Abstract
Background
Despite advancements in chronic myeloid leukemia (CML) therapy with tyrosine kinase inhibitors (TKIs), resistance and intolerance remain significant challenges. Leukemia stem cells (LSCs) and TKI-resistant cells rely on altered mitochondrial metabolism and oxidative phosphorylation. Targeting rewired energy metabolism and inducing non-apoptotic cell death, along with the release of damage-associated molecular patterns (DAMPs), can enhance therapeutic strategies and immunogenic therapies against CML and prevent the emergence of TKI-resistant cells and LSC persistence.
Methods
Transcriptomic analysis was conducted using datasets of CML patients' stem cells and healthy cells. DNA damage was evaluated by fluorescent microscopy and flow cytometry. Cell death was assessed by trypan blue exclusion test, fluorescent microscopy, flow cytometry, colony formation assay, and in vivo Zebrafish xenografts. Energy metabolism was determined by measuring NAD+ and NADH levels, ATP production rate by Seahorse analyzer, and intracellular ATP content. Mitochondrial fitness was estimated by measurements of mitochondrial membrane potential, ROS, and calcium accumulation by flow cytometry, and morphology was visualized by TEM. Bioinformatic analysis, real-time qPCR, western blotting, chemical reaction prediction, and molecular docking were utilized to identify the drug target. The immunogenic potential was assessed by high mobility group box (HMGB)1 ELISA assay, luciferase-based extracellular ATP assay, ectopic calreticulin expression by flow cytometry, and validated by phagocytosis assay, and in vivo vaccination assay using syngeneic C57BL/6 mice.
Results
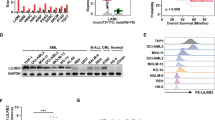
Transcriptomic analysis identified metabolic alterations and DNA repair deficiency signatures in CML patients. CML patients exhibited enrichment in immune system, DNA repair, and metabolic pathways. The gene signature associated with BRCA mutated tumors was enriched in CML datasets, suggesting a deficiency in double-strand break repair pathways. Additionally, poly(ADP-ribose) polymerase (PARP)1 was significantly upregulated in CML patients’ stem cells compared to healthy counterparts. Consistent with the CML patient DNA repair signature, treatment with the methylated indolequinone MAC681 induced DNA damage, mitochondrial dysfunction, calcium homeostasis disruption, metabolic catastrophe, and necroptotic-like cell death. In parallel, MAC681 led to PARP1 degradation that was prevented by 3-aminobenzamide. MAC681-treated myeloid leukemia cells released DAMPs and demonstrated the potential to generate an immunogenic vaccine in C57BL/6 mice. MAC681 and asciminib exhibited synergistic effects in killing both imatinib-sensitive and -resistant CML, opening new therapeutic opportunities.
Conclusions
Overall, increasing the tumor mutational burden by PARP1 degradation and mitochondrial deregulation makes CML suitable for immunotherapy.
Similar content being viewed by others
Introduction
Tyrosine kinase inhibitors (TKIs) represent the frontline treatment against chronic myeloid leukemia (CML), characterized by the t(9:22) chromosomal translocation, resulting in the breakpoint cluster region-Abelson (BCR-ABL)1 fusion oncogene with constitutively activated Abl tyrosine kinase activity. Despite TKI efficacy, resistant mutants turn CML into a controllable yet persisting chronic disease, interfering with the patient’s quality of life [1,2,3]. Moreover, one-third of newly diagnosed patients develop primary or secondary resistance toward approved TKIs, and clinical relapse occurs [4]. Imatinib resistance originates from BCR-ABL-dependent and -independent mechanisms, including leukemia stem cell (LSC) treatment insensitivity. LSCs and TKI/therapy-resistant cells depend on mitochondrial metabolism and oxidative phosphorylation (OXPHOS) to meet their energy demands [5, 6], in contrast to mature CML cells or normal hematopoietic stem cells (HSCs).
The goal of therapeutic approaches for CML treatment is to eliminate LSCs by identifying and targeting genetic alterations and molecular pathways that contribute to their survival and resistance to apoptosis. Efforts must be undertaken to develop combination therapies targeting tyrosine kinases with novel drug candidates that specifically target genes or molecules necessary for the survival mechanisms of CML LSCs [7]. Accordingly, identifying and targeting rewired energy metabolism is a relevant strategy for develo** non-ABL-related therapeutic options for treating TKI-resistant CML patients, preventing the appearance of TKI-resistant cells, and LSC persistence.
Nicotinamide adenine dinucleotide (NAD) is a substrate for NAD+-dependent enzymes and a major coenzyme in bioenergetic processes. The NAD+/NADH ratio modulates metabolic processes, including oxidative phosphorylation (OXPHOS), tricarboxylic acid (TCA) cycle, fatty acid oxidation (FAO), and glycolysis. NADH serves as a central hydride donor to mitochondrial ATP synthesis. Maintaining NAD+ levels and the NAD+/NADH ratio is crucial for mitochondrial function, ATP production, and homeostasis. Hence, NAD metabolism is a master regulator linking metabolic processes to OXPHOS [8]. Most cells replenish their NAD+ pool primarily via the NAD+ salvage pathway by recycling nicotinamide (NAM), generated by NAD+-consuming enzymes predominantly by poly (ADP-ribose) polymerase 1 (PARP1) and sirtuin (SIRT)1 [9, 18].
In addition to the cytotoxic activity, whether by apoptosis or controlled necrosis, the activation of the immune-mediated destruction of tumor cells is deeply involved in the curative effect of a given anticancer agent. There is a growing interest in promoting anti-tumor immune responses as a potential alternative strategy for cancer treatment [19]. Accordingly, the induction of immunogenic cell death (ICD) is an attractive approach to fighting cancer malignancies [20]. Pharmacological mediators of ICD stimulate the host’s immune system via the release of damage‐associated molecular patterns (DAMPs), eventually triggering a persistent immune response [21].
We investigated a TKI-independent, immunogenic cell death mechanism triggered by a novel, methylated indolequinone (MAC681). This compound targets PARP1 degradation, exacerbating mitochondrial dysfunction and inducting a non-canonical, necroptotic-like cell death. We used MAC681-treated, dying, or dead cells to demonstrate immunogenic vaccination against myeloid leukemia.
Materials and methods
Computational analysis of public CML datasets
This study utilized several publicly available datasets, which were processed as follows:
-
A)
GSE5550 [22]: 17 patients’ bone marrow samples sorted for CD34+ hematopoietic stem and progenitor cells, 8 healthy donors, and 9 untreated CML patients in chronic phase were downloaded from GEO. The samples were processed and normalized using the robust Multi-Array Average (RMA) expression measure from the “affy” package in R [23]. All the probes were analyzed separately; if multiple probes representing the same gene were present, only the most significant probe was kept.
-
B)
GSE97562 [24] and GSE47927 [25]: A total of 19 patients’ bone marrow samples sorted for CD34+, 8 healthy donors, and 11 untreated CML patients in chronic phase raw CEL files were downloaded from GEO (Supp. Table S1A). Samples were processed and normalized using the RMA expression measure from the “affy” package in R [23]. To correct for the batch effect between datasets, we employed the combat function of the "sva" R package [26].
-
C)
The Microarray Innovations in LEukemia (MILE) study, GSE13159 [27]: A total of 66 untreated CML patients’ bone marrow mononuclear cells raw CEL files were downloaded from GEO. Samples were processed and normalized using the robust RMA expression measure from the “affy” package in R [23]. Samples were divided into 4 groups based on the quartiles of PARP1 expression. The high and low quartiles were subsequently used for the differential expression analysis.
-
D)
The Cancer Cell Line Encyclopedia (CCLE) dataset: Gene expression transcript per million (TPM) normalized reads from all the cancer cell lines (n = 1165) were downloaded from the Depmap portal website (https://depmap.org/portal/download/all/) [28].
Differential expression analysis
To identify differentially expressed genes (DEGs) between healthy and CML CD34+ cells or between CML patients with high and low expression of PARP1, we utilized the “limma” R package [29]. We applied a threshold of |log2(FC)|> 0.3 and a false discovery rate (FDR) < 0.05. The resulting DEGs were visualized using volcano and generated using the “ggplot2” R package [30] (https://ggplot2.tidyverse.org).
Pathways and enrichment analysis
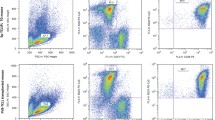
We performed functional enrichment analysis using GO terms, GSE analysis (GSEA), and Reactome pathways to analyze the identified DEGs comprehensively. GO terms and GSEA analysis were performed using “fgsea” and “msigdb” R packages. Reactome enrichments were performed on the web-based application [31] (https://reactome.org/). Pathways were considered significant when the FDR ≤ 0.05. We re-grouped each pathway for its higher hierarchical annotation in Reactome and calculated a weighted percentage of pathways enriched. Genes belonging to the NAD+ metabolome pathway were collected from **ing on the nucleus (24 h), a hallmark of necroptosis, are seen (top panel). Mitochondrial alterations (lower panel) induced by MAC681 are also visualized. G Time-dependent induction of mitochondrial membrane potential loss (left panel) and cell death induction (right panel) by MAC681 (5 µM). MMP was analyzed after incubation with MitoTracker Red (100 nM) by flow cytometry. Mitochondrial oxidative phosphorylation uncoupler CCCP (50 µM, 20 min) was used as a positive control. Flow cytometry analyzed cell death induction by propidium iodide (PI) uptake after 15 min staining with PI (0.5 µg/mL). DMSO (50%, 1 min) was used as a positive control (one-way ANOVA, Dunnett's multiple comparisons test *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). H Cells were treated with MAC681 (5 µM, 6 h) and analyzed by fluorescence microscopy after staining with Rhod2-AM (Rhod, 2.5 µM), MitoTracker Green (MTG, 100 nM), and Hoechst. The intensity of the Rhod2-AM signal was quantified in regions stained with MitoTracker Green, and the co-occurrence and correlation were quantified by Manders’, Spearman’s, and Pearson’s coefficients. Double positive cells were analyzed, and the average of three independent experiments was shown (double-sided unpaired t-test ***p ≤ 0.001). I Cytosolic Ca2+ levels of cells treated with MAC681 (5 µM) for indicated time points, stained by Fluo4-AM, were measured by flow cytometry (double-sided unpaired t-test *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). J Cytosolic Ca2+ levels of cells treated with MAC681 (5 µM) for indicated time points in the presence of EGTA (650 µM), followed by TSG (10 nM) treatment, were measured by Fluo4-AM every 15 s by flow cytometry, (AUC, double-sided unpaired t-test **p ≤ 0.01, ***p ≤ 0.001). K Colony formation assay with K-562 cells treated with MAC681 at the indicated concentrations. Images are representative of three independent experiments (One-way ANOVA, Dunnett's multiple comparisons test **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). L In vivo zebrafish xenografts model of tumor mass formation after injection of fluorescent CellTracker™ CM-DiI Dye-stained K-562 cells pre-treated for the indicated time. M MAC681 decreases the viability of CML patients’ PBMCs in dose- and time-dependent manners. CML primary cells were treated ex-vivo with indicated concentrations of MAC681, and cell viability was evaluated after 4, 12, and 24 h by trypan blue exclusion test. The histogram corresponds to the mean ± SD (n = 2 CML patients). N Analysis of ALDH activity in K-562 cells. Cells were treated with MAC681 (5 µM, 12 h) or Imatinib (0.25 µM, 24 h), and the ALDH inhibitor diethylaminobenzaldehyde (DEAB) was used to distinguish cell subpopulations with low and high ALDH activity. Representative dot plots showing the percentage of ALDH-positive cells and corresponding quantifications (right panel) of three independent experiments are presented (left panel) (one-way ANOVA, Tukey’s multiple comparisons test *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001)
Considering the upregulation of PARP1 in CML patients' datasets and the upregulation of metabolic pathways in patients expressing high levels of PARP1, we wondered if such a strong induction of DNA damage could result in the PARP-dependent depletion of total cellular NAD+ levels. Indeed, after MAC681 treatment, we detected an early, time-dependent decrease in cellular NAD+. MAC681 treatment reduced NAD+ by 20% in 30 min, 59% in 1 h, and 87% in 2 h. NAD+ depletion was followed by a reduction in NADH levels. An overall time-dependent decrease in the NAD+/NADH ratio was detected (Fig. 2B). Moreover, Hoechst/PI nuclear morphology analysis indicated that MAC681 induced necrotic cell death in K-562 cells and that the PARP1 inhibitor, 3ABA rescued cells from MAC681-induced lethality (Fig. 2C).
As metabolic pathways, including OXPHOS, rely on NAD availability, PARP1 activation promotes a rapid decrease in mitochondrial oxygen consumption [40]. We detected a significantly decreased OCR in MAC681-treated cells compared to the control (Suppl. Fig. S2A). Moreover, in K-562 cells, the basal ATP production rate relied more on oxidative phosphorylation than glycolysis (61% versus 39%, respectively) (Sup. Fig. S2B). Within two hours, MAC681 treatment induced a 43% reduction in the total ATP production rate compared to untreated cells (Fig. 2D). Furthermore, the MAC681-induced decrease in the ATP production rate was mainly due to decreased mitochondrial ATP production rate (Fig. 2D). Subsequently, after 4 h of treatment, we observed a 53.2% reduction in total intracellular ATP levels. After 8 h, ATP levels were reduced by 92% compared to controls (Fig. 2E).
The loss of NAD+ precedes the induction of mitochondrial depolarization and mitochondrial outer membrane permeability transition [41]. TEM analysis confirmed time-dependent mitochondrial alterations after MAC681 treatment, disrupting mitochondrial cristae after 4 h and mitochondrial swelling after 8 h (Fig. 2F, lower panel). In addition, the cells adopted a necroptotic phenotype (Fig. 2F, upper panel). In agreement with morphological changes and ATP loss, MAC681 triggered a time-dependent loss of mitochondrial membrane potential (MMP) (Fig. 2G, left panel). The MMP was strongly reduced at 4 h and 8 h of treatment, although cells remained viable, considering the minimal PI staining intake (Fig. 2G, right panel).
Since mitochondrial calcium overload could have triggered these mitochondrial morphological alterations, we assessed calcium accumulation in mitochondria by co-localizing Rhod2-AM mitochondrial Ca2+ dye with MitoTracker green after 6 h of treatment. Our results indicate that MAC681 induced Ca2+ accumulation in 67% of the mitochondria (Fig. 2H and Sup. Fig. S2C). To further analyze the perturbation of calcium homeostasis, we assessed the effect of MAC681 on cytosolic Ca2+ accumulation after Fluo4-AM staining. We detected an immediate increase in cytosolic Ca2+ at 2 h, which decreased after 4 h and returned to basal values at 8 h. At 10 h, we observed a secondary increase in cytosolic Ca2+ (Fig. 2I).
To further ascertain the origin of the observed cytosolic Ca2+ accumulation, we used EGTA to chelate extracellular Ca2+ influx. Under these conditions, adding the sarcoplasmic/endoplasmic reticulum Ca2+-ATPase (SERCA) inhibitor thapsigargin (TSG) led to a rapid accumulation of cytosolic Ca2+ in control cells. Interestingly, in MAC681-treated cells for 2 h, the addition of TSG significantly reduced the increase, which was abolished at 4 h (Fig. 2J). These data show that MAC681 rapidly depleted endoplasmic reticulum Ca2+ in line with the observed initial cytosolic Ca2+ increase.
PARP1 activation after DNA damage contributes to mitochondrial Ca2+ dysregulation and subsequent calpain activation [42, 43] to promote truncation and translocation of apoptosis-inducing factor (AIF) from the mitochondria to the nucleus. In MAC681-treated cells, AIF co-localized with nuclear Hoechst staining (Sup. Fig. S2D-E) compared to untreated controls. Moreover, we observed that calpain inhibitor PD 150606 prevented the PARP1 degradation (Sup. Fig. S2F), suggesting the involvement of calpain activation during MAC681-induced metabolic catastrophe.
The translational potential of MAC681 as an antileukemic agent in vitro and in vivo
Considering the observed antimetabolic potential of the methylated indolequinone MAC681, we assessed its effects on the viability in four human leukemia cells (K-562, K-562R, Jurkat, and U-937) (Sup. Fig. S2G). To assess differential toxicity, we treated the non-cancerous, proliferating cell line RPMI1788 representing healthy B lymphocytes and PBMCs obtained from healthy donors with increasing concentrations of MAC681. RPMI 1788 cells were 65-fold and PBMCs 51-fold less affected compared to K-562 cells when treated at 10 µM after 72 h (Sup. Fig. S2H).
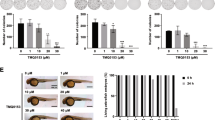
Considering the above-mentioned antileukemic effect of MAC681 at low µM concentrations, we investigated the effect of MAC681 by CFA. Results showed a dose-dependent decrease in the colony formation capacity of three chronic myeloid leukemia cell lines (K-562, KBM-5, Meg-01) and two acute myeloid leukemia cell lines (U-937, HL-60) (Fig. 2K and Sup. Fig. S2I-J). Next, we extended CFA results toward a zebrafish xenograft model to provide a more solid basis for the translational value of MAC681. In vivo data showed time-dependent inhibition of tumor mass formation (Fig. 2L and Sup. Fig. S2K). Moreover, we treated ex-vivo primary CML cells from two CML patients with MAC681. As seen in Fig. 2M, the MAC681 efficiently reduced the viability of CML patients’ PBMCs in dose- and time-dependent manners. We next investigated if MAC681 could target ALDH-positive cells. High expression of ALDH has been reported for normal and cancer stem and progenitor cells of various lineages, including hematopoietic cells [44, 45]. Elevated ALDH activity is an established marker for the identification of hematopoietic cells with stem-like characteristics. Our data shown in Fig. 2N provide evidence that MAC681 (5 µM, 12 h) is targeting preferentially the ALDH-positive population, decreasing the amount of ALDH bright cells by 93.3%. Interestingly, imatinib treatment (0.25 µM) at 24 h reduced the ALDH-positive population only by 42.4%. Our data validated MAC681 as an attractive antileukemic agent for further mechanistic investigations.
Off-target prevention of MAC681-induced cell death by the PARP inhibitor 3-aminobenzamide
As we previously detected a protective effect of the PARP inhibitor 3ABA against MAC681-induced cell death (Fig. 2C), we next assessed the effect of the 3ABA pre-treatment on the PARP1 protein level. Surprisingly, we observed the disappearance of full-length PARP1 (116 kDa) without any cleaved fragments after 8 h of MAC681 treatment alone. 3ABA pre-treatment prevented this PARP1 reduction (Fig. 3A). Hence, we analyzed the kinetics of the atypical MAC681-induced PARP1 reduction (Fig. 3B). We used an additional antibody able to recognize both apoptotic as well as necrotic PARP1 cleaved fragments (Supp. Fig. S3A). However, we did not detect any cleavage fragments either. In addition, we observed that MAC681 induced a time-dependent downregulation of PARP1 mRNA expression levels, which is prevented in the presence of 3ABA (Supp. Fig. S3B). Next, we verified whether the disappearance of full-length PARP1 might be related to its low mRNA levels after MAC681 treatment. Accordingly, we analyzed the stability of PARP1 in the presence of cycloheximide that blocks the elongation phase of protein synthesis (Supp. Fig. S3C). We showed that the stability of PARP1 protein remains unchanged up to 24 h in the presence of cycloheximide, proving that the PARP1 protein degradation observed after MAC681 treatment is independent of its effects on PARP1 mRNA levels.
Off-target protective effect of 3ABA in MAC681-treated cells. A Western blot analysis of PARP1 protein levels in K-562 cells treated with MAC681 (5 µM), 3ABA (5 mM), or their combination after 4 h. B Kinetic analysis of the PARP1 protein levels in K-562 cells treated with MAC681 at 5 µM. β-actin was used as a loading control. Etoposide 100 µM (VP16) was used as a positive control. C Closed-view binding mode of MAC681 in the catalytic pocket of PARP1. The well-known catalytic residues of PARP1 are presented in a white stick model with nitrogen, oxygen, and fluorine colored blue, orange, and cyan, respectively. MAC681 is presented as a magenta stick. D Intracellular ATP content in K-562 cells treated with MAC681 (5 µM), 3ABA (5 mM), or their combination normalized to control cells measured by CellTiter-Glo at indicated time points (one-way ANOVA, Tukey's multiple comparisons test ****p ≤ 0.0001). E Effects of MAC681 (5 µM), 3ABA (5 mM), or their combination on mitochondrial membrane potential (MMP) analyzed by flow cytometry with MitoTracker Red staining (left panel) at indicated time points. MFI represents median fluorescence intensity. Cell death was assessed simultaneously with propidium iodide (right panel) (one-way ANOVA, Šídák's multiple comparisons test *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). F Quantification of γH2AX nuclear foci formation by flow cytometry in K-562 cells treated with MAC681 (5 µM), 3ABA (5 mM), or their combination (one-way ANOVA, Dunnett's multiple comparisons test ****p ≤ 0.0001). Results correspond to the mean ± SD of three independent experiments. G Chemical structures of MAC681 and 3ABA are recognized as suitable reaction partners that can give rise to 5-position or 6-position adducts. H The 5-position and 6-position adducts bind to PARP1 in the same mode as 3ABA and MAC681 (left panel). Stabilization energies of docking compounds to PARP1 are shown (right panel). Compound 2US was used as a control for docking experiments
This type of atypical degradation could be explained by the direct interaction of MAC681 with PARP1 protein, a hypothesis we validated by computational docking. The control docking showed that 2, 3-dihydrobenzofuran-7-carboximade derivative (2US) was docked into the catalytic region of PARP1 in good agreement location with the actual binding mode in the crystal structure of PARP1 in complex with 2US (PDB ID: 40QA) (Supp. Fig. S3D, upper panel). We demonstrated that MAC681 binds with high affinity into the catalytic region of the PARP1 structure in the presence of damaged DNA (Supp. Fig. S3D, lower panel, MAC681 in magenta). MAC681 was located in active sites of the PARP1 with stabilization energies of − 7.1 kcal/mol. We closely observed that the aromatic ring of the bound MAC681 is surrounded by well-known inhibitor-binding residues such as His862, Tyr889, Tyr896, Phe897, and Tyr907 (Fig. 3C).
We further focused on the protective role of 3ABA. We assessed its effects on MAC681-induced ATP depletion and mitochondrial membrane potential (MMP) loss leading to cell death as analyzed by PI intake. 3ABA alone did not affect cellular ATP levels. On the other hand, 3ABA pre-treatment restored ATP levels that were depleted after the MAC681 treatment (Fig. 3D). Similarly, 3ABA alone did not affect MMP. It partially reverted the loss of MMP compared to cells treated with MAC681 alone (Fig. 3E left panel). Moreover, 3ABA rescued the cells from dying, as shown by PI intake (Fig. 3E right panel).
3ABA prevented the DNA damage induced by MAC681 not only in K-562 cells (Fig. 3F) but also in U-937, MOLM-13, MV4-11, and OCI-AML3 cell lines (Supp. Fig. S3E left panel) where MAC681 induced typical apoptotic nuclear fragmentation patterns (Supp. Fig. S3E right panel). 3ABA prevented observed nuclear morphology changes in all cell models. Concomitantly, 3ABA counteracted the characteristic apoptotic fragmentation of PARP1 protein caused by MAC681 treatment in U-937 cells (Supp. Fig. S3F). Interestingly, this protection (of cell death induction and PARP1 fragmentation) by 3ABA could not be observed in apoptotic U-937 cells treated with etoposide (Supp. Fig. S3F), demonstrating the specificity of a combination involving MAC681 and 3ABA. These data indicate that the protective effect of 3ABA in MAC681 treated cells could be independent of its primary PARP1-inhibition function and instead mediated by its so far not-described off-target effect.
The chemical structures of MAC681 and 3ABA are recognized as suitable reaction partners. Based on our previous results, the chemical structure of 3ABA seemed to be directly involved in the protective effect. Thus, we hypothesized that indolequinone MAC681 could act as a Michael acceptor, bearing the α,β-unsaturated bond, reacting with the aniline nucleophile of 3ABA (Michael donor), able to attack the position next to the -OCH3 of the quinone ring of MAC681. The above-mentioned Michael reaction would result in a hypothetical 3ABA-MAC681-5-position adduct (Fig. 3G). On the other hand, the reaction between 3ABA and MAC681 can also give rise to the 6-position adduct due to the loss of the methoxy group with the addition of the amine and the 2,4,6-trifluorophenol part of the molecule either intact (Fig. 3G) or lost (not shown). Using mass spectrometry, we confirmed the presence of a 6-position adduct when cells were treated with a combination of MAC681 and 3ABA for 6 h (Supp. Fig. S3G). Moreover, the 3ABA washout completely abrogates the protective effect of 3ABA detected without washout (Suppl. Fig. S3H). In the washout group, cells pre-treated with 3ABA and treated with MAC681 for 4 h were losing ATP at the same extent as cells treated with MAC681 alone (S3H upper left panel). The loss of the protection by 3ABA on early effects mediated by MAC681 was confirmed by the late outcome on cell concentration (S3H upper middle panel) and viability (S3H upper right panel). In addition, we confirmed that the cells in the washout group (3ABA + MAC681) were dying by the same cell death modality as cells treated with MAC681 alone, as shown by identical nuclear morphology patterns (S3H lower panel).
Indolequinones' DNA damaging potential is linked to their redox-cycling activity or alkylation potential. Using dichlorofluorescein and redox sensor red probes, we confirmed that MAC681 did not induce any reactive oxygen species (ROS) (Supp. Fig. S3I) and thus did not act as a redox cycler. Considering its alkylation activity, MAC681 could be transformed into a DNA alkylation agent through bioreductive activation by two electron reductases, such as NAD(P)H quinone dehydrogenase (NQO)1 or NQO2, via mechanisms of iminium salt (Supp. Fig. S3J) [46] or quinone methide formation (Supp. Fig. S3K) [47]. Analysis of CML patient datasets showed that CML patients have significantly higher expression of NQO1 or NQO2 or both than their healthy counterparts (Supp. Fig. S3L). Moreover, based on the CCLE dataset, K-562 cells have the highest expression of NQO2 and the second highest expression of NQO1 within the CML cell lines available (Supp. Fig. S3M).
Considering this, the above-mentioned hybrid compounds (5-position and 6-position adducts) would reduce the overall DNA damaging activity of MAC681 and thus confer 3ABA-mediated protection against MAC681-induced controlled necrosis triggered by excessive DNA damage.
In addition, docking results revealed that 3ABA and both hybrid compounds (3ABA-MAC681-5-position adduct and 3ABA-MAC681-6-position adduct) docked well into the similar PARP1 catalytic region where MAC681 did. Interestingly, both hybrid compounds have bound to PARP1 tighter than MAC681 or 3ABA alone, with stabilization energies of -9.4 and -8.5 kcal/mol, respectively (Fig. 3H).
MAC681-induced cell death has immunogenic potential
Our initial analysis of transcriptomic data (GSE5550) showed a significant down-regulation of genes encoding proteins involved in adaptive and innate immune responses in CML patients (Fig. 4A). Especially genes leading the NES score, belonging to antigen processing and presentation and interferon-gamma signaling categories are down-regulated (Fig. 4A). As the attenuation of these essential immune recognition mechanisms reduces the capacity of the patient’s immune effectors to recognize and eliminate the accumulating CML blasts, we were interesting to characterize further the MAC681-induced necrotic cell death observed in K-562 cells from the immunogenic perspective. We measured the emission of ICD-related DAMPs, including calreticulin exposure, extracellular ATP secretion, and HMGB1 release. As soon as 10 min and up to 2 h after the treatment, MAC681-treated cells reached between 2- to fourfold increase in ATP secretion compared to untreated controls (Fig. 4B). Additionally, we determined that 10 h of 5 µM MAC681 treatment doubled the exposure of CRT at the cell surface compared to the control (Fig. 4C). Besides intrinsic cellular mechanisms, the translocation of calreticulin is also mediated by soluble factors that operate in an autocrine/paracrine manner. Accordingly, IL-8 can be hyperactivated by ICD-inducers to facilitate the immunogenic exposure of calreticulin [48]. Hence, we measured the human IL-8 concentrations in cell culture supernatants and showed that 5 µM of MAC681 significantly induced IL-8 release at 8 and 10 h of treatment compared to control (1.4- and 1.3-fold increase, respectively) (Fig. 4D). Timeframe of IL-8 release is in line with calreticulin cell surface exposure in K-562 cells. In addition, we also provided evidence showing that at 8 h, MAC681 induced IL-8 mRNA expression in a dose-dependent manner in K-562 cells with a 6.4-fold increase at 5 µM (Supp. Fig. S4A).
MAC681 has an immunogenic potential in myeloid disorders. A GSEA plots showing genes linked to innate and adaptive immune response, antigen processing and presentation, and interferon-gamma signaling have negative NES scores in CML CD34+ cells (ES: enrichment Score, NES: normalized enrichment score). B Extracellular release of ATP into the supernatant induced by 5 µM of MAC681 at indicated time-points in K-562 cells was measured in a luciferase-based assay (double-sided unpaired t-test ***p ≤ 0.001, ****p ≤ 0.0001). C Ectopic calreticulin expression in K-562 cells was investigated by flow cytometry after 10 h of MAC681 treatment at 5 µM. ISO corresponds to isotypic control. Quantification (left panel) and representative experiment (right panel) (double-sided unpaired t-test *p ≤ 0.05, MFI = median fluorescence intensity). D Time-dependent release of IL-8 induced by MAC681 at 5 µM assessed by ELISA (double-sided unpaired t-test *p ≤ 0.05, **p ≤ 0.01). E Time-dependent release of HMGB1 after treatment with 5 µM of MAC681 assessed by ELISA in K-562 cells supernatants (double-sided unpaired t-test *p ≤ 0.05, ***p ≤ 0.001). F Phagocytosis of MAC681-treated (5 µM, 16 h) red-stained K-562 CML cells by green-stained J774A1 murine macrophages assessed by fluorescent microscopy. Murine macrophages were co-cultured with CML cells for 2 h. Representative pictures are shown on the left, and quantification on the right (double-sided unpaired t-test **p ≤ 0.01). G Kinetic analysis of tumor volume (left panel) in syngeneic immunocompetent C57BL/6 mice and the quantification of the overall response using the AUC. Tumor weight (middle panel) was measured after sacrifice, and excised tumors were visualized (right panel). “D” corresponds to days after inoculation with living C1498 cells (one-way ANOVA, Tukey's multiple comparisons test ***p ≤ 0.001, ****p ≤ 0.0001A). H Spleen weight (left panel) and spleen length (middle panel) were assessed after mice sacrifice (one-way ANOVA, Tukey's multiple comparisons test *p ≤ 0.05). Pictures of spleen excised from C57BL/6 mice are visualized (right panel). Error bars represent the SD of at least three independent experiments
Furthermore, we showed that necrosis-inducing concentration triggered significant HMGB1 release starting from 4 h and peaking at 24 h in MAC681-treated K-562 cells (Fig. 4E). Moreover, we showed that MAC681-induced HMGB1 release not only in CML cells but also in OCI-AML3 and C1498 AML cell lines (Supp. Fig. S4B-C). As the emission of DAMPs by dying cells plays a crucial role in recruiting immune effectors, we co-cultured J774A1 macrophages with MAC681 pre-treated K-562 cells. Fluorescence microscopy showed a 160-fold increase in the engulfment of red-stained MAC681-treated K-562 cells by green-stained macrophages (Fig. 4F).
However, neither morphological assessments of dying cells nor the release of the above-mentioned biochemical correlates, alone or in combination, can predict with complete certainty the induction of bona fide ICD [49]. The gold-standard approach to proving the immunogenicity of the cellular demise induced by a specific agent relies on vaccination experiments in immunocompetent mice. Hence, we treated syngeneic murine myeloid C1498 cancer cells with the MAC681 at different concentrations for 24 h and evaluated cell death induction by flow cytometry after annexin V/PI staining (Supp. Fig. S4D). Next, syngeneic immunocompetent C57BL/6 mice were vaccinated with dying cancer cells treated either with 5 µM MAC681 or a combination of 2.5 µM cytarabine (CTB) and 0.5 µM doxorubicin (DOX), a known induction therapy, into the right flanks. Control mice (non-vaccinated) were given 1 × PBS only. After one week, all mice were challenged with living C1498 cells into the contralateral flank of the mice, and tumor growth was monitored until the tumor volume reached > 600 mm3 in the non-vaccinated control group. We showed that tumor volume and weight were significantly reduced in the MAC681-treated group compared to the control group (Fig. 4G). Moreover, the MAC681 effect on tumor growth was similar to the impact of the induction therapy (CTB + DOX). In addition, we assessed the effect of MAC681 vaccination on spleen morphology and revealed that MAC681 significantly reduced spleen weight but did not impact its length (Fig. 4H). Altogether, these data provide strong evidence supporting the immunogenic potential of indolequinone MAC681.
Synergistic potential of MAC681 in combination with asciminib in imatinib-sensitive and -resistant human leukemia
Despite improvements in CML therapy by develo** efficient TKIs suppressing BCR-ABL activity, one in five patients develops resistance to TKIs. Moreover, 22% of de novo CML- patients in the chronic phase become intolerant (JALSG CML212 study), requiring discontinuation of the therapy because of adverse side effects. Accordingly, combining TKIs with non-ABL-related therapeutic strategies is required to prevent the onset and overcome the appearance of TKI-resistant clones or diminish the side effects of TKIs. Hence, we investigated the effect of the combination of MAC681 with the ATP-non-competitive, allosteric TKI asciminib in multi-TKI-resistant and sensitive K-562 cell models up to 72 h. We used the Chou-Talalay algorithm to determine the combination index and fraction-affected values. Our results in K-562R cells showed a synergistic effect of 3 µM MAC681 with 1, 10, and 30 µM of asciminib at any given time point (Fig. 5A). In K-562 sensitive cells, multiple combinations showed synergistic effects during 24 h, 48 h, and 72 h (Fig. 5B).
Synergistic effects of MAC681 with asciminib in K-562 and K-562R cells. Heatmap showing the inhibition of cell viability analyzed by trypan blue exclusion test in K-562R [A] and K-562 [B] cells by different combinatory treatments at 24 h, 48 h, and 72 h. The fraction affected (Fa) scale is shown as a sidebar, where 0 = no inhibitory effect and 1 = 100% inhibition of cell viability. The combinatory index was calculated by Compusyn software and is indicated within each quadrant (CI < 1 synergism, CI = 1 additive effect, CI > 1 antagonism)