Abstract
Background
As one of the most common malignancies, esophageal cancer has two subtypes, squamous cell carcinoma and adenocarcinoma, arising from distinct cells-of-origin. Distinguishing cell-type-specific molecular features from cancer-specific characteristics is challenging.
Results
We analyze whole-genome bisulfite sequencing data on 45 esophageal tumor and nonmalignant samples from both subtypes. We develop a novel sequence-aware method to identify large partially methylated domains (PMDs), revealing profound heterogeneity at both methylation level and genomic distribution of PMDs across tumor samples. We identify subtype-specific PMDs that are associated with repressive transcription, chromatin B compartments and high somatic mutation rate. While genomic locations of these PMDs are pre-established in normal cells, the degree of loss is significantly higher in tumors. We find that cell-type-specific deposition of H3K36me2 may underlie genomic distribution of PMDs. At a smaller genomic scale, both cell-type- and cancer-specific differentially methylated regions (DMRs) are identified for each subtype. Using binding motif analysis within these DMRs, we show that a cell-type-specific transcription factor HNF4A maintains the binding sites that it generates in normal cells, while establishing new binding sites cooperatively with novel partners such as FOSL1 in esophageal adenocarcinoma. Finally, leveraging pan-tissue single-cell and pan-cancer epigenomic datasets, we demonstrate that a substantial fraction of cell-type-specific PMDs and DMRs identified here in esophageal cancer are actually markers that co-occur in other cancers originating from related cell types.
Conclusions
These findings advance our understanding of DNA methylation dynamics at various genomic scales in normal and malignant states, providing novel mechanistic insights into cell-type- and cancer-specific epigenetic regulations.
Similar content being viewed by others
Background
Ranking seventh in cancer incidence and sixth in mortality worldwide, esophageal carcinoma is highly aggressive and its patients have poor outcomes, with a 5-year survival rate lower than 20% [1, 2]. Esophageal cancer comprises two major histologic subtypes: squamous cell carcinoma (ESCC) and adenocarcinoma (EAC). These two subtypes have distinct clinical characteristics. ESCC occurs predominantly in the upper and mid-esophagus; EAC is prevalent in the lower esophagus near the gastroesophageal junction (GEJ) and is associated with the precursor lesion known as Barrett’s esophagus (BE). Biologically, ESCC arises from the squamous epithelial cells and has common features with other squamous cell carcinomas (SCC), such as head and neck SCC (HNSCC). In comparison, EAC has columnar cell features and shares many characteristics with tubular gastrointestinal adenocarcinomas. In particular, EAC is almost indistinguishable from GEJ adenocarcinoma in terms of genomic, biological and clinical features.
Epigenetically, multiple studies have reported molecular changes in esophageal cancer, especially at the DNA methylation level [3,4,5,25]. We chose esophageal cancer as the disease model considering that the two subtypes are developed from distinct cell-of-origins, and we hypothesized that characterization of their methylome profiles might reveal cell-type- and cancer-specific methylation changes, together with underlying epigenetic mechanisms.
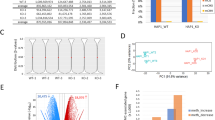
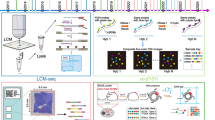
Identification of PMDs in esophageal samples by a sequence-aware multi-model PMD caller (MMSeekR). A A graphic model of the present study design. B Dot plots showing average methylation levels for all CpGs across the whole genome, CpGs within CGI promoters, common PMDs, SINE, LINE, and LTR in different samples. The annotations from Takai et al. [26] were used for CGI methylation quantification. C Development of a new PMD caller. The MethylSeekR α score measures the distribution of methylation levels in sliding windows with 201 consecutive CpGs across the genome. α score < 1 corresponds to a polarized distribution towards a high or low methylation level (that is, HMDs), while α score ≥ 1 corresponds to the distribution towards intermediate methylation levels (that is, PMDs). PCC shows the correlation between the predicted hypomethylation score based on a NN model, and the actual methylation level. A strong negative correlation indicates regions favoring PMDs, while weak/null correlation favors HMDs. D PCA analysis of 45 esophageal samples using the top 5000 most variable 30-kb tiles for the three PMD callers. E, F Representative windows showing PMDs successfully identified by MMSeekR but failed to be detected by either MethPipe (E) or MethylSeekR (F)
Results
Development of a novel sequence-aware calling method to identify PMDs
To characterize the esophageal cancer methylome, we analyzed WGBS profiles of 45 esophageal samples from two different cancer subtypes and their corresponding nonmalignant tissues [27] (Fig. 1A, Additional file 1: Fig. S1A). All of the nonmalignant esophageal squamous (NESQ) tissues showed high inter-sample correlation despite that they were from two different cohorts (Additional file 1: Fig. S1B and Additional file 2: Table S1). To analyze the overall methylation pattern, we first investigated the methylation level at various genomic domains (Fig. 1B). As anticipated, both global hypomethylation (especially in common PMDs, defined as shared PMDs identified from 40 different cancer types [19]), and CGI promoter hypermethylation were observed in tumor samples. EAC tumors harbored notably higher methylation levels in CGI promoters than ESCC tumors, in line with TCGA results showing that gastrointestinal adenocarcinoma had higher frequency of CGI hypermethylation than cancers from most other tissues [28]. Interestingly, most NGEJ tissues showed higher CGI promoter methylation levels than NESQ tissues, and usually even higher than ESCC tumor samples. Similar to EAC, BE samples (a recognized precursor lesion of EAC) were reported to have a hypermethylation pattern at CGI promoters [7]. Since our NGEJ tissues were pathologically confirmed as inflammatory tissues but devoid of apparent BE, this result suggests that CGI hypermethylation may occur in inflamed GEJ. Interestingly, CGI hypermethylation has been observed in long-term-cultured colon organoids and cells upon prolonged exposure to cigarette smoke extract [39]. B Different PMD categories were identified based on the frequency and overlap between the two esophageal cancer types. C Line plots showing average methylation levels for different PMD categories in esophageal tumors, where each line represents one sample. D Similar line plot patterns were observed using TCGA methylation datasets, showing the mean and standard deviation across samples. Each row in the heatmap below shows an individual sample. E Bar plots showing the percentage of WGBS PMDs overlap** with chromatin B compartments, which were predicted using TCGA methylation datasets and analyzed by minfi package. Methylation datasets in D and E are from the TCGA ESCA HM450k arrays, including 91 ESCC and 75 EAC samples. F Somatic mutation rates based on WGS in the indicated studies, calculated separately for each of the WGBS PMD categories. EAC WGS datasets: 276 samples; ESCC WGS datasets: 508 samples
We also correlated the methylation levels of subtype-specific PMDs to each of risk factors and clinicopathological parameters using HM450k datasets from the TCGA ESCA project. None of these factors, including age, smoking history, alcohol consumption, lymph node metastasis, and clinical stage, had significant impact on subtype-PMDs (Additional file 1: Fig. S3H). Another independent WGBS dataset (PRJNA523898, n = 42) again confirmed that there was no association between ESCC-specific PMDs with either age, clinical stage, or lymph node metastasis (Additional file 1: Fig. S3I).
At the transcription level, PMDs are reported to be less transcriptionally active than HMDs. We confirmed that subtype-specific PMDs were associated with low levels of gene expression specifically in the corresponding subtypes (Fig. 3A, B). To explore the biological implication of subtype-specific PMDs, we performed Cistrome-GO analysis using genes which were under-expressed in the subtype-specific PMD regions, finding that biological processes characteristic for the other subtype were enriched and repressed (Fig. 3C, D). Specifically, pathways of cornification, keratinocyte differentiation, and epidermis development, which are central to squamous cell differentiation and function, were enriched and inactive in EAC-specific PMDs (Fig. 3C). For example, many keratinocyte-specific genes were clustered within EAC-specific PMDs (Fig. 3E, left panel) and downregulated in EAC tumors (Fig. 3F). On the other hand, pathways important for gastrointestinal cell function, such as digestive system process, intestinal absorption, lipid metabolic process, and O − glycan processing, were enriched and suppressed in ESCC-specific PMDs (Fig. 3D). The right panel of Fig. 3E shows as an example that SLC2A2, which contributes to digestive system process and absorption, was located in ESCC-specific PMDs and downregulated in ESCC samples (Fig. 3F). These results suggest that subtype-specific PMDs contain inactive genes which are associated with cell-type-specific functions.
Subtype-specific PMDs control cell-type-specific functions. A, B In both EAC (A) and ESCC (B), genes covered by PMDs are expressed at lower levels than those in non-PMDs in a cancer-specific manner. C, D Cistrome-GO enrichment analyses using either EAC-specific (C) or ESCC-specific (D) PMDs and the downregulated genes within them. The top 15 most significant pathways are shown, and the number of genes enriched in each pathway is shown on the right. E Two representative genome windows showing the methylation profiles of EAC-specific (left panel) and ESCC-specific PMDs (right panel). CGI regions were masked using the annotation from Irizarry et al. [39]. F Volcano plots showing that genes residing within genome domains in E are downregulated in corresponding cancer subtypes. The differentially expressed genes were identified with the average expression level (FPKM) ≥ 0.1, adjusted p-value < 0.05 and absolute fold-change > 2. Expression RNA-seq in C, D, F are from the TCGA ESCA project, including 76 ESCC and 78 EAC samples
H3K36me2 is inversely associated with PMDs in a cell-type-specific manner
Both H3K36me2 and H3K36me3 were observed to recruit DNA methyltransferases (DNMT3A [40] and DNMT3B [41], respectively) to maintain DNA methylation levels in large chromatin domains. H3K36me3 is enriched in gene bodies of active transcripts, while H3K36me2 covers larger multi-gene domains. Indeed, we have previously shown that the deposition of H3K36me3 is inversely associated with PMD distribution [19]. Here, we further hypothesized that H3K36me2 also contributed to maintaining DNA methylation levels, and the histone modification by this mark might affect the genomic distribution of PMDs and HMDs. To test this, we performed H3K36me2 ChIP-seq in both EAC and ESCC cell lines. Indeed, shared HMDs (purple line) showed high H3K36me2 intensity in both cell types, while shared PMDs (yellow line) exhibited the lowest signals (Fig. 4A). EAC-specific PMDs (red line) had low H3K36me2 levels in EAC cells but high H3K36me2 levels in ESCC cells. The reciprocal pattern was observed in ESCC-specific PMDs (blue line). For example, H3K36me2 signals were undetectable in an EAC-specific PMD covering the loci of XR_945002.2 and XR_945004.2 in EAC cells, but were strong in ESCC (Fig. 4B, right panel). On the other hand, shared HMDs such as the one covering the VSP8 gene were decorated highly with H3K36me2 in both cell types (Fig. 4B, left panel).
The H3K36me2 mark is inversely associated with PMDs in a cell-type-specific manner. A Aggregation plots of H3K36me2 ChIP-seq levels in esophageal cancer cell lines across four different PMD categories: shared PMDs, EAC-specific PMDs, ESCC-specific PMDs, shared HMDs. B Representative genomic loci showing H3K36me2 signal from ChIP-seq, and subtype-specific PMDs from WGBS data. CGI regions were masked using the annotation from Irizarry et al. [39]. C Aggregation plots of H3K36me2 ChIP-seq levels in HNSCC cell lines across four different PMD categories. H3K36me2 ChIP-seq datasets were obtained from GSE149670. H3K36me2 signals in A, C were normalized by the CPM ratio of IP over Input in 5-kb windows
To further verify these results, we interrogated public H3K36me2 ChIP-seq data from HNSCC cell lines (squamous cancer highly similar to ESCC in terms of cell-of-origin and epigenome). Indeed, a similar pattern of H3K36me2 distribution to ESCC was observed in Cal27 and Det562 HNSCC cells. Specifically, both shared PMDs and ESCC-specific PMDs harbored low signals in HNSCC cell lines, while high H3K36me2 levels were found in HMDs and EAC-specific PMDs (Fig. 4C). However, FaDu appeared to be an outlier, showing invariably high levels across different regions (Fig. 4C), which warrants further investigation. Together, these results demonstrate a prominent depletion of H3K36me2 mark in PMDs in a cell-type-specific manner, which is likely owing to the finding that H3K36me2 promotes the maintenance of DNA methylation by recruiting DNMT3A.
Subtype-specific differentially methylated regions (DMRs) in esophageal cancer
We next sought to investigate differentially methylated regions (DMRs) at small genomic scales, given their direct roles in transcriptional regulation. However, our above results suggest an overwhelming, global effect of PMD hypomethylation in tumor samples, which can strongly affect the calling of focal DMRs. Indeed, PCA analysis of the most variable CpGs genome-wide revealed that PC1, the most significant component, was clearly driven by methylation loss at PMDs (Additional file 1: Fig. S4A).
To factor out the effect of PMD hypomethylation, we masked any PMD found within two-thirds of either EAC or ESCC samples (Additional file 1: Fig. S4B). We re-performed the PCA analysis, finding that the two cancer subtypes were completely separated by PC1, which was the most significant component and accounted for 42.2% of the total methylation variance (Additional file 1: Fig. S4C, left panel). In addition, nonmalignant and tumor samples were separated along PC2, and all NESQ samples were clustered closely together despite being generated from two different cohorts. Notably, this approach removed most correlation with the global methylation level (Additional file 1: Fig. S4C, right panel). Thus, it is critical to remove the effects of global hypomethylation when investigating cancer-associated methylation features outside PMDs.
We next identified DMRs between EAC and ESCC samples within the PMD-subtracted genome described above (~ 46.5% of the genome). Under the cutoff of q value < 0.05 and absolute delta methylation change > 0.2, a total of 7734 DMRs were hypomethylated in EAC and 5470 in ESCC (Fig. 5A). As expected, hypomethylated DMRs (hypoDMRs) had low average methylation levels in corresponding subtypes (Additional file 1: Fig. S4D-E). The majority of DMRs were about 1–2 kb long and located mostly in intronic and intergenic regions (Fig. 5B), similar to that of the random background (Additional file 1: Fig. S4F). To investigate the epigenomic characteristics of hypoDMRs, we systematically evaluated the chromatin accessibility at these regions, using the ATAC-seq data from the TCGA [42] and H3K27ac ChIP-seq data from previous studies [43,Methods” section