Abstract
Background
Genome editing has been considered as powerful tool in agricultural fields. However, genome editing progress in cattle has not been fast as in other mammal species, for some disadvantages including long gestational periods, single pregnancy, and high raising cost. Furthermore, technically demanding methods such as microinjection and somatic cell nuclear transfer (SCNT) are needed for gene editing in cattle. In this point of view, electroporation in embryos has been risen as an alternative.
Results
First, editing efficiency of our electroporation methods were tested for embryos. Presence of mutation on embryo was confirmed by T7E1 assay. With first combination, mutation rates for MSTN and PRNP were 57.6% ± 13.7% and 54.6% ± 13.5%, respectively. In case of MSTN/BLG, mutation rates were 83.9% ± 23.6% for MSTN, 84.5% ± 18.0% for BLG. Afterwards, the double-KO embryos were transferred to surrogates and mutation rate was identified in resultant calves by targeted deep sequencing. Thirteen recipients were transferred for MSTN/PRNP, 4 calves were delivered, and one calf underwent an induction for double KO. Ten surrogates were given double-KO embryos for MSTN/BLG, and four of the six calves that were born had mutations in both genes.
Conclusions
These data demonstrated that production of genome edited cattle via electroporation of RNP could be effectively applied. Finally, MSTN and PRNP from beef cattle and MSTN and BLG from dairy cattle have been born and they will be valuable resources for future precision breeding.
Similar content being viewed by others
Introduction
Genome editing has played a powerful role in various fields. In the same line with other fields, in livestock, genome editing tools such as ZFN, TALENs, and CRISPR/Cas9, has been applied to disease resistance, climate response, allergy free and increasing productivity [1,2,3,4,5,1) and in MSTN, where its phenoty** can be easily detected, it began to be observed (Fig. 3). Other phenoty** will be analyzed later because BLG should be analyzed after pregnancy and for PRNP, its test should be evaluated in the continuously follow up.
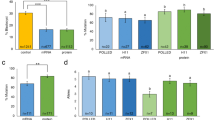
As summarized in Fig. 2, there was a unique fact in the genoty** analysis. In previous study, the MSTN mutated cattle showed only −12 bps [1]. However, in this study, although another gene delivery was applied into zygotes via RNP, −12 bps genoty** was dominantly identified as well, and new genoty** (1, 25 bps insertion and 3 bps deletion) was observed. In more details, no more than 2 genoty** was identified in calves with MSTN mutations (Fig. 2). However, three genoty** were observed in PRNP locus, and 3 or more mutations were observed in BLG except for one mutated calf. According to our results so far, −12 bp is dominantly observed in MSTN knockout cattle through microinjection or electroporation on IVF embryos. In the future, we will continue to monitor the results of knockout genoty** in vivo results through various gene loci. And we found very low percentage of off-target mutation in #5 (Additional file 3). After the offspring become at over than 12 months old, the genoty** and off-target analysis in their germ cells will be followed.
One interesting point is about double knockout event. In dairy breed calves, in which MSTN and BLG were targeted, there were high double knockout efficiency in randomly selected single cell colony analysis. However, in beef breed MSTN/PRNP KO calf, double knockout event in the offspring was very low while its event was comparable with that of MSTN/BLG in blastocyst stage. Because there is only one offspring with MSTN/PRNP KO, it is difficult to determine whether this occurrence was caused by specific target gene locus or breed/individual predisposition. We intend to conduct relevant research in the near future.
Recently, a genome edited organism without any recombinant DNA integration can be classified as non-GMO in many countries [9]. For instance, in Japan, MSTN edited fish is approved to be edible as a food chain [21] and in USA, low-risk determination was made for marketing genome-edited cattle [22]. In the case of animals, safety analysis is not necessary, and their health can be considered an indicator of animal product safety [23]. We were able to find no special complications or significant changes in blood analysis when we assessed the health of animals born as double knockouts, indicating that they are currently growing healthily. As a result, it is expected that the safety of meat and milk from these sources will be unaffected in the future.
Conclusions
In conclusion, these data demonstrated that genome editing on bovine embryos via electroporation of RNP could be effectively applied and proved. Finally, beef cattle with MSTN and PRNP mutation and dairy cattle with MSTN and BLG mutation have been born and they will be valuable resources for future precise breeding.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- BLG:
-
Beta-lactoglobulin
- Cas9:
-
CRISPR associated protein 9
- CRISPR:
-
Clustered regularly interspaced short palindromic repeats
- GMO:
-
Genetically modified organism
- IVF:
-
In vitro fertilization
- KO:
-
Knock-out
- MSTN:
-
Myostatin gene
- PRNP:
-
Prion protein gene
- RNP:
-
Ribonucleoprotein
- SCNT:
-
Somatic cell nuclear transfer
- sgRNA:
-
Single guide RNA
- T7E1:
-
T7 endonuclease I
- TALEN:
-
Transcription activator-like effector nucleases
- TALP:
-
Tyrode’s albumin lactate pyruvate
- ZFN:
-
Zinc-finger nuclease
References
Gim GM, Kwon DH, Eom KH, Moon JH, Park JH, Lee WW, et al. Production of MSTN-mutated cattle without exogenous gene integration using CRISPR-Cas9. Biotechnol J. 2022;17(7):e2100198.
Lee K, Uh K, Farrell K. Current progress of genome editing in livestock. Theriogenology. 2020;150:229–35.
Luo J, Song Z, Yu S, Cui D, Wang B, Ding F, et al. Efficient generation of myostatin (MSTN) biallelic mutations in cattle using zinc finger nucleases. PLoS ONE. 2014;9(4):e95225.
Proudfoot C, Carlson DF, Huddart R, Long CR, Pryor JH, King TJ, et al. Genome edited sheep and cattle. Transgenic Res. 2015;24:147–53.
Su F, Wang Y, Liu G, Ru K, Liu X, Yu Y, et al. Generation of transgenic cattle expressing human beta-defensin 3 as an approach to reducing susceptibility to Mycobacterium bovis infection. FEBS J. 2016;283(5):776–90.
Wang K, Ouyang H, **e Z, Yao C, Guo N, Li M, et al. Efficient generation of myostatin mutations in pigs using the CRISPR/Cas9 system. Sci Rep. 2015;5:16623.
Whitworth KM, Rowland RRR, Ewen CL, Trible BR, Kerrigan MA, Cino-Ozuna AG, et al. Gene-edited pigs are protected from porcine reproductive and respiratory syndrome virus. Nat Biotechnol. 2016;34(1):20–2.
Wu H, Wang Y, Zhang Y, Yang M, Lv J, Liu J, et al. TALE nickase-mediated SP110 knockin endows cattle with increased resistance to tuberculosis. Proc Natl Acad Sci U S A. 2015;112(13):E1530–9.
Van Eenennaam AL, Wells KD, Murray JD. Proposed U.S. regulation of gene-edited food animals is not fit for purpose. npj Sci Food. 2019;3:3.
Gouveia C, Huyser C, Egli D, Pepper MS. Lessons learned from somatic cell nuclear transfer. Int J Mol Sci. 2020;21(7):2314.
Thuan NV, Kishigami S, Wakayama T. How to improve the success rate of mouse cloning technology. J Reprod Dev. 2010;56(1):20–30.
Yum SY, Lee SJ, Kim HM, Choi WJ, Park JH, Lee WW, et al. Efficient generation of transgenic cattle using the DNA transposon and their analysis by next-generation sequencing. Sci Rep. 2016;6:27185.
Gim GM, Uhm KH, Kwon DH, Kim MJ, Jung DJ, Kim H, et al. Germline transmission of MSTN knockout cattle via CRISPR-Cas9. Theriogenology. 2022;192:22–7.
Camargo LSA, Owen JR, Van Eenennaam AL, Ross PJ. Efficient one-step knockout by electroporation of ribonucleoproteins into zona-intact bovine embryos. Front Genet. 2020;11:570069.
Chen S, Lee B, Lee AYF, Modzelewski AJ, He L. Highly efficient mouse genome editing by CRISPR ribonucleoprotein rlectroporation of zygotes. J Biol Chem. 2016;291(28):14457–67.
Guschin DY, Waite AJ, Katibah GE, Miller JC, Holmes MC, Rebar EJ. A rapid and general assay for monitoring endogenous gene modification. Methods Mol Biol. 2010;649:247–56.
Wang JY, Doudna JA. CRISPR technology: A decade of genome editing is only the beginning. Science. 2023;379(6629):eadd8643.
Platani M, Sokefun O, Bassil E, Apidianakis Y. Genetic engineering and genome editing in plants, animals and humans: Facts and myths. Gene. 2023;856:147141.
Van Eenennaam AL. Application of genome editing in farm animals: cattle. Transgenic Res. 2019;28(Suppl 2):93–100.
Yum SY, Youn KY, Choi WJ, Jang G. Development of genome engineering technologies in cattle: from random to specific. J Anim Sci Biotechnol. 2018;9:16.
Japan embraces CRISPR-edited fish. Nat Biotechnol. 2022;40:10. https://doi.org/10.1038/s41587-021-01197-8.
FDA news. FDA makes low-risk determination for marketing of products from genome-edited beef cattle after safety review. 2022. https://www.fda.gov/news-events/press-announcements/fda-makes-low-risk-determination-marketing-products-genome-edited-beef-cattle-after-safety-review.
Trott JF, Young AE, McNabb BR, Yang X, Bishop TF, Van Eenennaam AL. Animal health and food safety analyses of six offspring of a genome-edited Hornless bull. GEN Biotechnology. 2022;1(2):192–206.
Acknowledgements
We thank the members of the Goo Jang lab and Gyeonggi-do Animal Hygiene Testing Center for their valuable comments and the technical support.
Funding
This study was financially supported by the National Research Foundation of Korea (NRF-2021R1A5A1033157 for SRC program: 382 Comparative medicine Disease Research Center; NRF-2021R1F1A105195313), the Research Institute of Veterinary Science, the BK21 Four for Future Veterinary Medicine Leading Education and Research Center, and a Seoul National University (SNU) grant (#550e2020005).
Author information
Authors and Affiliations
Contributions
GJ devised the experiment. GMG and JHL produced and analyzed embryos for mutation. DJJ, DHK and JKY prepared surrogates and WWL transferred embryos. GMG, KHE and DHK collected the samples and analyzed mutation with WJS, which was supervised over by SYY. GMG and EKH merged whole data and prepared the manuscript which was edited by GJ. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Institutional Animal Care and Use Committee (SNU-180403–1) and performed under the guideline of Seoul National University. All animal practices such as blood collection or embryo transfer was carried out by a veterinarian.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Supplementary Information
Additional file 1
. List of guide RNA and detecting PCR primer sequences for each target genes.
Additional file 2
. List of primer sequences for deep sequencing of each target genes and off-target sites.
Additional file 3.
Off-target effect detection for gene edited calves.
Additional file 4.
Blood analysis of calves with gene mutation.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Gim, GM., Eom, KH., Kwon, DH. et al. Generation of double knockout cattle via CRISPR-Cas9 ribonucleoprotein (RNP) electroporation. J Animal Sci Biotechnol 14, 103 (2023). https://doi.org/10.1186/s40104-023-00902-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40104-023-00902-8