Abstract
Background
Archaea perform critical roles in the microbiome system, including utilizing hydrogen to allow for enhanced microbiome member growth and influencing overall host health. With the majority of microbiome research focusing on bacteria, the functions of archaea are largely still under investigation. Understanding methanogenic functions during the host lifetime will add to the limited knowledge on archaeal influence on gut and host health. In our study, we determined lifelong archaea dynamics, including detection and methanogenic functions, while assessing global, temporal and host distribution of our novel archaeal metagenome-assembled genomes (MAGs). We followed 7 monogastric swine throughout their life, from birth to adult (1–156 days of age), and collected feces at 22 time points. The samples underwent gDNA extraction, Illumina sequencing, bioinformatic quality and assembly processes, MAG taxonomic assignment and functional annotation. MAGs were utilized in downstream phylogenetic analysis for global, temporal and host distribution in addition to methanogenic functional potential determination.
Results
We generated 1130 non-redundant MAGs, representing 588 unique taxa at the species level, with 8 classified as methanogenic archaea. The taxonomic classifications were as follows: orders Methanomassiliicoccales (5) and Methanobacteriales (3); genera UBA71 (3), Methanomethylophilus (1), MX-02 (1), and Methanobrevibacter (3). We recovered the first US swine Methanobrevibacter UBA71 sp006954425 and Methanobrevibacter gottschalkii MAGs. The Methanobacteriales MAGs were identified primarily during the young, preweaned host whereas Methanomassiliicoccales primarily in the adult host. Moreover, we identified our methanogens in metagenomic sequences from Chinese swine, US adult humans, Mexican adult humans, Swedish adult humans, and paleontological humans, indicating that methanogens span different hosts, geography and time. We determined complete metabolic pathways for all three methanogenic pathways: hydrogenotrophic, methylotrophic, and acetoclastic. This study provided the first evidence of acetoclastic methanogenesis in archaea of monogastric hosts which indicated a previously unknown capability for acetate utilization in methanogenesis for monogastric methanogens. Overall, we hypothesized that the age-associated detection patterns were due to differential substrate availability via the host diet and microbial metabolism, and that these methanogenic functions are likely crucial to methanogens across hosts. This study provided a comprehensive, genome-centric investigation of monogastric-associated methanogens which will further improve our understanding of microbiome development and functions.
Similar content being viewed by others
Introduction
The gastrointestinal system contains countless microorganisms spanning multiple kingdoms performing diverse functions. Archaea, bacteria, viruses, and fungi work in concert and competition to acquire nutrients and space [1]. The focus of previous gut microbiome research has predominantly been on the identification and function of bacteria [2, 3]. However, archaea have been demonstrated to be equally important members of the gastrointestinal microbiome [4]. Methanogenic archaea, or archaea which carry out methanogenesis, perform crucial roles in the gut [4, 5]. Yet, current research has not indicated how methanogenic gut functions change throughout the lifetime of monogastric hosts [6, 7]. With limited research on archaea, and even more minimal analysis on methanogenic functions, we are lacking an in-depth understanding of gastrointestinal associated methanogens, especially our comprehension of methanogen influence on gut and host health throughout host stages of life. By investigating monogastric associated methanogens with a longitudinal approach, we are adding essential knowledge to the limited understanding of monogastric methanogens.
While some beneficial and detrimental associations of archaea to host health have been reported, overall the role of archaea in health and disease is still under investigation [5]. To date, archaea have been associated with a few illnesses, primarily gastrointestinal disorders such as constipation [5, 8, 9] and obesity [5, 10]. Conversely, archaea have also been associated with beneficial attributes. For example, archaea metabolize trimethylamine (TMA), which is thought to decrease cardiovascular disease [4, 5]. This research has prompted further evaluation of archaea members as a probiotic for cardiovascular health [5, 11]. Moreover, archaea allow continued microbial metabolism, growth and action by lowering hydrogen gut levels [5]. Archaea’s role of hydrogen utilization is especially important in the gut where microorganisms work in concert within the shared gut-microbiome system. However, with limited prior research, there is a critical need to understand the role of gastrointestinal archaea in health and sickness via hydrogen metabolism.
Overall, archaea are classified into four superphyla: Euryarchaeota, Asgard, TACK (Thaumarchaeota, Aigarchaeota, Crenarchaeota and Korarchaeota), and DPANN (Diapherotrites, Parvarchaeota, Aenigmarchaeota, Nanoarchaeota, and Nanohaloarchaeota) [12]. To date, Asgard archaea have not been indicated as methanogens [12], and TACK and DPANN have only been identified in non-host associated environmental sites [12,13,14]. Therefore, currently known host-associated gut methanogens fall within the seven orders of Euryarchaeota: Methanobacteriales, Methanococcales, Methanomicrobiales, Methanosarcinales, Methanocellales, Methanopyrales, Methanomassiliicoccales [15,16,17,18]. These Euryarchaeota orders are obligate anaerobes which perform methanogenesis to conserve energy for ATP production, where methane is a byproduct [15, 19]. Actions immediately following methanogenesis generate an ion gradient which is coupled with ATP production [20, 21].
Given the necessity for ATP production, it is unsurprising that historically, studies have primarily relied on the methanogenic gene methyl-coenzyme M reductase A (mcrA) or 16S rRNA for identification of gut-associated methanogens [7, 22,23,24]. McrA has been identified in all methanogens to date, as the protein performs a critical role in the final methane production step of methanogenesis [24, 25]. While prior research was heavily reliant on targeted PCR methodologies, we are in-large missing gene centric methanogenic understanding, from complete genetic sequencing, of gut-associated methanogens [26]. Functional methanogen studies become even more profound when evaluated in a longitudinal approach, especially when following the same hosts. In doing so, we can determine lifetime gut methanogen dynamics and host implications. Currently, studies which evaluate longitudinal methanogen dynamics typically involve ruminant hosts, such as cows, sheep, goats, and deer [27]. At the time of publication, we could not find a longitudinal study of methanogen genomes (i.e. not marker studies such as 16S rRNA or mcrA) following the same monogastrics hosts throughout their lifetime, highlighting the crucial need for such metagenomic longitudinal evaluations [6, 7]. Without this knowledge, we cannot determine lifetime dynamics of archaea, and how their methanogenic function may be related to age-associated factors, such as diet and host development.
Host-associated archaeal methanogens have been linked to various conditions of health and disease. Most archaea-centric intestinal microbiome studies have been conducted on a single time point in the lifetime of the host. Using molecular and cultural approaches, intestinal archaea have been identified in many hosts, including: humans, swine, horses, rats, birds, fish, and kangaroos [28]. Overall, these analyses reported that the most common methanogens in the gut are members of the Methanobacteriales and Methanomassiliicoccales orders [28]. However, little is known about the presence and distribution of archaea through the lifetime of the swine. There is also a lack of data on the functions of the archaea in the swine gut. Overall, this knowledge gap has hindered the identification of factors that influence the diversity, abundance, and functions of archaea in the swine. In this study, we evaluated methanogen abundance and functions of 7 monogastric swine hosts over their lifetime at 22 timepoints from birth through adulthood (ages 1–156 days). We recovered 8 methanogenic archaea metagenome-assembled genomes (MAGs) that exhibited differential colonization patterns in the host at different ages. While distribution of methanogens across multiple hosts has been previously demonstrated, we recovered the first US swine Methanobrevibacter UBA71 sp006954425 and Methanobrevibacter gottschalkii MAGs [28]. Moreover, we attributed methanogenic functional potential to our age-associated archaea, and identified the first evidence of acetoclastic methanogenesis in monogastric-associated archaea, found in our Methanomassiliicoccales MAGs, indicating a previously unknown capability of monogastric methanogens to utilize acetate in energy acquisition. Alternatively, we attributed hydrogenotrophic methanogenesis, where carbon dioxide (CO2) is utilized, in the Methanobacteriales. We surmised that the age-associated detection patterns were due to differential substrate availability, which was highly influenced by diet. Altogether, we provided a comprehensive, genome-centric investigation of monogastric-associated archaea to further our understanding of microbiome development and function.
Results and discussion
Taxonomic classification of gut metagenome-assembled genomes
To broadly sample gut-associated microorganisms of the swine host across different age-associated growth stages, we obtained ~ 5.8 × 109 paired-end reads from Illumina NovaSeq sequencing data of 112 swine fecal samples (Fig. 1 and Additional file 1: Table S3). After quality trimming, we generated ~ 5.2 × 109 paired-end reads. The resulting 3 co-assemblies contained ~ 9.4 × 106 contigs that described approximately ~ 3.6 × 1010 nucleotides and ~ 3.7 × 107 genes. Using a combination of automatic and manual binning strategies, with thresholds of > 70% complete and < 10% redundancy, resulted in 4,556 metagenome-assembled genomes (MAGs). We further removed redundancy by selecting a single representative for each set of genomes that shared an average nucleotide identity (ANI) of greater than 95%, resulting in 1,130 final non-redundant MAGs (nr-MAGs) (Additional file 1: Table S3). Among the nr-MAGs, we recovered an average of 203 ± 187 contigs, with an average N50 of 32,737 ± 35,205. The resolved nr-MAGs had completion values of 87.9% ± 8.6% and redundancy values of 3.2% ± 2.6%. The genomic lineages for archaeal and bacterial nr-MAGs based on domain-specific single-copy core genes resolved to 20 phyla (2 archaea phyla and 18 bacterial phyla) and 588 species (5 archaea species and 582 bacterial species). We could also assign 88.4% of the bacterial and archaeal nr-MAGs to their genera.
Resolved archaeal MAGs are genetically and phylogenetically similar to diverse hosts and geographic disbursed archaea
Among the 1,130 nr-MAGs that we resolved, our genomic collection also included 8 archaea nr-MAGs (hereafter known as archaea-MAGs; Ar-1 through Ar-8; Table 1; Additional file 1: Table S3). We observed that our resolved archaea-MAGs harbored genes which encoded for critical methyl-coenzyme M reductase (mcrABG) proteins required for methanogenesis, including mcrA which is typically utilized for methanogen classification [29, 30] (Additional files 1: Tables S3 and 2: Table S4). To our best knowledge, these MAGs represent the first genomic evidence of putative methanogens differential colonization pattern of the monogastric gut. The resolved methanogen MAGs had an average genome size of 1.4 Mbp, 1,573 KEGG gene annotations, 1,535 COG gene annotations, and a GC content ranging from 31 to 56% (Table 1; Additional file 1: Table S3). We resolved 7 of the methanogen MAGs to the species level with one archaea-MAG resolving to the genus level (Table 1; Additional file 1: Table S3). Our resolved archaea-MAGs were assigned to the following orders: Methanomassiliicoccales (5) and Methanobacteriales (3). Moreover, the genera were as follows: UBA71 (3), Methanomethylophilus (1), MX-02 (1), and Methanobrevibacter (3).
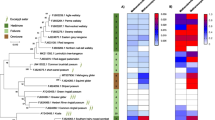
We downloaded 95 Methanomassiliicoccales and 97 Methanobacteriales genomes to investigate the phylogenetic relationship of our resolved archaea-MAGs (Fig. 2; Additional file 3: Fig. S1). We showed that our methanogen populations had close phylogenetic relationships with archaea from geographically distinct mammalian hosts, suggesting high similarities in gene functions in archaea among diverse host species. Given similarities amongst such diverse host species with diverse digestive systems, we hypothesize these close genetic relatives of our resolved archaea-MAGs might be more ubiquitous in a wider range of hosts that are currently discussed. We noticed Ar-4 clustered, as expected, with 6 Methanomethylophilus alvus strains: 5 from human gut samples and 1 from swine (MAG221) (Fig. 2A) [31,32,33,34,35,36]. Ar-7 clustered with 4 MX-02 sp006954405. United Kingdom strain 10 [37] and Chinese strain MAG014 [34] have been identified as swine-originating, whereas B5_69.fa and B45_maxbin.030.fa were from humans [38]. Interestingly, clustering on the same branch (B5_69.fa and B45_maxbin.030.fa) are archaea isolated from South African adult humans [38]. Ar-1 was in the same branch with archaea from Tibetan pig MAG098 [34]; Ar-2 with Chinese roe deer RGIG3983 [28].
We recovered from our study novel archaeal genomes that were previously unidentified in US swine. We were able to resolve and obtain the genomic information, to the best of our knowledge, of the first swine-associated Methanobrevibacter UBA71 sp006954425 and Methanobrevibacter gottschalkii MAGs. The methanogenic archaea family Methanobacteriales has been identified in many previous swine studies, with the majority of these studies utilizing 16S sequencing and/or real-time PCR identification [34, 37, 52,53,54,55,56,57,58,59,60,61]. Still, there is a lack of understanding of the Methanobacteriales in terms of genomic studies, and the Methanomassiliicoccales order collectively in general. Up to this moment, only three swine Methanomassiliicoccales MAGs (Methanomethylophilus alvus, MX-02 sp006954405, and Methanobrevibacter smithii) have been identified [34, 37, 61]. Thus, adding our highly resolved novel archaea-MAGs to the repertoire of swine-associated microbial populations will aid in understanding swine archaea, including functions, host associations (such as age, health status, sex, etc.), and global distribution.
Prevalence of archaeal MAGs and variants at distinct host ages
Assessing the abundance of the methanogens in different growth stages of the swine host provided an opportunity to investigate the association between the host-associated methanogens and the different conditions faced by the swine as they grow. Our genome-centric metagenome analyses revealed two dominant orders of archaea—Methanobacteriales and Methanomassiliicoccales. We showed that resolved methanogen MAGs were differentially detected at different growth stages of the swine, but does the environment affect the functional niche specificity between these two orders of archaea?
The heatmap shown in Fig. 3A provides a graphical summary of the changes in detection for the archaea-MAGs. Detection was defined as the proportion of a given contig in the MAG that is covered at least 1X. Hierarchical clustering grouped the archaea-MAGs into three clusters based on detection: A (top cluster; Ar-1 through Ar-4), B (middle cluster; Ar-5 and Ar-6), and C (bottom cluster; Ar-7 and Ar-8). We observed that Cluster A contained only Methanomassiliicoccales MAGs, while Cluster B contained 2 Methanobacteriales MAGs, and Cluster C one of each order. Cluster A archaea-MAGs were primarily identified in the final stage of growth adult hosts. Conversely, Cluster B methanogens were primarily identified in preweaning hosts. Finally, Cluster C archaea were identified throughout the host lifetime. Further support of distinct archaea-MAGs detection was supported through archaeal-MAG relative abundances (Additional file 4: Fig. S2).
A Detection (portion of MAG with at least 1X read coverage) heatmap of archaea-MAGs (rows) across all individual sample metagenomes (columns) with MAG taxonomy and stage annotation (Preweaning [P]; nursery [N]; growth adult [G]). B Single-nucleotide variant (SNV) analysis of Ar-7 and Ar-8 where box colors indicate competing nucleotides and stage is indicated along the bottom
We noticed the majority of archaea-MAGs detection values increased closely after a stage transition (preweaning to nursery and nursery to growth adult), suggesting that stage transition changes, including diet, housing, and stress, can lead to changes in microbiome composition [62]. Although, exactly how these changes impact archaea is relatively understudied, as most research evaluates bacteria, and therefore archaea-stage dynamics are a topic for future research [63, 64].
We investigated methanogen variants, and found the majority of variation occurred in periods when other archaea were dominating (preweaning and growth adult; Additional file 5: Table S5). We performed single-nucleotide variant (SNV) analysis on our two archaea-MAGs that showed continuous detection throughout the host lifetime (Cluster C MAGs: Ar-7 and Ar-8; Fig. 3B). We attributed the majority of variances to the unweaned host, and fewer variances were identified in the growth adult. Interestingly, the number of variant populations were highest during times where other archaea-MAGs were predominantly identified (Fig. 3B). We hypothesized that the variation found in the growth adult host could indicate a competitive microbial environment, while fewer variants as compared to the earlier growth stages could be due to a largely already developed gut microbiome. In a competitive gut microbiome system, it is beneficial to have genetic diversity which translates to increased functional diversity [65]. A similar competitive environment and SNV diversity was demonstrated in the human gut bacterial community [65]. Comparatively, as the gut developed and microbes established with focused functions, the variation decreased when humans reached 2 years of age [65]. Human preweaning gut development is similar to the development of the swine preweaning gut, albeit swine is relatively faster [66]. The conditions which encouraged the increased variation in the growth adult in our study could have been a change of diet, host stress, or other host-associated and environmental conditions [67].
While we demonstrated differing archaea and SNV association with age, we were primarily interested in methanogen function. We hypothesized methanogenic function influenced our resolved methanogen MAGs’ ability to establish in the microbiome at different host stages through energy acquisition via host diet. Phylogenetic similarity in archaea across geography and hosts prompted an investigation into whether our archaea-MAGs were identified in other hosts of similar developmental ages, and therefore similar archaeal functions.
Methanogens span host species, millennia, and geographic distance
We wanted to further demonstrate not only global and host distribution, but also temporal, or across time, identification of our methanogens beyond genetic similarity, as illustrated in our phylogenetic analyses. We mapped metagenomic sequencing reads from young and aged hosts to our archaea-MAGs from the following hosts: swine (n = 16) [68], humans (n = 429) [69, 70], mice (n = 60) [71], chicken (n = 71) [72], and cattle (n = 34) [73] (Fig. 4; Additional files 6: Tables S6 and 7: Table S7). Our archaea-MAGs were identified in older humans and varying aged swine metagenomes, but not in the chicken, mice or cattle metagenomes. We also demonstrated evidence of our archaea-MAGs in the ancient human gut and global distribution. Altogether we determined within a host species, archaeal age-association appeared to be similar, but some archaea span multiple host species, and for millennia [28]. We hypothesized differential archaeal function may be essential to the gut microbiome of many modern and ancient monogastric hosts.
A Detection (portion of MAG with at least 1 read coverage) heatmap of previously published swine metagenomes [68] mapped to this publication’s archaeal MAGs (Preweaning [P]; nursery [N]; growth adult [G]). B Detection box plots of previously published human metagenomes [69, 70] mapped to our archaeal MAGs (“Adult” from Mexican humans; “Paleo” from present day US and Mexico; all remaining groups from Sweden)
We determined our swine-associated methanogens were not present in poultry, mice and ruminant host metagenomes (Additional file 7: Table S7). Prescence of methanogens was determined when detection (portion of MAG with at least 1X read coverage) was greater than 0.25. This threshold aligns with previous genome-resolved metagenomic research and eliminated false-positive signals in read recruitment results [74,75,76]. Given the drastic differences in the ruminant digestive system compared to the monogastric gut, we were not surprised that our swine archaea-MAGs were not found in cattle from the United States (US) State of Pennsylvania. Although not identified consistently in all cattle, Methanobrevibacter smithii [77,78,79,80] and Methanobrevibacter gottschalkii [81,82,83,84] have been associated with the cow digestive tract. Similarly, UBA71 has been identified in adult chickens previously [85]. Given that similar taxonomic methanogens are present in cattle and chickens, we hypothesized the methanogens of these hosts were genetically distinct from the methanogens we identified in swine. Additionally, since the methanogens we identified were not consistently detected across our aging hosts, it was very probable that other metagenomes from these host populations could contain our methanogens. Future research is necessary to evaluate how distinct methanogen members function individually and collectively within the microbiome system to influence gut health in different host species.
Interestingly, we could only find a singular example of archaea attributed to the mouse gut: Methanomassiliicoccaceae DTU008 [86]. Remaining attempts, encompassing more than 1,000 metagenomes, proved unsuccessful in identifying mice gut archaea [87,88, Our study design and sample collection occurred as previously described [136]. We collected fecal samples from 7 swine over 22 timepoints, ranging in swine age from 1 to 156 days across three developmental stages: preweaning (P), nursery (N), and growth adult (G) (Fig. 1, Additional file 8: Table S1). Swine were born and raised at the Kansas State University Swine Teaching and Research Center. Swine originated from the same farrowing group, and were weaned between 18 and 20 days of age, depending on day of birth. We stored fecal samples at − 80 °C until DNA extraction. We extracted total genomic DNA from fecal samples utilizing the E.Z.N.A.® Stool DNA Kit (Omega Bio-tek Inc.; Norcross, GA), following the manufacturer protocols. We then quantified the extracted genomic DNA with a Nanodrop and Qubit™ (dsDNA BR Assay Kit [Thermo Fisher; Waltham, MA]) for DNA quality and concentration. We stored extracted DNA at − 80 °C until library preparation and sequencing. DNA libraries were generated for a total of 112 samples with Nextera DNA Flex (Illumina, Inc.; San Diego, CA). Resulting libraries were then visualized on a Tapestation 4200 (Agilent; Santa Clara, CA) and size-selected using the BluePippin (Sage Science; Beverly, MA). The final library pool of 112 samples was quantified on the Kapa Biosystems (Roche Sequencing; Pleasanton, CA) qPCR protocol, and sequenced on the Illumina NovaSeq S1 chip (Illumina, Inc.; San Diego, CA) with a 2 × 150 bp paired-end sequencing strategy. We utilized the ‘anvi-run-workflow’ program to run a combined bioinformatics workflow in anvi’o v.7.1 (https://anvio.org/install/) [137, 138], with a co-assembling strategy. The workflow used Snakemake to implement numerous tasks including: short-read quality filtering, assembly, gene calling, functional annotation, hidden Markov model search, metagenomic read-recruitment and binning [139]. Briefly, we processed sequencing reads using anvi’o’s ‘iu-filer-quality-minoche’ program, which removed low-quality reads following criteria outlined in Minoche et al. [140]. The resulting quality-control reads were termed “metagenome” per sample. We organized the samples into 3 metagenomic groups based on the developmental stages (P, N, G), and used anvi’o’s MEGAHIT v1.2.9 to co-assemble quality-filtered short reads into longer contiguous sequences (contigs) [137, 141]. The following methods were then utilized in anvi’o to further process the contigs: (1) ‘anvi-gen-contigs-database’ to compute k-mer frequencies and identify open reading frames (ORFs) using Prodigal v2.6.3 [137, 142]; (2) ‘anvi-run-hmms’ to annotate bacterial and archaeal single-copy, core genes with default single-copy genes and taxonomy of genomes [143] as defined by the The Genome Taxonomy Database (GTDB) [144] database (Archaea_76, Bacteria_71, Protista_83, and Ribosomal RNAs) [145] using HMMER v.3.2.1 [137, 146]; (3) ‘anvi-run-ncbi-cogs’ to annotate ORFs with NCBI’s Clusters of Orthologous Groups (COGs; https://www.ncbi.nlm.nih.gov/research/cog) [147]; and (4) ‘anvi-run-kegg-kofams’ to annotate ORFs from KOfam HMM databases of KEGG orthologs (https://www.genome.jp/kegg/) [148]. We mapped metagenomic short reads to contigs in anvi’o with Bowtie2 v2.3.5 [149], and we then converted map**s to BAM files with samtools v1.9 [137, 150, 151]. We used the anvi’o ‘anvi-profile’ program to profile BAM files with a minimum contig length of 1000 bp. Next, we combined profiles with ‘anvi-merge’ into a single anvi’o profile for downstream analyses. We grouped contigs into bins with ‘anvi-cluster-contigs’ and CONCOCT v1.1.0 [152]. We manually processed bins with ‘anvi-refine’ using bin tetranucleotide frequency and coverage across samples [137, Data analyses We used the “detection” criteria (> 0.25) for downstream statistical analyses. We downloaded metagenomes from swine [68], humans [69, 70], mice [71], chicken [72], and cattle [73], and performed map** to the non-redundant archaea-MAGs according to specifications above (Additional file 9: Table S2). We used RStudio v1.3.1093 [158] (https://www.rstudio.com/products/rstudio/) to visualize MAGs detection patterns using: pheatmap (pretty heatmaps) v1.0.12 [159], ggplot2 v3.3.5 (https://ggplot2.tidyverse.org/) [160], forcats v0.5.1 (https://forcats.tidyverse.org/) [161], dplyr v1.0.8 (https://dplyr.tidyverse.org/) [162], and ggpubr v0.4.0 (https://CRAN.R-project.org/package=ggpubr) [163]. We utilized the RASTtk Genome Annotation Service on PATRIC v3.6.12 (https://patricbrc.org/) and anvi’o COG annotations for metabolic function analyses [164, 165]. We used the comparative pathway tool in PATRIC to predict the metabolic pathways of our resolved non-redundant MAGs. We used ‘anvi-compute-genome-similarity’ to calculate average nucleotide identity (ANI) with a subset of samples (n = 21) from previously published human-associated archaea [16]. We obtained similar genomes that were deposited in public databases and performed phylogenetic analyses of our non-redundant MAGs in PATRIC [165]. Parameters were set as follows: 100 genes, 10 max allowed deletions, and 10 max allowed duplications. We constructed phylogenetic trees for our MAGs with 192 closely related genomes, using the amino acid and nucleotide sequences from the global protein families database. RAxML program was used to construct the trees based on pairwise differences between the aligned protein families of the selected sequences. Our final figures were edited in Inkscape v1.2.1 [166].Materials and methods
Study design, sample collection and DNA extraction
Metagenomic sequencing and ‘omics workflow
Availability of data and materials
We uploaded our metagenome raw sequencing data to the SRA under NCBI BioProject PRJNA798835. All other analyzed data, in the form of databases and fasta files, and bioinformatic scripts are accessible at figshare https://doi.org/10.6084/m9.figshare.20431713.
References
Coyte KZ, Rakoff-Nahoum S. Understanding competition and cooperation within the mammalian gut microbiome. Curr Biol. 2019;29:R538–44.
Shanahan F, Ghosh TS, O’Toole PW. The healthy microbiome-what is the definition of a healthy gut microbiome? Gastroenterology. 2021;160:483–94.
Aldars-García L, Chaparro M, Gisbert JP. Systematic review: the gut microbiome and its potential clinical application in inflammatory bowel disease. Microorganisms. 2021;9:5.
Gaci N, Borrel G, Tottey W, O’Toole PW, Brugère J-F. Archaea and the human gut: new beginning of an old story. World J Gastroenterol. 2014;20:16062–78.
Nkamga VD, Henrissat B, Drancourt M. Archaea: essential inhabitants of the human digestive microbiota. Human Microbiome Journal. 2017;3:1–8.
Kim JY, et al. The human gut archaeome: identification of diverse haloarchaea in Korean subjects. Microbiome. 2020;8:114.
Wampach L, et al. Colonization and succession within the human gut microbiome by archaea, bacteria, and microeukaryotes during the first year of life. Front Microbiol. 2017;8:738.
Pimentel M, et al. Methane production during lactulose breath test is associated with gastrointestinal disease presentation. Dig Dis Sci. 2003;48:86–92.
Ghoshal UC, Srivastava D, Verma A, Misra A. Slow transit constipation associated with excess methane production and its improvement following rifaximin therapy: a case report. J Neurogastroenterol Motil. 2011;17:185–8.
Samuel BS, Gordon JI. A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism. Proc Natl Acad Sci U S A. 2006;103:10011–6.
Brugère J-F, et al. Archaebiotics: proposed therapeutic use of archaea to prevent trimethylaminuria and cardiovascular disease. Gut Microbes. 2014;5:5–10.
Wang Y, et al. A methylotrophic origin of methanogenesis and early divergence of anaerobic multicarbon alkane metabolism. Sci Adv. 2021;7:1453.
Guy L, Ettema TJG. The archaeal “TACK” superphylum and the origin of eukaryotes. Trends Microbiol. 2011;19:580–7.
Vigneron A, Cruaud P, Lovejoy C, Vincent WF. Genomic evidence of functional diversity in DPANN archaea, from oxic species to anoxic vampiristic consortia. ISME Commun. 2022;2:1–10.
Oren A. Euryarchaeota. eLS 1–17 Preprint at https://doi.org/10.1002/9780470015902.a0004243.pub3 (2019).
Chibani CM, et al. A catalogue of 1,167 genomes from the human gut archaeome. Nat Microbiol. 2022;7:48–61.
Houshyar Y, Massimino L, Lamparelli LA, Danese S, Ungaro F. Going beyond bacteria: uncovering the role of archaeome and mycobiome in inflammatory bowel disease. Front Physiol. 2021;12:783295.
Yang K, et al. IDDF2021-ABS-0130 dysbiosis of gut archaea in obesity recovered after bariatric surgery. Gut. 2021;70:A46–A46.
Buan NR. Methanogens: pushing the boundaries of biology. Emerg Top Life Sci. 2018;2:629–46.
Gemperli AC, Dimroth P, Steuber J. Sodium ion cycling mediates energy coupling between complex I and ATP synthase. Proc Natl Acad Sci U S A. 2003;100:839–44.
Grüber G, Manimekalai MSS, Mayer F, Müller V. ATP synthases from archaea: the beauty of a molecular motor. Biochim Biophys Acta. 2014;1837:940–52.
Roswall J, et al. Developmental trajectory of the healthy human gut microbiota during the first 5 years of life. Cell Host Microbe. 2021;29:765-776.e3.
Hanachi M, et al. Longitudinal and comparative analysis of gut microbiota of tunisian newborns according to delivery mode. Front Microbiol. 2022;13:780568.
Saengkerdsub S, Ricke SC. Ecology and characteristics of methanogenic archaea in animals and humans. Crit Rev Microbiol. 2014;40:97–116.
Berghuis BA, et al. Hydrogenotrophic methanogenesis in archaeal phylum Verstraetearchaeota reveals the shared ancestry of all methanogens. Proc Natl Acad Sci U S A. 2019;116:5037–44.
Poretsky R, Rodriguez-R LM, Luo C, Tsementzi D, Konstantinidis KT. Strengths and limitations of 16S rRNA gene amplicon sequencing in revealing temporal microbial community dynamics. PLoS ONE. 2014;9:e93827.
Beauchemin KA, Ungerfeld EM, Eckard RJ, Wang M. Review: Fifty years of research on rumen methanogenesis: lessons learned and future challenges for mitigation. Animal. 2020;14:s2–16.
Thomas CM, Desmond-Le Quéméner E, Gribaldo S, Borrel G. Factors sha** the abundance and diversity of the gut archaeome across the animal kingdom. Nat Commun. 2022;13:3358.
Steinberg LM, Regan JM. mcrA-targeted real-time quantitative PCR method to examine methanogen communities. Appl Environ Microbiol. 2009;75:4435–42.
Qian L, et al. MCycDB: a curated database for comprehensively profiling methane cycling processes of environmental microbiomes. Mol Ecol Resour. 2022;22:1803–23.
PATRIC B13_60.fa. https://patricbrc.org/view/Genome/2774294.14.
PATRIC ERR321646-bin.11. https://patricbrc.org/view/Genome/1291540.8.
Guillaume B, et al. Genome sequence of “candidatus methanomethylophilus alvus” Mx1201, a methanogenic archaeon from the human gut belonging to a seventh order of methanogens. J Bacteriol. 2012;194:6944–5.
Zhou S, et al. Characterization of metagenome-assembled genomes and carbohydrate-degrading genes in the gut microbiota of tibetan pig. Front Microbiol. 2020;11:595066.
PATRIC MGYG-HGUT-02456. https://patricbrc.org/view/Genome/1291540.6.
PATRIC Mx-05. https://patricbrc.org/view/Genome/1291540.4.
Deng F, et al. Weaning time affects the archaeal community structure and functional potential in pigs. Front Microbiol. 2022;13:845621.
Tamburini FB, et al. Short- and long-read metagenomics of urban and rural South African gut microbiomes reveal a transitional composition and undescribed taxa. Nat Commun. 2022;13:926.
**e F, et al. An integrated gene catalog and over 10,000 metagenome-assembled genomes from the gastrointestinal microbiome of ruminants. Microbiome. 2021;9:137.
Rinke C, et al. Validation of picogram- and femtogram-input DNA libraries for microscale metagenomics. PeerJ. 2016;4:e2486.
PATRIC YE315. https://patricbrc.org/view/Genome/1609968.3.
Peng X, et al. Genomic and functional analyses of fungal and bacterial consortia that enable lignocellulose breakdown in goat gut microbiomes. Nat Microbiol. 2021;6:499–511.
Holman DB, Gzyl KE, Mou KT, Allen HK. Weaning age and its effect on the development of the swine gut microbiome and resistome. MSystems. 2021;6:e0068221.
Samuel BS, et al. Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc Natl Acad Sci U S A. 2007;104:10643–8.
PATRIC A54. https://patricbrc.org/view/Genome/1860156.3.
PATRIC MGYG-HGUT-04687. https://patricbrc.org/view/Genome/2173.814.
PATRIC ERR1297821-bin.15. https://patricbrc.org/view/Genome/2173.815.
Bowerman KL, et al. Disease-associated gut microbiome and metabolome changes in patients with chronic obstructive pulmonary disease. Nat Commun. 2020;11:5886.
Lou YC, et al. Infant gut strain persistence is associated with maternal origin, phylogeny, and functional potential including surface adhesion and iron acquisition. bioRxiv; 2021. https://doi.org/10.1101/2021.01.26.428340.
PATRIC ATCC 35061. https://patricbrc.org/view/Genome/420247.28.
Prins RA, Geelen MJH. Rumen characteristics of red deer, fallow deer, and roe deer. J Wildl Manage. 1971;35:673–80.
Yeast-Derived β-1,3-Glucan Substrate Significantly Increased the Diversity of Methanogens During In vitro Fermentation of Porcine Colonic Digesta. J Integr Agric 12, 2229–2234 (2013).
Luo Y, et al. Dietary pea fiber increases diversity of colonic methanogens of pigs with a shift from Methanobrevibacter to Methanomassiliicoccus-like genus and change in numbers of three hydrogenotrophs. BMC Microbiol. 2017;17:17.
Mi J, Peng H, Wu Y, Wang Y, Liao X. Diversity and community of methanogens in the large intestine of finishing pigs. BMC Microbiol. 2019;19:83.
Mao S-Y, Yang C-F, Zhu W-Y. Phylogenetic analysis of methanogens in the pig feces. Curr Microbiol. 2011;62:1386–9.
Luo Y-H, et al. Lean breed Landrace pigs harbor fecal methanogens at higher diversity and density than obese breed Erhualian pigs. Archaea. 2012;2012:605289.
Su Y, Bian G, Zhu Z, Smidt H, Zhu W. Early methanogenic colonisation in the faeces of Meishan and Yorkshire piglets as determined by pyrosequencing analysis. Archaea. 2014;2014:547908.
Federici S, et al. Archaeal microbiota population in piglet feces shifts in response to weaning: Methanobrevibacter smithii is replaced with Methanobrevibacter boviskoreani. FEMS Microbiol Lett. 2015;362:64.
Cao Z, et al. Effect of dietary fiber on the methanogen community in the hindgut of Lantang gilts. Animal. 2016;10:1666–76.
Bin P, et al. Effects of different levels of methionine on sow health and plasma metabolomics during late gestation. Food Funct. 2018;9:4979–88.
Crossfield M, et al. Archaeal and bacterial metagenome-assembled genome sequences derived from pig feces. Microbiol Resour Announc. 2022;11:e0114221.
Aluthge ND, et al. BOARD INVITED REVIEW: the pig microbiota and the potential for harnessing the power of the microbiome to improve growth and health. J Anim Sci. 2019;97:3741–57.
Hoffmann C, et al. Archaea and fungi of the human gut microbiome: correlations with diet and bacterial residents. PLoS ONE. 2013;8:e66019.
Patil Y, Gooneratne R, Ju X-H. Interactions between host and gut microbiota in domestic pigs: a review. Gut Microbes. 2020;11:310–34.
Vatanen T, et al. Genomic variation and strain-specific functional adaptation in the human gut microbiome during early life. Nat Microbiol. 2019;4:470–9.
Chen L, et al. The maturing development of gut microbiota in commercial piglets during the weaning transition. Front Microbiol. 2017;8:1688.
Qin Y, et al. Combined effects of host genetics and diet on human gut microbiota and incident disease in a single population cohort. Nat Genet. 2022;54:134–42.
Chen C, et al. Expanded catalog of microbial genes and metagenome-assembled genomes from the pig gut microbiome. Nat Commun. 2021;12:1106.
Wibowo MC, et al. Reconstruction of ancient microbial genomes from the human gut. Nature. 2021;594:234–9.
Bäckhed F, et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe. 2015;17:690–703.
Jovel J, et al. Metagenomics versus metatranscriptomics of the murine gut microbiome for assessing microbial metabolism during inflammation. Front Microbiol. 2022;13:829378.
Segura-Wang M, Grabner N, Koestelbauer A, Klose V, Ghanbari M. Genome-resolved metagenomics of the chicken gut microbiome. Front Microbiol. 2021;12:726923.
Haley BJ, Kim S-W, Salaheen S, Hovingh E, Van Kessel JAS. Differences in the microbial community and resistome structures of feces from preweaned calves and lactating dairy cows in commercial dairy herds. Foodborne Pathog Dis. 2020;17:494–503.
Watson AR, et al. Metabolic independence drives gut microbial colonization and resilience in health and disease. Genome Biol. 2023;24:78.
Delmont TO. Binning giant viruses and their close relatives with anvi’o. Meren Lab https://merenlab.org/2022/01/03/giant-viruses/.
Metapangenomics of Rothia and H. parainfluenzae. Daniel Utter (2023). https://dutter.github.io/projects/oral_metapan.
Wright A-DG, Auckland CH, Lynn DH. Molecular diversity of methanogens in feedlot cattle from Ontario and Prince Edward Island. Canada Appl Environ Microbiol. 2007;73:4206–10.
Zhou M, Hernandez-Sanabria E, Guan LL. Characterization of variation in rumen methanogenic communities under different dietary and host feed efficiency conditions, as determined by PCR-denaturing gradient gel electrophoresis analysis. Appl Environ Microbiol. 2010;76:3776–86.
Danielsson R, Schnürer A, Arthurson V, Bertilsson J. Methanogenic population and CH4 production in Swedish dairy cows fed different levels of forage. Appl Environ Microbiol. 2012;78:6172–9.
Skillman LC, Evans PN, Strömpl C, Joblin KN. 16S rDNA directed PCR primers and detection of methanogens in the bovine rumen. Lett Appl Microbiol. 2006;42:222–8.
Hernandez-Sanabria E, et al. Influence of sire breed on the interplay among rumen microbial populations inhabiting the rumen liquid of the progeny in beef cattle. PLoS ONE. 2013;8:e58461.
Malik PK, et al. Comparison of enteric methane yield and diversity of ruminal methanogens in cattle and buffaloes fed on the same diet. PLoS ONE. 2021;16:e0256048.
McCabe MS, et al. Illumina MiSeq phylogenetic amplicon sequencing shows a large reduction of an uncharacterised succinivibrionaceae and an increase of the methanobrevibacter gottschalkii clade in feed restricted cattle. PLoS ONE. 2015;10:e0133234.
Miller TL, Lin C. Description of Methanobrevibacter gottschalkii sp. Nov., Methanobrevibacter thaueri sp. nov., Methanobrevibacter woesei sp. nov. and Methanobrevibacter wolinii sp. nov. Int J Syst Evol Microbiol. 2002;52:819–22.
Gilroy R, et al. Extensive microbial diversity within the chicken gut microbiome revealed by metagenomics and culture. PeerJ. 2021;9:e10941.
Bowerman KL, et al. Effects of laboratory domestication on the rodent gut microbiome. ISME Commun. 2021;1:1–14.
Beresford-Jones BS, et al. The Mouse Gastrointestinal Bacteria Catalogue enables translation between the mouse and human gut microbiotas via functional map**. Cell Host Microbe. 2022;30:124-138.e8.
Zhu J, et al. An expanded gene catalog of mouse gut metagenomes. mSphere. 2021;6:10.
**ao L, et al. A catalog of the mouse gut metagenome. Nat Biotechnol. 2015;33:1103–8.
Kieser S, Zdobnov EM, Trajkovski M. Comprehensive mouse microbiota genome catalog reveals major difference to its human counterpart. PLoS Comput Biol. 2022;18:e1009947.
Bergamaschi M, et al. Gut microbiome composition differences among breeds impact feed efficiency in swine. Microbiome. 2020;8:110.
Massacci FR, et al. Late weaning is associated with increased microbial diversity and Faecalibacterium prausnitzii abundance in the fecal microbiota of piglets. Anim Microbiome. 2020;2:2.
de la Cuesta-Zuluaga J, Spector TD, Youngblut ND, Ley RE. Genomic insights into adaptations of trimethylamine-utilizing methanogens to diverse habitats, including the human gut. mSystems. 2021;6:e009390.
Mihajlovski A, Doré J, Levenez F, Alric M, Brugère J-F. Molecular evaluation of the human gut methanogenic archaeal microbiota reveals an age-associated increase of the diversity. Environ Microbiol Rep. 2010;2:272–80.
Camara A, et al. Clinical evidence of the role of Methanobrevibacter smithii in severe acute malnutrition. Sci Rep. 2021;11:5426.
Birgel D, et al. Methanogenesis produces strong 13C enrichment in stromatolites of Lagoa Salgada, Brazil: a modern analogue for Palaeo-/Neoproterozoic stromatolites? Geobiology. 2015;13:245–66.
Wang J-X, **e W, Zhang YG, Meador TB, Zhang CL. Evaluating production of Cyclopentyl Tetraethers by marine group II Euryarchaeota in the Pearl River Estuary and Coastal South China Sea: potential impact on the TEX86 paleothermometer. Front Microbiol. 2017;8:2077.
Orsi WD, et al. Climate oscillations reflected within the microbiome of Arabian Sea sediments. Sci Rep. 2017;7:6040.
Cavalazzi B, et al. Cellular remains in a ~3.42-billion-year-old subseafloor hydrothermal environment. Sci Adv. 2021;7:3963.
More KD, et al. Subseafloor Archaea reflect 139 kyrs of paleodepositional changes in the northern Red Sea. Geobiology. 2021;19:162–72.
Baxter BK. Great Salt Lake microbiology: a historical perspective. Int Microbiol. 2018;21:79–95.
Yang J, et al. Sedimentary archaeal amoA gene abundance reflects historic nutrient level and salinity fluctuations in Qinghai Lake. Tibetan Plateau Sci Rep. 2015;5:18071.
Pearson A, et al. Factors controlling the distribution of archaeal tetraethers in terrestrial hot springs. Appl Environ Microbiol. 2008;74:3523–32.
Neubeck A, et al. Microbial community structure in a serpentine-hosted abiotic gas seepage at the chimaera ophiolite, Turkey. Appl Environ Microbiol. 2017;83:e03430.
Weyrich LS, et al. Neanderthal behaviour, diet, and disease inferred from ancient DNA in dental calculus. Nature. 2017;544:357–61.
KEGG PATHWAY: Methane metabolism + Reference pathway. https://www.genome.jp/pathway/map00680.
Kaster A-K, Moll J, Parey K, Thauer RK. Coupling of ferredoxin and heterodisulfide reduction via electron bifurcation in hydrogenotrophic methanogenic archaea. Proc Natl Acad Sci U S A. 2011;108:2981–6.
Yan Z, Ferry JG. Electron bifurcation and confurcation in Methanogenesis and Reverse Methanogenesis. Front Microbiol. 2018;9:1322.
Berg IA, et al. Autotrophic carbon fixation in archaea. Nat Rev Microbiol. 2010;8:447–60.
KEGG MODULE: M00567. https://www.genome.jp/module/M00567.
Goldman AD, Leigh JA, Samudrala R. Comprehensive computational analysis of Hmd enzymes and paralogs in methanogenic Archaea. BMC Evol Biol. 2009;9:199.
Gottschalk G, Thauer RK. The Na+-translocating methyltransferase complex from methanogenic archaea. Biochim Biophys Acta. 2001;1505:28–36.
Thauer RK, Kaster A-K, Seedorf H, Buckel W, Hedderich R. Methanogenic archaea: ecologically relevant differences in energy conservation. Nat Rev Microbiol. 2008;6:579–91.
Söllinger A, Urich T. Methylotrophic methanogens everywhere — physiology and ecology of novel players in global methane cycling. Biochem Soc Trans. 2019;47:1895–907.
Muñoz-Tamayo R, et al. Hydrogenotrophic methanogens of the mammalian gut: Functionally similar, thermodynamically different-a modelling approach. PLoS ONE. 2019;14:e0226243.
Dong M, et al. In vitro methanol production from methyl coenzyme M using the Methanosarcina barkeri MtaABC protein complex. Biotechnol Prog. 2017;33:1243–9.
Hoeppner A, et al. Structure of the corrinoid:coenzyme M methyltransferase MtaA from Methanosarcina mazei. Acta Crystallogr D Biol Crystallogr. 2012;68:1549–57.
Muñoz-Velasco I, et al. Methanogenesis on early stages of life: ancient but not primordial. Orig Life Evol Biosph. 2018;48:407–20.
Hinderberger D, et al. Coordination and binding geometry of methyl-coenzyme M in the red1m state of methyl-coenzyme M reductase. J Biol Inorg Chem. 2008;13:1275–89.
Zhang J-W, et al. Newly discovered Asgard archaea Hermodarchaeota potentially degrade alkanes and aromatics via alkyl/benzyl-succinate synthase and benzoyl-CoA pathway. ISME J. 2021;15:1826–43.
Lang K, et al. New mode of energy metabolism in the seventh order of methanogens as revealed by comparative genome analysis of “Candidatus methanoplasma termitum.” Appl Environ Microbiol. 2015;81:1338–52.
Zhang C-J, Pan J, Liu Y, Duan C-H, Li M. Genomic and transcriptomic insights into methanogenesis potential of novel methanogens from mangrove sediments. Microbiome. 2020;8:94.
Evans PN, et al. An evolving view of methane metabolism in the Archaea. Nat Rev Microbiol. 2019;17:219–32.
MetaCyc EC 6.2.1.1. https://biocyc.org/META/NEW-IMAGE?type=REACTION&object=ACETATE--COA-LIGASE-RXN.
Grossart H-P, Frindte K, Dziallas C, Eckert W, Tang KW. Microbial methane production in oxygenated water column of an oligotrophic lake. Proc Natl Acad Sci U S A. 2011;108:19657–61.
Lavergne C, et al. Temperature differently affected methanogenic pathways and microbial communities in sub-Antarctic freshwater ecosystems. Environ Int. 2021;154:106575.
Kotsyurbenko OR, et al. Acetoclastic and hydrogenotrophic methane production and methanogenic populations in an acidic West-Siberian peat bog. Environ Microbiol. 2004;6:1159–73.
Martínez-Álvaro M, et al. Identification of complex rumen microbiome interaction within diverse functional niches as mechanisms affecting the variation of methane emissions in Bovine. Front Microbiol. 2020;11:659.
Ferry JG. Fundamentals of methanogenic pathways that are key to the biomethanation of complex biomass. Curr Opin Biotechnol. 2011;22:351–7.
Mand TD, Metcalf WW. Energy conservation and hydrogenase function in Methanogenic Archaea, in particular the genus Methanosarcina. Microbiol Mol Biol Rev. 2019;83:10.
Fernandes J, et al. Age, dietary fiber, breath methane, and fecal short chain fatty acids are interrelated in Archaea-positive humans. J Nutr. 2013;143:1269–75.
Hoedt EC, et al. Culture- and metagenomics-enabled analyses of the Methanosphaera genus reveals their monophyletic origin and differentiation according to genome size. ISME J. 2018;12:2942–53.
Tajima K, Aminov R. Structure and function of a nonruminant gut: a porcine model. In: Puniya AK, Singh R, Kamra DN, editors. Rumen microbiology: from evolution to revolution. Springer India; 2015. p. 47–75.
Hoedt EC, et al. Differences down-under: alcohol-fueled methanogenesis by archaea present in Australian macropodids. ISME J. 2016;10:2376–88.
Meziti A, et al. The reliability of metagenome-assembled genomes (MAGs) in representing natural populations: insights from comparing mags against isolate genomes derived from the same Fecal sample. Appl Environ Microbiol. 2021;87:e02593.
Feehan, B. et al. Stability and volatility shape the gut bacteriome and mycobiome dynamics in a pig model. bioRxiv (2022). https://doi.org/10.1101/2022.02.02.478893.
Eren AM, et al. Community-led, integrated, reproducible multi-omics with anvi’o. Nat Microbiol. 2021;6:3–6.
Shaiber A, et al. Functional and genetic markers of niche partitioning among enigmatic members of the human oral microbiome. Genome Biol. 2020;21:292.
Mölder F, et al. Sustainable data analysis with Snakemake. F1000Res. 2021;10:33.
Minoche AE, Dohm JC, Himmelbauer H. Evaluation of genomic high-throughput sequencing data generated on Illumina HiSeq and genome analyzer systems. Genome Biol. 2011;12:R112.
Li D, Liu C-M, Luo R, Sadakane K, Lam T-W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinf. 2015;31:1674–6.
Hyatt D, et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinf. 2010;11:119.
Parks DH, et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36:996–1004.
Oren A, et al. Proposal to include the rank of phylum in the international code of nomenclature of prokaryotes. Int J Syst Evol Microbiol. 2015;65:4284–7.
hmm-source [artifact]. Anvi’o Home https://anvio.org/help/7/artifacts/hmm-source/.
HMMER. http://hmmer.org/.
COG - NCBI. https://www.ncbi.nlm.nih.gov/research/cog.
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Langmead B, Wilks C, Antonescu V, Charles R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics. 2019;35:421–32.
Danecek P, et al. Twelve years of SAMtools and BCFtools. Gigascience. 2021;10:8.
Alneberg J, et al. Binning metagenomic contigs by coverage and composition. Nat Methods. 2014;11:1144–6.
Alneberg J, et al. CONCOCT: clustering contigs on coverage and composition. ar**v [q-bio.GN] (2013).
Murat Eren A, et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ. 2015;3:e1319.
Pritchard L, Glover RH, Humphris S, Elphinstone JG, Toth IK. Genomics and taxonomy in diagnostics for food security: soft-rotting enterobacterial plant pathogens. Anal Methods. 2016;8:12–24.
Lee M. Anvi’o “views” demystified. Meren Lab https://merenlab.org/2017/05/08/anvio-views/.
Quince C, et al. DESMAN: a new tool for de novo extraction of strains from metagenomes. Genome Biol. 2017;18:181.
RStudio Team. RStudio: integrated development environment for R. RStudio. https://www.rstudio.com/ (2020).
pheatmap function - RDocumentation. https://www.rdocumentation.org/packages/pheatmap/versions/1.0.12/topics/pheatmap.
Villanueva RAM, Chen ZJ. ggplot2: elegant graphics for data analysis. Measurement. 2019;17:160–7.
Wickham H. Forcats: tools for working with categorical variables (Factors) (2022).
Wickham H, François R, Henry L, Müller K. dplyr: a grammar of data manipulation (2022).
ggplot2 Based Publication Ready Plots. https://rpkgs.datanovia.com/ggpubr/index.html.
Brettin T, et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep. 2015;5:8365.
Davis JJ, et al. The PATRIC bioinformatics resource center: expanding data and analysis capabilities. Nucleic Acids Res. 2020;48:D606–12.
Developers IW. Inkscape. https://inkscape.org/.
Acknowledgements
Our team is very grateful to the large number of individuals and organizations which assisted us in performing this research. We thank members of the Kansas State University swine team (Frank Martin, Mark Nelson, Duane Baughman, and Julia Holen) for aiding in the sample collection. Gratitude is also extended to the University of Kansas Medical Center Genome Sequencing Facility for their expertise and assistance in sequencing including: Clark Bloomer, Dr. Veronica Cloud, Rosanne Skinner, and Yafen Niu.
Funding
We greatly appreciate assistance from the following sources: Kansas State University Interdepartmental Genetics Program (fellowship for Brandi Feehan), Global Food Systems Seed Grant Program, Kansas Intellectual and Developmental Disabilities Research Center (NIH U54 HD 090216), the Molecular Regulation of Cell Development and Differentiation—COBRE (P30 GM122731-03)—the NIH S10 High-End Instrumentation Grant (NIH S10OD021743) and the Frontiers CTSA Grant (UL1TR002366) at the University of Kansas Medical Center, Kansas City, KS 66160.
Author information
Authors and Affiliations
Contributions
BF, RG and STML designed the study. Sample collection was performed by BF. BF, VD, KR, KW, and SP completed DNA extraction and Nanodrop and Qubit quality analysis. BF and QR performed anvi’o and PATRIC bioinformatic analyses. BF and STML attributed biological relevance, wrote the manuscript, prepared figures, and Additional files. BF and STML performed major manuscript and figure refinement while remaining authors contributed to lighter refinement. MC provided valuable insights which aided in study progression and manuscript preparation. All authors read, contributed to manuscript revision, and approved the submitted version.
Corresponding author
Ethics declarations
Ethical approval and consent to participate
Pigs were managed according to the Kansas State University Institutional Animal Care and Use Committee (IACUC) approved protocol #4036, and methods are reported according to ARRIVE guidelines. The authors also confirmed that all methods were performed in accordance with relevant guidelines and regulations, and we affirmed that all methods were approved by Kansas State University.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S3.
Anvi’o results from initial bins and resulting redundant and nonredundant MAGs, including taxonomic classification detailing archaeal MAGs and PATRIC genome IDs [137].
Additional file 2: Table S4.
PATRIC RAStk and COGG gene annotations for methanogen MAGs.
Additional file 3: Fig. S1.
Complete, non-collapsed, Methanobacteriales and Methanomassiliicoccales phylogenetic trees with statistics.
Additional file 4: Fig. S2:
Relative abundance of archaea-MAGs (rows) across all individual sample metagenomes (columns) (Preweaning [P]; nursery [N]; growth adult [G]).
Additional file 5: Table S5.
Single nucleotide variant (SNV) results.
Additional file 6: Table S6.
Metadata from metagenomes utilized in map** to our archaeal MAGs (swine [62], human infant [64], human adult and palaeofaeces [63], mice [65], chicken [66], and cattle [67]).
Additional file 7: Table S7.
Detection results from metagenome map** to archaea-MAGs.
Additional file 8: Table S1.
Demographics (diet, birth date, housing group, etc.) of swine hosts and dams, and sample metadata (swine age, host ID, stage and general health information, etc.).
Additional file 9: Table S2.
Sequencing and assembly analysis including: metagenomic read counts initially obtained and assembly statistics according to co-assembly stage.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Feehan, B., Ran, Q., Dorman, V. et al. Novel complete methanogenic pathways in longitudinal genomic study of monogastric age-associated archaea. anim microbiome 5, 35 (2023). https://doi.org/10.1186/s42523-023-00256-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s42523-023-00256-6