Abstract
Ubiquitination is a versatile post-translational modification (PTM), which regulates diverse fundamental features of protein substrates, including stability, activity, and localization. Unsurprisingly, dysregulation of the complex interaction between ubiquitination and deubiquitination leads to many pathologies, such as cancer and neurodegenerative diseases. The versatility of ubiquitination is a result of the complexity of ubiquitin (Ub) conjugates, ranging from a single Ub monomer to Ub polymers with different length and linkage types. To further understand the molecular mechanism of ubiquitination signaling, innovative strategies are needed to characterize the ubiquitination sites, the linkage type, and the length of Ub chain. With advances in chemical biology tools, computational methodologies, and mass spectrometry, protein ubiquitination sites and their Ub chain architecture have been extensively revealed. The obtained information on protein ubiquitination helps to crack the molecular mechanism of ubiquitination in numerous pathologies. In this review, we summarize the recent advances in protein ubiquitination analysis to gain updated knowledge in this field. In addition, the current and future challenges and barriers are also reviewed and discussed.
Similar content being viewed by others
Introduction
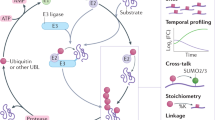
The covalent attachment of ubiquitin (Ub) to protein substrate (ubiquitination) is an important post-translational modification (PTM) that regulates diverse cellular functions [1,2,3,4]. Ub is a small and highly conserved 76-residue protein in eukaryotes. The C-terminal glycine of Ub (G76) is covalently attached to its substrate protein, which is regulated in a cascade order of E1 Ub-activating enzymes, E2 Ub-conjugating enzymes, and E3 Ub ligases (Fig. 1A) [5, 6]. Ub is reversibly removed from the substrate through Ub hydrolases known as a family of proteins named deubiquitinases (DUBs) [7]. Maintenance of cellular ubiquitination homeostasis is fulfilled by the orchestrated interplay of 2 E1 enzymes, ~ 40 E2 enzymes, more than 1000 E3 ligases, and approximately 100 DUBs encoded by the human genome [8,9,10]. Aberrance of ubiquitin-related enzymes’ activity leads to the dysregulation of protein ubiquitination, promoting the pathogenesis of numerous diseases, such as cancer, neurodegenerative diseases, and so on [11,12,13,63]. This IAA-induced artifact can now be avoided by using less reactive chloroacetamide or by lowering the reaction temperature and concentration of iodoacetamide [64]. In addition, the profiling of K-ε-GG peptides using antibodies may exhibit slight peptide sequence bias [65].
UbiSite antibody-based approach to profiling ubiquitination sites
To overcome some drawbacks associated with the diGly approach, Akimov et al. recently generated an antibody, UbiSite, which recognized the C-terminal 13 amino acids (ESTLHLVLRLRGG) of Ub after Lys-C digestion (Fig. 3B). By combining UbiSite-based enrichment and pre-fractionation, the authors identified over 63,000 unique ubiquitination sites on 9200 proteins in human Hep-G2 and Jurkat cell lines after proteasomal inhibitor treatment by MS analysis [66]. In addition, 104 N-terminal ubiquitination sites were identified in their results which were ignored by the diGly approach. As the only available tool with the ability to enrich peptides from protein N-terminal ubiquitination, the UbiSite antibody-based approach might be a useful approach to studying the function and regulation of N-terminal ubiquitination [67].
Firstly, due to the specific C-terminal sequence (ESTLHLVLRLRGG) of Ub, the UbiSite antibody significantly improves specificity toward ubiquitinated peptides by reducing the non-specific enrichment of Ub-like proteins such as NEDD8 and ISG15. Secondly, UbiSite enables enrichment of protein N-terminal ubiquitination, which advances the understanding of protein N-terminal ubiquitination. Hoverer, the sequence ESTLHLVLRLRGG is quite long on the ubiquitinated peptides, resulting in more extra fragments during MS/MS analysis, which will impair the identification efficiency of protein ubiquitination using Lys-C digestion only [45]. Therefore, extra enzymatic digestion will benefit the protein profiling for the UbiSite approach [45].
Insights into N-terminal ubiquitination using the antibody toolkit
Conjugation of Ub to lysine ε-amino group is the most common type of ubiquitination. Besides, the α-amino group of protein N-terminus has also been identified as non-canonical ubiquitinated targets [68, 69]. However, current approaches, except the UbiSite approach, mainly focus on lysine ubiquitination profiling, there are limited approaches to analyzing the N-terminal ubiquitination, which hampers the functional study of protein N-terminal ubiquitination. To identify protein N-terminal ubiquitination, three approaches for profiling protein N-terminal ubiquitination, anti-GGX mAbs based approach, UbiSite based approach, and StUbEx PLUS approach, were reported [66, 70, 71]. For example, Davies et al. generated four monoclonal antibodies (anti-GGX mAbs) that selectively recognize tryptic peptides with an N-terminal diGly remnant rather than the K-ε-GG group, realizing the specific enrichment and identification of protein N-terminal ubiquitination [70]. UBE2W is the only E2 Ub conjugating enzyme reported to regulate the protein N-terminal ubiquitination. The authors used anti-GGX mAbs to enrich and analyze UBE2W regulated N-terminal ubiquitination events [72]. They identified 152 unique N-terminal ubiquitination sites derived from 109 endogenous proteins. Of the 152 unique N-terminal ubiquitination sites, 32 sites are reported as the potential substrates of UBE2W, demonstrating that the anti-GGX mAbs-based approach is qualified for protein N-terminal ubiquitination profiling.
Among the three methods for analyzing N-terminal ubiquitination, the anti-GGX mAbs-based approach is designed for selectively enriching tryptic peptides with an N-terminal diglycine remnant rather than a diglycine remnant on lysine. However, the UbiSite antibody specifically recognizes the ubiquitin 13-residue remnant on N-terminus and Lys residue after Lys-C digestion. Different antibodies show some bias toward some preferent sequences. In addition, StUbEx PLUS (see below) is an antibody-free approach that is different from the above antibody-based approaches, showing no bias toward lysine and N-terminal ubiquitination. That’s why only a small part of N-terminal ubiquitination sites identified by the anti-GGX mAbs-based approach were overlapped with the ubiquitination sites identified by UbiSite and StUbEx PLUS approaches [66, 70, 71]. This result nicely highlights the limitations and complementary of the current approaches.
Antibody-free approaches to profiling ubiquitination sites
Antibody-based strategies are the most widely used approaches to systematically profiling protein ubiquitination at the peptide level. However, antibody-based approaches to profiling protein ubiquitination entail some shortcomings such as (a) the bias toward the amino acid sequence surrounding the ubiquitination sites and (b) expensive antibodies limit the widespread application. As an alternative to antibody-based approaches, Gevaert et al. reported an antibody-free approach, termed Ub COmbined FRActional DIagonal Chromatography (COFRADIC) (Fig. 3C), for enriching and identifying protein ubiquitination at the peptide level [73, 74]. In brief, all primary amino groups were blocked by acetylation at the protein level, followed by USP2 catalytic core domain (USP2cc) incubation to hydrolysis Ub from the ubiquitinated proteins and reintroduce of the free ε-amine groups at the ubiquitination sites. To the free ε-amines, a glycine linked to a hydrophobic tert-butyloxycarbonyl (BOC) group was attached, which was further used to enrich the peptides via two reverse-phase HPLC (RP-HPLC) runs before and after TFA-based removal of BOC groups. Gevaert et al. used this approach to profiling protein ubiquitination in native human Jurkat cell lysate and in Arabidopsis thaliana, resulting in the identification of over 7500 endogenous ubiquitination sites on 3300 different proteins and 3009 endogenous ubiquitination sites on 1607 proteins, respectively.
Another reported antibody-free approach is StUbEx PLUS (Fig. 3D). Based on the Stable Tagged Ub Exchange (StUbEx) strategy which was used to enrich ubiquitinated proteins, Akimov et al. modified StUbEx strategy, StUbEx PLUS, to specifically enriched the ubiquitinated peptides [71]. The authors built a StUbEx PLUS system to insert His-tag between serine 65 and threonine 66 in the recombinant Ub. After proteolytic digestion with specific lysine cleavage enzymes, the tag was still attached to the ubiquitination sites and enriched by Ni-NTA beads. Since Ub-like protein doesn’t carry His-tag, the interference from Neddylation and ISGylation were also avoided here. Using the StUbEx PLUS strategy, 41,589 unique ubiquitination sites on 7762 proteins were identified in U-2 OS cells.
The antibody-free approach showed a powerful ability to enrich and profile protein ubiquitination, easier handling and cheaper antibody-free approaches for protein ubiquitination profiling are very much needed to overcome the drawbacks of antibody-based approaches. Recently, we proposed an antibody-free approach, termed AFUP, to profiling protein ubiquitination by selectively clicking the ubiquitinated lysine, resulting in 7103 ubiquitination site identification with high confidence in 5 mg HeLa lysates (in preparation).
Antibody-free-based strategies play a vital role in the studies of protein ubiquitination, which can overcome some drawbacks, for example, bias toward to sequence surrounding the ubiquitination sites and high cost. Even though antibody-free approaches enable global identification of protein ubiquitination, there still exist some shortcomings. First, the dataset identified by antibody-free approaches is quite smaller than the dataset identified by antibody-based approaches, suggesting that more effective antibody-free approaches are needed to expand the depth of the protein ubiquitination pool. Second, the Ub-COFRADIC approach is complicated and time-consuming. Third, StUbEx PLUS or other approaches like StUbEx PLUS require a tagged Ub which changes the structure of Ub, introducing some artifacts and limiting its application in animal or patient tissues. Therefore, an antibody-free approach, with high throughput, convenient and effective identification of protein ubiquitination, is urgently needed.
Insights into the ubiquitinated proteins at the level of architecture
Ub chains with distinct topologies regulate the protein stability, protein–protein interaction, or protein localization in eukaryotic cells, and thus play vital roles in multifunctional signals [75]. The approaches discussed above mainly reveal the ubiquitinated substrates and ubiquitination sites, which are not able to reveal the Ub chain architecture. There are several possibilities to detect the architectures of Ub chains. For example, linkage-specific antibodies which specifically recognize Ub linkages are used to identify the distinct topologies of Ub chains [8]. Currently, MS-based proteomics is categorized into bottom-up proteomics (BUP), middle-down proteomics (MDP), and top-down proteomics (TDP) based on the analytes (Fig. 4). Bottom-up is a traditional strategy of digesting proteins into small peptides for LC–MS analysis with high throughput. However, complete digestion leads to the inability to distinguish the protein isoforms. Middle-down is a restricted digestion proteomics strategy to generate longer peptides for LC–MS analysis, resulting in analyzing a wider range of peptide fragments. Compared with bottom-up proteomics and middle-down proteomics, Top-down proteomics strategy does not need digestion, but directly analyzes the intact protein by LC–MS to gain a comprehensive characterization of the analyzed protein. In this section, we mainly discuss the MS-based approaches to map** the topologies of Ub chains (Fig. 5).
Schematic workflow of bottom-up proteomics, middle-down proteomics, and top-down proteomics. The protein structure is downloaded from National Center for Biotechnology Information (MMDB ID: 209664, PDB ID: 7KW7) [104].
Approaches to getting insights into the architecture of Ub chains. A Schematic diagram of the BUP strategy of UbiCRest to characterize the substrate ubiquitin chain type. B Schematic diagram of the MDP strategy of Ub-clip** to characterize the substrate ubiquitin architecture. C Schematic diagram of TDP strategy to characterize the substrate ubiquitin architecture
BUP is the conventional approach to analysis. A disadvantage of the BUP strategy is the loss of architectural information on polyUb or branched chains upon trypsin digestion. Therefore, whole-cell K-ε-GG analysis does not reveal the topological information of Ub chain attached to the substrates. To reveal the information of Ub chains, linkage-specific antibodies, Affimers, or binding domains are firstly used to enrich a specific chain type and the BUP strategy is further used to identify the linkage-specific substrates [38, 39, 56, 76]. There are several reviews that summarize the application of linkage-specific chain enrichment strategies to study the structure topologies of polyUb chains [28, 49, 77,78,79]. Map** the topologies of Ub chains requires the detection of Ub molecules modified by other Ub molecules. Ohtake et al. used a mutated Ub in which the arginine 54 was replaced by an alanine. This mutant enables the discrimination between branched K48/K63 linkages and unbranched linkages [80]. The authors revealed that the Ub chain branched at K48 and K63 regulated nuclear factor κb (NF-κB) signaling. Ub chain restriction (UbiCRest) is another approach to analyzing Ub chain architecture (Fig. 5A), in which substrates (ubiquitinated proteins or polyUb chains) are treated with a panel of linkage-specific DUBs in parallel reactions [81]. Several issues should be considered when using the UbiCRest approach. First, some DUBs may be non-specific for unexpected Ub chains, leading to cross specificity. Second, many DUBs can’t hydrolyze long (n > 4) chains from the substrates [82]. Third, simple UbiCRest is unable for heterotypic chain analysis [83,84,85]. In addition to the methods described above, absolute quantification of ubiquitin (Ub-AQUA) by MS is a standard way to detect the multiformity of ubiquitin linkages that are covalently attached to the protein substrate. By synthesizing isotope-labeled internal standard-tryptic peptides corresponding to mono-ubiquitin and poly-ubiquitin chains bound to cyclin B1, Kirkpatrick et al. revealed that cyclin B1 was modified by complex ubiquitin chain architecture linked through Lys63, Lys11 and Lys48 [86]. By the combination of affinity chromatography and protein standard absolute quantification (PSAQ) mass spectrometry, Kaiser et al. developed a strategy, termed ubiquitin-PSAQ, to quantify cellular concentrations of ubiquitin species by spiking stable isotope-labeled free ubiquitin and ubiquitin conjugates into the lysates [87]. The authors used ubiquitin-PSAQ to measure the concentrations of ubiquitin types in both cell lines as well as mouse and human brain tissue and found that the concentrations of different ubiquitin types varied significantly in different samples. Therefore, BUD-based Ub-AQUA strategies will play vital roles in determining the ubiquitin topology of specific substrates and the concentrations of ubiquitin types of the whole proteome.
As an alternative, the MDP approach has been the most widely used in determining branched chains. Utilizing limited tryptic digestion, the MDP strategy has been applied in determining the abundance of branched Ub chains and detecting the specific linkages, such as K6/K48 and K29/K48 linkages [88,89,90,91]. Considering the high activity of trypsin towards Lys and Arg residues, it is difficult to control the process of tryptic digestion in the MDP strategy. Ub-clip** is a kind of MDP strategy (Fig. 5B), which uses an engineered viral protease, Lbpro*. Lbpro* is created by mutating the 102 Leu to Trp of Lbpro, a foot and mouth disease leader protease, to enable the preferable cleavage towards all types of diubiquitin. The cleavage of Lbpro* happens after Arg74 of Ub and leaves the signature C-terminal diGly attached to modified residue [92]. Swatek et al. used Ub-clip** to quantify branch-point Ub and surprisingly found that about 10–20% of Ub chains seemed to exist as branched type. The authors also showed that PINK1/PARKIN-mediated mitophagy predominantly exploited mono- and short-chain polyUb [93].
Compared to BUP and MDP, TDP may be an ideal platform for the analysis of proteomes bearing Ub chains of different lengths, linkages, and architectures (Fig. 5C) [28]. The major challenge of TDP is that its gas-phase dissociation produces overlap** and low signal-to-noise (S/N) fragments with increasing molecular weight [94]. Therefore, to better characterization of Ub chains by TDP, it’s important to optimize the instrumental parameters [95]. Lee et al. used a TDP strategy that utilized electron-transfer/collision-induced dissociation (ETciD) activation to achieve extensive fragmentation to facilitate the characterization of chain topography and lysine linkage sites, such as K48 linkage and K63 linkage [96]. Thus, while TDP has been used to analyze the topologies of some well-defined ubiquitylated proteins, the application of TDP in the analysis of complex, heterogenous mixtures has not been realized. With advantages in sample preparation and analytical approaches, TDP will play a vital role in analyzing the topologies of Ub chains for a complex system.
MS-based proteomics plays an important role in the identification of Ub chains and linkages. However, it should be noted that strategies for detecting branched chains by MS usually include an enrichment step to increase the recovery of substrates before digestion or direct analysis.
Prediction of protein ubiquitination via computational algorithms
The current methodologies for systematically analyzing protein ubiquitination can be divided into two categories: MS-based strategies and computational strategies. MS-based experimental strategies are often expensive, labor-intensive, and time-consuming. Compared with MS-based strategies, prediction protein ubiquitination using a variety of machine-learning methods can provide simple and rapid research solutions, and provide valuable information for further laboratory studies [97].
Over the past decade, researchers have achieved great success in applying different feature extraction methods to predict protein ubiquitination sites, such as machine-learning algorithms. These computational approaches predict new ubiquitination sites by learning sequence context characteristics of ubiquitination sites of the experimentally verified ubiquitination sites. UbiPred was the first tool reported by Tung et al. for predicting ubiquitination sites, which was implemented by using a Support Vector Machine (SVM) with 31 informative physicochemical features selected from published amino acid indices [98, 99]. The authors utilized UbiPred and identified 23 ubiquitylation sites, which were further validated. In 2018, He et al. reported a multimodal deep architecture to identify the ubiquitination sites and evaluated their method on the available database PLMD, leading to 66.4% specificity, 66.7% sensitivity, and 66.43% accuracy [100]. Recently, Wang et al. reported a new method, named HUbipPred, which utilized the binary encoding and physicochemical properties of amino acids as training input and integrated two kinds of neural networks to build the model [101]. HUbiPred greatly improved the prediction accuracy compared to previous predictors such as DeepUbi and hCKSAAP_UbSite. At present, more and more bioinformatics tools have been developed for predicting protein ubiquitination sites. We refer the readers to a series of excellent reviews discussing the different methods, predictive algorithms, functionality, and properties for predicting protein ubiquitination [97, 102, 103].
Since there is no universal algorithm that can accurately predict protein ubiquitination sites, the fusion of multiple computational methods may be an effective method to comprehensively predict protein ubiquitination sites. In addition, a large amount of protein ubiquitination data has been rapidly accumulated in the last decade. As expected, the improvement of the ubiquitination sequence logo is consistent with the increased sensitivity of the recently developed predictor [103]. Because computational methodology always introduces false-positive results, verification of the prediction results through experimental methods is required.
Conclusions
Protein ubiquitination is one of the most difficult PTMs to be identified due to its large size, low abundance, and dynamic regulation. To identify protein ubiquitination, a diversity of enrichment approaches to ubiquitination at multiple levels have been developed, including the protein level and the peptide level (Table 1). Since MS analysis has become the most powerful tool to precisely identify PTMs, the developed enrichment approaches combined with the advanced MS enabled the identification of tens of thousands of ubiquitination sites corresponding to thousands of ubiquitinated proteins. Considering the fact that MS-based experimental methods are often expensive, labor-intensive, and time-consuming, bioinformatics approaches and tools based on machine learning from the reported ubiquitination dataset have recently been developed for predicting protein ubiquitination sites. However, these tools are constructed based on different training libraries, prediction algorithms, functionality, and features, complicating their utilities and applications. Despite various limitations, it is now possible for researchers to analyze thousands of ubiquitination sites using experimental methods or predicting approaches. However, the prediction of ubiquitination sites by machine learning underlies the complex nature of Ub chain topologies. Because this approach only predicts the ubiquitination sites rather than the topologies of Ub chains, experimental methods are the only way to get insights into the architecture of Ub chains.
The complexity of ubiquitination stems from the ability to form the polymerization with different length (number of Ub molecules), linkage, and overall architecture. Conformation of the polyUb plays a vital role in regulating the function of substrates in diverse physiological and pathological processes. Although a series of techniques have been developed to detect branched Ub chains, none of them can reveal the topology and length of branched chains. Methodology breakthrough is desperately needed to provide systematic insights into the overall architecture of Ub chain, such as the exact structure of different linkages. Top-down proteomics (TDP) may be a promising tool to map the structure of Ub chains bearing different lengths, linkages, and architectures. However, how to solve the S/N ratio of the ubiquitinated substrates is a great challenge.
In summary, protein ubiquitination analysis has made significant progress over the past decade. However, significant challenges remain in this area. Promising methods and dedicated databases will help us untangle the complexities of ubiquitination and facilitate the discovery of biomarkers associated with abnormal protein ubiquitination in a variety of diseases.
Availability of data and materials
Not applicable.
Abbreviations
- PTM:
-
Post-translational modification
- Ub:
-
Ubiquitin
- DUBs:
-
Deubiquitinases
- M:
-
Methionine
- K:
-
Lysine
- R:
-
Arginine
- UBDs:
-
Ubiquitin binding domains
- LT:
-
Large tumor
- MS:
-
Mass spectrometry
- TUBEs:
-
Tandem-repeated Ub-binding entities
- TR-TUBEs:
-
Trypsin-resistant tandem-repeated Ub-binding entities
- ThUBD:
-
Tandem hybrid ubiquitin binding domains
- AQUA:
-
Absolute quantification
- diGly:
-
Diglycine
- LFASP:
-
Large-scale filter-aided sample preparation
- DIA:
-
Data independent acquisition
- IAA:
-
Iodoacetamide
- COFRADIC:
-
COmbined FRActional DIagonal Chromatography
- USP2cc:
-
USP2 catalytic core domain
- BOC:
-
Tert-butyloxycarbonyl
- StUbEx:
-
Stable Tagged Ub Exchange
- BUP:
-
Bottom-up proteomics
- UbiCRest:
-
Ubiquitin chain restriction
- MDP:
-
Middle-down proteomics
- TDP:
-
Top-down proteomics
- S/N:
-
Signal-to-noise
References
Xu G, Jaffrey SR. Proteomic identification of protein ubiquitination events. Biotechnol Genet Eng Rev. 2013;29:73–109.
Mabb AM. Historical perspective and progress on protein ubiquitination at glutamatergic synapses. Neuropharmacology. 2021;196:108690.
Li X, Gong L, Gu H. Regulation of immune system development and function by Cbl-mediated ubiquitination. Immunol Rev. 2019;291(1):123–33.
Zhou X, Sun SC. Targeting ubiquitin signaling for cancer immunotherapy. Signal Transduct Target Ther. 2021;6(1):16.
Santonico E. Old and new concepts in ubiquitin and NEDD8 recognition. Biomolecules. 2020;10(4):566.
Hershko A, Ciechanover A. The ubiquitin system. Annu Rev Biochem. 1998;67:425–79.
Mevissen TET, Komander D. Mechanisms of deubiquitinase specificity and regulation. Annu Rev Biochem. 2017;86:159–92.
Huang Q, Zhang X. Emerging roles and research tools of atypical ubiquitination. Proteomics. 2020;20(9):e1900100.
Snyder NA, Silva GM. Deubiquitinating enzymes (DUBs): regulation, homeostasis, and oxidative stress response. J Biol Chem. 2021;297(3):101077.
van Wijk SJ, Timmers HT. The family of ubiquitin-conjugating enzymes (E2s): deciding between life and death of proteins. FASEB J. 2010;24(4):981–93.
Lee J, et al. Genome-scale CRISPR screening identifies cell cycle and protein ubiquitination processes as druggable targets for erlotinib-resistant lung cancer. Mol Oncol. 2021;15(2):487–502.
Aprile-Garcia F, et al. Nascent-protein ubiquitination is required for heat shock-induced gene downregulation in human cells. Nat Struct Mol Biol. 2019;26(2):137–46.
Li T, et al. Dural effects of oxidative stress on cardiomyogenesis via Gata4 transcription and protein ubiquitination. Cell Death Dis. 2018;9(2):246.
**ao Y, et al. Roles of protein ubiquitination in inflammatory bowel disease. Immunobiology. 2020;225(6):152026.
Kulathu Y, Komander D. Atypical ubiquitylation—the unexplored world of polyubiquitin beyond Lys48 and Lys63 linkages. Nat Rev Mol Cell Biol. 2012;13(8):508–23.
Yau R, Rape M. The increasing complexity of the ubiquitin code. Nat Cell Biol. 2016;18(6):579–86.
Swatek KN, Komander D. Ubiquitin modifications. Cell Res. 2016;26(4):399–422.
Radley EH, et al. The “dark matter” of ubiquitin-mediated processes: opportunities and challenges in the identification of ubiquitin-binding domains. Biochem Soc Trans. 2019;47(6):1949–62.
Fennell LM, Rahighi S, Ikeda F. Linear ubiquitin chain-binding domains. FEBS J. 2018;285(15):2746–61.
Dikic I, Wakatsuki S, Walters KJ. Ubiquitin-binding domains—from structures to functions. Nat Rev Mol Cell Biol. 2009;10(10):659–71.
Hicke L, Schubert HL, Hill CP. Ubiquitin-binding domains. Nat Rev Mol Cell Biol. 2005;6(8):610–21.
Kim HJ, et al. Crosstalk between HSPA5 arginylation and sequential ubiquitination leads to AKT degradation through autophagy flux. Autophagy. 2021;17(4):961–79.
Tsuchiya H, Endo A, Saeki Y. Multi-step ubiquitin decoding mechanism for proteasomal degradation. Pharmaceuticals (Basel). 2020;13(6):128.
Li Y, et al. Stabilization of p18 by deubiquitylase CYLD is pivotal for cell cycle progression and viral replication. NPJ Precis Oncol. 2021;5(1):14.
Ha J, et al. The deubiquitinating enzyme USP20 regulates the TNFalpha-induced NF-kappaB signaling pathway through stabilization of p62. Int J Mol Sci. 2020;21(9):3116.
Morgan EL, Chen Z, Van Waes C. Regulation of NFkappaB signalling by ubiquitination: a potential therapeutic target in head and neck squamous cell carcinoma? Cancers (Basel). 2020;12(10):2877.
Chen RH, Chen YH, Huang TY. Ubiquitin-mediated regulation of autophagy. J Biomed Sci. 2019;26(1):80.
Deol KK, Strieter ER. The ubiquitin proteoform problem. Curr Opin Chem Biol. 2021;63:95–104.
Shi R, et al. SNX27-driven membrane localisation of OTULIN antagonises linear ubiquitination and NF-kappaB signalling activation. Cell Biosci. 2021;11(1):146.
Majolee J, et al. The interplay of Rac1 activity, ubiquitination and GDI binding and its consequences for endothelial cell spreading. PLoS ONE. 2021;16(7):e0254386.
Do EK, et al. Kap1 regulates the self-renewal of embryonic stem cells and cellular reprogramming by modulating Oct4 protein stability. Cell Death Differ. 2021;28(2):685–99.
Ortiz LE, Pham AM, Kwun HJ. Identification of the merkel cell polyomavirus large tumor antigen ubiquitin conjugation residue. Int J Mol Sci. 2021;22(13):7169.
Mishra V. Affinity tags for protein purification. Curr Protein Pept Sci. 2020;21(8):821–30.
Peng J, et al. A proteomics approach to understanding protein ubiquitination. Nat Biotechnol. 2003;21(8):921–6.
Akimov V, et al. StUbEx: stable tagged ubiquitin exchange system for the global investigation of cellular ubiquitination. J Proteome Res. 2014;13(9):4192–204.
Danielsen JM, et al. Mass spectrometric analysis of lysine ubiquitylation reveals promiscuity at site level. Mol Cell Proteomics. 2011;10(3):M110003590.
Denis NJ, et al. Tryptic digestion of ubiquitin standards reveals an improved strategy for identifying ubiquitinated proteins by mass spectrometry. Proteomics. 2007;7(6):868–74.
Newton K, et al. Ubiquitin chain editing revealed by polyubiquitin linkage-specific antibodies. Cell. 2008;134(4):668–78.
Matsumoto ML, et al. Engineering and structural characterization of a linear polyubiquitin-specific antibody. J Mol Biol. 2012;418(3–4):134–44.
Nakayama Y, et al. Identification of linear polyubiquitin chain immunoreactivity in tau pathology of Alzheimer’s disease. Neurosci Lett. 2019;703:53–7.
Hjerpe R, Rodriguez MS. Efficient approaches for characterizing ubiquitinated proteins. Biochem Soc Trans. 2008;36(Pt 5):823–7.
Tan F, et al. Proteomic analysis of ubiquitinated proteins in normal hepatocyte cell line Chang liver cells. Proteomics. 2008;8(14):2885–96.
Hjerpe R, et al. Efficient protection and isolation of ubiquitylated proteins using tandem ubiquitin-binding entities. EMBO Rep. 2009;10(11):1250–8.
Xolalpa W, et al. Isolation of the ubiquitin-proteome from tumor cell lines and primary cells using TUBEs. Methods Mol Biol. 2016;1449:161–75.
Hu Z, et al. Proteomic approaches for the profiling of ubiquitylation events and their applications in drug discovery. J Proteomics. 2021;231:103996.
Silva GM, Vogel C. Mass spectrometry analysis of K63-ubiquitinated targets in response to oxidative stress. Data Brief. 2015;4:130–4.
Mata-Cantero L, et al. New insights into host-parasite ubiquitin proteome dynamics in P. falciparum infected red blood cells using a TUBEs-MS approach. J Proteomics. 2016;139:45–59.
Silva GM, Finley D, Vogel C. K63 polyubiquitination is a new modulator of the oxidative stress response. Nat Struct Mol Biol. 2015;22(2):116–23.
Mattern M, et al. Using ubiquitin binders to decipher the ubiquitin code. Trends Biochem Sci. 2019;44(7):599–615.
Yoshida Y, et al. A comprehensive method for detecting ubiquitinated substrates using TR-TUBE. Proc Natl Acad Sci USA. 2015;112(15):4630–5.
Gao Y, et al. Enhanced purification of ubiquitinated proteins by engineered tandem hybrid ubiquitin-binding domains (ThUBDs). Mol Cell Proteomics. 2016;15(4):1381–96.
**ao W, et al. Specific and unbiased detection of polyubiquitination via a sensitive non-antibody approach. Anal Chem. 2020;92(1):1074–80.
Scott D, et al. Mass spectrometry insights into a tandem ubiquitin-binding domain hybrid engineered for the selective recognition of unanchored polyubiquitin. Proteomics. 2016;16(14):1961–9.
Kadimisetty K, et al. Tandem ubiquitin binding entities (TUBEs) as tools to explore ubiquitin-proteasome system and PROTAC drug discovery. Methods Mol Biol. 2021;2365:185–202.
Back S, et al. Site-Specific K63 ubiquitinomics provides insights into translation regulation under stress. J Proteome Res. 2019;18(1):309–18.
Michel MA, et al. Ubiquitin linkage-specific affimers reveal insights into K6-linked ubiquitin signaling. Mol Cell. 2017;68(1):233-246.e5.
Xu G, Paige JS, Jaffrey SR. Global analysis of lysine ubiquitination by ubiquitin remnant immunoaffinity profiling. Nat Biotechnol. 2010;28(8):868–73.
Wagner SA, et al. A proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol Cell Proteomics. 2011;10(10):M111013284.
Kim W, et al. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol Cell. 2011;44(2):325–40.
Casanovas A, et al. Large-scale filter-aided sample preparation method for the analysis of the ubiquitinome. Anal Chem. 2017;89(7):3840–6.
**ao W, et al. Ac-LysargiNase complements trypsin for the identification of ubiquitinated sites. Anal Chem. 2019;91(24):15890–8.
Hansen FM, et al. Data-independent acquisition method for ubiquitinome analysis reveals regulation of circadian biology. Nat Commun. 2021;12(1):254.
Nielsen ML, et al. Iodoacetamide-induced artifact mimics ubiquitination in mass spectrometry. Nat Methods. 2008;5(6):459–60.
Sylvestersen KB, Young C, Nielsen ML. Advances in characterizing ubiquitylation sites by mass spectrometry. Curr Opin Chem Biol. 2013;17(1):49–58.
Wagner SA, et al. Proteomic analyses reveal divergent ubiquitylation site patterns in murine tissues. Mol Cell Proteomics. 2012;11(12):1578–85.
Akimov V, et al. UbiSite approach for comprehensive map** of lysine and N-terminal ubiquitination sites. Nat Struct Mol Biol. 2018;25(7):631–40.
Kuehnel K. Catch and identify your prey. Nat Chem Biol. 2018;14(9):831.
Ciechanover A, Ben-Saadon R. N-terminal ubiquitination: more protein substrates join in. Trends Cell Biol. 2004;14(3):103–6.
Ciechanover A. N-terminal ubiquitination. Methods Mol Biol. 2005;301:255–70.
Davies CW, et al. Antibody toolkit reveals N-terminally ubiquitinated substrates of UBE2W. Nat Commun. 2021;12(1):4608.
Akimov V, et al. StUbEx PLUS-a modified stable tagged ubiquitin exchange system for peptide level purification and in-depth map** of ubiquitination sites. J Proteome Res. 2018;17(1):296–304.
Scaglione KM, et al. The ubiquitin-conjugating enzyme (E2) Ube2w ubiquitinates the N terminus of substrates. J Biol Chem. 2013;288(26):18784–8.
Walton A, et al. It’s time for some “Site”-seeing: novel tools to monitor the ubiquitin landscape in Arabidopsis thaliana. Plant Cell. 2016;28(1):6–16.
Stes E, et al. A COFRADIC protocol to study protein ubiquitination. J Proteome Res. 2014;13(6):3107–13.
Haakonsen DL, Rape M. Branching out: improved signaling by heterotypic ubiquitin chains. Trends Cell Biol. 2019;29(9):704–16.
Lopitz-Otsoa F, et al. Integrative analysis of the ubiquitin proteome isolated using Tandem Ubiquitin Binding Entities (TUBEs). J Proteomics. 2012;75(10):2998–3014.
Kliza K, Husnjak K. Resolving the complexity of ubiquitin networks. Front Mol Biosci. 2020;7:21.
Mendes ML, Fougeras MR, Dittmar G. Analysis of ubiquitin signaling and chain topology cross-talk. J Proteomics. 2020;215:103634.
Yau RG, et al. Assembly and function of heterotypic ubiquitin chains in cell-cycle and protein quality control. Cell. 2017;171(4):918-933.e20.
Ohtake F, et al. The K48–K63 branched ubiquitin chain regulates NF-kappaB signaling. Mol Cell. 2016;64(2):251–66.
Hospenthal MK, Mevissen TET, Komander D. Deubiquitinase-based analysis of ubiquitin chain architecture using Ubiquitin Chain Restriction (UbiCRest). Nat Protoc. 2015;10(2):349–61.
Schaefer JB, Morgan DO. Protein-linked ubiquitin chain structure restricts activity of deubiquitinating enzymes. J Biol Chem. 2011;286(52):45186–96.
Emmerich CH, et al. Activation of the canonical IKK complex by K63/M1-linked hybrid ubiquitin chains. Proc Natl Acad Sci USA. 2013;110(38):15247–52.
Hospenthal MK, Freund SM, Komander D. Assembly, analysis and architecture of atypical ubiquitin chains. Nat Struct Mol Biol. 2013;20(5):555–65.
Nakasone MA, et al. Mixed-linkage ubiquitin chains send mixed messages. Structure. 2013;21(5):727–40.
Kirkpatrick DS, et al. Quantitative analysis of in vitro ubiquitinated cyclin B1 reveals complex chain topology. Nat Cell Biol. 2006;8(7):700–10.
Kaiser SE, et al. Protein standard absolute quantification (PSAQ) method for the measurement of cellular ubiquitin pools. Nat Methods. 2011;8(8):691–6.
Rana A, Ge Y, Strieter ER. Ubiquitin chain enrichment middle-down mass spectrometry (UbiChEM-MS) reveals cell-cycle dependent formation of Lys11/Lys48 branched ubiquitin chains. J Proteome Res. 2017;16(9):3363–9.
Crowe SO, et al. Ubiquitin chain enrichment middle-down mass spectrometry enables characterization of branched ubiquitin chains in cellulo. Anal Chem. 2017;89(8):4428–34.
Deol KK, Eyles SJ, Strieter ER. Quantitative middle-down MS analysis of Parkin-mediated ubiquitin chain assembly. J Am Soc Mass Spectrom. 2020;31(5):1132–9.
Valkevich EM, et al. Middle-down mass spectrometry enables characterization of branched ubiquitin chains. Biochemistry. 2014;53(30):4979–89.
Swatek KN, et al. Irreversible inactivation of ISG15 by a viral leader protease enables alternative infection detection strategies. Proc Natl Acad Sci USA. 2018;115(10):2371–6.
Swatek KN, et al. Insights into ubiquitin chain architecture using Ub-clip**. Nature. 2019;572(7770):533–7.
Kline JT, et al. Sequential ion-ion reactions for enhanced gas-phase sequencing of large intact proteins in a Tribrid Orbitrap mass spectrometer. J Am Soc Mass Spectrom. 2021;32(9):2334–45.
Geis-Asteggiante L, Lee AE, Fenselau C. Analysis of the topology of ubiquitin chains. Methods Enzymol. 2019;626:323–46.
Lee AE, et al. Preparing to read the ubiquitin code: top-down analysis of unanchored ubiquitin tetramers. J Mass Spectrom. 2016;51(8):629–37.
He W, Wei L, Zou Q. Research progress in protein posttranslational modification site prediction. Brief Funct Genomics. 2018;18(4):220–9.
Tung CW, Ho SY. Computational identification of ubiquitylation sites from protein sequences. BMC Bioinform. 2008;9:310.
Kawashima S, et al. AAindex: amino acid index database, progress report 2008. Nucleic Acids Res. 2008;36(Database issue):D202–5.
He F, et al. Large-scale prediction of protein ubiquitination sites using a multimodal deep architecture. BMC Syst Biol. 2018;12(Suppl 6):109.
Wang X, Yan R, Wang Y. Computational identification of human ubiquitination sites using convolutional and recurrent neural networks. Mol Omics. 2021;17:948–55.
Chen Z, et al. Towards more accurate prediction of ubiquitination sites: a comprehensive review of current methods, tools and features. Brief Bioinform. 2015;16(4):640–57.
Wang L, Zhang R. Towards computational models of identifying protein ubiquitination sites. Curr Drug Targets. 2019;20(5):565–78.
Wang RY, et al. Structure of Hsp90-Hsp70-Hop-GR reveals the Hsp90 client-loading mechanism. Nature. 2022;601(7893):460–4.
Acknowledgements
We thank Qing Zhang, and the members of the **aofei Zhang laboratory for their discussions.
Funding
Work in Zhang lab is supported by Grants from the National Key R&D Program of China (2020YFE0202200), the Science and Technology Planning Project of Guangdong Province (2018B030306047, 2020B1212060052), Guangzhou Science and Technology Project (201904010469) and Guangzhou Regenerative Medicine and Health Guangdong Laboratory project (2018GZR110104003). Mingwei Sun is supported by grant from the National Natural Science Foundation (22004021) and the GuangDong Basic and Applied Basic Research Foundation (2020A1515110119).
Author information
Authors and Affiliations
Contributions
Writing, MWS and XFZ.; supervision, XFZ. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Sun, M., Zhang, X. Current methodologies in protein ubiquitination characterization: from ubiquitinated protein to ubiquitin chain architecture. Cell Biosci 12, 126 (2022). https://doi.org/10.1186/s13578-022-00870-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13578-022-00870-y