Abstract
Background
The biological function of lncRNA ELF3-AS1 remains largely unknown in cancers. The cause of SNAI2 overexpression in tumor metastasis remains largely unclear. The molecular mechanisms underlying the high co-expression of antisense lncRNAs and adjacent protein-coding genes remains unclear.
Methods
RNA-seq, CHIP and dual-luciferase reporter assay were performed to identify lncRNAs regulated by SNAI2. MicroRNA-seq and RNA-seq studies were conducted to reveal the biological function of ELF3-AS1 in GC. RNA pulldown and CHIRP assays were conducted to identify the protein that interacts with ELF3-AS1.
Results
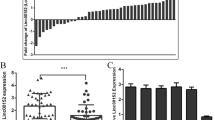
A total of 123 lncRNAs were identified to be regulated by SNAI2 in GC by RNA sequencing. The ELF3 gene and antisense lncRNA ELF3-AS1 were both transcriptionally repressed by SNAI2 or SNAI1. Down-regulation of ELF3-AS1 and ELF3 predicted poor prognosis in GC. Nuclear localized lncRNA ELF3-AS1 negatively regulated GC cell cycle progression via suppressing G1/S transition and histone synthesis. ELF3-AS1 mainly inhibited GC metastasis by repressing SNAI2 signaling. Additionally, ELF3-AS1 modulated ELF3 mRNA stability by RNA-RNA interaction. The RNA duplexes formed by ELF3 mRNA and lncRNA ELF3-AS1 directly interacted with the double-stranded RNA (dsRNA) binding protein complex ILF2/ILF3 (NF45/NF90). In turn, the ILF2/ILF3 complex dynamically regulated the expression of ELF3-AS1 and ELF3 by affecting the dsRNA stability.
Conclusions
The SNAI2-ELF3-AS1 feedback loop regulates ELF3 expression at transcriptional and post-transcriptional levels and drives gastric cancer metastasis by maintaining SNAI2 overexpression. The ILF2/ILF3 complex plays a critical role in regulating dsRNA stability. In addition, our work provides a direct evidence that head-to-head antisense lncRNAs can share promoters with neighboring coding genes, which make their expression subject to similar transcriptional regulation, leading to high co-expression.
Similar content being viewed by others
Background
Gastric cancer (GC) is the third most common cause of cancer-related deaths [1, 2]. Although treatments for GC have been greatly improved, the survival remains poor due to the inability to diagnose this cancer in early stage [3, 4]. The most common route of GC metastasis is lymph node metastasis, followed by peritoneal dissemination metastasis and liver metastasis [5, 6]. Approximately one-third of GC patients are diagnosed at an advanced stage with metastasis, and 4–14% have metastatic disease to the liver [7, 9B), suggesting ILF2/ILF3 complex may play a critical role in modulating dsRNA stability. Interestingly, a recent publication by Mahale and colleagues provided very similar evidence for the role of antisense RNA (IER3-AS1) and sense RNA (IER3 )[35]. Based on RNA-seq analysis, they found that activation of FGF2/FGFR signaling greatly enhanced the mRNA levels of IER3 and IER3-AS1. IER3 and IER3-AS1 formed RNA duplex and regulated their mRNA stability each other by interacting with HNRNPK protein [35].
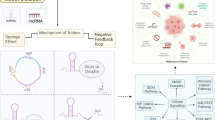
The working model of the SNAI2-ELF3-AS1 feedback loop. A ELF3-AS1 and ELF3 shared promoters and were transcriptionally repressed by SNAI1/2. B The molecular mechanisms underlying how ELF3-AS1 regulates ELF3 mRNA stability. C The molecular mechanisms of the biological role of ELF3-AS1 in GC. D The working model of SNAI2-ELF3-AS1 feedback loop in driving GC progression
The biological effects of transcriptional repression of ELF3-AS1 by SNAI2 remains unknown. SNAI2 is a rapid-turnover protein [36]. Recently, Kang et al. had reported that SNAI2 protein turnover was regulated by the ubiquitin-proteasome system (UPS) [12, 13]. However, in theory, the regulation of SNAI2 protein turnover should be not only at the (post-) translational level, but also at the (post-) transcriptional level. Herein, our finding strongly indicated that the SNAI2-repressed lncRNA ELF3-AS1 played an essential role in maintaining SNAI2 mRNA stability. Knockdown of ELF3-AS1 results in decreased expression levels of miRNAs targeting SNAI2, upregulation of SNAI2 mRNA and protein, and activation of downstream signaling of SNAI2. Additionally, the overall survival analysis based on the TCGA data showed that ELF3-AS1 and SNAI2 possessed opposite prognoses in pan-cancer (Figure S5). These findings highlighted that SNAI2 achieves self-overexpression by transcriptionally repressing ELF3-AS1. Once SNAI2 is overexpressed, it can transcriptionally repress ELF3-AS1 expression, thereby maintaining self-overexpression state in tumor metastasis. On the other hand, the downregulation of lncRNA ELF3-AS1 promoted GC cell proliferation by accelerating the G1/S transition and increasing histone-coding gene expression (Fig 9C).
Conclusions
In summary, a novel double-negative feedback loop between SNAI2 and lncRNA ELF3-AS1 was identified in GC. The SNAI2-ELF3-AS1 feedback loop drives GC metastasis by continuously activating SNAI2 signaling and regulating ELF3 expression at transcriptional and post-transcriptional levels (Fig 9D). In GC, SNAI2 was overexpressed, resulting in decreased expression level of ELF3 and ELF3-AS1. In turn, ELF3-AS1 downregulation further drives tumor progression by continuously activating SNAI2 signaling and promoting cell proliferation, thereby leading to a poor prognosis in GC (Fig 9D).
Availability of data and materials
The datasets supporting the conclusions of this article are included within the article and its additional files.
Abbreviations
- GC:
-
Gastric cancer
- TCGA:
-
The Cancer Genome Atlas
- UTR:
-
Untranslated region
- EMT:
-
epithelial-mesenchymal transition
- TPM:
-
transcripts per million
- qRT-PCR:
-
quantitative reverse transcription Polymerase Chain Reaction
- GEO:
-
Gene Expression Omnibus
- FPKM:
-
Fragments Per Kilobase of exon model per Million mapped fragments
- STAD:
-
Stomach adenocarcinoma
- HNSC:
-
Head and Neck squamous cell carcinoma
- KIRC:
-
Kidney renal clear cell carcinoma
- ESCA:
-
Esophageal carcinoma
- READ:
-
Rectum adenocarcinoma
- LIHC:
-
Liver hepatocellular carcinoma
- LUSC:
-
Lung squamous cell carcinoma
- CHIRP:
-
Chromatin Isolation by RNA Purification
- dsRNA:
-
Double-stranded RNA
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2021. Ca-Cancer J Clin. 2021;71(1):7–33.
Sakai H, Kawakami H, Teramura T, Onodera Y, Somers E, Furuuchi K, et al. Folate receptor alpha increases chemotherapy resistance through stabilizing MDM2 in cooperation with PHB2 that is overcome by MORAb-202 in gastric cancer. Clin Transl Med. 2021;11(6)e454.
Wang XH, Jiang ZH, Yang HM, Zhang Y, Xu LH. Hypoxia-induced FOXO4/LDHA axis modulates gastric cancer cell glycolysis and progression. Clin Transl Med. 2021;11(1):e279.
Wang J, Zhang M, Hu X, She J, Sun R, Qin S, et al. miRNA-194 predicts favorable prognosis in gastric cancer and inhibits gastric cancer cell growth by targeting CCND1. FEBS Open Bio. 2021;11(7):1814–26.
Van Cutsem E, Sagaert X, Topal B, Haustermans K, Prenen H. Gastric cancer. Lancet. 2016;388(10060):2654–64.
Gotoda T, Yanagisawa A, Sasako M, Ono H, Nakanishi Y, Shimoda T, et al. Incidence of lymph node metastasis from early gastric cancer: estimation with a large number of cases at two large centers. Gastric Cancer. 2000;3(4):219–25.
Mervic L. Time course and pattern of metastasis of cutaneous melanoma differ between men and women. Plos One. 2012;7(3):e32955.
Chen Q, Ge X, Zhang Y, **a H, Yuan D, Tang Q, et al. Plasma miR-122 and miR-192 as potential novel biomarkers for the early detection of distant metastasis of gastric cancer. Oncol Rep. 2014;31(4):1863–70.
Wang J, Zhang M, Hu X, She J, Sun R, Qin S, et al. MiRNA-194 predicts favorable prognosis in gastric cancer and inhibits gastric cancer cell growth by targeting CCND1: FEBS Open Bio; 2021.
Li D, Cheng P, Wang J, Qiu X, Zhang X, Xu L, et al. IRF6 is directly regulated by ZEB1 and ELF3, and predicts a favorable prognosis in gastric cancer. Front Oncol. 2019;9:220.
Qin S, Wang Z, Huang C, Huang P, Li D. Serine protease PRSS23 drives gastric cancer by enhancing tumor associated macrophage infiltration via FGF2. Front Immunol. 2022;13.
Fan HJ, Wang XX, Li WY, Shen MH, Wei Y, Zheng HQ, et al. ASB13 inhibits breast cancer metastasis through promoting SNAI2 degradation and relieving its transcriptional repression of YAP. Genes Dev. 2020;34(19-20):1359–72.
Li WY, Shen MH, Jiang YZ, Zhang RN, Zheng HQ, Wei Y, et al. Deubiquitinase USP20 promotes breast cancer metastasis by stabilizing SNAI2. Genes Dev. 2020;34(19-20):1310–5.
Du F, Li XW, Feng WB, Qiao CY, Chen J, Jiang MZ, et al. SOX13 promotes colorectal cancer metastasis by transactivating SNAI2 and c-MET. Oncogene. 2020;39(17):3522–40.
Olmeda D, Montes A, Moreno-Bueno G, Flores JM, Portillo F, Cano A. Snai1 and Snai2 collaborate on tumor growth and metastasis properties of mouse skin carcinoma cell lines. Oncogene. 2008;27(34):4690–701.
Fan L, Lei H, Zhang S, Peng Y, Fu C, Shu G, et al. Non-canonical signaling pathway of SNAI2 induces EMT in ovarian cancer cells by suppressing miR-222-3p transcription and upregulating PDCD10. Theranostics. 2020;10(13):5895.
Peng L, Fu J, Chen Y, Ming Y, He H, Zeng S, et al. Transcription factor SNAI2 exerts pro-tumorigenic effects on glioma stem cells via PHLPP2-mediated Akt pathway. Cell Death Dis. 2022;13(6):1–9.
Li DD, Wang JJ, Zhang MX, Hu XH, She JJ, Qiu XM, et al. LncRNA MAGI2-AS3 Is Regulated by BRD4 and Promotes Gastric Cancer Progression via Maintaining ZEB1 Overexpression by Sponging miR-141/200a. Mol Ther-Nucl Acids. 2020;19:109–23.
Li D, Xu M, Wang Z, Huang P, Huang C, Chen Z, et al. The EMT-induced lncRNA NR2F1-AS1 positively modulates NR2F1 expression and drives gastric cancer via miR-29a-3p/VAMP7 axis. Cell Death Dis. 2022;13(1):1–10.
Li D, She J, Hu X, Zhang M, Sun R, Qin S. The ELF3-regulated lncRNA UBE2CP3 is over-stabilized by RNA–RNA interactions and drives gastric cancer metastasis via miR-138-5p/ITGA2 axis. Oncogene. 2021;40(35):5403–15.
Davis MC, Kesthely CA, Franklin EA, MacLellan SR. The essential activities of the bacterial sigma factor. Can J Microbiol. 2017;63(2):89–99.
Ali MM, Akhade VS, Kosalai ST, Subhash S, Statello L, Meryet-Figuiere M, et al. PAN-cancer analysis of S-phase enriched lncRNAs identifies oncogenic drivers and biomarkers. Nat Commun. 2018;9(1):883.
Statello L, Ali MM, Reischl S, Mahale S, Kosalai ST, Huarte M, et al. The DNA damage inducible lncRNA SCAT7 regulates genomic integrity and topoisomerase 1 turnover in lung adenocarcinoma. 2021;3(1):zcab002.
Zhang Z, Nong L, Chen M-L, Gu X-L, Zhao W-W, Liu M-H, et al. LncRNA ELF3-AS1 Promotes Non-Small Cell Lung Cancer Cell Invasion and Migration by Downregulating miR-212. Cancer Biother Radiopharm. 2022;37(2):119–24.
Luo S, Lu JY, Liu L, Yin Y, Chen C, Han X, et al. Divergent lncRNAs Regulate Gene Expression and Lineage Differentiation in Pluripotent Cells. Cell Stem Cell. 2016;18(5):637–52.
Chen LL. Linking Long Noncoding RNA Localization and Function. Trends Biochem Sci. 2016;41(9):761–72.
Satyanarayana A, Hilton MB, Kaldis P. p21 inhibits Cdk1 in the absence of Cdk2 to maintain the G1/S phase DNA damage checkpoint. Mol Biol Cell. 2008;19(1):65–77.
Fiaschi-Taesch NM, Salim F, Kleinberger J, Troxell R, Cozar-Castellano I, Selk K, et al. Induction of Human beta-Cell Proliferation and Engraftment Using a Single G1/S Regulatory Molecule, cdk6. Diabetes. 2010;59(8):1926–36.
Wen X, Liu X, Mao YP, Yang XJ, Wang YQ, Zhang PP, et al. Long non-coding RNA DANCR stabilizes HIF-1alpha and promotes metastasis by interacting with NF90/NF45 complex in nasopharyngeal carcinoma. Theranostics. 2018;8(20):5676–89.
Guan DY, Altan-Bonnet N, Parrott AM, Arrigo CJ, Li Q, Khaleduzzaman M, et al. Nuclear factor 45 (NF45) is a regulatory subunit of complexes with NF90/110 involved in mitotic control. Mol Cell Biol. 2008;28(14):4629–41.
Oh SC, Sohn BH, Cheong JH, Kim SB, Lee JE, Park KC, et al. Clinical and genomic landscape of gastric cancer with a mesenchymal phenotype. Nat Commun. 2018;9(1):1777.
Cristescu R, Lee J, Nebozhyn M, Kim KM, Ting JC, Wong SS, et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat Med. 2015;21(5):449–56.
Bartl J, Zanini M, Bernardi F, Forget A, Blümel L, Talbot J, et al. The HHIP-AS1 lncRNA promotes tumorigenicity through stabilization of dynein complex 1 in human SHH-driven tumors. Nat Commun. 2022;13(1):1–15.
Bryzghalov O, Szczesniak MW, Makalowska I. Retroposition as a source of antisense long non-coding RNAs with possible regulatory functions. Acta Biochim Pol. 2016;63(4):825–33.
Mahale S, Setia M, Prajapati B, Subhash S, Yadav MP, Thankaswamy Kosalai S, et al. HnRNPK maintains single strand RNA through controlling double-strand RNA in mammalian cells. Nat Commun. 2022;13(1):1–20.
Zheng H, Kang Y. Multilayer control of the EMT master regulators. Oncogene. 2014;33(14):1755–63.
Acknowledgements
We are very grateful to Dr. Jiwei Li and Dr. Zhaohui Ji (Lifegenes Biotechnology, Shanghai, China) for contributing to the RNA-Seq analysis. We are very grateful to Dr. Guoxi Huang (BioNanoSearching Biotechnology, Guangzhou, China) for contributing to RNA pulldown and RIP analysis.
Funding
This study was supported by grants from the National Natural Science Foundation of China (82203829, 82273451 and 81802375); Hubei Provincial Natural Science Foundation (2022CFB for DL and SQ), the Faculty Development Grants from Hubei University of Medicine (2020QDJZR024 to CH and 2020QDJZR012 to PH) and the Grants of Open-Ended Design Project from Hubei Key Laboratory of Embryonic Stem Cell Research (no. 2021ESOF021) and the Advantages Discipline Group (Medicine) Project in Higher Education of Hubei Province (2022XKQT2).
Author information
Authors and Affiliations
Contributions
QS conceived and designed the study. QS and LD wrote the paper. LD and SL performed most of the experiments. ZX, HP, HC and CZ carried out initial data analyses and performed partial of the experiments. All authors contributed to drafting the manuscript. All authors have read and approved the final submitted manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not available.
Consent for publication
All authors agreed on the manuscript.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Li, D., Shen, L., Zhang, X. et al. LncRNA ELF3-AS1 inhibits gastric cancer by forming a negative feedback loop with SNAI2 and regulates ELF3 mRNA stability via interacting with ILF2/ILF3 complex. J Exp Clin Cancer Res 41, 332 (2022). https://doi.org/10.1186/s13046-022-02541-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13046-022-02541-9