Abstract
The human bone marrow mesenchymal stem cells (hBMSCs) undergo intense osteogenic differentiation, a crucial bone formation mechanism. Evidence from prior studies suggested an association between long noncoding RNAs (lncRNAs) and the osteogenic differentiation of hBMSCs. However, precise roles and molecular mechanisms are still largely unknown. In this work, we report for the first time that lncRNA KCNMA1 antisense RNA 1 (KCNMA1-AS1) plays a vital role in regulating hBMSCs’ osteogenic differentiation. Here, it was observed that the KCNMA1-AS1 expression levels were significantly upregulated during osteogenic differentiation. In addition, KCNMA1-AS1 overexpression enhanced in vitro osteogenic differentiation of hBMSCs and in vivo bone formation, whereas knockdown of KCNMA1-AS1 resulted in the opposite result. Additionally, the interaction between KCNMA1-AS1 and mothers against decapentaplegic homolog 9 (SMAD9) was confirmed by an RNA pull-down experiment, mass spectrometry, and RIP assay. This interaction regulated the activation of the SMAD9 signaling pathway. Moreover, rescue assays demonstrated that the inhibitor of the SMAD9 signaling pathway reversed the stimulative effects on osteogenic differentiation of hBMSCs by KCNMA1-AS1 overexpression. Altogether, our results stipulate that KCNMA1-AS1 promotes osteogenic differentiation of hBMSCs via activating the SMAD9 signaling pathway and can serve as a biomarker and therapeutic target in treating bone defects.
Similar content being viewed by others
Introduction
The bone tissue structure of the oral and maxillofacial region is an important basis for supporting the facial shape. However, maxillofacial bone defects due to trauma, tumor resection, infection, and congenital malformation can cause not only facial deformities, but also difficulty in chewing, swallowing, speaking, and breathing. This could affect a patient’s physiological and psychological well-being [1,2,3]. Nowadays, autologous bone grafts are commonly employed as the standard procedure to repair bone defects. Yet, the disadvantages such as secondary injury of the donor site, limited supply, and poor prognosis in patients with osteoporosis are prominent [4, 5]. Although synthetic bone can be an alternative, its limitations include immune rejection, infection, and low capacity for osteoinduction and osteo-integration [6, 7]. Lately, bone tissue engineering has been the focus and is receiving extensive attention and research. Bone tissue engineering is a combination of seed cells, specific osteogenic factors, and biological scaffold materials aiming to create a reconstructed tissue with a similar function as a natural bone tissue; it is anticipated to offer a successful procedure for bone reconstruction [8,9,10]. Among the various types of cells, mesenchymal stem cells (MSCs) are generally preferred for bone repair. Human bone marrow mesenchymal stem cells (hBMSCs), as adult stem cells, have promising potential as seed cells in bone tissue engineering owing to their ability to self-renew, pluripotency, and mild immunogenicity [11,12,13]. Osteogenic differentiation of MSCs primarily includes cell differentiation, cell maturation, extracellular matrix formation, and mineralization. This process can be regulated by growth factors belonging to the Wnt family and bone morphogenetic protein (BMP) family, transcription factors that include β-catenin, runt-related transcription factor 2 (RUNX2), osterix (OSX), and certain microRNAs (miRNAs) [14, 15].
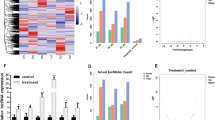
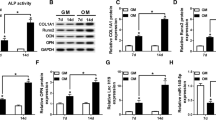
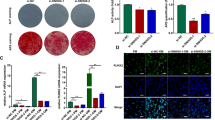
In some recent studies, long noncoding RNAs (lncRNAs) are reported to regulate how hBMSCs differentiate into osteoblasts [16, 27, 40]. KCNMA1-AS1 was previously shown to possess oncogenic activity and promote the proliferation of ovarian cancer cells [25]. No relevant reports to date have reported KCNMA1-AS1 to have any effect on osteogenic differentiation. In the current study, the increased expression level of KCNMA1-AS1 was observed during the osteogenic differentiation of hBMSCs. Accordingly, the prospective function of KCNMA1-AS1 in regulating osteogenic differentiation was investigated during this work. Gain/loss function assays were performed with lentiviral transfection models in subsequent experiments, which revealed that KCNMA1-AS1 overexpression promoted osteogenesis of hBMSCs both in vivo and in vitro. In contrast,KCNMA1-AS1 deficiency gave rise to the opposite outcome. These results indicated a positive influence of KCNMA1-AS1 on osteogenic differentiation taking place in the hBMSCs.
Further, the underlying osteogenic differentiation mechanism involving KCNMA1-AS1 was investigated using FISH assay and the results indicated that KCNMA1-AS1 was mainly localized in the nuclei of hBMSCs. The function of lncRNAs depends on subcellular localization. Nuclear lncRNAs have been found to influence gene expressions by interacting with transcription factors and proteins in diverse biological processes [41]. For instance, the interaction between lncRNA HOTTIP and WDR5 results in the activation of the Wnt/β-catenin pathway, which enhances the osteogenic differentiation of BMSCs [42]. LncRNA MEG3 regulates chondrogenic differentiation by inhibiting TRIB2, which is achieved by binding with EZH2 [43] as well as LINC02273; it is associated with hnRNPL, which promotes metastasis of breast cancer by increasing AGR2 transcription [44]. Furthermore, using RNA pull-down assay, mass spectrometry, and RIP assay, we were able to detect a tight junction between KCNMA1-AS1 and SMAD9.
Belonging to the transforming growth factor beta (TGF-β) superfamily, mothers against decapentaplegic (SMAD) proteins, which have found eight members in mammals, have been divided into three subgroups, namely, common SMAD (Co-SMAD or SMAD4), receptor-regulated SMADs or R-SMADs, and inhibitory SMADs or I-SMADs. SMAD9 (previously known as SMAD8), being an R-SMAD, is a prominent transcription factor. Also, SMAD9, along with SMAD1 and SMAD5, is phosphorylated and activated directly by BMP type I receptor to form a heterotrimeric complex with SMAD4; these complexes take part in the regulation of target genes and proteins [45,46,47]. It is important to note that the SMAD9 signaling pathway plays a significant role in regulating bone development as described before [48,49,50,51]. Rare lncRNAs have also been reported to be associated with the SMAD9 signaling pathway during osteogenic differentiation, in which lncRNA SNHG5 [52] is an example. Surprisingly, in this study, we discovered that KCNMA1-AS1 overexpression triggered the phosphorylation of SMAD9 both in vivo and in vitro activating the SMAD9 signaling cascade, whereas KCNMA1-AS1 interference attenuated the expression of p-SMAD9, thereby inhibiting the SMAD9 signaling pathway. To verify whether KCNMA1-AS1 impacts the osteogenic differentiation of hBMSCs by activating the SMAD9 signaling pathway, we performed rescue experiments using LDN193189 as a repressor of the SMAD9 signaling pathway. The expression levels of p-SMAD9 and osteogenic specific markers tested by western blot suggested that they were remarkably lowered in hBMSCs treated with KCNMA1-AS1 overexpression and LDN193189 compared to those treated with KCNMA1-AS1 overexpression and DMSO. In addition, the application of LDN193189 abrogated the promotional effects of KCNMA1-AS1 overexpression on ALP activity and mineralized calcium nodules deposition. These findings proved that KCNMA1-AS1 regulates osteogenic differentiation by modulating the SMAD9 signaling pathway. However, the effect of KCNMA1-AS1 on bone regeneration certainly needs to be elucidated further using bone defect models.
The present research showed that lncRNA KCNMA1-AS1 promoted osteogenic differentiation of HBMSCs by targeting the SMAD signaling pathway. LncRNA KCNMA1-AS1 was previously found to be implicated in the progression and migration of epithelial ovarian cancer by promoting proliferation, migration and inhibiting apoptosis [53]. Some other lncRNAs also exhibit a concerning duality - promoting osteoblast differentiation while also fueling cancer progression. Taking an example of LncRNA HOTTIP, it was found to enhance human osteogenic BMSCs differentiation via the activation of the Wnt/β-catenin signalling pathway [42], the activation of which induces enhanced epithelial-mesenchymal transition in cancer metastasis [54]. LncRNA HOTTIP also promoted pancreatic cancer cell proliferation, survival and migration [55]. Taking another example, LncRNA SNHG5 was shown to promote the osteogenic differentiation of BMSCs via the miR-212-3p/GDF5/SMAD pathway. SMAD signaling collaborates with oncogenic pathways like Wnt, NF-kB, and Notch to fuel epithelial-to-mesenchymal transition (EMT) and invasion. LncRNA SNHG5 was also found to play a tumor-promoting role in many cancer types including nasopharyngeal carcinoma [56], hepatocellular carcinoma [57], and cervical cancer [58]. The complex, context-specific functionality of these lncRNAs presents challenges for exploiting their bone regenerative potential while avoiding cancer promotion. There are a few key considerations when assessing whether lncRNAs implicated in both osteogenic differentiation and cancer progression could realistically be used as therapy for bone defects. The complexity of lncRNAs functioning divergently in different cell contexts, their specific roles in driving cancer progression, challenges with targeted delivery, and safety risks of potentially promoting tumors all pose significant barriers to clinical use. Innovative delivery methods, genetic screening, combination therapies with other bone anabolics, short-term treatment and extensive preclinical testing in bone defects models may help mitigate concerns, but far more research is needed. Close collaboration between bone biology and oncology experts will be critical to determine if any lncRNAs implicated in cancer may realistically be harnessed safely and effectively for bone defect therapies. Overall, the cancer risk poses a significant hurdle, and extensive further study is required before this novel genetic targeting approach could be clinically viable.
In summary, for the first time, we have reported here that lncRNA KCNMA1-AS1 has augmented expression during osteogenic differentiation. The overexpression of KCNMA1-AS1 results in enhanced hBMSCs’ osteogenic differentiation. Our research revealed a novel mechanism where the KCNMA1-AS1 was found to regulate osteogenic differentiation of hBMSCs via the SMAD9 signaling pathway (Fig. 7), which provides a novel perspective on translational research in bone tissue engineering.
Data Availability
The data used and analyzed during the current study are available from the corresponding author upon reasonable request.
References
Tissue-engineered autologous. grafts for facial bone reconstruction | Science Translational Medicine [Internet]. [cited 2023 Sep 14]. Available from: https://www.science.org/doi/abs/10.1126/scitranslmed.aad5904.
Kinoshita Y, Maeda H. Recent developments of functional scaffolds for craniomaxillofacial bone tissue engineering applications. Sci World J. 2013;2013.
Cells | Free Full-Text. | Craniofacial Bone Tissue Engineering: Current Approaches and Potential Therapy [Internet]. [cited 2023 Sep 14]. Available from: https://www.mdpi.com/2073-4409/10/11/2993.
Grayson WL, Bunnell BA, Martin E, Frazier T, Hung BP, Gimble JM. Stromal cells and stem cells in clinical bone regeneration. Nat Rev Endocrinol. 2015;11(3):140–50.
Park JJ, Rochlin DH, Parsaei Y, Shetye PR, Witek L, Leucht P et al. Bone tissue Engineering Strategies for Alveolar Cleft: review of preclinical results and guidelines for Future studies. Cleft Palate Craniofac J. 2022;10556656221104954.
Kawecki F, Clafshenkel WP, Fortin M, Auger FA, Fradette J. Biomimetic tissue-engineered bone substitutes for maxillofacial and craniofacial repair: the potential of cell sheet technologies. Adv Healthc Mater. 2018;7(6):1700919.
Valtanen RS, Yang YP, Gurtner GC, Maloney WJ, Lowenberg DW. Synthetic and bone tissue engineering graft substitutes: what is the future? Injury. 2021;52:72–7.
Wan Z, Zhang P, Liu Y, Lv L, Zhou Y. Four-dimensional bioprinting: current developments and applications in bone tissue engineering. Acta Biomater. 2020;101:26–42.
Laird NZ, Acri TM, Tingle K, Salem AK. Gene-and RNAi-activated scaffolds for bone tissue engineering: current progress and future directions. Adv Drug Deliv Rev. 2021;174:613–27.
Maia FR, Bastos AR, Oliveira JM, Correlo VM, Reis RL. Recent approaches towards bone tissue engineering. Bone. 2022;154:116256.
Gu J, Wang B, Wang T, Zhang N, Liu H, Gui J et al. Effects of cartilage progenitor cells, bone marrow mesenchymal stem cells and chondrocytes on cartilage repair as seed cells: an in vitro study. Drug Des Devel Ther. 2022;1217–30.
Yoon JY, Mandakhbayar N, Hyun J, Yoon DS, Patel KD, Kang K, et al. Chemically-induced osteogenic cells for bone tissue engineering and Disease modeling. Biomaterials. 2022;289:121792.
Zha K, Tian Y, Panayi AC, Mi B, Liu G. Recent advances in enhancement strategies for osteogenic differentiation of mesenchymal stem cells in bone tissue engineering. Front Cell Dev Biol. 2022;10:824812.
Fakhry M, Hamade E, Badran B, Buchet R, Magne D. Molecular mechanisms of mesenchymal stem cell differentiation towards osteoblasts. World J Stem Cells. 2013;5(4):136.
Salhotra A, Shah HN, Levi B, Longaker MT. Mechanisms of bone development and repair. Nat Rev Mol Cell Biol. 2020;21(11):696–711.
Yang Q, Jia L, Li X, Guo R, Huang Y, Zheng Y, et al. Long noncoding RNAs: new players in the osteogenic differentiation of bone marrow-and adipose-derived mesenchymal stem cells. Stem Cell Rev Rep. 2018;14:297–308.
Guo Q, Guo Q, **ao Y, Li C, Huang Y, Luo X. Regulation of bone marrow mesenchymal stem cell fate by long non-coding RNA. Bone. 2020;141:115617.
Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;136(4):629–41.
Wu H, Yang L, Chen LL. The diversity of long noncoding RNAs and their generation. Trends Genet. 2017;33(8):540–52.
Non-coding. RNA networks in cancer | Nature Reviews Cancer [Internet]. [cited 2023 Sep 14]. Available from: https://www.nature.com/articles/nrc.2017.99.
Qian X, Zhao J, Yeung PY, Zhang QC, Kwok CK. Revealing lncRNA structures and interactions by sequencing-based approaches. Trends Biochem Sci. 2019;44(1):33–52.
Ashrafizadeh M, Zarrabi A, Mostafavi E, Aref AR, Sethi G, Wang L, et al. Non-coding RNA-based regulation of inflammation. Semin Immunol. 2022;59:101606.
Chew CL, Conos SA, Unal B, Tergaonkar V. Noncoding RNAs: master regulators of inflammatory signaling. Trends Mol Med. 2018;24(1):66–84.
NAIL. : an evolutionarily conserved lncRNA essential for licensing coordinated activation of p38 and NFκB in colitis | Gut [Internet]. [cited 2023 Sep 14]. Available from: https://gut.bmj.com/content/70/10/1857.abstract.
Ma Z, Wang YY, **n HW, Wang L, Arfuso F, Dharmarajan A, et al. The expanding roles of long non-coding RNAs in the regulation of cancer stem cells. Int J Biochem Cell Biol. 2019;108:17–20.
Silva AM, Moura SR, Teixeira JH, Barbosa MA, Santos SG, Almeida MI. Long noncoding RNAs: a missing link in osteoporosis. Bone Res. 2019;7(1):10.
Zhang N, Hu X, He S, Ding W, Wang F, Zhao Y, et al. LncRNA MSC-AS1 promotes osteogenic differentiation and alleviates osteoporosis through sponging microRNA-140–5p to upregulate BMP2. Biochem Biophys Res Commun. 2019;519(4):790–6.
Tan YT, Lin JF, Li T, Li JJ, Xu RH, Ju HQ. LncRNA-mediated posttranslational modifications and reprogramming of energy metabolism in cancer. Cancer Commun. 2021;41(2):109–20.
Shi L, Yang Y, Li M, Li C, Zhou Z, Tang G, et al. LncRNA IFITM4P promotes immune Escape by up-regulating PD-L1 via dual mechanism in oral carcinogenesis. Mol Ther. 2022;30(4):1564–77.
Zhang H, Xu R, Li B, **n Z, Ling Z, Zhu W, et al. LncRNA NEAT1 controls the lineage fates of BMSCs during skeletal aging by impairing mitochondrial function and pluripotency maintenance. Cell Death Differ. 2022;29(2):351–65.
Lanzillotti C, De Mattei M, Mazziotta C, Taraballi F, Rotondo JC, Tognon M, et al. Long non-coding RNAs and microRNAs interplay in osteogenic differentiation of mesenchymal stem cells. Front Cell Dev Biol. 2021;9:646032.
Liu J, Yao Y, Huang J, Sun H, Pu Y, Tian M, et al. Comprehensive analysis of lncRNA-miRNA-mRNA networks during osteogenic differentiation of bone marrow mesenchymal stem cells. BMC Genomics. 2022;23(1):425.
Long Y, Wang X, Youmans DT, Cech TR. How do lncRNAs regulate transcription? Sci Adv. 2017;3(9):eaao2110.
Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22(2):96–118.
Li G, Yun X, Ye K, Zhao H, An J, Zhang X, et al. Long non-coding RNA-H19 stimulates osteogenic differentiation of bone marrow mesenchymal stem cells via the microRNA-149/SDF-1 axis. J Cell Mol Med. 2020;24(9):4944–55.
Yin J, Zheng Z, Zeng X, Zhao Y, Ai Z, Yu M, et al. lncRNA MALAT1 mediates osteogenic differentiation of bone mesenchymal stem cells by sponging miR-129-5p. PeerJ. 2022;10:e13355.
Wang CG, Hu YH, Su SL, Zhong D. LncRNA DANCR and miR-320a suppressed osteogenic differentiation in osteoporosis by directly inhibiting the Wnt/β-catenin signaling pathway. Exp Mol Med. 2020;52(8):1310–25.
Weng W, Di S, **ng S, Sun Z, Shen Z, Dou X, et al. Long non-coding RNA DANCR modulates osteogenic differentiation by regulating the miR-1301-3p/PROX1 axis. Mol Cell Biochem. 2021;476:2503–12.
Chen Q, Wang M, Wu S. The lncRNA MCF2L-AS1 controls osteogenic differentiation by regulating miR-33a. Cell Cycle. 2020;19(9):1059–65.
Ju C, Liu R, Zhang YW, Zhang Y, Zhou R, Sun J, et al. Mesenchymal stem cell-associated lncRNA in osteogenic differentiation. Biomed Pharmacother. 2019;115:108912.
LNCcation. : lncRNA localization and function | Journal of Cell Biology | Rockefeller University Press [Internet]. [cited 2023 Sep 14]. Available from: https://rupress.org/jcb/article/220/2/e202009045/211695/LNCcation-lncRNA-localization-and-functionlncRNA.
LncRNA HOTTIP enhances human osteogenic. BMSCs differentiation via interaction with WDR5 and activation of Wnt/β-catenin signalling pathway - ScienceDirect [Internet]. [cited 2023 Sep 14]. Available from: https://www.sciencedirect.com/science/article/abs/pii/S0006291X20303004.
You D, Yang C, Huang J, Gong H, Yan M, Ni J. Long non-coding RNA MEG3 inhibits chondrogenic differentiation of synovium-derived mesenchymal stem cells by epigenetically inhibiting TRIB2 via methyltransferase EZH2. Cell Signal. 2019;63:109379.
**u B, Chi Y, Liu L, Chi W, Zhang Q, Chen J, et al. LINC02273 drives Breast cancer Metastasis by epigenetically increasing AGR2 transcription. Mol Cancer. 2019;18:1–20.
Derynck R, Zhang YE. Smad-dependent and smad-independent pathways in TGF-β family signalling. Nature. 2003;425(6958):577–84.
Massagué J, Seoane J, Wotton D. Smad transcription factors. Genes Dev. 2005;19(23):2783–810.
McDonald GL, Wang M, Hammond CL, Bergen DJ. Pharmacological manipulation of early zebrafish skeletal development shows an important role for Smad9 in control of skeletal progenitor populations. Biomolecules. 2021;11(2):277.
Song B, Estrada KD, Lyons KM. Smad signaling in skeletal development and regeneration. Cytokine Growth Factor Rev. 2009;20(5–6):379–88.
Lee JS, Kim ME, Seon JK, Kang JY, Yoon TR, Park YD, et al. Bone-forming peptide-3 induces osteogenic differentiation of bone marrow stromal cells via regulation of the ERK1/2 and Smad1/5/8 pathways. Stem Cell Res. 2018;26:28–35.
** L, Zhang YF, Zhao ZJ, Pan DS, Liang W. Prunella vulgaris L protects glucocorticoids-induced osteogenesis inhibition in bone marrow mesenchymal stem cells through activating the smad pathway. Eur Rev Med Pharmacol Sci. 2020;24(10):5691–6.
Yang W, Zhu W, Yang Y, Guo M, Qian H, Jiang W, et al. Exosomal mir-100-5p inhibits osteogenesis of hBMSCs and angiogenesis of HUVECs by suppressing the BMPR2/Smad1/5/9 signalling pathway. Stem Cell Res Ther. 2021;12:1–16.
Han Y, Yang Q, Huang Y, Jia L, Zheng Y, Li W. Long non-coding RNA SNHG5 promotes the osteogenic differentiation of bone marrow mesenchymal stem cells via the miR-212-3p/GDF5/SMAD pathway. Stem Cell Res Ther. 2022;13(1):1–20.
KCNMA1-AS. 1 attenuates apoptosis of epithelial Ovarian cancer cells and serves as a risk factor for poor prognosis of epithelial Ovarian cancer.
Fodde R, Brabletz T. Wnt/β-catenin signaling in cancer stemness and malignant behavior. Curr Opin Cell Biol. 2007;19(2):150–8.
The long non. -coding RNA HOTTIP enhances pancreatic cancer cell proliferation, survival and migration - PMC [Internet]. [cited 2023 Sep 14]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4484423/.
LncRNA SNHG5 promotes nasopharyngeal carcinoma. progression by regulating miR-1179/HMGB3 axis | BMC Cancer | Full Text [Internet]. [cited 2023 Sep 14]. Available from: https://bmccancer.biomedcentral.com/articles/https://doi.org/10.1186/s12885-020-6662-5.
LncRNA SNHG5 promotes the proliferation. and cancer stem cell-like properties of HCC by regulating UPF1 and Wnt-signaling pathway | Cancer Gene Therapy [Internet]. [cited 2023 Sep 14]. Available from: https://www.nature.com/articles/s41417-022-00456-3.
LncRNA SNHG5 promotes. cervical cancer progression by regulating the miR-132/SOX4 pathway: Autoimmunity: Vol 54, No 2 [Internet]. [cited 2023 Sep 14]. Available from: https://www.tandfonline.com/doi/abs/https://doi.org/10.1080/08916934.2020.1864731.
Acknowledgements
Nothing to declare.
Funding
We appreciate the research funding provided by the National Science Foundation of China (Grant No.: 81670950).
Author information
Authors and Affiliations
Contributions
“ZM took the lead in writing the manuscript and worked out almost all of the technical details. JL, XJ, WG, WW, and SL conceived the presented idea, planned and carried out the experiments, analyzed the data, as well as reviewed and edited the manuscript. SL, GS, HX, and JZ conceived the study, supervised the project, and were in charge of overall direction and planning. All authors provided critical feedback and helped shape the research, analysis and manuscript.“
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Mai, Z., Liu, J., Jiang, X. et al. Long noncoding RNA KCNMA1-AS1 promotes osteogenic differentiation of human bone marrow mesenchymal stem cells by activating the SMAD9 signaling pathway. Biol Direct 18, 81 (2023). https://doi.org/10.1186/s13062-023-00425-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13062-023-00425-2