Abstract
Background
This study aimed to develop a radiogenomic prognostic prediction model for colorectal cancer (CRC) by investigating the biological and clinical relevance of intratumoural heterogeneity.
Methods
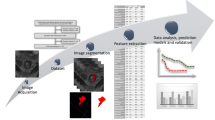
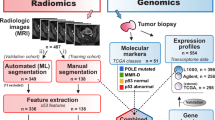
This retrospective multi-cohort study was conducted in three steps. First, we identified genomic subclones using unsupervised deconvolution analysis. Second, we established radiogenomic signatures to link radiomic features with prognostic subclone compositions in an independent radiogenomic dataset containing matched imaging and gene expression data. Finally, the prognostic value of the identified radiogenomic signatures was validated using two testing datasets containing imaging and survival information collected from separate medical centres.
Results
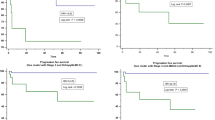
This multi-institutional retrospective study included 1601 patients (714 females and 887 males; mean age, 65 years ± 14 [standard deviation]) with CRC from 5 datasets. Molecular heterogeneity was identified using unsupervised deconvolution analysis of gene expression data. The relative prevalence of the two subclones associated with cell cycle and extracellular matrix pathways identified patients with significantly different survival outcomes. A radiogenomic signature-based predictive model significantly stratified patients into high- and low-risk groups with disparate disease-free survival (HR = 1.74, P = 0.003). Radiogenomic signatures were revealed as an independent predictive factor for CRC by multivariable analysis (HR = 1.59, 95% CI:1.03–2.45, P = 0.034). Functional analysis demonstrated that the 11 radiogenomic signatures were predominantly associated with extracellular matrix and immune-related pathways.
Conclusions
The identified radiogenomic signatures might be a surrogate for genomic signatures and could complement the current prognostic strategies.
Similar content being viewed by others
Background
Colorectal cancer (CRC) is the third leading cause of cancer-related deaths worldwide. Despite recent advancements in therapeutic techniques, the 5-year overall survival (OS) for this malignancy is only approximately 50% [1]. Therefore, there is an urgent need to develop prognostic biomarkers for improving CRC treatment. Substantial research has demonstrated that CRC is a heterogeneous disease with distinct molecular features and clinical responses [2,3,4]. An accurate understanding of the biological properties of CRC heterogeneity is essential for precise treatment, prediction of clinical prognosis, and the development of molecular subtype-specific targeted drugs.
Intratumour heterogeneity (ITH) is a hallmark of cancer that drives tumour evolution and disease progression. Increased ITH has been linked to a higher chance of recurrence, regardless of cancer type or treatment [5]. Therefore, exploration of ITH is helpful for the development of accurate prognostic tools. Previous studies have shown that the ITH of CRC can be characterised by massive parallel sequencing data [6,7,8]. Recent studies on CRC subtypes have employed unsupervised clustering to classify whole-genome expression profiles derived from bulk tumours. This unsupervised method has been effectively applied to a number of malignancies [9] but is less effective for mixtures with unknown compositions and noise. The deconvolution approach is an alternative unsupervised method that can estimate the underlying subclones of genomics in complex tissues to better understand tumour heterogeneity and predict prognosis [10].
Numerous studies on gene signature biomarkers have been published because of the advent of sequencing technology. However, their clinical applications are relatively limited. Current gene expression profiling methods are expensive, time-consuming, invasive, and require tumour biopsies for tissue extraction. Therefore, it was unavailable for all the patients. In contrast, radiomic biomarkers do not incur any additional expenses, because medical imaging is a routine part of the clinical decision-making process. Unlike biopsies, medical imaging is noninvasive and can provide information about the entire tumour phenotype, including ITH. Multiple studies have reported an association between radiomic characteristics and underlying gene expression patterns.
Radiogenomics explores the association between radiomic features and genomic characteristics, with the aim of revealing relevant features that reflect the underlying biological functions most related to clinical phenotypes. Numerous studies have established the viability of radiogenomics for identifying intrinsic molecular subtypes and gene expression profiles in cancers such as ovarian cancer [5B, C). A robust predictive model was constructed by establishing a link between genomics and radiomics. In the process of applying the model, only image data is required in the absence of genomic data, which dramatically lowers the threshold for clinical application of the model. Currently, imaging examinations are routinely used for tumour diagnosis and therapy decisions. Utilising images as input data for prognostic prediction models does not significantly increase healthcare expenditure. Furthermore, imaging examinations are noninvasive and can be repeated at various times. CT-based radiogenomic signatures allow us to forecast patient prognosis and ITH prior to surgery.
Owing to the construction of a link between genomics and radiomics, the model is substantially more interpretable. Imaging characteristics have been related to CRC outcomes, such as treatment response, lymph node metastasis, local recurrence, and survival [28,29,30], but their biological underpinnings remain unclear. In the present study, we did not introduce relevant prior information but identified four CRC genomic subclones by analysing a large number of gene expression profiles using a fully unsupervised deconvolution strategy. According to our study, tumours with a low proportion of cell cycle subclones and a high proportion of extracellular matrix subclones were associated with a shorter survival rate. Among the signalling pathways within the cell cycle subclone, the G1/S transition and cell cycle checkpoint pathways likely reflect the DNA damage response and can be exploited for prognosis [31]. Cell cycle checkpoints can repair DNA and prevent further damage by detecting damaged DNA and temporarily halting the cell cycle progression. Cell cycle dysregulation can lead to abnormal cell proliferation and apoptosis, and is responsible for tumourigenesis. Defects in cell cycle checkpoints may be a cause of genomic instability in tumours [32]. Therefore, abnormalities in cell cycle pathways have prognostic significance in CRC. The ECM subclone is another subclone strongly associated with prognosis. It is reported that ECM remodelling is associated with CRC carcinogenesis and progression [33, 34]. As a major component of the tumour microenvironment, the ECM plays a crucial role in tumour progression and treatment response. Chakravarthy et al. built a signature that linked extracellular matrix genes to immune evasion and immunotherapy failure [35]. Eleven radiomic characteristics were chosen for our model, the majority of which were enriched in ECM- and immune-related pathways, which are well-known prognosis-related pathways. This suggests that the prognostic value of these radiomic signatures has a biological foundation. These morphological textures and spatial features are inseparable from the gene- and cell-level characteristics. Machine learning helps us better understand the biology behind these morphological textures and spatial features. Using this three-step methodology, we created a prognostic prediction model that provides an entry point for elucidating the underlying molecular mechanisms.
Our study had several limitations. First, this was a retrospective study, which led to inevitable disadvantages. In follow-up research, these findings should be validated by prospective studies to reduce the bias caused by uncontrollable factors in retrospective studies. Second, our genomic development dataset and corresponding testing dataset were obtained from public databases. However, cohorts 3 and 4 came from local medical centres and provided in-house data. CT scans from different machines at different centres better validate the robustness and clinical usability of the model. Third, the regions of interest are manually annotated, and this process is time-consuming and tedious. We are currently investigating more robust semi-automatic annotation methods [36] to address this issue.
Conclusions
In conclusion, we conducted an integrative analysis of genomics and radiomics to dissect ITH and build models for predicting the prognosis of patients with CRC. The unsupervised deconvolution method for genomic subclone identification provides a new perspective for exploring tumour heterogeneity. Radiogenomic signatures can be independent prognostic biomarkers and may serve as surrogates for genomic signatures. This integrative analysis of the radiogenomic strategy shows great promise for understanding ITH, and can be extended to other cancers to help patients who might benefit from precise clinical treatment.
Availability of data and materials
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- CRC:
-
Colorectal cancer
- ITH:
-
Intratumor heterogeneity
- OS:
-
Overall survival
- GSEA:
-
Gene set enrichment analysis
- SYSU6H:
-
The Sixth Affiliated Hospital of Sun Yat-sen University
- KSH:
-
The First People's Hospital of Kashi Prefecture
- DFS:
-
Disease-free survival
- ECM:
-
Extracellular Matrix
References
Siegel RL, Miller KD, Goding Sauer A, Fedewa SA, Butterly LF, Anderson JC, Cercek A, Smith RA, Jemal A. Colorectal cancer statistics, 2020. CA Cancer J Clin. 2020;70:145–64.
Roepman P, Schlicker A, Tabernero J, Majewski I, Tian S, Moreno V, Snel MH, Chresta CM, Rosenberg R, Nitsche U, et al. Colorectal cancer intrinsic subtypes predict chemotherapy benefit, deficient mismatch repair and epithelial-to-mesenchymal transition. Int J Cancer. 2014;134:552–62.
Guinney J, Dienstmann R, Wang X, de Reyniès A, Schlicker A, Soneson C, Marisa L, Roepman P, Nyamundanda G, Angelino P, et al. The consensus molecular subtypes of colorectal cancer. Nat Med. 2015;21:1350–6.
Schlicker A, Beran G, Chresta CM, McWalter G, Pritchard A, Weston S, Runswick S, Davenport S, Heathcote K, Castro DA, et al. Subtypes of primary colorectal tumors correlate with response to targeted treatment in colorectal cell lines. BMC Med Genomics. 2012;5:66.
Li Z, Seehawer M, Polyak K. Untangling the web of intratumour heterogeneity. Nat Cell Biol. 2022;24:1192–201.
Budinska E, Popovici V, Tejpar S, D’Ario G, Lapique N, Sikora KO, Di Narzo AF, Yan P, Hodgson JG, Weinrich S. Gene expression patterns unveil a new level of molecular heterogeneity in colorectal cancer. J Pathol. 2013;231:63–76.
Sadanandam A, Lyssiotis CA, Homicsko K, Collisson EA, Gibb WJ, Wullschleger S, Ostos LCG, Lannon WA, Grotzinger C, Del Rio M. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat Med. 2013;19:619–25.
Marisa L, de Reynies A, Duval A, Selves J, Gaub MP, Vescovo L, Etienne-Grimaldi MC, Schiappa R, Guenot D, Ayadi M, et al. Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLoS Med. 2013;10: e1001453.
Wang X, Markowetz F. Dissecting cancer heterogeneity–an unsupervised classification approach. Int J Biochem Cell Biol. 2013;45:2574–9.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12:453–7.
Martins FC, de Santiago I, Trinh A, **an J, Guo A, Sayal K, Jimenez-Linan M, Deen S, Driver K, Mack M. Combined image and genomic analysis of high-grade serous ovarian cancer reveals PTEN loss as a common driver event and prognostic classifier. Genome Biol. 2014;15:1–15.
Cooper LA, Kong J, Gutman DA, Dunn WD, Nalisnik M, Brat DJ. Novel genotype-phenotype associations in human cancers enabled by advanced molecular platforms and computational analysis of whole slide images. Lab Invest. 2015;95:366–76.
Fan M, **a P, Clarke R, Wang Y, Li L. Radiogenomic signatures reveal multiscale intratumour heterogeneity associated with biological functions and survival in breast cancer. Nat Commun. 2020;11:4861.
Wu J, Cui Y, Sun X, Cao G, Li B, Ikeda DM, Kurian AW, Li R. Unsupervised clustering of quantitative image phenotypes reveals breast cancer subtypes with distinct prognoses and molecular pathways. Clin Cancer Res. 2017;23:3334–42.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci. 2005;102:15545–50.
Smith JJ, Deane NG, Wu F, Merchant NB, Zhang B, Jiang A, Lu P, Johnson JC, Schmidt C, Bailey CE. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology. 2010;138:958–68.
Chen L, Wu C-T, Wang N, Herrington DM, Clarke R, Wang Y. debCAM: a bioconductor R package for fully unsupervised deconvolution of complex tissues. Bioinformatics. 2020;36:3927–9.
Monti S, Tamayo P, Mesirov J, Golub T. Consensus clustering: a resampling-based method for class discovery and visualization of gene expression microarray data. Mach Learn. 2003;52:91–118.
Wang K, Duan X, Gao F, Wang W, Liu L, Wang X. Dissecting cancer heterogeneity based on dimension reduction of transcriptomic profiles using extreme learning machines. PLoS ONE. 2018;13: e0203824.
Kassambara A, Kosinski M, Biecek P, Fabian S: Package ‘survminer’. Drawing Survival Curves using ‘ggplot2’(R package version 03 1) 2017.
KeK T. Cell-cycle-dependent regulation of DNA replication and its relevance to cancer pathology. J Pathol. 2005;205:123–9.
Henke E, Nandigama R, Ergün S. Extracellular matrix in the tumor microenvironment and its impact on cancer therapy. Front Mol Biosci. 2020;6:160.
Gao F, Wang W, Tan M, Zhu L, Zhang Y, Fessler E, Vermeulen L, Wang X. DeepCC: a novel deep learning-based framework for cancer molecular subtype classification. Oncogenesis. 2019;8:1–12.
Vitale I, Shema E, Loi S, Galluzzi L. Intratumoral heterogeneity in cancer progression and response to immunotherapy. Nat Med. 2021;27:212–24.
Miranda A, Hamilton PT, Zhang AW, Pattnaik S, Becht E, Mezheyeuski A, Bruun J, Micke P, de Reynies A, Nelson BH. Cancer stemness, intratumoral heterogeneity, and immune response across cancers. Proc Natl Acad Sci U S A. 2019;116:9020–9.
Li H, Courtois ET, Sengupta D, Tan Y, Chen KH, Goh JJL, Kong SL, Chua C, Hon LK, Tan WS, et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat Genet. 2017;49:708–18.
Kyrochristos ID, Roukos DH. Comprehensive intra-individual genomic and transcriptional heterogeneity: Evidence-based Colorectal Cancer Precision Medicine. Cancer Treat Rev. 2019;80:101894.
Huang YQ, Liang CH, He L, Tian J, Liang CS, Chen X, Ma ZL, Liu ZY. Development and validation of a radiomics nomogram for preoperative prediction of lymph node metastasis in colorectal cancer. J Clin Oncol. 2016;34:2157–64.
Lovinfosse P, Polus M, Van Daele D, Martinive P, Daenen F, Hatt M, Visvikis D, Koopmansch B, Lambert F, Coimbra C, et al. FDG PET/CT radiomics for predicting the outcome of locally advanced rectal cancer. Eur J Nucl Med Mol Imaging. 2018;45:365–75.
Liu Z, Meng X, Zhang H, Li Z, Liu J, Sun K, Meng Y, Dai W, **e P, Ding Y, et al. Predicting distant metastasis and chemotherapy benefit in locally advanced rectal cancer. Nat Commun. 2020;11:4308.
Mauri G, Arena S, Siena S, Bardelli A, Sartore-Bianchi A. The DNA damage response pathway as a land of therapeutic opportunities for colorectal cancer. Ann Oncol. 2020;31:1135–47.
Solier S, Zhang YW, Ballestrero A, Pommier Y, Zoppoli G. DNA damage response pathways and cell cycle checkpoints in colorectal cancer: current concepts and future perspectives for targeted treatment. Curr Cancer Drug Targets. 2012;12:356–71.
Zhong ME, Chen Y, **ao Y, Xu L, Zhang G, Lu J, Qiu H, Ge W, Wu B. Serum extracellular vesicles contain SPARC and LRG1 as biomarkers of colon cancer and differ by tumour primary location. EBioMedicine. 2019;50:211–23.
Levi-Galibov O, Lavon H, Wassermann-Dozorets R, Pevsner-Fischer M, Mayer S, Wershof E, Stein Y, Brown LE, Zhang W, Friedman G, et al. Heat shock factor 1-dependent extracellular matrix remodeling mediates the transition from chronic intestinal inflammation to colon cancer. Nat Commun. 2020;11:6245.
Chakravarthy A, Khan L, Bensler NP, Bose P, De Carvalho DD. TGF-beta-associated extracellular matrix genes link cancer-associated fibroblasts to immune evasion and immunotherapy failure. Nat Commun. 2018;9:4692.
Gao F, Hu M, Zhong ME, Feng S, Tian X, Meng X, Ni-Jia-Ti MY, Huang Z, Lv M, Song T, et al. Segmentation only uses sparse annotations: Unified weakly and semi-supervised learning in medical images. Med Image Anal. 2022;80:102515.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (82102475, MZ), China Postdoctoral Science Foundation (2021T140769, MZ; 2021TQ0382, XD), Guangdong Basic and Applied Basic Research Foundation (2020A1515110489, MZ) and the National Key Clinical Discipline.
Funding
This study was supported by the National Natural Science Foundation of China (82102475, MZ), China Postdoctoral Science Foundation (2021T140769, MZ; 2021TQ0382, XD), Guangdong Basic and Applied Basic Research Foundation (2020A1515110489, MZ) and the National Key Clinical Discipline. The funders had no role in the study design, data collection, data analysis and interpretation, manuscript preparation, or decision to publish.
Author information
Contributions
Conception and design: XD, MZ, development of methodology: XD, FG. Acquisition of data: MZ, XD, MN, HQ, DX, DC, CL, ZH, QZ. Analysis and interpretation of data: MZ, XD, MN. Writing, review, and/or revision of the manuscript: MZ, XD, MN. Administrative, technical, or material support: FG, XW. Study supervision: FG, XW. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Ethics approval for the retrospective review of imaging and clinical data was received from the local ethics committees. The requirement for written informed consent was waived.
Consent for publication
All authors read and approved the final version of the manuscript.
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Figure S1.
Radiomic features used to predict the prognosis of the CRC risk groups. (A)Example of patients in the low risk with radiomics feature (SurfaceVolmeRation) value of 0.82 and in the high-risk group with value of 0.29. The regions of interest (ROI) for tumour(red) are shown. (B) Boxplot of SurfaceVolmeRation value within radiogenomics dataset for the low risk and high risk radiomics groups. Table S1. Selected imaging feature description.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Zhong, ME., Duan, X., Ni-jia-ti, Mydl. et al. CT-based radiogenomic analysis dissects intratumor heterogeneity and predicts prognosis of colorectal cancer: a multi-institutional retrospective study. J Transl Med 20, 574 (2022). https://doi.org/10.1186/s12967-022-03788-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12967-022-03788-8