mares A, Smith IED, Eunhwa K, Campos SA, Miah AH, et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat Chem Biol. 2015;11(8):611–7. https://doi.org/10.1038/nchembio.1858.
Martinez Molina D, Nordlund P. The cellular thermal shift assay: a novel biophysical assay for in situ drug target engagement and mechanistic biomarker studies. Annu Rev Pharmacol Toxicol. 2016;56(1):141–61. https://doi.org/10.1146/annurev-pharmtox-010715-103715.
Wu S, Jiang Y, Hong Y, Chu X, Zhang Z, Tao Y, et al. BRD4 PROTAC degrader ARV-825 inhibits T-cell acute lymphoblastic leukemia by targeting “Undruggable” Myc-pathway genes. Cancer Cell Int. 2021;21(1):230. https://doi.org/10.1186/s12935-021-01908-w.
Gabay M, Li Y, Felsher DW. MYC activation is a hallmark of cancer initiation and maintenance. Cold Spring Harb Perspect Med. 2014;4(6):a014241. https://doi.org/10.1101/cshperspect.a014241.
Burke MR, Smith AR, Zheng G. Overcoming cancer drug resistance utilizing PROTAC technology. Front Cell Dev Biol. 2022; 10. https://doi.org/10.3389/fcell.2022.872729.
Pettersson M, Crews CM. PROteolysis TArgeting Chimeras (PROTACs) - past, present and future. Drug Discov Today Technol. 2019;31:15–27. https://doi.org/10.1016/j.ddtec.2019.01.002.
Banik SM, Pedram K, Wisnovsky S, Ahn G, Riley NM, Bertozzi CR. Lysosome-targeting chimaeras for degradation of extracellular proteins. Nature. 2020;584(7820):291–7. https://doi.org/10.1038/s41586-020-2545-9.
Pei J, Wang G, Feng L, Zhang J, Jiang T, Sun Q, et al. Targeting lysosomal degradation pathways: new strategies and techniques for drug discovery. J Med Chem. 2021;64(7):3493–507. https://doi.org/10.1021/acs.jmedchem.0c01689.
Ding Y, Fei Y, Lu B. Emerging new concepts of degrader technologies. Trends Pharmacol Sci. 2020;41(7):464–74. https://doi.org/10.1016/j.tips.2020.04.005.
Takahashi D, Moriyama J, Nakamura T, Miki E, Takahashi E, Sato A, et al. AUTACs: cargo-specific degraders using selective autophagy. Mol Cell. 2019;76(5):797-810.e10. https://doi.org/10.1016/j.molcel.2019.09.009.
Cotton AD, Nguyen DP, Gramespacher JA, Seiple IB, Wells JA. Development of antibody-based PROTACs for the degradation of the cell-surface immune checkpoint protein PD-L1. J Am Chem Soc. 2021;143(2):593–8. https://doi.org/10.1021/jacs.0c10008.
Tan L, Gray NS. When kinases meet PROTACs: when kinases meet PROTACs †. Chin J Chem. 2018;36(10):971–7. https://doi.org/10.1002/cjoc.201800293.
Ferguson FM, Gray NS. Kinase inhibitors: the road ahead. Nat Rev Drug Discov. 2018;17(5):353–77. https://doi.org/10.1038/nrd.2018.21.
Hines J, Gough JD, Corson TW, Crews CM. Posttranslational protein knockdown coupled to receptor tyrosine kinase activation with phosphoPROTACs. Proc Natl Acad Sci U S A. 2013;110(22):8942–7. https://doi.org/10.1073/pnas.1217206110.
Henning RK, Varghese JO, Das S, Nag A, Tang G, Tang K, et al. Degradation of Akt using protein-catalyzed capture agents: DEGRADATION OF AKT USING PCC AGENTS. J Pept Sci. 2016;22(4):196–200. https://doi.org/10.1002/psc.2858.
Sun Y, Zhao X, Ding N, Gao H, Gao H, Wu Y, et al. PROTAC-induced BTK degradation as a novel therapy for mutated BTK C481S induced ibrutinib-resistant B-cell malignancies. Cell Res. 2018;28(7):779–81. https://doi.org/10.1038/s41422-018-0055-1.
del Mar Noblejas-López M, Nieto-Jiménez C, Burgos M, Gómez-Juárez M, Montero JC, Esparís-Ogando A, et al. Activity of BET-proteolysis targeting chimeric (PROTAC) compounds in triple negative breast cancer. J Exp Clin Cancer Res. 2019;38(1):383–383. https://doi.org/10.1186/s13046-019-1387-5.
Hendriks RW, Yuvaraj S, Kil LP. Targeting Bruton’s tyrosine kinase in B cell malignancies. Nat Rev Cancer. 2014;14(4):219–32. https://doi.org/10.1038/nrc3702.
Pan Z, Scheerens H, Li S-J, Schultz BE, Sprengeler PA, Burrill LC, et al. Discovery of selective irreversible inhibitors for Bruton’s tyrosine kinase. ChemMedChem. 2007;2(1):58–61. https://doi.org/10.1002/cmdc.200600221.
Honigberg LA, Smith AM, Sirisawad M, Verner E, Loury D, Chang B, et al. The Bruton tyrosine kinase inhibitor PCI-32765 blocks B-cell activation and is efficacious in models of autoimmune disease and B-cell malignancy. Proc Natl Acad Sci U S A. 2010;107(29):13075–80. https://doi.org/10.1073/pnas.1004594107.
Woyach JA, Furman RR, Liu T-M, Ozer HG, Zapatka M, Ruppert AS, et al. Resistance mechanisms for the Bruton’s tyrosine kinase inhibitor ibrutinib. N Engl J Med. 2014;370(24):2286–94. https://doi.org/10.1056/NEJMoa1400029.
Haertle L, Barrio S, Munawar U, Han S, Zhou X, Vogt C, et al. Cereblon enhancer methylation and IMiD resistance in multiple myeloma. Blood. 2021;138(18):1721–6. https://doi.org/10.1182/blood.2020010452.
Lazarian G, Yin S, ten Hacken E, Sewastianik T, Uduman M, Font-Tello A, et al. A hotspot mutation in transcription factor IKZF3 drives B cell neoplasia via transcriptional dysregulation. Cancer Cell. 2021;39(3):380-393.e8. https://doi.org/10.1016/j.ccell.2021.02.003.
Robbins DW, Noviski M, Rountree R, Tan M, Brathaban N, Ingallinera T, et al. Nx-5948, a selective degrader of BTK with activity in preclinical models of hematologic and brain malignancies. Blood. 2021;138(Supplement 1):2251. https://doi.org/10.1182/blood-2021-147473.
Huang P, Chandra V, Rastinejad F. Structural overview of the nuclear receptor superfamily: insights into physiology and therapeutics. Annu Rev Physiol. 2010;72:247–72. https://doi.org/10.1146/annurev-physiol-021909-135917.
Pollock JA, Wardell SE, Parent AA, Parent AA, Stagg DB, Ellison SJ, et al. Inhibiting androgen receptor nuclear entry in castration-resistant prostate cancer. Nat Chem Biol. 2016;12(10):795–801. https://doi.org/10.1038/nchembio.2131.
Salami J, Alabi SB, Willard RR, Vitale NJ, Wang J, et al. Androgen receptor degradation by the proteolysis-targeting chimera ARCC-4 outperforms enzalutamide in cellular models of prostate cancer drug resistance. Commun Biol. 2018;1(1):100–100. https://doi.org/10.1038/s42003-018-0105-8.
Heinlein CA, Chang C. Androgen receptor in prostate cancer. Endocr Rev. 2004;25(2):276–308. https://doi.org/10.1210/er.2002-0032.
Sanford M. Enzalutamide: a review of its use in metastatic, castration-resistant prostate cancer. Drugs. 2013;73(15):1723–32. https://doi.org/10.1007/s40265-013-0129-9.
Palmbos PL, Hussain M. Non-castrate metastatic prostate cancer: have the treatment options changed? Semin Oncol. 2013;40(3):337–46. https://doi.org/10.1053/j.seminoncol.2013.04.007.
Han X, Zhao L, **ang W, Qin C, Miao B, McEachern D, et al. Strategies toward discovery of potent and orally bioavailable proteolysis targeting chimera degraders of androgen receptor for the treatment of prostate cancer. J Med Chem. 2021;64(17):12831–54. https://doi.org/10.1021/acs.jmedchem.1c00882.
**ang W, Zhao L, Han X, Qin C, et al. Discovery of ARD-2585 as an exceptionally potent and orally active PROTAC degrader of androgen receptor for the treatment of advanced prostate cancer. J Med Chem. 2021;64(18):13487–509. https://doi.org/10.1021/acs.jmedchem.1c00900.
Tong CWS, Wu M, Cho WCS, To KKW. Recent advances in the treatment of breast cancer. Front Oncol. 2018;8:227. https://doi.org/10.3389/fonc.2018.00227.
Anderson WF, Katki HA, Rosenberg PS. Incidence of breast cancer in the United States: current and future trends. J Natl Cancer Inst. 2011;103(18):1397–402. https://doi.org/10.1093/jnci/djr257.
Nilsson S, Koehler KF, Gustafsson J-Å. Development of subtype-selective oestrogen receptor-based therapeutics. Nat Rev Drug Discov. 2011;10(10):778–92. https://doi.org/10.1038/nrd3551.
Jia M, Dahlman-Wright K, Gustafsson J-Å. Estrogen receptor alpha and beta in health and disease. Best Pract Res Clin Endocrinol Metab. 2015;29(4):557–68. https://doi.org/10.1016/j.beem.2015.04.008.
Wang Y, Tang S-C. The race to develop oral SERDs and other novel estrogen receptor inhibitors: recent clinical trial results and impact on treatment options. Cancer Metastasis Rev. 2022. https://doi.org/10.1007/s10555-022-10066-y.
Howell A, Sapunar F. Fulvestrant revisited: efficacy and safety of the 500-mg dose. Clin Breast Cancer. 2011;11(4):204–10. https://doi.org/10.1016/j.clbc.2011.02.002.
Osborne CK, Wakeling A, Nicholson RI. Fulvestrant: an oestrogen receptor antagonist with a novel mechanism of action. Br J Cancer. 2004;90(Suppl 1):S2-6. https://doi.org/10.1038/sj.bjc.6601629.
Robertson JFR, Harrison M. Fulvestrant: pharmacokinetics and pharmacology. Br J Cancer. 2004;90(Suppl 1):S7-10. https://doi.org/10.1038/sj.bjc.6601630.
Robertson JFR, Lindemann J, Garnett S, Anderson E, Nicholson RI, Kuter I, et al. A good drug made better: the fulvestrant dose-response story. Clin Breast Cancer. 2014;14(6):381–9. https://doi.org/10.1016/j.clbc.2014.06.005.
Mottamal M, Kang B, Peng X, Wang G. From pure antagonists to pure degraders of the estrogen receptor: evolving strategies for the same target. ACS Omega. 2021;6(14):9334–43. https://doi.org/10.1021/acsomega.0c06362.
Cyrus K, Wehenkel M, Choi E-Y, Lee H, Swanson HI, Kim KB. Jostling for position: optimizing linker location in the design of estrogen receptor-targeting PROTACs. ChemMedChem. 2010;5(7):979–85. https://doi.org/10.1002/cmdc.201000146.
Kargbo RB. PROTAC-mediated degradation of estrogen receptor in the treatment of cancer. ACS Med Chem Lett. 2019;10(10):1367–9. https://doi.org/10.1021/acsmedchemlett.9b00397.
Tecalco-Cruz AC, Zepeda-Cervantes J, Ramírez-Jarquín JO, Rojas-Ochoa A. Proteolysis-targeting chimeras and their implications in breast cancer. Explor Target Anti-Tumor Ther. 2021;2(6):496–510. https://doi.org/10.37349/etat.2021.00060.
Qin H, Zhang Y, Lou Y, Pan Z, Song F, et al. Overview of PROTACs targeting the estrogen receptor: achievements for biological and drug discovery. Curr Med Chem. 2021; 28. https://doi.org/10.2174/0929867328666211110101018.
Qi S-M, Dong J, Xu Z-Y, Cheng X-D, Zhang W, Zhang W-D, et al. PROTAC: an effective targeted protein degradation strategy for cancer therapy. Front Pharmacol. 2021;12:692574. https://doi.org/10.3389/fphar.2021.692574.
Liu J, Chen H, Kaniskan HÜ, **e L, Chen X, ** J, et al. TF-PROTACs enable targeted degradation of transcription factors. J Am Chem Soc. 2021;143(23):8902–10. https://doi.org/10.1021/jacs.1c03852.
Vaquerizas JM, Kummerfeld SK, Teichmann SA, Luscombe NM. A census of human transcription factors: function, expression and evolution. Nat Rev Genet. 2009;10(4):252–63. https://doi.org/10.1038/nrg2538.
**ao X, Li BX, Mitton B, Ikeda A, Sakamoto KM. Targeting CREB for cancer therapy: friend or foe. Curr Cancer Drug Targets. 2010;10(4):384–91. https://doi.org/10.2174/156800910791208535.
Darnell JE. Transcription factors as targets for cancer therapy. Nat Rev Cancer. 2002;2(10):740–9. https://doi.org/10.1038/nrc906.
Bushweller JH. Targeting transcription factors in cancer - from undruggable to reality. Nat Rev Cancer. 2019;19(11):611–24. https://doi.org/10.1038/s41568-019-0196-7.
Tammineni P, Anugula C, Mohammed F, Anjaneyulu M, Larner AC, et al. The import of the transcription factor STAT3 into mitochondria depends on GRIM-19, a component of the electron transport chain. J Biol Chem. 2013;288(7):4723–32. https://doi.org/10.1074/jbc.m112.378984.
Banerjee K, Resat H. Constitutive activation of STAT3 in breast cancer cells: a review. Int J Cancer. 2016;138(11):2570–8. https://doi.org/10.1002/ijc.29923.
Johnson DE, O’Keefe RA, Grandis JR. Targeting the IL-6/JAK/STAT3 signalling axis in cancer. Nat Rev Clin Oncol. 2018;15(4):234–48. https://doi.org/10.1038/nrclinonc.2018.8.
Bai L, Zhou H, Xu R, Zhao Y, Chinnaswamy K, McEachern D, et al. A potent and selective small-molecule degrader of STAT3 achieves complete tumor regression in vivo. Cancer Cell. 2019;36(5):498-511.e17. https://doi.org/10.1016/j.ccell.2019.10.002.
Debnath B, Xu S, Neamati N. Small molecule inhibitors of signal transducer and activator of transcription 3 (Stat3) protein. J Med Chem. 2012;55(15):6645–68. https://doi.org/10.1021/jm300207s.
Pallandre J-R, Borg C, Rognan D, Boibessot T, Luzet V, Yesylevskyy S, et al. Novel aminotetrazole derivatives as selective STAT3 non-peptide inhibitors. Eur J Med Chem. 2015;103:163–74. https://doi.org/10.1016/j.ejmech.2015.08.054.
Yang J, Stark GR. Roles of unphosphorylated STATs in signaling. Cell Res. 2008;18(4):443–51. https://doi.org/10.1038/cr.2008.41.
Heppler LN, Frank DA. Inhibit versus destroy: are PROTAC degraders the solution to targeting STAT3? Cancer Cell. 2019;36(5):459–61. https://doi.org/10.1016/j.ccell.2019.10.010.
Zoppi V, Hughes SJ, Maniaci C, Testa A, Gmaschitz T, Wieshofer C, et al. Iterative design and optimization of initially inactive proteolysis targeting chimeras (PROTACs) identify VZ185 as a potent, fast, and selective von Hippel-Lindau (VHL) based dual degrader probe of BRD9 and BRD7. J Med Chem. 2019;62(2):699–726. https://doi.org/10.1021/acs.jmedchem.8b01413.
Alabi S, Jaime-Figueroa S, Yao Z, Gao Y, Hines J, Samarasinghe KTG, et al. Mutant-selective degradation by BRAF-targeting PROTACs. Nat Commun. 2021;12(1):920. https://doi.org/10.1038/s41467-021-21159-7.
Cheng M, Yu X, Lu K, **e L, Wang L, Meng F, et al. Discovery of potent and selective epidermal growth factor receptor (EGFR) bifunctional small-molecule degraders. J Med Chem. 2020;63(3):1216–32. https://doi.org/10.1021/acs.jmedchem.9b01566.
Wang M, Lu J, Wang M, Yang C-Y, Wang S. Discovery of SHP2-D26 as a first, potent, and effective PROTAC degrader of SHP2 protein. J Med Chem. 2020;63(14):7510–28. https://doi.org/10.1021/acs.jmedchem.0c00471.
He Y, Zhang X, Chang J, Kim H-N, Zhang P, Wang Y, et al. Using proteolysis-targeting chimera technology to reduce navitoclax platelet toxicity and improve its senolytic activity. Nat Commun. 2020;11(1):1996. https://doi.org/10.1038/s41467-020-15838-0.
Yang X, Wang Z, Pei Y, Song N, Xu L, Feng B, et al. Discovery of thalidomide-based PROTAC small molecules as the highly efficient SHP2 degraders. Eur J Med Chem. 2021;218:113341. https://doi.org/10.1016/j.ejmech.2021.113341.
Donoghue C, Cubillos-Rojas M, Gutierrez-Prat N, Sanchez-Zarzalejo C, Verdaguer X, Riera A, et al. Optimal linker length for small molecule PROTACs that selectively target p38α and p38β for degradation. Eur J Med Chem. 2020;201:112451. https://doi.org/10.1016/j.ejmech.2020.112451.
Hines J, Lartigue S, Dong H, Qian Y, Crews CM. MDM2-recruiting PROTAC offers superior, synergistic antiproliferative activity via simultaneous degradation of BRD4 and stabilization of p53. Cancer Res. 2019;79(1):251–62. https://doi.org/10.1158/0008-5472.CAN-18-2918.
Schiemer J, Horst R, Meng Y, Montgomery JI, Xu Y, Feng X, et al. Snapshots and ensembles of BTK and cIAP1 protein degrader ternary complexes. Nat Chem Biol. 2021;17(2):152–60. https://doi.org/10.1038/s41589-020-00686-2.
Zhang X, He Y, Zhang P, Budamagunta V, et al. Discovery of IAP-recruiting BCL-XL PROTACs as potent degraders across multiple cancer cell lines. Eur J Med Chem. 2020;199:112397. https://doi.org/10.1016/j.ejmech.2020.112397.
Meng F, Xu C, Park K-S, Kaniskan HÜ, Wang GG, ** J. Discovery of a first-in-class degrader for nuclear receptor binding SET domain protein 2 (NSD2) and Ikaros/Aiolos. J Med Chem. 2022;65(15):10611–25. https://doi.org/10.1021/acs.jmedchem.2c00807.
Bricelj A, Steinebach C, Kuchta R, Gütschow M, Sosič I. E3 ligase ligands in successful PROTACs: an overview of syntheses and linker attachment points. Front Chem. 2021;9:707317. https://doi.org/10.3389/fchem.2021.707317.
Cao C, He M, Wang L, He Y, Rao Y. Chemistries of bifunctional PROTAC degraders. Chem Soc Rev. 2022;51(16):7066–114. https://doi.org/10.1039/D2CS00220E.
Lai AC, Toure M, Hellerschmied D, Salami J, Jaime-Figueroa S, Eunhwa K, et al. Modular PROTAC design for the degradation of oncogenic BCR-ABL. Angew Chem. 2016;55(2):807–10. https://doi.org/10.1002/anie.201507634.
Maple HJ, Clayden N, Baron A, Stacey C, Felix R. Develo** degraders: principles and perspectives on design and chemical space. MedChemComm. 2019;10(10):1755–64. https://doi.org/10.1039/C9MD00272C.
Han X, Wang C, Qin C, **ang W, Fernandez-Salas E, Yang C-Y, et al. Discovery of ARD-69 as a highly potent proteolysis targeting chimera (PROTAC) degrader of androgen receptor (AR) for the treatment of prostate cancer. J Med Chem. 2019;62(2):941–64. https://doi.org/10.1021/acs.jmedchem.8b01631.
Kargbo RB. PROTAC-mediated degradation of Bruton’s tyrosine kinase as a therapeutic strategy for cancer. ACS Med Chem Lett. 2021;12(5):688–9. https://doi.org/10.1021/acsmedchemlett.1c00178.
He S, Dong G, Cheng J, Wu Y, Sheng C. Strategies for designing proteolysis targeting chimaeras (PROTACs). Med Res Rev. 2022;42(3):1280–342. https://doi.org/10.1002/med.21877.
Han X, Sun Y. Strategies for the discovery of oral PROTAC degraders aimed at cancer therapy. Cell Rep Phys Sci. 2022;3(10):101062. https://doi.org/10.1016/j.xcrp.2022.101062.
Cyrus K, Wehenkel M, Choi E-Y, Han H-J, Lee H, Swanson H, et al. Impact of linker length on the activity of PROTACs. Mol Biosyst. 2011;7(2):359–64. https://doi.org/10.1039/c0mb00074d.
Bemis TA, La Clair JJ, Burkart MD. Unraveling the role of linker design in proteolysis targeting chimeras. J Med Chem. 2021;64(12):8042–52. https://doi.org/10.1021/acs.jmedchem.1c00482.
Naro Y, Darrah K, Deiters A. Optical control of small molecule-induced protein degradation. J Am Chem Soc. 2020;142(5):2193–7. https://doi.org/10.1021/jacs.9b12718.
Liu J, Peng Y, Wei W. Light-controllable PROTACs for temporospatial control of protein degradation. Front Cell Dev Biol. 2021;9:678077. https://doi.org/10.3389/fcell.2021.678077.
Pfaff P, Samarasinghe KTG, Crews CM, Carreira EM. Reversible spatiotemporal control of induced protein degradation by bistable photoPROTACs. ACS Cent Sci. 2019;5(10):1682–90. https://doi.org/10.1021/acscentsci.9b00713.
Xue G, Wang K, Zhou D, Zhong H, Pan Z. Light-induced protein degradation with photocaged PROTACs. J Am Chem Soc. 2019;141(46):18370–4. https://doi.org/10.1021/jacs.9b06422.
Liu J, Chen H, Ma L, He Z, Wang D, Liu Y, et al. Light-induced control of protein destruction by opto-PROTAC. Sci Adv. 2020;6(8):eaay5154. https://doi.org/10.1126/sciadv.aay5154.
Lebraud H, Wright DJ, Johnson CN, Heightman TD. Protein degradation by in-cell self-assembly of proteolysis targeting chimeras. ACS Cent Sci. 2016;2(12):927–34. https://doi.org/10.1021/acscentsci.6b00280.
Zheng S, Tan Y, Wang Z, Li C, Zhang Z, Sang X, et al. Accelerated rational PROTAC design via deep learning and molecular simulations. Nat Mach Intell. 2022;4(9):739–48. https://doi.org/10.1038/s42256-022-00527-y.
Riching KM, Schwinn MK, Vasta JD, Robers MB, Machleidt T, Urh M, et al. CDK family PROTAC profiling reveals distinct kinetic responses and cell cycle-dependent degradation of CDK2. SLAS Discov Adv Life Sci R D. 2021;26(4):560–9. https://doi.org/10.1177/2472555220973602.
Kargbo RB. PROTAC compounds targeting TRK for use in cancer therapeutics. ACS Med Chem Lett. 2020;11(6):1090–1. https://doi.org/10.1021/acsmedchemlett.0c00235.
Burslem GM, Schultz AR, Bondeson DP, Eide CA, Savage Stevens SL, Druker BJ, et al. Targeting BCR-ABL1 in chronic myeloid leukemia by PROTAC-mediated targeted protein degradation. Cancer Res. 2019;79(18):4744–53. https://doi.org/10.1158/0008-5472.CAN-19-1236.
Adhikari B, Bozilovic J, Diebold M, Schwarz JD, Hofstetter J, Schröder M, et al. PROTAC-mediated degradation reveals a non-catalytic function of AURORA-A kinase. Nat Chem Biol. 2020;16(11):1179–88. https://doi.org/10.1038/s41589-020-00652-y.
Pang X-J, Liu X-J, Liu Y, Liu W-B, Li Y-R, Yu G-X, et al. Drug discovery targeting focal adhesion kinase (FAK) as a promising cancer therapy. Molecules. 2021;26(14):4250. https://doi.org/10.3390/molecules26144250.
Burslem GM, Song J, Chen X, Chen X, Chen X, Hines J, et al. Enhancing antiproliferative activity and selectivity of a FLT-3 inhibitor by proteolysis targeting chimera conversion. J Am Chem Soc. 2018;140(48):16428–32. https://doi.org/10.1021/jacs.8b10320.
Hyun S, Shin D. Chemical-mediated targeted protein degradation in neurodegenerative diseases. Life. 2021;11(7):607. https://doi.org/10.3390/life11070607.
Chu T-T, Gao N, Li Q-Q, Chen P-G, Yang X-F, Chen Y-X, et al. Specific knockdown of endogenous tau protein by peptide-directed ubiquitin-proteasome degradation. Chem Biol. 2016;23(4):453–61. https://doi.org/10.1016/j.chembiol.2016.02.016.
Fiorillo A, Morea V, Colotti G, Ilari A. Huntingtin ubiquitination mechanisms and novel possible therapies to decrease the toxic effects of mutated Huntingtin. J Pers Med. 2021;11(12):1309. https://doi.org/10.3390/jpm11121309.
Harding R, Tong Y, Tong Y. Proteostasis in Huntington’s disease: disease mechanisms and therapeutic opportunities. Acta Pharmacol Sin. 2018;39(5):754–69. https://doi.org/10.1038/aps.2018.11.
Tomoshige S, Nomura S, Ohgane K, Hashimoto Y, Ishikawa M. Discovery of Small Molecules that Induce the Degradation of Huntingtin. Angew Chem Int Ed. 2017;56(38):11530–33. https://doi.org/10.1002/anie.201706529.
Chaudhary D, Robinson S, Romero DL. Recent advances in the discovery of small molecule inhibitors of interleukin-1 receptor-associated kinase 4 (IRAK4) as a therapeutic target for inflammation and oncology disorders. J Med Chem. 2015;58(1):96–110. https://doi.org/10.1021/jm5016044.
Nunes JP, McGonagle GA, Eden J, Kiritharan G, et al. Targeting IRAK4 for degradation with PROTACs. ACS Med Chem Lett. 2019;10(7):1081–5. https://doi.org/10.1021/acsmedchemlett.9b00219.
Zhang J, Fu L, Shen B, Liu Y, Wang W, Cai X, et al. Assessing IRAK4 functions in ABC DLBCL by IRAK4 kinase inhibition and protein degradation. Cell Chem Biol. 2020;27(12):1500-1509.e13. https://doi.org/10.1016/j.chembiol.2020.08.010.
Nguyen HCB, Adlanmerini M, Hauck AK, Lazar MA. Dichotomous engagement of HDAC3 activity governs inflammatory responses. Nature. 2020;584(7820):286–90. https://doi.org/10.1038/s41586-020-2576-2.
Fischer F, Alves Avelar LA, Murray L, Kurz T. Designing HDAC-PROTACs: lessons learned so far. Future Med Chem. 2022;14(3):143–66. https://doi.org/10.4155/fmc-2021-0206.
Dokmanovic M, Clarke C, Marks PA. Histone deacetylase inhibitors: overview and perspectives. Mol Cancer Res MCR. 2007;5(10):981–9. https://doi.org/10.1158/1541-7786.MCR-07-0324.
Cao F, de Weerd S, Chen D, Zwinderman MRH, van der Wouden PE, Dekker FJ. Induced protein degradation of histone deacetylases 3 (HDAC3) by proteolysis targeting chimera (PROTAC). Eur J Med Chem. 2020;208:112800. https://doi.org/10.1016/j.ejmech.2020.112800.
**ao Y, Wang J, Zhao LY, Chen X, Zheng G, Zhang X, et al. Discovery of histone deacetylase 3 (HDAC3)-specific PROTACs. Chem Commun Camb Engl. 2020;56(68):9866–9. https://doi.org/10.1039/d0cc03243c.
de Wispelaere M, Du G, Donovan KA, Zhang T, Eleuteri NA, Yuan JC, et al. Small molecule degraders of the hepatitis C virus protease reduce susceptibility to resistance mutations. Nat Commun. 2019;10(1):3468. https://doi.org/10.1038/s41467-019-11429-w.
Martinez-Ortiz W, Zhou M-M. Could PROTACs protect us from COVID-19? Drug Discov Today. 2020;25(11):1894–6. https://doi.org/10.1016/j.drudis.2020.08.007.
Desantis J, Mercorelli B, Celegato M, Croci F, Bazzacco A, Baroni M, et al. Indomethacin-based PROTACs as pan-coronavirus antiviral agents. Eur J Med Chem. 2021;226:113814. https://doi.org/10.1016/j.ejmech.2021.113814.
Al-Horani RA, Kar S. Potential anti-SARS-CoV-2 therapeutics that target the post-entry stages of the viral life cycle: a comprehensive review. Viruses. 2020;12(10):E1092. https://doi.org/10.3390/v12101092.
Vane JR. Inhibition of prostaglandin synthesis as a mechanism of action for aspirin-like drugs. Nature New Biol. 1971;231(25):232–5. https://doi.org/10.1038/newbio231232a0.
Gordon DE, Hiatt J, Bouhaddou M, Rezelj VV, Ulferts S, Braberg H, et al. Comparative host-coronavirus protein interaction networks reveal pan-viral disease mechanisms. Science. 2020;370(6521):eabe9403. https://doi.org/10.1126/science.abe9403.
Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature. 2020;583(7816):459–68. https://doi.org/10.1038/s41586-020-2286-9.
Terracciano R, Preianò M, Fregola A, Pelaia C, Montalcini T, Savino R. Map** the SARS-CoV-2-host protein-protein interactome by affinity purification mass spectrometry and proximity-dependent biotin labeling: a rational and straightforward route to discover host-directed anti-SARS-CoV-2 therapeutics. Int J Mol Sci. 2021;22(2):E532. https://doi.org/10.3390/ijms22020532.
Lu X-Y, Shi X-J, Hu A, Wang J-Q, Ding Y, Jiang W, et al. Feeding induces cholesterol biosynthesis via the mTORC1-USP20-HMGCR axis. Nature. 2020;588(7838):479–84. https://doi.org/10.1038/s41586-020-2928-y.
Luo G, Li Z, Lin X, Li X, Chen Y, ** K, et al. Discovery of an orally active VHL-recruiting PROTAC that achieves robust HMGCR degradation and potent hypolipidemic activity in vivo. Acta Pharm Sin B. 2021;11(5):1300–14. https://doi.org/10.1016/j.apsb.2020.11.001.
Li M-X, Yang Y, Zhao Q, Wu Y, Song L, Yang H, et al. Degradation versus inhibition: development of proteolysis-targeting chimeras for overcoming statin-induced compensatory upregulation of 3-Hydroxy-3-methylglutaryl coenzyme A reductase. J Med Chem. 2020;63(9):4908–28. https://doi.org/10.1021/acs.jmedchem.0c00339.
Li Z, Lin Y, Song H, Qin X, Yu Z, Zhang Z, et al. First small-molecule PROTACs for G protein-coupled receptors: inducing 1A-adrenergic receptor degradation. Acta Pharm Sin B. 2020;10(9):1669–79. https://doi.org/10.1016/j.apsb.2020.01.014.
Zamanakou M, Germenis AE, Karanikas V. Tumor immune escape mediated by indoleamine 2,3-dioxygenase. Immunol Lett. 2007;111(2):69–75. https://doi.org/10.1016/j.imlet.2007.06.001.
Si J, Shi X, Sun S, Zou B, Li Y, An D, et al. Hematopoietic progenitor kinase1 (HPK1) mediates T cell dysfunction and is a druggable target for T cell-based immunotherapies. Cancer Cell. 2020;38(4):551-566.e11. https://doi.org/10.1016/j.ccell.2020.08.001.
Cantrill C, Chaturvedi P, Rynn C, Petrig Schaffland J, Walter I, Wittwer MB. Fundamental aspects of DMPK optimization of targeted protein degraders. Drug Discov Today. 2020;25(6):969–82. https://doi.org/10.1016/j.drudis.2020.03.012.
Zhang L, Riley-Gillis B, Vijay P, Shen Y. Acquired resistance to BET-PROTACs (proteolysis-targeting chimeras) caused by genomic alterations in core components of E3 ligase complexes. Mol Cancer Ther. 2019;18(7):1302–11. https://doi.org/10.1158/1535-7163.MCT-18-1129.
Ottis P, Palladino C, Thienger P, Britschgi A, Heichinger C, Berrera M, et al. Cellular resistance mechanisms to targeted protein degradation converge toward impairment of the engaged ubiquitin transfer pathway. ACS Chem Biol. 2019;14(10):2215–23. https://doi.org/10.1021/acschembio.9b00525.
Schapira M, Calabrese MF, Bullock AN, Crews CM. Targeted protein degradation: expanding the toolbox. Nat Rev Drug Discov. 2019;18(12):949–63. https://doi.org/10.1038/s41573-019-0047-y.
An S, Fu L. Small-molecule PROTACs: an emerging and promising approach for the development of targeted therapy drugs. EBioMedicine. 2018;36:553–62. https://doi.org/10.1016/j.ebiom.2018.09.005.
Miles LE. Properties, variants, and applications of the immunoradiometric assay method. Ric Clin Lab. 1975;5(1):59–72. https://doi.org/10.1007/BF02910016.
Gao H, Sun X, Rao Y. PROTAC technology: opportunities and challenges. ACS Med Chem Lett. 2020;11(3):237–40. https://doi.org/10.1021/acsmedchemlett.9b00597.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (22277086, 22007070), the Science and Technology Department of Sichuan Province (2022ZDZX0028, 2022NSFSC1409), 1.3.5 Project for Disciplines of Excellence, West China Hospital, Sichuan University (ZYJC21075) and the Post-Doctor Research Project, West China Hospital, Sichuan University (2021HXBH006).
Code availability
Not applicable.
Author information
Authors and Affiliations
Contributions
All authors contributed substantially to this work. Zi Liu wrote and prepared the original draft. Mingxing Hu, Yu Yang, Haoxuan Zhou, Chengyali Liu and Hongqun Ma wrote, reviewed, and edited the manuscript. Chenghao Du, Yuanwei Chen, Lei Fan, Youling Gong and Yongmei **e supervised and revised the study. All authors approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This article does not involve animal and human experiments.
Consent for publication
Not applicable.
Competing interests
Author Yuanwei Chen, Lei Fan, Hongqun Ma are employees in Hinova Pharmaceuticals Inc., but has no potential relevant financial or non-financial interests to disclose. The other authors have no conflicts of interest to declare.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Z., Hu, M., Yang, Y. et al. An overview of PROTACs: a promising drug discovery paradigm. Mol Biomed 3, 46 (2022). https://doi.org/10.1186/s43556-022-00112-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s43556-022-00112-0