Abstract
The need and desire to understand the environment, especially the quality of one’s local water and air, has continued to expand with the emergence of the digital age. The bottleneck in understanding the environment has switched from being able to store all of the data collected to collecting enough data on a broad range of contaminants of environmental concern. Nanomaterial enabled sensors represent a suite of technologies developed over the last 15 years for the highly specific and sensitive detection of environmental contaminants. With the promise of facile, low cost, field-deployable technology, the ability to quantitatively understand nature in a systematic way will soon be a reality. In this review, we first introduce nanosensor design before exploring the application of nanosensors for the detection of three classes of environmental contaminants: pesticides, heavy metals, and pathogens.
Similar content being viewed by others
Background
Nanomaterial enabled sensors are an exciting technology that provide exquisite detection, on the nanomolar to sub-picomolar level, of environmental contaminants [1,2,3,4,5]. Interest in these sensors stems from their potential for facile, in-field contaminant detection without the need for expensive lab equipment. Many past reviews in this area have grouped sensors based on the signal transduction method [2,3,4,5], nanoparticle backbone [7,8,9,10], or contaminant class [1, 11, 12], thus leaving one important paradigm virtually untouched: classifying sensors based on the analyte(s) of interest. Because environmental scientists and engineers are often interested in determining if a specific contaminant exists at a field site and if its concentration is above the regulatory limit, there was a need to organize a review based upon the detection of specific contaminants. This review has been developed to address these concerns. First, we summarize the general concepts underlying a nano-enabled sensor and then discuss recent developments in nanomaterial enabled detection of nine specific analytes: two pesticides, four metals, and three pathogens. A nearly infinite number of chemicals of environmental concern exist and although it would be impossible to outline all of them, the fundamental nanosensor designs can be seen in the examples outlined within the review. For the reader interested in nanosensors for pharmaceutical detection we direct them to the work of Nagaraj et al. [13] and the reviews of Sanvicens et al. [14] and Cristea et al. on antibiotic detection [15].
Introduction
Nanomaterial enabled sensors consist of three components: a nanomaterial(s), a recognition element that provides specificity, and a signal transduction method that provides a means of relaying the presence of the analyte (Fig. 1). These components are not necessarily distinct entities within a sensor, but every nanosensor can be characterized on the basis of these three divisions. Sensors can be designed to detect a single analyte or multiple analytes, termed multiplex detection. In addition to detecting an analyte by producing a signal, a ‘turn-on’ or ‘off/on’ sensor, some of the sensors described below are based on a ‘turn-off’ or ‘on/off’ mechanism, where-by a decrease in signal indicates the presence of an analyte.
Nanosensor design schematic. First, a class and subsequently a specific contaminant of interest is selected (i). The contaminants discussed in this review are denoted with an asterisk. Next, the number of analytes to be detected by the sensor is chosen (ii) and then the probe is designed. A nanoprobe consists of two core elements, a signal transduction method and at least one nanomaterial, and may also include a recognition element (iii). Ultimately, the sensor deployment format is selected (iv)
Nanomaterials
Nanomaterials have enabled advances in sensor design such as miniaturization, portability, and rapid signal response times. High surface area to volume ratios and facile surface functionalization make nanomaterials highly sensitive to changes in surface chemistry thus enabling nanosensors to achieve extremely low detection limits. In some cases, the enhanced sensitivity of nano-enabled sensors is due to the fact that nanomaterials are of a similar size as the analyte of interest (e.g., metal ions, pathogens, biomolecules, antibodies, DNA) and are thus capable of interrogating previously unreachable matrices [4]. We briefly introduce three different general nanomaterial classes: quantum dots (QDs), metal nanoparticles, and carbonaceous nanomaterials.
Quantum dots
QDs are semiconductor nanocrystals with a typical composition MX where M is commonly cadmium (Cd) or zinc (Zn) and X is selenium (Se), sulfur (S), or tellurium (Te). QDs are often coated by a second MX alloy, a shell, to create core/shell QDs with highly tuned properties. Common QDs employed in sensor applications include: CdSe [16], CdSe/ZnS [17,18,19], CdTe [20,34]. The choice of cap** agent depends on the desired function and nanoparticle composition. The interested reader is referred to recent reviews by Saha et al. [7] and Wei et al. [35] for additional details on gold enabled sensors.
A range of nanostructured metal oxides (NMOs) have been explored for sensing applications. NMOs include: iron oxides, titanium oxides, zirconium oxides, cerium oxides, zinc oxides, and tin oxides. Magnetic iron oxides, such as magnetite (Fe3O4) and maghemite (γ-Fe3O4), have low toxicity, are economically friendly, and can be easily functionalized with ligands, antibodies, and other cap** agents [36]. One important allure of magnetic NPs arises from their use in facilitated separation processes and remediation applications [12]. Titanium dioxide, TiO2, has also been embraced in nano-sensor design [37, 38], but it is most typically used and studied for its photocatalytic properties.
Carbon-based nanomaterials
Carbon nanotubes (CNTs) and graphene are often employed in nano-enabled sensors because of their large surface area, excellent electrical conductivity, high thermal conductivity and mechanical strength [39]. One recent application of these nanomaterials has been their use to increase the sensitivity of glassy carbon electrodes (GCE) for electrochemical sensing [40, 9, 72]. Magnetic relaxation switches have been used to detect nucleic acids (DNA and mRNA), proteins [73] and viruses [74] among other targets.
Analytes
As defined at the outset of this review, a wide variety of different analytes can be detected by nanomaterial-based sensors. In this portion of the review, we focus explicitly on the applications of nanosensors towards detection of pesticides, metals, and pathogens.
Pesticides
There is great interest in detection of pesticides given their widespread use, their toxicity, and their proclivity for bioaccumulation. Currently, over 800 active ingredients, in 100 different substance classes are present in commercial pesticides [75]; we summarize the major pesticide classes in Table 1. Organophosphorus (OP), carbamates, neonicotinoids, and triazines are the dominant classes and to date have been the focus of nano-enabled pesticide detection. Liu et al. [75], Verma et al. [76], Aragay et al. [1], Evtugyn et al. [60] and Pang et al. [77], provide detailed reviews of pesticide detection techniques. In this section, a brief background on pesticide detection will be followed by a discussion of recent advances.
Organophosphates
Pesticides are often designed to impact a specific enzyme; many forms of pesticide detection are based on observing and monitoring this enzyme either directly or indirectly. Organophosphate and carbamate pesticides inhibit the production of acetylcholinesterase (AChE) an enzyme that catalyzes the hydrolysis of acetylcholine, a neurotransmitter [78, 79]. The fundamental reaction is shown in Eq. 1.
A class of rapid and sensitive electrochemical sensors has been developed around the immobilization of AChE on a solid electrode surface [11] and Ullah et al. [102].
Mercury
The negative neurological effects of mercury exposure to humans have driven extensive investigation into the geochemical cycling and detection of this element [103]. A major focus of mercury (HgII) nanosensor development has been the production of DNA-based probes [47,48,49,50, 104,105,106]. Thymine–thymine (T–T) base-mismatches in DNA are significantly stabilized in the presence of HgII [104] due to the formation of metal base pairs [107]. Two major types of oligonucleotide mercury probes have been reported in the literature: G-quadruplexes [48, 49], which unfold, and nearly complementary single strands, which hybridize [106]. A growing number of mercury sensors are being constructed using multiple nano-elements, such as the mercury sandwich assay described by Liu et al. [50]. In this assay, magnetic silica spheres encapsulated in a gold shell and Raman labeled gold nanoparticles were functionalized with complementary DNA sequences that contained five mismatched thymine sites, Fig. 5. The DNA sequences were chosen such that the binding energy between the complementary aspects of the strands was insufficient to allow them to fully hybridize. In the presence of mercury, full hybridization occurred thus decreasing the inter-probe spacing and creating a plasmonic hotspot. Owing to the magnetic particle cores, the nanoprobes could be easily recovered with an external magnet and subsequently recycled.

(Reprinted with permission from Liu et al. [50]. Copyright 2014 American Chemical Society)
Schematic of SERS-active system for HgII ion detection. Schematic illustration of the SERS-active system for HgII ion detection based on T–Hg–T bridges using DNA-Au NPs and DNA-MSS@Au NPs
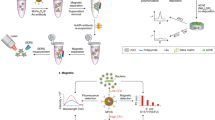
Thiol mediated assays for mercury detection have been described in the literature for a variety of nanoparticles such as gold [108,Cadmium The body of work on nano-enabled sensors for cadmium (Cd) detection is less robust than that for mercury and lead, but detection limits on the order of nano-molar have been reported. A variety of nanomaterials have been explored including QDs [22, 118], single wall carbon nanotubes (SWCNT) [119], and antimony nanoparticles [120]. Gui et al. [22] described an off/on-sensor fluorescence sensor for CdII detection. Photoluminescent CdTe/CdS QDs were first quenched (i.e., turned-off), by ammonium pyrrolidine dithiocarbamate (APDC) due to the partial loss of the Cd–thiol surface layer and subsequent surface passivation. Introduced cadmium ions displaced the APDC from the QD surface and restored the photoluminescence (PL); thus, turning the sensor on. The sensor was highly selective for CdII, a threefold increase was seen in the PL intensity, and a limit of detection of 6 nM was determined. Gui et al. [118] enhanced the accuracy of their CdII detection device by creating a ratiometric sensor. In this sensor, the fluorescence of two different chromophores was measured in order to minimize the error introduced by fluctuation in the photoluminescence of the QDs. To limit interactions between the QDs and the secondary dye, the CdTe QD cores were coated with a polymer, polyethylenimine (PEI), prior to conjugation with fluorescein isothiocyanate (FITC). The QDs were then quenched using sulfur (S2−) while the FITC signal was maintained. Again, upon introduction of cadmium the sensor was turned on and the photoluminescence was restored. The limit of detection was slightly higher for this sensor compared to the same groups initial report, 12 nM vs. 6 nM, but was linear across a much larger range, 0.1–15 µM compared with 0.1–2 µM. High chromium (Cr) absorption in vivo can result in various diseases, including fibro-proliferative diseases, airway hypersensitivity, lung cancer, nasal cancer, and other types of tumors [121]. Multiple immunoassays have been described for the detection of chromium [45, 46], but they are all based on the work of Liu et al. [46]. In pursuit of an immunochromatographic assay (ICA), Liu et al. developed novel anti-CrIII-EDTA monoclonal antibodies (McAb). Chromium ions are too small to elicit an immune response and thus they were mixed with the highly effective bifunctional chelating agent, isothiocyanobenzyl-EDTA, and conjugated to the carrier protein bovine serum albumin (BSA) before being introduced to mice from which the antibodies were ultimately extracted. The immunoassay dipstick was composed of the three main parts: (i) a conjugation pad that was dosed with the anti-Cr-EDTA antibodies; (ii) a test line that contained the analyte of interest, Cr-EDTA, and; (iii) a control line that contained goat- anti-mouse antibodies. To run a sample, liquid is introduced to the dipstick and travels into the conjugation pad where the probes are brought into solution. For a negative sample, the free antibody probes bind to the test line, whereas in a positive sample no probes will bind as all antibody sites are already occupied and thus no signal is produced at the test line. The antibodies at the control line will capture any probes in the solution even those that are bound to the target of interest and is use to verify that capillary action wicked the solution through the whole length of the dipstick. The ultimate result of Liu et al. was an assay with a visual limit of detection of 50 ng/mL and an analysis time of < 5 min. Ever since John Snow’s 1854 revelation that cholera was spread through the consumption of contaminated water, waterborne pathogen detection has been a key area of research. The World Health Organization (WHO) recognizes twelve bacteria, eight viruses, seven protozoa, and two helminths as pathogens of significance in drinking water supplies, as outlined in Table 2 [6]. Pathogen detection methods typically focus on: (i) whole analyte (cell) detection or detection of a representative epitope on the cell membrane; (ii) genetic material detection; or (iii) pathogenic product (e.g., toxin) detection. For the sake of brevity, herein we confine our discussion to the detection of Vibrio cholerae and the toxin it produces, cholera toxin, Legionella pneumophila, which was responsible for greater than 50% of the waterborne disease outbreaks between 2011 and 2012 [122], and Pseudomonas aeruginosa, which the WHO recently classified as a critical pathogen in light of the proliferation of antimicrobial resistant species [123]. For expanded reviews we refer the reader to the works of Kumar et al. [124] and Mocan et al. [125]. Cholera, the infamous disease that spawned germ theory is now virtually unknown in the United States, but it continues to pose a major disease burden around the world with an estimated 1.3–4.0 million cases of cholera a year leading to between 21,000 and 143,000 deaths [126]. Cholera is an acute diarrhoeal disease caused by the ingestion of contaminated water or food containing the bacterium Vibrio cholerae. In the intestines, the bacteria colonize the mucosa and begin to secrete cholera toxin (CT), which initiates the disease symptoms [127]. Nanosensors have been fabricated to detect both Vibrio cholerae [128, 129] and CT, but the majority of the literature has focused on detection of CT subunit B (CT-B) [130,131,132,133,134] because the subunit induces cellular uptake of the toxin and not all V. cholerae isolates are toxigenic [135]. Label-based detection of CT can be achieved using antibodies, ganglioside GM1 (the binding site of CT), or β-galactose, a sugar with strong affinity towards CT. Ahn et al. [130] provide a nice summary of CT-B detection and reported a fluorescence resonance energy transfer (FRET) based method with a theoretical detection limit of 280 pM. In FRET, fluorescence from QDs is quenched, and the energy is transferred by another particle such as a gold nanoparticle. The quenching is inhibited in the presence of the target. Specifically, the cholera toxin binds to the β-galactose modified gold nanoparticles prohibiting the binding of the QDs. Named for the famous 1976 outbreak at the American Legion, Legionnaires’ disease is a pneumonia like disease caused by the bacterium Legionella pneumophila. Under specific conditions, the bacterium can flourish in building (premise) plum** and infect people when they inhale aerosols containing the infective agent. Two approaches have been presented in the literature for nano-enabled Legionella detection: whole organism detection [136, 137] and DNA detection [138,139,140]. Martin el al. [136] developed a whole organism sensor that combined a sandwich immunoassay for bacterial capture with amperometric transduction. Magnetic nanoparticles were modified with poly(dopamine) (pDA) and ultimately functionalized with specific capture antibodies (C-Ab) to create MNPs@pDA-C-Ab probes. After incubation with the sample, a second detector antibody labeled with horseradish peroxidase was introduced and a magnetic field was used to capture the immunocomplexes on a screen-printed carbon electrode (SPCE). The authors found the assay to be specific for Legionella, but they needed a preconcentration step in order to achieve a LOD below the reference of 100 colony-forming units (CFU) L−1. However, with a runtime of < 3 h, compared to 10 days for the standard approach, and a LOD of 10 CFU mL−1, the sensor has the potential to be used as a rapid first screening method for highly contaminated water systems. In a recent report, Melaine et al. [139] described the multiplex detection of 16S rRNA from Legionella, Pseudomonas aeruginosa (discussed below) and Salmonella typhimurium. A DNA microarray with capture DNA specific for each target was assembled on a surface plasmon resonance imaging (SPRi) substrate, e.g., gold coated nanoprisms. Upon hybridization of the DNA with isolated 16S rRNA a change in the reflectivity signal was observed, as shown in the bottom of Fig. 6. To extend the dynamic range of detection and enhance sensitivity, gold nanoparticles functionalized with a detection probe were introduced to the sample and ultimately RNA at concentrations as low as 10 pg mL−1 were detected. (Adapted with permission from Melaine et al. [139]. Copyright 2017 American Chemical Society) A schematic of multiplex RNA detection using surface plasmon resonance imaging (SPRi). A schematic of multiplex RNA detection using surface plasmon resonance imaging (SPRi). RNA fragments are first extracted from bacteria of interest (a). A biochip functionalized with three specific capture probes (CP) and a negative control probe (NP), each demarcated in a unique color (b (i)) is shown to exhibit no change in reflectivity (c (i)). Upon introduction to the RNA (b (ii)), there is an increase in single (c (ii)). Finally, gold nanoparticles functionalized with the detection probe (GNP-DP) are introduced and shown to enhance the change in reflectivity An opportunistic pathogen, Pseudomonas aeruginosa can be found in sources such as feces, soil, water, and sewage with the most important route of exposure being skin (dermal) contact with contaminated water or tools. Similar to Legionella, P. aeruginosa can colonize premise plumbing and has been associated with outbreaks of nosocomial infections in hospitals [141]. Most of the detection schemes reported for P. aeruginosa focus on whole pathogen detection [142,143,144,145,146] with the work of Melanie et al. [139], discussed above, on 16s rRNA detection being an outlier. In addition, to oligonucleotide recognition elements [139, 142,143,144], antibodies [145, 147] and bacteriophages [146] have also been used for specific detection of P. aeruginosa. The first P. aeruginosa aptamer was discovered by Wang et al. [148] in 2011 and subsequently has been used in a range of sensors. The discussion that follows highlights two sensors that utilize optical transduction. Yoo et al. [142] and Hu et al. [144] fabricated nano-textured substrates to produce localized surface plasmon resonance (LSPR) chips (Fig. 7). Yoo et al. choose a three-step fabrication approach, first gold was deposited on a glass slide, silica nanoparticles were then deposited and then followed by the deposition of a second gold layer whereas Hu et al. opted for standard nanosphere lithography. The two groups also chose different methods to functionalize the sensor with Yoo et al. attaching the aptamers directly to the sensor surface via a gold-thiol bond. In contrast, Hu et al. used a polyethylene glycol (PEG) spacer to minimize steric hindrance for the aptamers with the goal of achieving a lower detection level. Hu et al. were successful at develo** a sensor with a linear response at low concentrations and a lower limit of detection, 10 CFU mL−1 vs. Yoo et al.’s 104 CFU mL−1. It should be noted that one of Yoo et al.’s goals was to create a low volume sensor and that their LOD was obtained in a 3 µL sample. (Reprinted with permission from Hu et al. [144]. Copyright 2018 American Chemical Society) Schematic of P. aeruginosa LSPR sensor chip (left). Sensor calibration curve, where error bar represents the standard deviation of all data points at a specific bacterial concentration (right).Chromium
Pathogens
\(Vibrio\; cholerae\) and cholera toxin
\(Legionella\; pneumophila\)

\(Pseudomonas\; aeruginosa\)

Conclusions
Nanosensor development for environmental contaminants is growing rapidly and, as described throughout this review, nanomaterials and recognition agents are continuously being combined in new and creative ways. The recent developments in sensor design aim to overcome the shortcomings of first-generation sensors such as nonspecific binding, particle size variation, nanoparticle aggregation, and nanoparticle stability. Questions of assay selectivity and sensitive in complex environmental matrices remains but a growing number of reports are using representative matrices to demonstrate the stability and selectivity of their sensors. The robustness of field deployable sensors is a must if individuals are going to be empowered to analyze their environment.
Abbreviations
- Abs:
-
antibodies
- AChE:
-
acetylcholinesterase
- AgNP:
-
silver nanoparticles
- APDC:
-
ammonium pyrrolidine dithiocarbamate
- AuNP or GNP:
-
gold nanoparticles
- AuNP/BC:
-
gold nanoparticle/bacteria cellulose
- BSA:
-
bovine serum albumin
- Cd:
-
cadmium
- CFU:
-
colony-forming unit
- CNT:
-
carbon nanotubes
- Cr:
-
chromium
- CS:
-
chitosan
- CT:
-
cholera toxin
- DNA:
-
deoxyribonucleic acid
- dNTP:
-
deoxyribonucleotide triphosphate
- DPV:
-
differential pulse voltammetry
- Fe3O4 :
-
magnetite
- FRET:
-
fluorescence resonance energy transfer
- FTIC:
-
fluorescein isothiocyanate
- GCE:
-
glassy carbon electrodes
- Hg:
-
mercury
- ICA:
-
immunochromatographic assay
- LOD:
-
fluorescence resonance energy transfer
- LSPR:
-
localized surface plasmon resonance
- mAbs or McAb:
-
monoclonal antibodies
- MB:
-
magnetic bead
- MNP:
-
magnetic nanoparticles
- MPA:
-
3-mercaptopropionic acid
- mRNA:
-
messenger RNA
- NMO:
-
nanostructured metal oxides
- NP:
-
nanoparticles
- OP:
-
organophosphorus pesticide
- pAbs:
-
polyclonal
- Pb:
-
lead
- PCR:
-
polymerase chain reaction
- PEG:
-
polyethylene glycol
- PEI:
-
polyethylenimine
- PL:
-
photoluminescence
- PPy:
-
polypyrrole
- QD:
-
quantum dot
- R6G:
-
rhodamine 6G
- rA:
-
ribonucleoside adenosine
- RCA:
-
rolling circle amplification
- rGO:
-
reduced graphene oxide
- RNA:
-
ribonucleic acid
- S:
-
sulfur
- Se:
-
selenium
- SEF:
-
surface enhanced fluorescence
- SELEX:
-
systematic evolution of ligands by exponential enrichment
- SERS:
-
surface enhanced Raman spectroscopy
- SPR:
-
surface plasmon resonance
- SPRi:
-
surface plasmon resonance imaging
- ssDNA:
-
single stranded DNA
- SWASV:
-
square wave anodic strip** voltammetry
- SWCNT:
-
single wall carbon nanotubes
- Te:
-
tellurium
- TGA:
-
thioglycolic acid
- TiO2 :
-
titanium dioxide
- TMB:
-
3,3,5,5-tetramethylbenzidine
- WHO:
-
World Health Organization
- Zn:
-
zinc
- γ-Fe3O4 :
-
maghemite
References
Aragay G, Pino F, Merkoci A. Nanomaterials for sensing and destroying pesticides. Chem Rev. 2012;112:5317–38.
Sadik OA, Aluoch AO, Zhou A. Status of biomolecular recognition using electrochemical techniques. Biosens Bioelectron. 2009;24:2749–65.
Bănică F-G. Nanomaterial applications in optical transduction. In: Chemical sensors and biosensors. Chichester: Wiley; 2012. p. 454–72.
Grieshaber D, MacKenzie R, Vörös J, Reimhult E. Electrochemical biosensors—sensor principles and architectures. Sensors. 2008;8:1400–58.
Liu G, Lin Y. Nanomaterial labels in electrochemical immunosensors and immunoassays. Talanta. 2007;74:308–17.
World Health Organization. Guidelines for drinking-water quality, vol. 1. Geneva: World Health Organization; 2008.
Saha K, Agasti SS, Kim C, Li X, Rotello VM. Gold nanoparticles in chemical and biological sensing. Chem Rev. 2012;112:2739–79.
Lu AH, Salabas EL, Schuth F. Magnetic nanoparticles: synthesis, protection, functionalization, and application. Angew Chem Int Ed Engl. 2007;46:1222–44.
Reddy LH, Arias JL, Nicolas J, Couvreur P. Magnetic nanoparticles: design and characterization, toxicity and biocompatibility, pharmaceutical and biomedical applications. Chem Rev. 2012;112:5818–78.
Xu Z, Dong J. Synthesis, characterization, and application of magnetic nanocomposites for the removal of heavy metals from industrial effluents. In: Shah V, editor. Emerging environmental technologies. Springer, Netherlands: Dordrecht; 2008. p. 105–48.
Li M, Gou H, Al-Ogaidi I, Wu N. Nanostructured sensors for detection of heavy metals: a review. ACS Sustain Chem Eng. 2013;1:713–23.
Mohmood I, Lopes C, Lopes I, Ahmad I, Duarte A, Pereira E. Nanoscale materials and their use in water contaminants removal—a review. Environ Sci Pollut Res. 2013;20:1239–60.
Nagaraj VJ, Jacobs M, Vattipalli KM, Annam VP, Prasad S. Nanochannel-based electrochemical sensor for the detection of pharmaceutical contaminants in water. Environ Sci Process Impacts. 2014;16:135–40.
Sanvicens N, Mannelli I, Salvador JP, Valera E, Marco MP. Biosensors for pharmaceuticals based on novel technology. TrAC Trends Anal Chem. 2011;30:541–53.
Cristea C, Tertis M, Galatus R. Magnetic nanoparticles for antibiotics detection. Nanomaterials. 2017;7:119.
Algarra M, Campos BB, Alonso B, Miranda MS, Martínez ÁM, Casado CM, Esteves da Silva JCG. Thiolated DAB dendrimers and CdSe quantum dots nanocomposites for Cd(II) or Pb(II) sensing. Talanta. 2012;88:403–7.
Li M, Zhou X, Guo S, Wu N. Detection of lead(II) with a “turn-on” fluorescent biosensor based on energy transfer from CdSe/ZnS quantum dots to graphene oxide. Biosens Bioelectron. 2013;43:69–74.
Luan W, Yang H, Wan Z, Yuan B, Yu X, Tu S-T. Mercaptopropionic acid capped CdSe/ZnS quantum dots as fluorescence probe for lead(II). J Nanopart Res. 2012;14:1–8.
Sung T-W, Lo Y-L. Highly sensitive and selective sensor based on silica-coated CdSe/ZnS nanoparticles for Cu2+ ion detection. Sens Actuators B Chem. 2012;165:119–25.
Chao MR, Chang YZ, Chen JL. Hydrophilic ionic liquid-passivated CdTe quantum dots for mercury ion detection. Biosens Bioelectron. 2013;42:397–402.
Gan TT, Zhang YJ, Zhao NJ, **ao X, Yin GF, Yu SH, Wang HB, Duan JB, Shi CY, Liu WQ. Hydrothermal synthetic mercaptopropionic acid stabled CdTe quantum dots as fluorescent probes for detection of Ag(+). Spectrochim Acta A Mol Biomol Spectrosc. 2012;99:62–8.
Gui R, An X, Su H, Shen W, Chen Z, Wang X. A near-infrared-emitting CdTe/CdS core/shell quantum dots-based OFF–ON fluorescence sensor for highly selective and sensitive detection of Cd2+. Talanta. 2012;94:257–62.
Liu X-Q, Liu Q, Cao S-H, Cai W-P, Weng Y-H, **e K-X, Li Y-Q. Directional surface plasmon-coupled emission of CdTe quantum dots and its application in Hg(II) sensing. Anal Methods. 2012;4:3956.
Wu H, Liang J, Han H. A novel method for the determination of Pb2+ based on the quenching of the fluorescence of CdTe quantum dots. Microchim Acta. 2007;161:81–6.
Wang YQ, Liu Y, He XW, Li WY, Zhang YK. Highly sensitive synchronous fluorescence determination of mercury(II) based on the denatured ovalbumin coated CdTe QDs. Talanta. 2012;99:69–74.
Koneswaran M, Narayanaswamy R. l-Cysteine-capped ZnS quantum dots based fluorescence sensor for Cu2+ ion. Sens Actuators B Chem. 2009;139:104–9.
Ke J, Li X, Shi Y, Zhao Q, Jiang X. A facile and highly sensitive probe for Hg(II) based on metal-induced aggregation of ZnSe/ZnS quantum dots. Nanoscale. 2012;4:4996–5001.
Link S, El-Sayed MA. Spectral properties and relaxation dynamics of surface plasmon electronic oscillations in gold and silver nanodots and nanorods. J Phys Chem B. 1999;103:8410–26.
Romo-Herrera JM, Alvarez-Puebla RA, Liz-Marzan LM. Controlled assembly of plasmonic colloidal nanoparticle clusters. Nanoscale. 2011;3:1304–15.
Sonntag MD, Klingsporn JM, Zrimsek AB, Sharma B, Ruvuna LK, Van Duyne RP. Molecular plasmonics for nanoscale spectroscopy. Chem Soc Rev. 2014;43:1230–47.
Chen Y-Y, Chang H-T, Shiang Y-C, Hung Y-L, Chiang C-K, Huang C-C. Colorimetric assay for lead ions based on the leaching of gold nanoparticles. Anal Chem. 2009;81:9433–9.
Lee Y-F, Nan F-H, Chen M-J, Wu H-Y, Ho C-W, Chen Y-Y, Huang C-C. Detection and removal of mercury and lead ions by using gold nanoparticle-based gel membrane. Anal Methods. 2012;4:1709.
Lok C-N, Ho C-M, Chen R, He Q-Y, Yu W-Y, Sun H, Tam P-H, Chiu J-F, Che C-M. Silver nanoparticles: partial oxidation and antibacterial activities. J Biol Inorg Chem. 2007;12:527–34.
Daniel M-C, Astruc D. Gold nanoparticles: assembly, supramolecular chemistry, quantum-size-related properties, and applications toward biology, catalysis, and nanotechnology. Chem Rev. 2004;104:293–346.
Wei H, Abtahi SH, Vikesland PJ. LSPR-based colorimetric and SERS sensors for environmental analysis. Environ Sci. 2015;2(2):120–35.
Hnaiein M, Hassen WM, Abdelghani A, Fournier-Wirth C, Coste J, Bessueille F, Leonard D, Jaffrezic-Renault N. A conductometric immunosensor based on functionalized magnetite nanoparticles for E. coli detection. Electrochem Commun. 2008;10:1152–4.
Filanovsky B, Markovsky B, Bourenko T, Perkas N, Persky R, Gedanken A, Aurbach D. Carbon electrodes modified with TiO2/metal nanoparticles and their application for the detection of trinitrotoluene. Adv Func Mater. 2007;17:1487–92.
Li H, Li J, Xu Q, Yang Z, Hu X. A derivative photoelectrochemical sensing platform for 4-nitrophenolate contained organophosphates pesticide based on carboxylated perylene sensitized nano-TiO2. Anal Chim Acta. 2013;766:47–52.
Yang W, Ratinac KR, Ringer SP, Thordarson P, Gooding JJ, Braet F. Carbon nanomaterials in biosensors: should you use nanotubes or graphene? Angew Chem Int Ed Engl. 2010;49:2114–38.
Fan H, Li Y, Wu D, Ma H, Mao K, Fan D, Du B, Li H, Wei Q. Electrochemical bisphenol A sensor based on N-doped graphene sheets. Anal Chim Acta. 2012;711:24–8.
Dong J, Fan X, Qiao F, Ai S, **n H. A novel protocol for ultra-trace detection of pesticides: combined electrochemical reduction of Ellman’s reagent with acetylcholinesterase inhibition. Anal Chim Acta. 2013;761:78–83.
Kim YA, Lee E-H, Kim K-O, Lee YT, Hammock BD, Lee H-S. Competitive immunochromatographic assay for the detection of the organophosphorus pesticide chlorpyrifos. Anal Chim Acta. 2011;693:106–13.
Zhao WW, Ma ZY, Yu PP, Dong XY, Xu JJ, Chen HY. Highly sensitive photoelectrochemical immunoassay with enhanced amplification using horseradish peroxidase induced biocatalytic precipitation on a CdS quantum dots multilayer electrode. Anal Chem. 2012;84:917–23.
Trilling AK, Beekwilder J, Zuilhof H. Antibody orientation on biosensor surfaces: a minireview. Analyst. 2013;138:1619–27.
Jiajie L, Hongwu L, Caifeng L, Qiangqiang F, Caihong H, Zhi L, Tianjiu J, Yong T. Silver nanoparticle enhanced Raman scattering-based lateral flow immunoassays for ultra-sensitive detection of the heavy metal chromium. Nanotechnology. 2014;25:495501.
Liu X, **ang JJ, Tang Y, Zhang XL, Fu QQ, Zou JH, Lin Y. Colloidal gold nanoparticle probe-based immunochromatographic assay for the rapid detection of chromium ions in water and serum samples. Anal Chim Acta. 2012;745:99–105.
Li M, Wang Q, Shi X, Hornak LA, Wu N. Detection of mercury(II) by quantum dot/DNA/gold nanoparticle ensemble based nanosensor via nanometal surface energy transfer. Anal Chem. 2011;83:7061–5.
Li T, Li B, Wang E, Dong S. G-quadruplex-based DNAzyme for sensitive mercury detection with the naked eye. Chem Commun. 2009. https://doi.org/10.1039/b903993g.
Long F, Zhu A, Shi H, Wang H, Liu J. Rapid on-site/in-situ detection of heavy metal ions in environmental water using a structure-switching DNA optical biosensor. Sci Rep. 2013;3:2308.
Liu M, Wang Z, Zong S, Chen H, Zhu D, Wu L, Hu G, Cui Y. SERS detection and removal of mercury(II)/silver(I) using oligonucleotide-functionalized core/shell magnetic silica Sphere@Au nanoparticles. ACS Appl Mater Interfaces. 2014;6:7371–9.
Ma J, Chen Y, Hou Z, Jiang W, Wang L. Selective and sensitive mercuric(II) ion detection based on quantum dots and nicking endonuclease assisted signal amplification. Biosens Bioelectron. 2013;43:84–7.
Zhang M, Yin BC, Tan W, Ye BC. A versatile graphene-based fluorescence “on/off” switch for multiplex detection of various targets. Biosens Bioelectron. 2011;26:3260–5.
Evtugyn GA, Budnikov HC, Nikolskaya EB. Sensitivity and selectivity of electrochemical enzyme sensors for inhibitor determination. Talanta. 1998;46:465–84.
Bies C, Lehr CM, Woodley JF. Lectin-mediated drug targeting: history and applications. Adv Drug Deliv Rev. 2004;56:425–35.
Rhoades RA, Pflanzer RG. Human physiology. Boston: Thomson Learning; 2002.
Ellington AA, Kullo IJ, Bailey KR, Klee GG. Antibody-based protein multiplex platforms: technical and operational challenges. Clin Chem. 2010;56:186–93.
Engvall E, Perlmann P. Enzyme-linked immunosorbent assay, Elisa. III quantitation of specific antibodies by enzyme-labeled anti-immunoglobulin in antigen-coated tubes. J Immunol. 1972;109:129–35.
Bordeaux J, Welsh AW, Agarwal S, Killiam E, Baquero MT, Hanna JA, Anagnostou VK, Rimm DL. Antibody validation. Biotechniques. 2010;48:197–209.
Huang CC, Huang YF, Cao Z, Tan W, Chang HT. Aptamer-modified gold nanoparticles for colorimetric determination of platelet-derived growth factors and their receptors. Anal Chem. 2005;77:5735–41.
Evtugyn GA. Biosensors for pesticides and foodborne pathogens. Portable biosensing of food toxicants and environmental pollutants., Sensors seriesBoca Raton: CRC Press; 2013. p. 605–80.
Famulok M, Mayer G. Aptamer modules as sensors and detectors. Acc Chem Res. 2011;44:1349–58.
Low SY, Hill JE, Peccia J. DNA aptamers bind specifically and selectively to (1→3)-β-d-glucans. Biochem Biophys Res Commun. 2009;378:701–5.
Hamula CLA, Zhang H, Li F, Wang Z, Le Chris X, Li X-F. Selection and analytical applications of aptamers binding microbial pathogens. TrAC Trends Anal Chem. 2011;30:1587–97.
Stoltenburg R, Reinemann C, Strehlitz B. SELEX—a (r)evolutionary method to generate high-affinity nucleic acid ligands. Biomol Eng. 2007;24:381–403.
Hoinka J, Berezhnoy A, Dao P, Sauna ZE, Gilboa E, Przytycka TM. Large scale analysis of the mutational landscape in HT-SELEX improves aptamer discovery. Nucleic Acids Res. 2015;43:5699–707.
Bănică F-G. What are chemical sensors? In: Chemical sensors and biosensors. Chichester: Wiley; 2012. p. 1–20.
Vikesland PJ, Wigginton KR. Nanomaterial enabled biosensors for pathogen monitoring—a review. Environ Sci Technol. 2010;44:3656–69.
Bae SW, Tan W, Hong J-I. Fluorescent dye-doped silica nanoparticles: new tools for bioapplications. Chem Commun. 2012;48:2270–82.
Sanvicens N, Pastells C, Pascual N, Marco MP. Nanoparticle-based biosensors for detection of pathogenic bacteria. TrAC Trends Anal Chem. 2009;28:1243–52.
Garcia-Aljaro C, Cella LN, Shirale DJ, Park M, Munoz FJ, Yates MV, Mulchandani A. Carbon nanotubes-based chemiresistive biosensors for detection of microorganisms. Biosens Bioelectron. 2010;26:1437–41.
Koets M, van der Wijk T, van Eemeren JTWM, van Amerongen A, Prins MWJ. Rapid DNA multi-analyte immunoassay on a magneto-resistance biosensor. Biosens Bioelectron. 2009;24:1893–8.
Kaittanis C, Naser SA, Perez JM. One-step, nanoparticle-mediated bacterial detection with magnetic relaxation. Nano Lett. 2006;7:380–3.
Zhang Y, Yang Y, Ma W, Guo J, Lin Y, Wang C. Uniform magnetic core/shell microspheres functionalized with Ni2+—iminodiacetic acid for one step purification and immobilization of his-tagged enzymes. ACS Appl Mater Interfaces. 2013;5:2626–33.
Perez JM, Simeone FJ, Saeki Y, Josephson L, Weissleder R. Viral-induced self-assembly of magnetic nanoparticles allows the detection of viral particles in biological media. J Am Chem Soc. 2003;125:10192–3.
Liu S, Zheng Z, Li X. Advances in pesticide biosensors: current status, challenges, and future perspectives. Anal Bioanal Chem. 2013;405:63–90.
Verma N, Bhardwaj A. Biosensor technology for pesticides—a review. Appl Biochem Biotechnol. 2015;175:3093–119.
Pang S, Yang T, He L. Review of surface enhanced Raman spectroscopic (SERS) detection of synthetic chemical pesticides. TrAC Trends Anal Chem. 2016;85:73–82.
Fukuto TR. Mechanism of action of organophosphorus and carbamate insecticides. Environ Health Perspect. 1990;87:245–54.
Stenersen J. Chemical pesticides mode of action and toxicology. Boca Raton: CRC Press; 2004.
Du D, Ye X, Cai J, Liu J, Zhang A. Acetylcholinesterase biosensor design based on carbon nanotube-encapsulated polypyrrole and polyaniline copolymer for amperometric detection of organophosphates. Biosens Bioelectron. 2010;25:2503–8.
Amine A, Mohammadi H, Bourais I, Palleschi G. Enzyme inhibition-based biosensors for food safety and environmental monitoring. Biosens Bioelectron. 2006;21:1405–23.
Yadavalli VK, Koh W-G, Lazur GJ, Pishko MV. Microfabricated protein-containing poly(ethylene glycol) hydrogel arrays for biosensing. Sens Actuators B Chem. 2004;97:290–7.
Nikolelis DP. Portable chemical sensors: weapons against bioterrorism. Dordrecht: Springer Netherlands; 2012.
Yang Y, Asiri AM, Du D, Lin Y. Acetylcholinesterase biosensor based on a gold nanoparticle-polypyrrole-reduced graphene oxide nanocomposite modified electrode for the amperometric detection of organophosphorus pesticides. Analyst. 2014;139:3055–60.
Yu G, Wu W, Zhao Q, Wei X, Lu Q. Efficient immobilization of acetylcholinesterase onto amino functionalized carbon nanotubes for the fabrication of high sensitive organophosphorus pesticides biosensors. Biosens Bioelectron. 2015;68:288–94.
Cui H-F, Wu W-W, Li M-M, Song X, Lv Y, Zhang T-T. A highly stable acetylcholinesterase biosensor based on chitosan-TiO2-graphene nanocomposites for detection of organophosphate pesticides. Biosens Bioelectron. 2018;99:223–9.
Pang S, Labuza TP, He L. Development of a single aptamer-based surface enhanced Raman scattering method for rapid detection of multiple pesticides. Analyst. 2014;139:1895–901.
Zhang C, Wang L, Tu Z, Sun X, He Q, Lei Z, Xu C, Liu Y, Zhang X, Yang J, et al. Organophosphorus pesticides detection using broad-specific single-stranded DNA based fluorescence polarization aptamer assay. Biosens Bioelectron. 2014;55:216–9.
Laurier KGM, Poets M, Vermoortele F, Cremer GD, Martens JA, Uji-i H, De Vos DE, Hofkens J, Roeffaers MBJ. Photocatalytic growth of dendritic silver nanostructures as SERS substrates. Chem Commun. 2012;48:1559–61.
Nie Y, Teng Y, Li P, Liu W, Shi Q, Zhang Y. Label-free aptamer-based sensor for specific detection of malathion residues by surface-enhanced Raman scattering. Spectrochim Acta Part A Mol Biomol Spectrosc. 2018;191:271–6.
Wang LM, Cai J, Wang YL, Fang QK, Wang SY, Cheng Q, Du D, Lin YH, Liu FQ. A bare-eye-based lateral flow immunoassay based on the use of gold nanoparticles for simultaneous detection of three pesticides. Microchim Acta. 2014;181:1565–72.
Jeschke P, Nauen R, Schindler M, Elbert A. Overview of the status and global strategy for neonicotinoids. J Agric Food Chem. 2011;59:2897–908.
Simon-Delso N, Amaral-Rogers V, Belzunces LP, Bonmatin JM, Chagnon M, Downs C, Furlan L, Gibbons DW, Giorio C, Girolami V, et al. Systemic insecticides (neonicotinoids and fipronil): trends, uses, mode of action and metabolites. Environ Sci Pollut Res Int. 2015;22:5–34.
Verdian A. Apta-nanosensors for detection and quantitative determination of acetamiprid—a pesticide residue in food and environment. Talanta. 2018;176:456–64.
Weerathunge P, Ramanathan R, Shukla R, Sharma TK, Bansal V. Aptamer-controlled reversible inhibition of gold nanozyme activity for pesticide sensing. Anal Chem. 2014;86:11937–41.
United States Environmental Protection Agency (USEPA) Atrazine Chemical Summary 2007. https://archive.epa.gov/region5/teach/web/pdf/atrazine_summary.pdf.
Liu X, Li W-J, Li L, Yang Y, Mao L-G, Peng Z. A label-free electrochemical immunosensor based on gold nanoparticles for direct detection of atrazine. Sens Actuators B Chem. 2014;191:408–14.
Yılmaz E, Özgür E, Bereli N, Türkmen D, Denizli A. Plastic antibody based surface plasmon resonance nanosensors for selective atrazine detection. Mater Sci Eng C. 2017;73:603–10.
Wei H, Vikesland PJ. pH-triggered molecular alignment for reproducible SERS detection via an AuNP/nanocellulose platform. Sci Rep. 2015;5:18131.
Yang S, Wu T, Zhao X, Li X, Tan W. The optical property of core-shell nanosensors and detection of atrazine based on localized surface plasmon resonance (LSPR) sensing. Sensors. 2014;14:13273.
Mohapatra S, Bera MK, Das RK. Rapid “turn-on” detection of atrazine using highly luminescent N-doped carbon quantum dot. Sens Actuators B Chem. 2018;263:459–68.
Ullah N, Mansha M, Khan I, Qurashi A. Nanomaterial-based optical chemical sensors for the detection of heavy metals in water: recent advances and challenges. TrAC Trends Anal Chem. 2018;100:155–66. https://doi.org/10.1016/j.trac.2018.01.002.
Selid P, Xu H, Collins EM, Striped Face-Collins M, Zhao JX. Sensing mercury for biomedical and environmental monitoring. Sensors. 2009;9:5446–59.
Miyake Y, Togashi H, Tashiro M, Yamaguchi H, Oda S, Kudo M, Tanaka Y, Kondo Y, Sawa R, Fujimoto T, et al. MercuryII-mediated formation of thymine–HgII–thymine base pairs in DNA duplexes. J Am Chem Soc. 2006;128:2172–3.
Gao ZF, Song WW, Luo HQ, Li NB. Detection of mercury ions (II) based on non-cross-linking aggregation of double-stranded DNA modified gold nanoparticles by resonance Rayleigh scattering method. Biosens Bioelectron. 2015;65:360–5.
Ono A, Togashi H. Highly selective oligonucleotide-based sensor for mercury(II) in aqueous solutions. Angew Chem Int Ed. 2004;43:4300–2.
Clever GH, Kaul C, Carell T. DNA–metal base pairs. Angew Chem Int Ed Engl. 2007;46:6226–36.
Kim Y-R, Mahajan RK, Kim JS, Kim H. Highly sensitive gold nanoparticle-based colorimetric sensing of mercury(II) through simple ligand exchange reaction in aqueous media. ACS Appl Mater Interfaces. 2009;2:292–5.
**aorong Y, Huixiang L, Juan X, Xuemei T, He H, Danbi T. A simple and cost-effective sensing strategy of mercury(II) based on analyte-inhibited aggregation of gold nanoparticles. Nanotechnology. 2011;22:275503.
Huang CC, Chang HT. Selective gold-nanoparticle-based “turn-on” fluorescent sensors for detection of mercury(II) in aqueous solution. Anal Chem. 2006;78:8332–8.
Chansuvarn W, Tuntulani T, Imyim A. Colorimetric detection of mercury(II) based on gold nanoparticles, fluorescent gold nanoclusters and other gold-based nanomaterials. TrAC Trends Anal Chem. 2015;65:83–96.
Alam A, Ravindran A, Chandran P, Sudheer Khan S. Highly selective colorimetric detection and estimation of Hg2+ at nano-molar concentration by silver nanoparticles in the presence of glutathione. Spectrochim Acta Part A Mol Biomol Spectrosc. 2015;137:503–8.
Goyer RA. Lead toxicity: from overt to subclinical to subtle health effects. Environ Health Perspect. 1990;86:177–81.
Kim H-K, Liu J, Li J, Nagraj N, Li M, Pavot CMB, Lu Y. Metal-dependent global folding and activity of the 8–17 DNAzyme studied by fluorescence resonance energy transfer. J Am Chem Soc. 2007;129:6896–902.
Tang S, Tong P, Li H, Tang J, Zhang L. Ultrasensitive electrochemical detection of Pb(2)(+) based on rolling circle amplification and quantum dots tagging. Biosens Bioelectron. 2013;42:608–11.
Li T, Wang E, Dong S. Lead(II)-induced allosteric G-quadruplex DNAzyme as a colorimetric and chemiluminescence sensor for highly sensitive and selective Pb2+ detection. Anal Chem. 2010;82:1515–20.
Gao C, Yu XY, Xu RX, Liu JH, Huang XJ. AlOOH-reduced graphene oxide nanocomposites: one-pot hydrothermal synthesis and their enhanced electrochemical activity for heavy metal ions. ACS Appl Mater Interfaces. 2012;4:4672–82.
Gui R, An X, Huang W. An improved method for ratiometric fluorescence detection of pH and Cd2+ using fluorescein isothiocyanate-quantum dots conjugates. Anal Chim Acta. 2013;767:134–40.
Sun D, **e X, Cai Y, Zhang H, Wu K. Voltammetric determination of Cd2+ based on the bifunctionality of single-walled carbon nanotubes-Nafion film. Anal Chim Acta. 2007;581:27–31.
Toghill KE, **ao L, Wildgoose GG, Compton RG. Electroanalytical determination of cadmium(II) and lead(II) using an antimony nanoparticle modified boron-doped diamond electrode. Electroanalysis. 2009;21:1113–8.
Gibb HJ, Lees PS, Pinsky PF, Rooney BC. Lung cancer among workers in chromium chemical production. Am J Ind Med. 2000;38:115–26.
Beer KD, Gargano JW, Roberts VA, Hill VR, Garrison LE, Kutty PK, Hilborn ED, Wade TJ, Fullerton KE, Yoder JS. Surveillance for waterborne disease outbreaks associated with drinking water-United States, 2011–2012. MMWR Morb Mortal Wkly Rep. 2015;64:842–8.
World Health Organization. WHO publishes list of bacteria for which new antibiotics are urgently needed. Geneva: WHO; 2017.
Kumar N, Hu Y, Singh S, Mizaikoff B. Emerging biosensor platforms for the assessment of water-borne pathogens. Analyst. 2018;143:359–73.
Mocan T, Matea CT, Pop T, Mosteanu O, Buzoianu AD, Puia C, Iancu C, Mocan L. Development of nanoparticle-based optical sensors for pathogenic bacterial detection. J Nanobiotechnology. 2017;15:25.
Ali M, Nelson AR, Lopez AL, Sack DA. Updated global burden of cholera in endemic countries. PLoS Negl Trop Dis. 2015;9:e0003832.
Finkelstein RA. Cholera, Vibrio cholerae O1 and O139, and other pathogenic vibrios. In: Baron S, editor. Medical microbiology. 4th ed. Galveston: University of Texas Medical Branch at Galveston; 1996.
Yu CY, Ang GY, Chua AL, Tan EH, Lee SY, Falero-Diaz G, Otero O, Rodríguez I, Reyes F, Acosta A, et al. Dry-reagent gold nanoparticle-based lateral flow biosensor for the simultaneous detection of Vibrio cholerae serogroups O1 and O139. J Microbiol Methods. 2011;86:277–82.
Tam PD, Thang CX. Label-free electrochemical immunosensor based on cerium oxide nanowires for Vibrio cholerae O1 detection. Mater Sci Eng C. 2016;58:953–9.
Ahn K-S, Lim KR, Jeong D, Lee BY, Kim KS, Lee W-Y. Fluorescence energy transfer inhibition bioassay for cholera toxin based on galactose-stabilized gold nanoparticles and amine-terminated quantum dots. Microchem J. 2016;124:9–14.
Khan SA, DeGrasse JA, Yakes BJ, Croley TR. Rapid and sensitive detection of cholera toxin using gold nanoparticle-based simple colorimetric and dynamic light scattering assay. Anal Chim Acta. 2015;892:167–74.
Zhang C-H, Liu L-W, Liang P, Tang L-J, Yu R-Q, Jiang J-H. Plasmon coupling enhanced raman scattering nanobeacon for single-step, ultrasensitive detection of cholera toxin. Anal Chem. 2016;88:7447–52.
Goldman ER, Clapp AR, Anderson GP, Uyeda HT, Mauro JM, Medintz IL, Mattoussi H. Multiplexed toxin analysis using four colors of quantum dot fluororeagents. Anal Chem. 2004;76:684–8.
Schofield CL, Field RA, Russell DA. Glyconanoparticles for the colorimetric detection of cholera toxin. Anal Chem. 2007;79:1356–61.
Azarian T, Ali A, Johnson JA, Jubair M, Cella E, Ciccozzi M, Nolan DJ, Farmerie W, Rashid MH, Sinha-Ray S, et al. Non-toxigenic environmental Vibrio cholerae O1 strain from Haiti provides evidence of pre-pandemic cholera in Hispaniola. Sci Rep. 2016;6:36115.
Martín M, Salazar P, Jiménez C, Lecuona M, Ramos MJ, Ode J, Alcoba J, Roche R, Villalonga R, Campuzano S, et al. Rapid Legionella pneumophila determination based on a disposable core–shell Fe3O4@poly(dopamine) magnetic nanoparticles immunoplatform. Anal Chim Acta. 2015;887:51–8.
Park J, You X, Jang Y, Nam Y, Kim MJ, Min NK, Pak JJ. ZnO nanorod matrix based electrochemical immunosensors for sensitivity enhanced detection of Legionella pneumophila. Sens Actuators B Chem. 2014;200:173–80.
Wu T-Y, Su Y-Y, Shu W-H, Mercado AT, Wang S-K, Hsu L-Y, Tsai Y-F, Chen C-Y. A novel sensitive pathogen detection system based on microbead quantum dot system. Biosens Bioelectron. 2016;78:37–44.
Melaine F, Saad M, Faucher S, Tabrizian M. Selective and high dynamic range assay format for multiplex detection of pathogenic Pseudomonas aeruginosa, Salmonella typhimurium, and Legionella pneumophila RNAs using surface plasmon resonance imaging. Anal Chem. 2017;89:7802–7.
Foudeh AM, Daoud JT, Faucher SP, Veres T, Tabrizian M. Sub-femtomole detection of 16s rRNA from Legionella pneumophila using surface plasmon resonance imaging. Biosens Bioelectron. 2014;52:129–35.
Falkinham JO III, Hilborn ED, Arduino MJ, Pruden A, Edwards MA. Epidemiology and ecology of opportunistic premise plumbing pathogens: Legionella pneumophila, Mycobacterium avium, and Pseudomonas aeruginosa. Environ Health Perspect. 2015;123:749.
Yoo SM, Kim D-K, Lee SY. Aptamer-functionalized localized surface plasmon resonance sensor for the multiplexed detection of different bacterial species. Talanta. 2015;132:112–7.
Jia F, Xu L, Yan W, Wu W, Yu Q, Tian X, Dai R, Li X. A magnetic relaxation switch aptasensor for the rapid detection of Pseudomonas aeruginosa using superparamagnetic nanoparticles. Microchim Acta. 2017;184:1539–45.
Hu J, Fu K, Bohn PW. Whole-cell Pseudomonas aeruginosa localized surface plasmon resonance aptasensor. Anal Chem. 2018;90(3):2326–32.
Krithiga N, Viswanath KB, Vasantha VS, Jayachitra A. Specific and selective electrochemical immunoassay for Pseudomonas aeruginosa based on pectin–gold nano composite. Biosens Bioelectron. 2016;79:121–9.
He Y, Wang M, Fan E, Ouyang H, Yue H, Su X, Liao G, Wang L, Lu S, Fu Z. Highly specific bacteriophage-affinity strategy for rapid separation and sensitive detection of viable Pseudomonas aeruginosa. Anal Chem. 2017;89:1916–21.
Ellairaja S, Krithiga N, Ponmariappan S, Vasantha VS. Novel pyrimidine tagged silver nanoparticle based fluorescent immunoassay for the detection of Pseudomonas aeruginosa. J Agric Food Chem. 2017;65:1802–12.
Wang K-Y, Zeng Y-L, Yang X-Y, Li W-B, Lan X-P. Utility of aptamer-fluorescence in situ hybridization for rapid detection of Pseudomonas aeruginosa. Eur J Clin Microbiol Infect Dis. 2011;30:273–8.
Authors’ contributions
MRW and PJV reviewed the literature. MRW drafted and wrote significant portions of the manuscript. MRW and PJV edited the manuscript. Both authors read and approved the final manuscript.
Acknowledgements
Support for MRW was provided by the Virginia Tech Graduate School through the Sustainable Nanotechnology Interdisciplinary Graduate Education Program (VTSuN IGEP).
Competing interests
The authors declare that they have no competing interests. The authors are entirely responsible for the content of the review of the opinions contained within it.
Availability of data and materials
Not applicable.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Funding
NSF and the Environmental Protection Agency under NSF Cooperative Agreement EF-0830093, Center for the Environmental Implications of Nano-Technology (CEINT).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Willner, M.R., Vikesland, P.J. Nanomaterial enabled sensors for environmental contaminants. J Nanobiotechnol 16, 95 (2018). https://doi.org/10.1186/s12951-018-0419-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12951-018-0419-1