Abstract
Graph Convolutional Neural Network (GCNN) is a popular class of deep learning (DL) models in material science to predict material properties from the graph representation of molecular structures. Training an accurate and comprehensive GCNN surrogate for molecular design requires large-scale graph datasets and is usually a time-consuming process. Recent advances in GPUs and distributed computing open a path to reduce the computational cost for GCNN training effectively. However, efficient utilization of high performance computing (HPC) resources for training requires simultaneously optimizing large-scale data management and scalable stochastic batched optimization techniques. In this work, we focus on building GCNN models on HPC systems to predict material properties of millions of molecules. We use HydraGNN, our in-house library for large-scale GCNN training, leveraging distributed data parallelism in PyTorch. We use ADIOS, a high-performance data management framework for efficient storage and reading of large molecular graph data. We perform parallel training on two open-source large-scale graph datasets to build a GCNN predictor for an important quantum property known as the HOMO-LUMO gap. We measure the scalability, accuracy, and convergence of our approach on two DOE supercomputers: the Summit supercomputer at the Oak Ridge Leadership Computing Facility (OLCF) and the Perlmutter system at the National Energy Research Scientific Computing Center (NERSC). We present our experimental results with HydraGNN showing (i) reduction of data loading time up to 4.2 times compared with a conventional method and (ii) linear scaling performance for training up to 1024 GPUs on both Summit and Perlmutter.
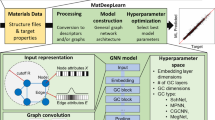
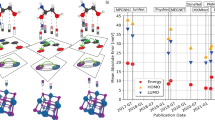
Similar content being viewed by others
Introduction
Drug discovery and molecular design rely heavily on predicting material properties directly from their atomic structure. A particular property of interest for molecular design is the energy gap between the highest occupied molecular orbital (HOMO) and lowest unoccupied molecular orbital (LUMO), known as the HOMO-LUMO gap. The HOMO-LUMO gap is a valid approximation for the lowest excitation energy of a molecule and is used to express its chemical reactivity. In particular, molecules that are more chemically reactive are characterized by a lower HOMO-LUMO gap. There are many physics-based computational approaches to compute the HOMO-LUMO gap of a molecule such as ab initio molecular dynamics (MD) [1, 2] and density-functional tight-binding (DFTB) [3]. While these methods have been instrumental in predictive materials science, they are extremely computationally expensive. The advent of deep learning (DL) models has provided alternative methodologies to produce fast and accurate predictions of material properties and hence enable rapid screening in the large search space to select material candidates with desirable properties [4,5,6,7].
In particular, graph convolutional neural network (GCNN) models are extensively used in material science to predict material properties from atomic information [1.
We have developed an extensible data loader module in HydraGNN that allows reading data from different storage formats. In this work, we evaluate the following three methods for loading data from training datasets.
-
Inline data loading: load SMILES strings written in CSV format into memory and then convert each SMILES into a graph object at every batch data loading. It has the smallest memory footprint.
-
Object data loading: convert all SMILES strings into graph objects and export them in a serialized format (e.g., Pickle) during a preprocessing phase. A process loads each data batch directly from the file system and unpacks into a memory.
-
ADIOS data loading: convert SMILES strings into graph objects mapped to ADIOS variables, and write them into an ADIOS file in a pre-processing step. Each process then loads its batch data during training in parallel along with other processes.
We will discuss performance comparisons in the next section.
Numerical results
In this section, we assess our development of DDP training in HydraGNN on two state-of-the-art DOE supercomputers, Summit and Perlmutter. We discuss the scalability of our approach, and compare the performance of different I/O backends for storing and reading graph data.
Setup
We perform our evaluation on two supercomputers of DOE. Both systems provide state-of-the-art GPU-based heterogeneous architectures.
Summit is a supercomputer at OLCF, one of DOE’s Leadership Computing Facilities (LCFs). Summit consists of about 4600 compute nodes. Each node has a hybrid architecture containing two IBM POWER9 CPUs and six NVIDIA Volta GPUs connected with NVIDIA’s high-speed NVLink. Each node contains 512 GB of DDR4 memory for CPUs and 96 GB of High Bandwidth Memory (HBM2) for GPUs. Summit nodes are connected in a non-blocking fat-tree topology using a dual-rail Mellanox EDR InfiniBand interconnection.
Perlmutter is a supercomputer at NERSC. Perlmutter consists of about 3000 CPU-only nodes and 1500 GPU-accelerated nodes. We use only the GPU-accelerated nodes in our work. Each GPU-accelerated node has an AMD EPYC 7763 (Milan) processor and four NVIDIA Ampere A100 GPUs connected to each other with NVLink-3. Each GPU node has 256 GB of DDR4 memory and 40 GB HBM2 per each GPU. All nodes in Perlmutter are connected with the HPE Cray Slingshot interconnect.
We demonstrate the performance of HydraGNN using two large-scale datasets, a previously published benchmark dataset for graph-based learning (PCQM4Mv2) [11, 44] and a custom dataset generated for this work (AISD HOMO-LUMO) [13]. For both datasets, molecule information is provided as SMILES strings. The PCQM4Mv2 consists of HOMO-LUMO gap values for about 3.3 million molecules. In total, 31 different types of atoms (i.e., H, B, C, N, O, F, Si, P, S, Cl, Ca,Ge,As, Se, Br, I, Mg, Ti, Ga, Zn, Ar, Be, He, Al, Kr, V, Na, Li, Cu, Ne, Ni) are involved in the dataset. The custom AISD HOMO-LUMO dataset was generated using molecular structures from previous work [45]. It is a collection of approximately 10.5 million molecules and contains 6 element types (i.e., H, C, N, O, S, and F).
For scalability tests, we use HydraGNN with 6 PNA [28] convolutional layers and 55 neurons per PNA layer. The model is trained by using the AdamW method [46] with a learning rate of 0.001, local batch size of 128, and maximum epochs set to 3. The training set for each of the NN represents 94% of the total dataset; the validation and test sets each represent \(1/3^{rd}\) and \(2/3^{rd}\) parts respectively of the remaining data. For the error convergence tests, the HydraGNN model uses 200 neurons per layer.
Scalability of DDP
We perform DDP training with HydraGNN for PCQM4Mv2 and AISD data on Summit at ORNL and Perlmutter at NERSC using multiple CPUs and GPUs. The number of graphs and size in the datasets are summarized in Table 2.
We measure the total training time for PCQM4Mv2 and AISD HOMO-LUMO datasets over three epochs. As discussed previously, each training consists of a data loading phase, followed by forward calculation, backward calculation, and optimizer update. We test the scalability of DDP by varying the number of nodes on each system, ranging from a single node up to 256 nodes on Summit and 128 nodes on Perlmutter, corresponding to using 1536 Volta GPUs and 512 A100 GPUs respectively. Figure 5 shows the result. The scaling plot (top) shows the averaged training time for PCQM4Mv2 and AISD HOMO-LUMO on each system with a varying number of nodes, and the detailed timings of each sub-function during the training on Summit are shown at the bottom.
We obtain near-linear scaling up to 1024 GPUs for both PCQM4Mv2 and AISD HOMO-LUMO data. As we further scale the workflow on Summit, the number of batches per GPU decreases, leading to sub-optimal utilization of GPU resources. As a result, we see a drop in speedup as we scale beyond 1024 GPUs on Summit. We expect similar scaling behavior on Perlmutter, but we were limited to using 128 nodes (i.e., 512 GPUs) for this work.
Comparing different I/O backends
Data loading takes a significant amount of time in training, as shown in Fig. 5, and hence is a crucial step in the overall workflow. We compare three different data loading methods—inline, object loading, and ADIOS data loading, as discussed in "Distributed data parallel training" section. Figure 6 presents time taken by the three methods for the PCQM4Mv2 data set on Summit. As expected, it outperforms the CSV data loading test case in which SMILES data is converted into a graph object for every molecule. ADIOS outperforms Pickle-based data loading by 4.2x on a single Summit node and 1.5x on 32 Summit nodes. To provide a complete picture of HydaGNN’s data processing, Fig. 7 shows the pre-processing performance in converting the PCQM4Mv2 data into ADIOS2 on Summit. ADIOS supports parallel writing and shows scalable performance as we add CPUs in parallel.
Accuracy
Next, we perform long-running HydraGNN training for the HOMO-LUMO gap prediction with PCQM4Mv2 and AISD HOMO-LUMO datasets until training converges.
Figures 8 and 9 show the prediction results for PCQM4Mv2 and AISD HOMO-LUMO datasets respectively. With PCQM4Mv2, we achieve a prediction error of around 0.10 and 0.12 eV, measured in mean absolute error (MAE), for the training and validation set, respectively. We note that PCQM4Mv2 is a public dataset released without test data for the purpose of maintaining the OGB-LSC LeaderboardsFootnote 2. The reported validation MAE from multiple models varies from 0.0857 to 0.1760 eV on the Leaderboard. The validation error of 0.12 eV in this work is within the accepted range. As for the AISD HOMO-LUMO dataset, it contains almost thrice as many molecules as the PCQM4Mv2 dataset. The MAE errors for training, validation, and test sets are 0.14 eV, which is similar to the PCQM4Mv2 dataset. Figure 10 shows the accuracy convergence on the AISD HOMO-LUMO dataset using different numbers of Summit GPUs. It shows that the HydraGNN training with 192 GPUs quickly converged in 0.3 h (wall time) to the similar accuracy level achieved by the 6 GPUs (a single Summit node) that took about 8.2 h.
We highlight that the convergence of the distributed GCNN training with 192 GPUs is deteriorated compared to the distributed training with only 6 GPUs. This is due to a well known numerical artifact that destabilizes the training of DL models at large scale and causes a performance drop because large scale DDP training is mathematically equivalent to large-batch training. In fact, processing data in large batches significantly reduces the stochastic oscillations of the stochastic optimizer used for DL training, thus making the DL training more likely to be trapped in steep local minima, which adversely affect generalization. Although the final accuracy of the GCNN training with 192 GPUs is slightly worse than the one obtained using 6 GPUs for training, we emphasize the significant advantage that HPC resources provide in speeding-up the training. Better accuracy can be obtained when training DL models at large scale by adaptively tuning the learning rate [47, 48] or by applying quasi-Newton accelerations [49], but this goes beyond the focus of our current work.
Conclusions and future work
In this paper, we present a computational workflow that performs DDP training to predict the HOMO-LUMO gap of molecules. We have implemented DDP in HydraGNN, a GCNN library developed at ORNL, which can utilize heterogeneous computing resources including CPUs and GPUs. For efficient storage and loading of large molecular data, we use the ADIOS high-performance data management framework. ADIOS helps reduce the storage footprint of large-scale graph structures as compared with commonly used methods, and provides an easy way to efficiently load data and distribute them amongst processes. We have conducted studies using two molecular datasets on the OLCF’s Summit and NERSC’s Perlmutter supercomputers. Our results show the near-linear scaling of HydraGNN for the test datasets up to 1024 GPUs. Additionally, we present the accuracy and convergence behavior of the distributed training with increasing number of GPUs.
Through efficiently managing large-scale datasets and training in parallel, HydraGNN provides an effective surrogate model for accurate and rapid screening of large chemical spaces for molecular design. Future work will be dedicated to integrating the scalable DDP training of HydraGNN in a computational workflow to perform molecular design.
Availability of data and materials
The AISD HOMO-LUMO dataset has been generated and analysed for this work. It is open-source and accessible at the following URL https://doi.ccs.ornl.gov/ui/doi/394. Our main code HydraGNN is openly available on Github: https://github.com/ORNL/HydraGNN
References
Car R, Parrinello M (1985) Unified approach for molecular dynamics and density-functional theory. Phys Rev Lett 55:2471–2474. https://doi.org/10.1103/PhysRevLett.55.2471
Marx D, Hutter J (2012) Ab Initio molecular dynamics. Basic theory and advanced methods. Cambridge University Press, New York
Sokolov M, Bold BM, Kranz JJ, Hofener S, Niehaus TA, Elstner M (2021) Analytical time-dependent long-range corrected density functional tight binding (TD-LC-DFTB) gradients in DFTB+: implementation and benchmark for excited-state geometries and transition energies. J Chem Theory Comput. 17(4):2266–2282
Gaultois MW, Oliynyk AO, Mar A, Sparks TD, Mulholland GJ, Meredig B (2016) Perspective: web-based machine learning models for real-time screening of thermoelectric materials properties. APL Mater. 4:053213. https://doi.org/10.1063/1.4952607
Lu S, Zhou Q, Ouyang Y, Guo Y, Li Q, Wang J (2018) Accelerated discovery of stable lead-free hybrid organic-inorganic perovskites via machine learning. Nat Commun. 9:3405. https://doi.org/10.1038/s41467-018-05761-w
Gómez-Bombarelli R (2016) Design of efficient molecular organic light-emitting diodes by a high-throughput virtual screening and experimental approach. Nat Mater 15:1120–1127. https://doi.org/10.1038/nmat4717
Xue D, Balachandran PV, Hogden J, Theiler J, Xue D, Lookman T (2016) Accelerated search for materials with targeted properties by adaptive design. Nat Commun 7:11241. https://doi.org/10.1038/nmat4717
**e T, Grossman JC (2018) Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties. Phys Rev Lett. 120(14):14530. https://doi.org/10.1103/PhysRevLett.120.145301
Chen C, Ye W, Zuo Y, Zheng C, Ong SP (2019) Graph networks as a universal machine learning framework for molecules and crystals. Chem Mater 31(9):3564–3572. https://doi.org/10.1021/acs.chemmater.9b01294
Reymond JL (2015) The chemical space project. Acc Chem Res 48(3):722–730. https://doi.org/10.1021/ar500432k
Hu W, Fey M, Zitnik M, Dong Y, Ren H, Liu B, Catasta M, Leskovec J (2020) Open graph benchmark: Datasets for machine learning on graphs. Advances in Neural Information Processing Systems 2020-Decem(NeurIPS), 1–34 ar**v:2005.00687
Hu W, Fey M, Ren H, Nakata M, Dong Y, Leskovec J (2021) OGB-LSC: A large-scale challenge for machine learning on graphs. ar**v preprint ar**v:2103.09430
Blanchard AE, Gounley J, Bhowmik D, Pilsun Y, Irle S AISD HOMO-LUMO. https://doi.org/10.13139/ORNLNCCS/1869409
Lupo Pasini M, Zhang P, Reeve ST, Choi JY (2022) Multi-task graph neural networks for simultaneous prediction of global and atomic properties in ferromagnetic systems. Mach Learn Sci Technol. 3(2):025007. https://doi.org/10.1088/2632-2153/ac6a51
Godoy WF, Podhorszki N, Wang R, Atkins C, Eisenhauer G, Gu J, Davis P, Choi J, Germaschewski K, Huck K et al (2020) ADIOS 2: the adaptable input output system. A framework for high-performance data management. SoftwareX 12:100561
Gilmer J, Schoenholz SS, Riley PF, Vinyals O, Dahl GE (2017) Neural message passing for quantum chemistry. In: International Conference on Machine Learning, pp. 1263–1272. PMLR
Choudhary K, DeCost B (2021) Atomistic line graph neural network for improved materials property predictions. NPJ Comput Mater 7(1):1–8
Nakamura T, Sakaue S, Fujii K, Harabuchi Y, Maeda S (2020) Iwata S Selecting molecules with diverse structures and properties by maximizing submodular functions of descriptors learned with graph neural networks. Sci Rep. 12:1124. https://doi.org/10.1021/acs.jcim.0c00687
Rahaman O, Gagliardi A (2020) Deep learning total energies and orbital energies of large organic molecules using hybridization of molecular fingerprints. J Chem Inf Model. 60(12):5971–5983. https://doi.org/10.1021/acs.jcim.0c00687
Ramakrishnan R, Dral PO, Rupp M, Von Lilienfeld OA (2014) Quantum chemistry structures and properties of 134 kilo molecules. Sci Data 1(1):1–7
Stuke A, Kunkel C, Golze D, Todorović M, Margraf JT, Reuter K, Rinke P, Oberhofer H (2020) Atomic structures and orbital energies of 61,489 crystal-forming organic molecules. Sci Data 7(1):1–11
Ying C, Cai T, Luo S, Zheng S, Ke G, He D, Shen Y, Liu T-Y (2021) Do transformers really perform badly for graph representation? In: Advances in Neural Information Processing Systems, vol. 34, pp. 28877–28888. https://proceedings.neurips.cc/paper/2021/file/f1c1592588411002af340cbaedd6fc33-Paper.pdf
Park W, Chang W-G, Lee D, Kim J, Hwang S-w (2022) GRPE: Relative positional encoding for graph transformer. In: ICLR2022 Machine Learning for Drug Discovery. https://openreview.net/forum?id=GNfAFN_p1d
Besta M, Hoefler T (2022) Parallel and distributed graph neural networks: an in-depth concurrency analysis. https://doi.org/10.48550/ARXIV.2205.09702
Folk M, Heber G, Koziol Q, Pourmal E, Robinson D (2011) An overview of the HDF5 technology suite and its applications. In: Proceedings of the EDBT/ICDT 2011 Workshop on Array Databases, pp. 36–47
Scarselli F, Gori M, Tsoi AC, Hagenbuchner M, Monfardini G (2009) The graph neural network model. IEEE Trans Neural Netw 20(1):61–80. https://doi.org/10.1109/TNN.2008.2005605
Defferrard M, Bresson X, Vandergheynst P (2016) Convolutional neural networks on graphs with fast localized spectral filtering. In: Lee, D., Sugiyama, M., Luxburg, U., Guyon, I., Garnett, R. (eds.) Advances in Neural Information Processing Systems, vol. 29. Curran Associates, Inc., Centre Convencions Internacional Barcelona, Barcelona Sain. https://proceedings.neurips.cc/paper/2016/file/04df4d434d481c5bb723be1b6df1ee65-Paper.pdf
Corso G, Cavalleri L, Beaini D, Liò P, Veličković P (2020) Principal neighbourhood aggregation for graph nets. Adv Neural Inf Process Syst 33:13260–13271
Hamilton WL, Ying R, Leskovec J (2017) Inductive representation learning on large graphs. In: Guyon, I., Luxburg, U.V., Bengio, S., Wallach, H., Fergus, R., Vishwanathan, S., Garnett, R. (eds.) Advances in Neural Information Processing Systems, vol. 30, pp. 1025–1035. Curran Associates, Inc., Long Beach Convention Center, Long Beach. https://proceedings.neurips.cc/paper/2017/file/5dd9db5e033da9c6fb5ba83c7a7ebea9-Paper.pdf
Lupo Pasini M, Reeve ST, Zhang P, Choi JY (2021) HydraGNN. Computer Software. https://doi.org/10.11578/dc.20211019.2. https://github.com/ORNL/HydraGNN
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, Killeen T, Lin Z, Gimelshein N, Antiga L et al (2019) Pytorch: an imperative style, high-performance deep learning library. Adv Neural Inf Process Syst. 32
Fey M, Lenssen JE (2019) Fast graph representation learning with PyTorch Geometric. In: ICLR Workshop on Representation Learning on Graphs and Manifolds
PyTorch Geometric. https://pytorch-geometric.readthedocs.io/en/latest/
Dominski J, Cheng J, Merlo G, Carey V, Hager R, Ricketson L, Choi J, Ethier S, Germaschewski K, Ku S et al (2021) Spatial coupling of gyrokinetic simulations, a generalized scheme based on first-principles. Phys Plasmas 28(2):022301
Merlo G, Janhunen S, Jenko F, Bhattacharjee A, Chang C, Cheng J, Davis P, Dominski J, Germaschewski K, Hager R et al (2021) First coupled GENE-XGC microturbulence simulations. Phys Plasmas 28(1):012303
Cheng J, Dominski J, Chen Y, Chen H, Merlo G, Ku S-H, Hager R, Chang C-S, Suchyta E, D’Azevedo E et al (2020) Spatial core-edge coupling of the particle-in-cell gyrokinetic codes GEM and XGC. Phys Plasmas 27(12):122510
Poeschel F, Godoy WF, Podhorszki N, Klasky S, Eisenhauer G, Davis PE, Wan L, Gainaru A, Gu J, Koller F et al (2021) Transitioning from file-based HPC workflows to streaming data pipelines with openPMD and ADIOS2. ar**v preprint ar**v:2107.06108
Wan L, Huebl A, Gu J, Poeschel F, Gainaru A, Wang R, Chen J, Liang X, Ganyushin D, Munson T et al (2021) Improving I/O performance for exascale applications through online data layout reorganization. IEEE Trans Parallel Distrib Syst 33(4):878–890
Wang D, Luo X, Yuan F, Podhorszki N (2017) A data analysis framework for earth system simulation within an in-situ infrastructure. J Comput Commun. 5(14)
Thompson AP, Aktulga HM, Berger R, Bolintineanu DS, Brown WM, Crozier PS, in ’t Veld PJ, Kohlmeyer A, Moore SG, Nguyen TD, Shan R, Stevens MJ, Tranchida J, Trott C, Plimpton SJ, (2022) LAMMPS—a flexible simulation tool for particle-based materials modeling at the atomic, meso, and continuum scales. Comp Phys Comm. 271:108171. https://doi.org/10.1016/j.cpc.2021.108171
OLCF Supercomputer Summit. https://www.olcf.ornl.gov/olcf-resources/compute-systems/summit/
Weininger D (1998) SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J Chem Inf Comput Sci. 28:31–36. https://doi.org/10.1021/ci00057a005
Nakata M, Shimazaki T (2017) PubChemQC project: a large-scale first-principles electronic structure database for data-driven chemistry. J Chem Inf Model 57(6):1300–1308. https://doi.org/10.1021/acs.jcim.7b00083
Blanchard AE, Gounley J, Bhowmik D, Shekar MC, Lyngaas I, Gao S, Yin J, Tsaris A, Wang F, Glaser J (2021) Language models for the prediction of SARS-CoV-2 inhibitors. Preprint at https://www.biorxiv.org/content/10.1101/2021.12.10.471928v1
Loshchilov I, Hutter F (2019) Decoupled weight decay regularization. In: 7th International Conference on Learning Representations, ICLR 2019. OpenReview.net, New Orleans, LA, USA. https://openreview.net/forum?id=Bkg6RiCqY7
You Y, Gitman I, Ginsburg B (2017) Large batch training of convolutional networks. ar**v:1708.03888 [cs.CV]. ar**v:1708.03888
You Y, Hseu J, Ying C, Demmel J, Keutzer K, Hsieh C-J (2019) Large-batch training for lstm and beyond. In: Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis. SC ’19. Association for Computing Machinery, New York, NY, USA. https://doi.org/10.1145/3295500.3356137
Pasini ML, Yin J, Reshniak V, Stoyanov MK (2022) Anderson acceleration for distributed training of deep learning models. In: SoutheastCon 2022, pp. 289–295. https://doi.org/10.1109/SoutheastCon48659.2022.9763953
Acknowledgements
Massimiliano Lupo Pasini thanks Dr. Vladimir Protopopescu for his valuable feedback in the preparation of this manuscript.
This manuscript has been authored by UT-Battelle, LLC under Contract No. DE-AC05-00OR22725 with the U.S. Department of Energy. The publisher, by accepting the article for publication, acknowledges that the U.S. Government retains a non-exclusive, paid up, irrevocable, world-wide license to publish or reproduce the published form of the manuscript, or allow others to do so, for U.S. Government purposes. The DOE will provide public access to these results in accordance with the DOE Public Access Plan (http://energy.gov/downloads/doe-public-access-plan).
Funding
This work was supported in part by the Office of Science of the Department of Energy and by the Laboratory Directed Research and Development (LDRD) Program of Oak Ridge National Laboratory. This research is sponsored by the Artificial Intelligence Initiative as part of the Laboratory Directed Research and Development (LDRD) Program of Oak Ridge National Laboratory, managed by UT-Battelle, LLC, for the US Department of Energy under contract DE-AC05-00OR22725. An award of computer time was provided by the OLCF Director’s Discretion Project program using OLCF awards CSC457 and MAT250 and the INCITE program. This work used resources of the Oak Ridge Leadership Computing Facility and of the Edge Computing program at the Oak Ridge National Laboratory, which is supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC05-00OR22725. This research used resources of the National Energy Research Scientific Computing Center (NERSC), a U.S. Department of Energy Office of Science User Facility located at Lawrence Berkeley National Laboratory, operated under Contract No. DE-AC02-05CH11231 using NERSC award ASCR-ERCAP-m4133.
Author information
Authors and Affiliations
Contributions
All authors contributed to the paper’s conception and design. JYC and KM developed the ADIOS schema, and designed the experiments for evaluating different data loaders in HydraGNN. JYC implemented the ADIOS and Pickle backends, performed experiments on Summit and Perlmutter, and analyzed results. PZ contributed to the development of HydraGNN, results analysis and paper writing. AB performed material preparation, data collection and analysis. MLP contributed to the development of core capabilities in HydraGNN and wrote the main narrative of the manuscript. All authors contributed by commenting on the previous versions of the manuscript and revisions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Choi, J.Y., Zhang, P., Mehta, K. et al. Scalable training of graph convolutional neural networks for fast and accurate predictions of HOMO-LUMO gap in molecules. J Cheminform 14, 70 (2022). https://doi.org/10.1186/s13321-022-00652-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13321-022-00652-1