Abstract
Pancreatic cancer (PC) is one of the most common malignancies. Surgical resection is a potential curative approach for PC, but most patients are unsuitable for operations when at the time of diagnosis. Even with surgery, some patients may still experience tumour metastasis during the operation or shortly after surgery, as precise prognosis evaluation is not always possible. If patients miss the opportunity for surgery and resort to chemotherapy, they may face the challenging issue of chemotherapy resistance. In recent years, liquid biopsy has shown promising prospects in disease diagnosis, treatment monitoring, and prognosis assessment. As a noninvasive detection method, liquid biopsy offers advantages over traditional diagnostic procedures, such as tissue biopsy, in terms of both cost-effectiveness and convenience. The information provided by liquid biopsy helps clinical practitioners understand the molecular mechanisms underlying tumour occurrence and development, enabling the formulation of more precise and personalized treatment decisions for each patient. This review introduces molecular biomarkers and detection methods in liquid biopsy for PC, including circulating tumour cells (CTCs), circulating tumour DNA (ctDNA), noncoding RNAs (ncRNAs), and extracellular vesicles (EVs) or exosomes. Additionally, we summarize the applications of liquid biopsy in the early diagnosis, treatment response, resistance assessment, and prognostic evaluation of PC.
Similar content being viewed by others
Introduction
Pancreatic cancer (PC) is one of the most common malignancies, and the number of PC cases has doubled over the past two decades. The incidence of PC varies significantly across regions and populations, with the highest rates observed in North America, Europe, and Australia [1,2,3]. Recent years have seen a rapid increase in deaths due to PC, which can be attributed to global population growth and age structure changes and is closely linked to social and economic development [4]. According to predictions, PC is expected to become the third leading cause of cancer-related deaths in the European Union [5]. By 2030, it is projected to overtake breast, prostate, and colorectal cancers and become the second leading cause of cancer-related deaths in the United States [6].
Pancreatic ductal adenocarcinoma (PDAC) is a primary histological subtype of PC, accounting for 90% of all cases [7, 8]. Surgical resection is one of the methods of a potential cure, but most PDAC patients are unsuitable for operations when they are diagnosed [9, 10]. Therefore, screening and diagnosis should be conducted as early as possible to ensure a positive outcome. The diagnosis of PDAC relies on endoscopic ultrasonography-guided fine needle aspiration (EUS-FNA), magnetic resonance imaging (MRI), and computed tomography (CT) [11,12,13,14]. However, there are some problems, such as invasiveness, high cost, and exposure of subjects to radiation [15, 16]. In addition, the molecular composition of tumours is complex and dynamic, and repeated endoscopy examinations create a significant burden on patients [17]. Although the potential role of diagnostic biomarkers of cancer is constantly evolving, reliable diagnostic biomarkers for PC are still lacking. For instance, carbohydrate antigen 19-9 (CA19-9) has been extensively studied as a biomarker for detecting PC [18]. However, due to the lack of specificity of CA19-9, it can be expressed in various liver and gallbladder diseases as well as other types of malignant tumours, and elevated levels can also occur in some benign obstructive diseases [19]. Fucosyltransferase 3 (also known as the Lewis gene) is the key enzyme involved in the biosynthesis of CA19-9. Approximately 5–10% of individuals are Lewis antigen-negative, which means they do not secrete or secrete very little CA19-9, and to some extent, this also hinders the diagnosis of PC [20]. Therefore, CA19-9 alone cannot offer a conclusive diagnosis and must be combined with different clinical presentations, imaging tests, and biomarkers [15, 21].
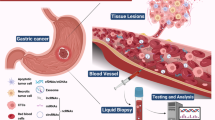
In recent years, liquid biopsy has garnered attention due to its advantages of lower invasiveness and the ability to continuously monitor cancer progression. While blood is considered the most critical biofluid for liquid biopsy (Fig. 1), other clinical samples, such as cerebrospinal fluid, saliva, ascites, pleural effusion, and urine, have also been used [22,23,24,25,26]. Different sample sources have unique characteristics, with the prevailing view suggesting that blood samples carry a richer molecular information profile. Although noninvasive samples such as stool, urine, and saliva may contain less biomarker information than blood, they can provide valuable information about the location of diseases. For instance, certain biomarkers in urine may be associated with kidney or bladder conditions [27], while stool biomarkers may be linked to digestive system disorders [28]. Currently, the potential targets of liquid biopsy are circulating tumour cells (CTCs), circulating tumour DNA (ctDNA), noncoding RNAs (ncRNAs), messenger RNAs (mRNAs), and extracellular vesicles (EVs), which can provide information about tumour genomics, transcriptomics, and proteomics.
Common samples, biomarkers, and clinical applications in liquid biopsy for pancreatic cancer. Blood is typically the most commonly used material in liquid biopsy, in addition to pancreatic juice, saliva, urine, and stool. Circulating tumour cells, circulating tumour DNA, noncoding RNAs, and extracellular vesicles are among the most common biomarkers. Liquid biopsy has a wide range of clinical applications, playing a crucial role in early diagnosis, treatment monitoring, and prognosis evaluation. Created with BioRender.com
Liquid biopsy exhibits high utility in the management of PC, with applications spanning early diagnosis, treatment strategies, drug resistance, recurrence monitoring, and prognosis assessment for PC patients. This review provides an overview of the biomarkers and detection methods utilized in liquid biopsy and their applications in the early diagnosis, treatment response, and prognosis evaluation of PC (Table 1). We also discuss the future trends of liquid biopsy and assess its limitations to improve current management strategies for patients.
Biomarkers and detection methods
Circulating tumour cells
CTCs, which detach from the primary tumour, can enter the circulatory system and travel through the bloodstream. However, the majority of CTCs die in the peripheral blood within 1 to 2.5 hours due to mechanical forces or immune system attacks. Nevertheless, a small fraction of CTCs can survive and initiate distant metastasis [67, 68]. Numerous metastatic precursors within CTCs increase the risk of tumour metastasis and recurrence [69,70,71]. According to most perspectives, CTCs are believed to exhibit specific differences from primary tumours despite originating from primary tumours. This heterogeneity leads to their detachment from the primary tumour and acquisition of epithelial-mesenchymal transition (EMT) characteristics, facilitating intravascular infiltration and enhancing their potential for metastasis [72, 73].
The analysis process of CTCs mainly involves three stages: enrichment, detection, and characterization. Most enrichment methods are applied based on the surface phenotype or physical properties of CTCs. The CellSearch system, developed using an antibody targeting epithelial cell adhesion molecule (EpCAM), is the sole CTC detection technology approved by the U.S. Food and Drug Administration due to its ability to detect CTCs expressing EpCAM [29, 74]. However, this strategy cannot detect cells with low EpCAM expression due to the potential loss of epithelial antigens during the EMT process [34]. Furthermore, the abundance of CTCs varies across different types of cancer; the CellSearch system is more suitable for tumours with higher CTC abundance [75]. In microfluidic devices, affinity-based separation methods can also be employed. Designing microfluidic devices with varying materials, sizes, and structures to manipulate blood flow patterns creates additional opportunities for interacting CTCs and antibodies [32, 57, 76]. Pahattuge et al. [77] introduced a modular microfluidic system called SMART-Chip. They demonstrated that the SMART-Chip platform could significantly reduce the processing time by more than 50% when handling blood samples obtained from patients with PDAC and colorectal cancer compared to manual sample processing. Furthermore, microfabricated porous membranes can be employed to filtrate and isolate CTCs due to their size, which is larger than that of normal blood cells [78, 79].
CTCs are primarily detected using protein expression, immunocytochemistry, and nucleic acid methods. Flow cytometry allows for the quantitative assessment and characterization of protein expression in CTCs, offering the advantage of evaluating multiple biomarkers to characterize CTCs comprehensively. However, it exhibits lower sensitivity for detecting rare populations of CTCs [80]. Immunohistochemical staining and immunofluorescence are commonly employed techniques for detection and characterization purposes. Immunofluorescence, in particular, enables the visual confirmation of protein expression and localization by fluorescent markers. In the conventional cytofluorimetry approach, isolation is achieved through the utilization of specific antibodies that recognize markers selected on CTCs. This method utilizes monoclonal antibodies specifically targeting antigens expressed by CTCs, which results in the exclusion of CTCs that do not express such antigens but are present in the circulation. Consequently, this presents a challenge in obtaining or develo** novel antibodies against specific targets [81]. The flexibility of immunofluorescence technology makes it a powerful tool for studying the protein expression of tumour cells. For instance, with the use of multimarker immunofluorescence panels, researchers can gain a more comprehensive understanding of the distribution and expression patterns of different CTC subtypes [82]. This not only aids in tumour classification and staging but also provides valuable insights for personalized therapy. In addition, CTCs can also be detected using techniques such as high-resolution image scanning, mutational analysis, and single-cell next-generation sequencing (scNGS). The molecular characteristics of CTCs were initially determined on enriched fractions, which provided limited information about tumour heterogeneity. In recent years, the rapidly advancing single-cell sequencing technology has become the preferred method for isolating individual CTCs and studying tumour heterogeneity. These technologies will facilitate the comprehensive characterization of CTCs at multiple omics and functional levels, enabling effective monitoring of the dynamic changes in tumour heterogeneity in individual cancer patients [83, 84].
Although the precise role of CTCs in tumour development remains elusive, they offer a valuable approach for obtaining comprehensive insights into tumours through liquid biopsy. In PC management, CTCs play a significant and beneficial role in patient diagnosis, prognostic evaluation, recurrence monitoring, and treatment decisions. In this regard, we have summarized the clinical applications of CTCs in various aspects of PC management in recent years (Table 2).
Circulating tumour DNA
Cell-free DNA (cfDNA) is a crucial genetic component found in the bloodstream; its origin primarily stems from apoptotic, necrotic, and actively secreted fragments originating from healthy, inflamed, and tumour tissue. These fragments are typically approximately 150–180 base pairs in length [97,98,99]. CtDNA represents a distinctive subset of cfDNA released into the blood by CTCs. Compared to cfDNA, ctDNA is present in relatively lower amounts in the bloodstream, constituting only 1% (or even less than 0.01%) of cfDNA [98,99,100]. Most ctDNA fragments have lengths ranging from 160 to 200 base pairs, and they are less influenced by intratumoural heterogeneity compared to tumour tissues [97, 101, 102]. Additionally, ctDNA has a half-life of approximately 15 minutes to 2.5 hours, which means that it serves as a real-time tumour biomarker. In contrast, traditional blood protein biomarkers usually take weeks to manifest, and ctDNA can dynamically reflect the status of a tumour at a specific moment [103, 104]. Furthermore, ctDNA carries tumour-related genomic information, such as gene expression levels, mutations, the methylation status, and microsatellite instability. Compared to traditional biopsy markers, ctDNA is an ideal biomarker, especially for the real-time monitoring of treatment effectiveness and prognosis assessment.
CtDNA detection includes ctDNA preparation, library construction, analysis, and data alignment. One aspect of ctDNA detection focuses on genetic mutations. Single-base mutations have the potential to activate oncogenes, disrupting the balance between oncogenes and tumour suppressor genes, thereby instigating tumorigenesis. Another aspect involves DNA methylation, which plays a role in tumour initiation that is similar to that of DNA mutations [105, 106]. Mutation detection is a vital component of the analysis. Due to the extremely low abundance of ctDNA, employing highly sensitive techniques for detecting tumour mutations is crucial. Conventional approaches rely on polymerase chain reaction (PCR), but recent advancements in PCR and sequencing technologies have paved the way for alternative methods, including quantitative PCR (qPCR), digital PCR (dPCR), droplet digital PCR (ddPCR), and next-generation sequencing (NGS). qPCR allows real-time monitoring of DNA amplification with higher speed, reproducibility, and quantification. NGS platforms offer several advantages, including the ability to screen for unknown mutations and structural and copy-number variations. dPCR and ddPCR involve partitioning DNA samples into thousands or even millions of separate compartments or droplets, effectively reducing background noise associated with traditional methods and enabling the detection of tumour DNA at a variant allele frequency (VAF) below 0.1% [107,108,109]. In recent years, integrated detection strategies combining gene editing techniques, functional enzymes, and nanomaterials have been developed to effectively increase the net content of mutation fragments, thereby facilitating the identification of target gene mutations within ctDNA [106]. There are various methods for DNA methylation detection. Whole-genome bisulfite sequencing (WGBS-seq) is considered the gold standard for DNA methylation analysis. It can identify partially methylated regions in cancer cells. However, the sensitivity of this method may be compromised by DNA degradation [26, 110].
KRAS mutations are the most prevalent genetic alteration in PC. They are present in over 90% of patients and are considered an early driving factor in PDAC [108]. Castells et al. [111] demonstrated that the presence of KRAS mutations in plasma DNA served as a highly specific molecular marker for diagnosis and prognosis in a PDAC cohort of 44 patients. However, it is essential to emphasize that previous cfDNA sequencing results have not only identified mutations known to exist in tumours but have also uncovered a multitude of variations that are absent in tumour tissues [112]. In particular, some patients undergoing chemotherapy may harbour minimal residual lesions composed of drug-resistant cells. In such cases, the mutations detected in cfDNA in the bloodstream do not exclusively originate from tumour cells. CfDNA may also carry mutations from other sources, including those induced by the disease state or treatment. Undoubtedly, KRAS mutations are one of the vital indicators for evaluating PDAC, and their role has received widespread attention as a primary focus in many PC studies (Table 3). However, overall, the application of ctDNA and mutation analysis in PDAC still requires further strategies to thoroughly assess this detection method.
Noncoding RNAs
NcRNAs were once perceived to have a limited impact on tumour initiation and progression due to their inability to encode proteins. However, emerging evidence has highlighted the essential regulatory functions of ncRNAs. In addition to their capacity to modulate gene and protein expression, ncRNAs actively participate in diverse tumorigenic processes, including EMT, autophagy, and apoptosis [122,123,124]. NcRNAs can be classified into two main categories based on their lengths: small noncoding RNAs (sncRNAs), with a length of less than 200 nucleotides, and long noncoding RNAs (lncRNAs), with a length exceeding 200 nucleotides [125, 126]. In addition, circRNAs, which are RNA molecules with a circular structure, have been recognized for their significant regulatory roles in gene expression, cell proliferation, cell differentiation, and disease development in recent years. SncRNAs encompass several subtypes, including microRNAs (miRNAs), small nucleolar RNAs, small nuclear RNAs, piwi-interacting RNAs, and tRNA-derived small RNAs [127]. Among them, miRNAs are the most extensively studied factors in cancer research, and liquid biopsy identifies miRNAs actively secreted by CTCs and tumour cells themselves [128, 129]. MiRNAs can influence genes, with thousands of miRNAs regulating approximately 60% of the genes. Their principal function involves binding to recognition sites in the 3' untranslated region, thereby reducing mRNA stability and suppressing gene expression [130, 131].
LncRNAs play a regulatory role in protein and miRNA functions and expression levels and contribute to chromatin remodelling [130, 73, 95, 96]. However, some research suggests that there may not be a significant difference in CTC counts between blood samples collected before and after chemotherapy, possibly due to variations in CTC identification and treatment strategies [29, 93, 197]. In addition to quantitative analyses, the molecular characteristics of CTCs are also frequently used to assess a patient's treatment response [198]. Some CTC measurement techniques enable genetic profiling of CTCs, allowing the detection of key gene mutations, such as those in KRAS, HER2, and TP53 [199,200,201]. Furthermore, programmed death ligand 1 (PD-L1) staining methods can be employed to evaluate the status of CTCs in patients receiving monoclonal antibody therapy, with PD-L1-negative CTC patients often achieving better treatment outcomes [202]. In most cases, CTCs express chemokine receptors, with CXC-motif chemokine receptor 4 (CXCR4) being the most commonly expressed receptor. Continuous monitoring of CXCR4 during treatment serves as a predictive biomarker, providing information to identify which patients are likely to benefit from treatment or develop resistance [203, 204]. Regarding drug sensitivity, Wu et al. [205] conducted a study wherein they collected CTCs from patients diagnosed with PDAC and expanded them ex vivo into organoids. The sensitivity of these organoids to nine drugs (GEM, 5-fluorouracil, erlotinib, irinotecan, olaparib, oxaliplatin, paclitaxel, palbociclib, and trametinib) was examined. A significant correlation was observed between the drug sensitivity of CTCs and clinical outcomes. This indicates that the drug sensitivity of CTCs holds the potential to predict therapeutic outcomes in PDAC, thus enabling the avoidance of ineffective treatments. CECs have been proposed as a potential tool to predict how patients will respond to antiangiogenic cancer therapies. However, it is important to recognize that their diverse phenotypes may exhibit different dynamics during the course of treatment. Given the unique characteristics of CECs and their crucial role in liquid biopsies, this avenue of research holds promise and warrants further exploration. A clinical trial focused on late-stage pancreatic cancer patients monitored CEC levels during neoadjuvant therapy and observed an overall increase in CECs in response to combination therapy that was attributed to chemotherapy-induced vascular damage exacerbating CEC release [206]. Furthermore, research concerning surgery, which is a common treatment method, has indicated that CEC levels typically decrease after tumour resection. This decline may result from the disruption of PDAC-derived growth factor recruitment of endothelial cells after tumour removal, subsequently reducing CEC levels [207].
The longitudinal assessment of ctDNA enables dynamic monitoring of disease trajectory, including treatment monitoring and the detection of minimal residual disease, and serves as an alternative biomarker for overall disease burden [208]. Tao et al. [209] conducted a study to examine the role of ctDNA in monitoring treatment response in a cohort of 17 PDAC patients who were treated with the FOLFIRINOX regimen (fluorouracil, irinotecan, and oxaliplatin). Among the 12 patients who responded to chemotherapy, 11 exhibited a reduction in the mutant allele fraction (MAF) of cfDNA. In contrast, the remaining 5 patients who developed chemotherapy resistance showed an increase in the ctDNA MAF during disease progression. These findings suggest that the levels of ctDNA partly reflect the tumour burden. In another study, Groot et al. [210] identified a substantial decrease in the probability of detecting ctDNA in the bloodstream of patients who underwent neoadjuvant chemotherapy compared to those who did not receive any preoperative chemotherapy (21% vs. 69%; p < 0.001). Although the practice of longitudinal ctDNA monitoring in PDAC cases remains limited, these studies underscore the potential of ctDNA as a crucial monitoring biomarker during the therapeutic course.
The emergence of chemoresistance presents formidable obstacles for nonsurgical candidates, thereby exacerbating their clinical predicament. Several miRNAs are considered key regulatory elements involved in acquiring chemoresistance in PDAC. Lu et al. [163] demonstrated that the expression of plasma miR-20a-5p in PDAC patients who exhibited resistance to GEM was markedly diminished compared to that in nonresistant patients (p < 0.01). The authors proposed that miR-20a-5p potentially regulates the expression of the RRM2 protein, thereby exerting an influence on the sensitivity of tumour cells to GEM. MiRNA levels have the potential to serve as informative indicators regarding disease progression, whether assessed before treatment initiation or during the treatment course. In a study by van der Sijde et al. [211], the elevated expression levels of serum miR-373-3p before FOLFIRINOX therapy was identified as a predictive factor for disease progression. Correspondingly, the reduced expression levels of miR-194-5p following a single cycle of FOLFIRINOX treatment indicated disease deterioration. LncRNA holds considerable importance in guiding the therapeutic approach for PC. Zhang et al. [215,216,217]. Although the CTC count is commonly used as the determining criterion in studies, distinguishing CTC subgroups can also reflect the tumour status to some extent. Certain CTC subgroups with specific phenotypes, for example, indicate the tendency of the tumour for metastasis [218, 219]. Semaan et al. [33] identified and characterized 4 CTC subpopulations that can be used for the clinical stratification of PC, providing a valuable perspective for applying liquid biopsy technologies in prognostic prediction. Some studies suggest that CTC distribution shows spatial heterogeneity and that portal venous blood may be a better option for assessing PDAC prognosis than peripheral venous blood [206].
Research on ctDNA primarily focuses on three KRAS mutations (G12D, G12V, and G12R) [226]; however, G12R has a lower detection rate than the other two mutations [227, 228]. Ako et al. [227] studied postoperative recurrence and overall survival in PDAC by analysing two KRAS mutations (G12D and G12V); they discovered that patients with both KRAS mutations had significantly lower disease-free survival. Interestingly, in a separate study, Guo et al. [216] specifically examined the influence of the G12D mutation on the prognosis of PDAC. Among 26 patients with PDAC, those with the KRAS G12D mutation had notably reduced overall survival (12.1 vs. 24.9 months, p < 0.001) and recurrence-free survival (6.3 vs. 17.4 months, p < 0.001) compared to those without the mutation. Notably, patients with the KRAS G12D mutation exhibited a distinct early recurrence trend and poorer clinical outcomes.
In previous studies, several reports have confirmed the predictive role of ncRNAs in PC. Most of these studies constructed multifactor prognostic risk models and performed survival analyses with these models [189, 229, In conclusion, the application of liquid biopsy in the clinical management of PC aligns with the concept of precision medicine. Biological samples obtained through noninvasive procedures can provide detailed information about various aspects of the tumour, which aids in monitoring tumour development and evaluating treatment responses. Moreover, this information assists clinical physicians in understanding the molecular mechanisms of tumour occurrence and development and providing more accurate and personalized treatment decisions for each patient. There are also some limitations, including low-sensitivity detection techniques, nonstandardized analysis workflows, and small sample sizes; these limitations are significant barriers of liquid biopsy. It cannot be denied that with the continuous advancement of technological methods and large-scale clinical trials, many biomarkers have begun to demonstrate their value, indicating broad prospects for their application. Liquid biopsy will become an indispensable technology for tumour diagnosis and treatment.Conclusion
Availability of data and materials
Not applicable.
Abbreviations
- PC:
-
Pancreatic cancer
- PDAC:
-
Pancreatic ductal adenocarcinoma
- EUS-FNA:
-
Endoscopic ultrasonography-guided fine needle aspiration
- MRI:
-
Magnetic resonance imaging
- CT:
-
Computed tomography
- CA19-9:
-
Carbohydrate antigen 19-9
- CTCs:
-
Circulating tumour cells
- ctDNA:
-
Circulating tumour DNA
- ncRNAs:
-
Noncoding RNAs
- mRNAs:
-
Messenger RNAs
- EVs:
-
Extracellular vesicles
- EMT:
-
Epithelial-mesenchymal transition
- EpCAM:
-
Epithelial cell adhesion molecule
- scNGS:
-
Single-cell next-generation sequencing
- cfDNA:
-
Cell-free DNA
- PCR:
-
Polymerase chain reaction
- qPCR:
-
Quantitative PCR
- dPCR:
-
Digital PCR
- ddPCR:
-
Droplet digital PCR
- NGS:
-
Next-generation sequencing
- VAF:
-
Variant allele frequency
- WGBS-seq:
-
Whole-genome bisulfite sequencing
- sncRNAs:
-
Small noncoding RNAs
- lncRNAs:
-
Long noncoding RNAs
- miRNAs:
-
MicroRNAs
- SERS:
-
Surface-enhanced Raman spectroscopy
- SPR:
-
Surface plasmon resonance
- CECs:
-
Circulating epithelial cells
- AUC:
-
Area under the curve
- exLR:
-
Extracellular vesicle long RNA
- GPC1:
-
Glypican-1
- GEM:
-
Gemcitabine
- PD-L1:
-
Programmed death ligand 1
- CXCR4:
-
CXC-motif chemokine receptor 4
- MAF:
-
Mutant allele fraction
- CHI3L1:
-
Chitinase 3-like-1
- FN1:
-
Fibronectin
- OS:
-
Overall survival
References
Ilic M, Ilic I. Epidemiology of pancreatic cancer. World J Gastroenterol. 2016;22:9694–705.
Raimondi S, Maisonneuve P, Lowenfels AB. Epidemiology of pancreatic cancer: an overview. Nat Rev Gastroenterol Hepatol. 2009;6:699–708.
Klein AP. Pancreatic cancer epidemiology: understanding the role of lifestyle and inherited risk factors. Nat Rev Gastroenterol Hepatol. 2021;18:493–502.
Wong MCS, Jiang JY, Liang M, Fang Y, Yeung MS, Sung JJY. Global temporal patterns of pancreatic cancer and association with socioeconomic development. Sci Rep. 2017;7:3165.
Ferlay J, Partensky C, Bray F. More deaths from pancreatic cancer than breast cancer in the EU by 2017. Acta Oncol. 2016;55:1158–60.
Rahib L, Smith BD, Aizenberg R, Rosenzweig AB, Fleshman JM, Matrisian LM. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014;74:2913–21.
Cid-Arregui A, Juarez V. Perspectives in the treatment of pancreatic adenocarcinoma. World J Gastroenterol. 2015;21:9297–316.
Lux A, Kahlert C, Grützmann R, Pilarsky C. c-Met and PD-L1 on Circulating Exosomes as Diagnostic and Prognostic Markers for Pancreatic Cancer. Int J Mol Sci. 2019;20(13):3305.
Neuzillet C, Tijeras-Raballand A, Bourget P, Cros J, Couvelard A, Sauvanet A, Vullierme MP, Tournigand C, Hammel P. State of the art and future directions of pancreatic ductal adenocarcinoma therapy. Pharmacol Ther. 2015;155:80–104.
Martin-Perez E, Domínguez-Muñoz JE, Botella-Romero F, Cerezo L, Matute Teresa F, Serrano T, Vera R. Multidisciplinary consensus statement on the clinical management of patients with pancreatic cancer. Clin Transl Oncol. 2020;22:1963–75.
Semaan A, Bernard V, Lee JJ, Wong JW, Huang J, Swartzlander DB, Stephens BM, Monberg ME, Weston BR, Bhutani MS, et al. Defining the Comprehensive Genomic Landscapes of Pancreatic Ductal Adenocarcinoma Using Real-World Endoscopic Aspiration Samples. Clin Cancer Res. 2021;27:1082–93.
Ashida R, Kitano M. Endoscopic ultrasound-guided tissue acquisition for pancreatic ductal adenocarcinoma in the era of precision medicine. Dig Endosc. 2022;34:1329–39.
Barat M, Marchese U, Pellat A, Dohan A, Coriat R, Hoeffel C, Fishman EK, Cassinotto C, Chu L, Soyer P. Imaging of Pancreatic Ductal Adenocarcinoma: An Update on Recent Advances. Can Assoc Radiol J. 2023;74:351–61.
Anta JA, Martínez-Ballestero I, Eiroa D, García J, Rodríguez-Comas J. Artificial intelligence for the detection of pancreatic lesions. Int J Comput Assist Radiol Surg. 2022;17:1855–65.
Zhang L, Sanagapalli S, Stoita A. Challenges in diagnosis of pancreatic cancer. World J Gastroenterol. 2018;24:2047–60.
Zhou H, Zhu L, Song J, Wang G, Li P, Li W, Luo P, Sun X, Wu J, Liu Y, et al. Liquid biopsy at the frontier of detection, prognosis and progression monitoring in colorectal cancer. Mol Cancer. 2022;21:86.
Driessens G, Beck B, Caauwe A, Simons BD, Blanpain C. Defining the mode of tumour growth by clonal analysis. Nature. 2012;488:527–30.
Misek DE, Patwa TH, Lubman DM, Simeone DM. Early detection and biomarkers in pancreatic cancer. J Natl Compr Canc Netw. 2007;5:1034–41.
Mann DV, Edwards R, Ho S, Lau WY, Glazer G. Elevated tumour marker CA19-9: clinical interpretation and influence of obstructive jaundice. Eur J Surg Oncol. 2000;26:474–9.
Guo M, Luo G, Lu R, Shi W, Cheng H, Lu Y, ** K, Yang C, Wang Z, Long J, et al. Distribution of Lewis and Secretor polymorphisms and corresponding CA19-9 antigen expression in a Chinese population. FEBS Open Bio. 2017;7:1660–71.
Marrelli D, Caruso S, Pedrazzani C, Neri A, Fernandes E, Marini M, Pinto E, Roviello F. CA19-9 serum levels in obstructive jaundice: clinical value in benign and malignant conditions. Am J Surg. 2009;198:333–9.
Erben V, Poschet G, Schrotz-King P, Brenner H. Comparing Metabolomics Profiles in Various Types of Liquid Biopsies among Screening Participants with and without Advanced Colorectal Neoplasms. Diagnostics (Basel). 2021;11(3):561.
Wu H, Ji H, Yang W, Zhang M, Guo Y, Li B, Wang J, Chen R, Chen Y, Wang X. Liquid biopsy using ascitic fluid and pleural effusion supernatants for genomic profiling in gastrointestinal and lung cancers. BMC Cancer. 2022;22:1020.
Wang Y, Springer S, Mulvey CL, Silliman N, Schaefer J, Sausen M, James N, Rettig EM, Guo T, Pickering CR, et al. Detection of somatic mutations and HPV in the saliva and plasma of patients with head and neck squamous cell carcinomas. Sci Transl Med. 2015;7:293ra104.
De Mattos-Arruda L, Mayor R, Ng CKY, Weigelt B, Martínez-Ricarte F, Torrejon D, Oliveira M, Arias A, Raventos C, Tang J, et al. Cerebrospinal fluid-derived circulating tumour DNA better represents the genomic alterations of brain tumours than plasma. Nat Commun. 2015;6:8839.
Nikanjam M, Kato S, Kurzrock R. Liquid biopsy: current technology and clinical applications. J Hematol Oncol. 2022;15:131.
Oshi M, Murthy V, Takahashi H, Huyser M, Okano M, Tokumaru Y, Rashid OM, Matsuyama R, Endo I, Takabe K. Urine as a Source of Liquid Biopsy for Cancer. Cancers (Basel). 2021;13(11):2652.
Zygulska AL, Pierzchalski P. Novel Diagnostic Biomarkers in Colorectal Cancer. Int J Mol Sci. 2022;23(2):852.
Yeo D, Bastian A, Strauss H, Saxena P, Grimison P, Rasko JEJ. Exploring the Clinical Utility of Pancreatic Cancer Circulating Tumor Cells. Int J Mol Sci. 2022;23(3):1671.
Khoja L, Backen A, Sloane R, Menasce L, Ryder D, Krebs M, Board R, Clack G, Hughes A, Blackhall F, et al. A pilot study to explore circulating tumour cells in pancreatic cancer as a novel biomarker. Br J Cancer. 2012;106:508–16.
Tao L, Su L, Yuan C, Ma Z, Zhang L, Bo S, Niu Y, Lu S, **u D. Postoperative metastasis prediction based on portal vein circulating tumor cells detected by flow cytometry in periampullary or pancreatic cancer. Cancer Manag Res. 2019;11:7405–25.
Nagrath S, Sequist LV, Maheswaran S, Bell DW, Irimia D, Ulkus L, Smith MR, Kwak EL, Digumarthy S, Muzikansky A, et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature. 2007;450:1235–9.
Semaan A, Bernard V, Kim DU, Lee JJ, Huang J, Kamyabi N, Stephens BM, Qiao W, Varadhachary GR, Katz MH, et al. Characterisation of circulating tumour cell phenotypes identifies a partial-EMT sub-population for clinical stratification of pancreatic cancer. Br J Cancer. 2021;124:1970–7.
Martini V, Timme-Bronsert S, Fichtner-Feigl S, Hoeppner J, Kulemann B. Circulating Tumor Cells in Pancreatic Cancer: Current Perspectives. Cancers (Basel). 2019;11(11):1659.
Yeo D, Kao S, Gupta R, Wahlroos S, Bastian A, Strauss H, Klemm V, Shrestha P, Ramirez AB, Costandy L, et al. Accurate isolation and detection of circulating tumor cells using enrichment-free multiparametric high resolution imaging. Front Oncol. 2023;13:1141228.
Sergeant G, Roskams T, van Pelt J, Houtmeyers F, Aerts R, Topal B. Perioperative cancer cell dissemination detected with a real-time RT-PCR assay for EpCAM is not associated with worse prognosis in pancreatic ductal adenocarcinoma. BMC Cancer. 2011;11:47.
Kulemann B, Rösch S, Seifert S, Timme S, Bronsert P, Seifert G, Martini V, Kuvendjiska J, Glatz T, Hussung S, et al. Pancreatic cancer: Circulating Tumor Cells and Primary Tumors show Heterogeneous KRAS Mutations. Sci Rep. 2017;7:4510.
Yu J, Gemenetzis G, Kinny-Köster B, Habib JR, Groot VP, Teinor J, Yin L, Pu N, Hasanain A, van Oosten F, et al. Pancreatic circulating tumor cell detection by targeted single-cell next-generation sequencing. Cancer Lett. 2020;493:245–53.
Ozkumur E, Shah AM, Ciciliano JC, Emmink BL, Miyamoto DT, Brachtel E, Yu M, Chen PI, Morgan B, Trautwein J, et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci Transl Med. 2013;5:179ra147.
Liao C-J, Hsieh C-H, Wang H-M, Chou W-P, Chiu T-K, Chang J-H, Chao AC, Wu M-H. Isolation of label-free and viable circulating tumour cells (CTCs) from blood samples of cancer patients through a two-step process: negative selection-type immunomagnetic beads and spheroid cell culture-based cell isolation. RSC Adv. 2017;7:29339–49.
Watanabe F, Suzuki K, Tamaki S, Abe I, Endo Y, Takayama Y, Ishikawa H, Kakizawa N, Saito M, Futsuhara K, et al. Longitudinal monitoring of KRAS-mutated circulating tumor DNA enables the prediction of prognosis and therapeutic responses in patients with pancreatic cancer. PLoS ONE. 2019;14:e0227366.
Mauger F, How-Kit A, Tost J. COLD-PCR Technologies in the Area of Personalized Medicine: Methodology and Applications. Mol Diagn Ther. 2017;21:269–83.
Semrad T, Barzi A, Lenz HJ, Hutchins IM, Kim EJ, Gong IY, Tanaka M, Beckett L, Holland W, Burich RA, et al. Pharmacodynamic separation of gemcitabine and erlotinib in locally advanced or metastatic pancreatic cancer: therapeutic and biomarker results. Int J Clin Oncol. 2015;20:518–24.
Toledano-Fonseca M, Cano MT, Inga E, Rodríguez-Alonso R, Gómez-España MA, Guil-Luna S, Mena-Osuna R, de la Haba-Rodríguez JR, Rodríguez-Ariza A, Aranda E. Circulating Cell-Free DNA-Based Liquid Biopsy Markers for the Non-Invasive Prognosis and Monitoring of Metastatic Pancreatic Cancer. Cancers (Basel). 2020;12(7):1754.
Cheng H, Liu C, Jiang J, Luo G, Lu Y, ** K, Guo M, Zhang Z, Xu J, Liu L, et al. Analysis of ctDNA to predict prognosis and monitor treatment responses in metastatic pancreatic cancer patients. Int J Cancer. 2017;140:2344–50.
Davis AA, Iams WT, Chan D, Oh MS, Lentz RW, Peterman N, Robertson A, Shah A, Srivas R, Wilson TJ, et al. Early Assessment of Molecular Progression and Response by Whole-genome Circulating Tumor DNA in Advanced Solid Tumors. Mol Cancer Ther. 2020;19:1486–96.
Pisanic TR 2nd, Athamanolap P, Poh W, Chen C, Hulbert A, Brock MV, Herman JG, Wang TH. DREAMing: a simple and ultrasensitive method for assessing intratumor epigenetic heterogeneity directly from liquid biopsies. Nucleic Acids Res. 2015;43:e154.
Li J, Wei L, Zhang X, Zhang W, Wang H, Zhong B, **e Z, Lv H, Wang X. DISMIR: Deep learning-based noninvasive cancer detection by integrating DNA sequence and methylation information of individual cell-free DNA reads. Brief Bioinform. 2021;22(6):bbab250.
Badowski C, He B, Garmire LX. Blood-derived lncRNAs as biomarkers for cancer diagnosis: the Good, the Bad and the Beauty. NPJ Precis Oncol. 2022;6:40.
Müller S, Raulefs S, Bruns P, Afonso-Grunz F, Plötner A, Thermann R, Jäger C, Schlitter AM, Kong B, Regel I, et al. Next-generation sequencing reveals novel differentially regulated mRNAs, lncRNAs, miRNAs, sdRNAs and a piRNA in pancreatic cancer. Mol Cancer. 2015;14:94.
Fu XL, Liu DJ, Yan TT, Yang JY, Yang MW, Li J, Huo YM, Liu W, Zhang JF, Hong J, et al. Analysis of long non-coding RNA expression profiles in pancreatic ductal adenocarcinoma. Sci Rep. 2016;6:33535.
Martino S, Tammaro C, Misso G, Falco M, Scrima M, Bocchetti M, Rea I, De Stefano L, Caraglia M. microRNA Detection via Nanostructured Biochips for Early Cancer Diagnostics. Int J Mol Sci. 2023;24(9):7762.
Wang X, Luo G, Zhang K, Cao J, Huang C, Jiang T, Liu B, Su L, Qiu Z. Hypoxic Tumor-Derived Exosomal miR-301a Mediates M2 Macrophage Polarization via PTEN/PI3Kγ to Promote Pancreatic Cancer Metastasis. Cancer Res. 2018;78:4586–98.
Xu YF, Xu X, Bhandari K, Gin A, Rao CV, Morris KT, Hannafon BN, Ding WQ. Isolation of extra-cellular vesicles in the context of pancreatic adenocarcinomas: Addition of one stringent filtration step improves recovery of specific microRNAs. PLoS ONE. 2021;16:e0259563.
Buenafe AC, Dorrell C, Reddy AP, Klimek J, Marks DL. Proteomic analysis distinguishes extracellular vesicles produced by cancerous versus healthy pancreatic organoids. Sci Rep. 2022;12:3556.
Stranska R, Gysbrechts L, Wouters J, Vermeersch P, Bloch K, Dierickx D, Andrei G, Snoeck R. Comparison of membrane affinity-based method with size-exclusion chromatography for isolation of exosome-like vesicles from human plasma. J Transl Med. 2018;16:1.
Chen K, Amontree J, Varillas J, Zhang J, George TJ, Fan ZH. Incorporation of lateral microfiltration with immunoaffinity for enhancing the capture efficiency of rare cells. Sci Rep. 2020;10:14210.
Ludwig AK, De Miroschedji K, Doeppner TR, Börger V, Ruesing J, Rebmann V, Durst S, Jansen S, Bremer M, Behrmann E, et al. Precipitation with polyethylene glycol followed by washing and pelleting by ultracentrifugation enriches extracellular vesicles from tissue culture supernatants in small and large scales. J Extracell Vesicles. 2018;7:1528109.
Zhao J, Guo M, Song Y, Liu S, Liao R, Zhang Y, Zhang Y, Yang Q, Gu Y, Huang X. Serum exosomal and serum glypican-1 are associated with early recurrence of pancreatic ductal adenocarcinoma. Front Oncol. 2022;12:992929.
Shao H, Im H, Castro CM, Breakefield X, Weissleder R, Lee H. New Technologies for Analysis of Extracellular Vesicles. Chem Rev. 2018;118:1917–50.
Li J, Li Y, Chen S, Duan W, Kong X, Wang Y, Zhou L, Li P, Zhang C, Du L, Wang C. Highly Sensitive Exosome Detection for Early Diagnosis of Pancreatic Cancer Using Immunoassay Based on Hierarchical Surface-Enhanced Raman Scattering Substrate. Small Methods. 2022;6:e2200154.
Tatischeff I, Larquet E, Falcón-Pérez JM, Turpin PY, Kruglik SG. Fast characterisation of cell-derived extracellular vesicles by nanoparticles tracking analysis, cryo-electron microscopy, and Raman tweezers microspectroscopy. J Extracell Vesicles. 2012;1. https://doi.org/10.3402/jev.v1i0.19179.
Li S, Ma Q. Electrochemical nano-sensing interface for exosomes analysis and cancer diagnosis. Biosens Bioelectron. 2022;214:114554.
Bellassai N, D’Agata R, Jungbluth V, Spoto G. Surface Plasmon Resonance for Biomarker Detection: Advances in Non-invasive Cancer Diagnosis. Front Chem. 2019;7:570.
Yu D, Li Y, Wang M, Gu J, Xu W, Cai H, Fang X, Zhang X. Exosomes as a new frontier of cancer liquid biopsy. Mol Cancer. 2022;21:56.
Lewis JM, Vyas AD, Qiu Y, Messer KS, White R, Heller MJ. Integrated Analysis of Exosomal Protein Biomarkers on Alternating Current Electrokinetic Chips Enables Rapid Detection of Pancreatic Cancer in Patient Blood. ACS Nano. 2018;12:3311–20.
Kowalik A, Kowalewska M, Góźdź S. Current approaches for avoiding the limitations of circulating tumor cells detection methods-implications for diagnosis and treatment of patients with solid tumors. Transl Res. 2017;185:58-84.e15.
Agashe R, Kurzrock R. Circulating Tumor Cells: From the Laboratory to the Cancer Clinic. Cancers (Basel). 2020;12(9):2361.
Castro-Giner F, Aceto N. Tracking cancer progression: from circulating tumor cells to metastasis. Genome Med. 2020;12:31.
Tayoun T, Faugeroux V, Oulhen M, Aberlenc A, Pawlikowska P, Farace F. CTC-Derived Models: A Window into the Seeding Capacity of Circulating Tumor Cells (CTCs). Cells. 2019;8(10):1145.
Aceto N, Toner M, Maheswaran S, Haber DA. En Route to Metastasis: Circulating Tumor Cell Clusters and Epithelial-to-Mesenchymal Transition. Trends Cancer. 2015;1:44–52.
Mavroudis D. Circulating cancer cells. Ann Oncol. 2010;21(Suppl 7):vii95-100.
Lin D, Shen L, Luo M, Zhang K, Li J, Yang Q, Zhu F, Zhou D, Zheng S, Chen Y, Zhou J. Circulating tumor cells: biology and clinical significance. Signal Transduct Target Ther. 2021;6:404.
Dirix L, Buys A, Oeyen S, Peeters D, Liègeois V, Prové A, Rondas D, Vervoort L, Mariën V, Laere SV, Vermeulen P. Circulating tumor cell detection: A prospective comparison between Cell Search® and RareCyte® platforms in patients with progressive metastatic breast cancer. Breast Cancer Res Treat. 2022;193:437–44.
Gruijs M, Zeelen C, Hellingman T, Smit J, Borm FJ, Kazemier G, Dickhoff C, Bahce I, de Langen J, Smit EF, et al. Detection of Circulating Tumor Cells Using the Attune NxT. Int J Mol Sci. 2022;24(1):21.
Rajta I, Huszánk R, Szabó AT, Nagy GU, Szilasi S, Fürjes P, Holczer E, Fekete Z, Járvás G, Szigeti M, et al. Tilted pillar array fabrication by the combination of proton beam writing and soft lithography for microfluidic cell capture: Part 1 Design and feasibility. Electrophoresis. 2016;37:498–503.
Pahattuge TN, Freed IM, Hupert ML, Vaidyanathan S, Childers K, Witek MA, Weerakoon-Ratnayake K, Park D, Kasi A, Al-Kasspooles MF, et al. System Modularity Chip for Analysis of Rare Targets (SMART-Chip): Liquid Biopsy Samples. ACS Sens. 2021;6:1831–9.
Lin HK, Zheng S, Williams AJ, Balic M, Groshen S, Scher HI, Fleisher M, Stadler W, Datar RH, Tai YC, Cote RJ. Portable filter-based microdevice for detection and characterization of circulating tumor cells. Clin Cancer Res. 2010;16:5011–8.
Hosokawa M, Yoshikawa T, Negishi R, Yoshino T, Koh Y, Kenmotsu H, Naito T, Takahashi T, Yamamoto N, Kikuhara Y, et al. Microcavity array system for size-based enrichment of circulating tumor cells from the blood of patients with small-cell lung cancer. Anal Chem. 2013;85:5692–8.
Lowes LE, Allan AL. Circulating Tumor Cells and Implications of the Epithelial-to-Mesenchymal Transition. Adv Clin Chem. 2018;83:121–81.
Couto-Cunha A, Jerónimo C, Henrique R. Circulating Tumor Cells as Biomarkers for Renal Cell Carcinoma: Ready for Prime Time? Cancers (Basel). 2022;15(1):287.
Lee DH, Yoon W, Lee A, Han Y, Byun Y, Kang JS, Kim H, Kwon W, Suh YA, Choi Y, et al. Multi-biomarker panel prediction model for diagnosis of pancreatic cancer. J Hepatobiliary Pancreat Sci. 2023;30:122–32.
Gao Y, Ni X, Guo H, Su Z, Ba Y, Tong Z, Guo Z, Yao X, Chen X, Yin J, et al. Single-cell sequencing deciphers a convergent evolution of copy number alterations from primary to circulating tumor cells. Genome Res. 2017;27:1312–22.
Keller L, Pantel K. Unravelling tumour heterogeneity by single-cell profiling of circulating tumour cells. Nat Rev Cancer. 2019;19:553–67.
Chen J, Wang H, Zhou L, Liu Z, Tan X. A combination of circulating tumor cells and CA199 improves the diagnosis of pancreatic cancer. J Clin Lab Anal. 2022;36:e24341.
Cheng H, He W, Yang J, Ye Q, Cheng L, Pan Y, Mao L, Chu X, Lu C, Li G, et al. Ligand-targeted polymerase chain reaction for the detection of folate receptor-positive circulating tumour cells as a potential diagnostic biomarker for pancreatic cancer. Cell Prolif. 2020;53:e12880.
Buscail E, Alix-Panabières C, Quincy P, Cauvin T, Chauvet A, Degrandi O, Caumont C, Verdon S, Lamrissi I, Moranvillier I, et al. High Clinical Value of Liquid Biopsy to Detect Circulating Tumor Cells and Tumor Exosomes in Pancreatic Ductal Adenocarcinoma Patients Eligible for Up-Front Surgery. Cancers (Basel). 2019;11(11):1656.
Ankeny JS, Court CM, Hou S, Li Q, Song M, Wu D, Chen JF, Lee T, Lin M, Sho S, et al. Circulating tumour cells as a biomarker for diagnosis and staging in pancreatic cancer. Br J Cancer. 2016;114:1367–75.
Sefrioui D, Blanchard F, Toure E, Beaussire L, Dolfus C, Perdrix A, Paresy M, Antonietti M, Iwanicki-Caron I, et al. Diagnostic value of CA19.9, circulating tumour DNA and circulating tumour cells in patients with solid pancreatic tumours. Br J Cancer. 2017;117:1017–25.
Zhu P, Liu HY, Liu FC, Gu FM, Yuan SX, Huang J, Pan ZY, Wang WJ. Circulating Tumor Cells Expressing Krüppel-Like Factor 8 and Vimentin as Predictors of Poor Prognosis in Pancreatic Cancer Patients. Cancer Control. 2021;28:10732748211027164.
Court CM, Ankeny JS, Sho S, Winograd P, Hou S, Song M, Wainberg ZA, Girgis MD, Graeber TG, Agopian VG, et al. Circulating Tumor Cells Predict Occult Metastatic Disease and Prognosis in Pancreatic Cancer. Ann Surg Oncol. 2018;25:1000–8.
Poruk KE, Blackford AL, Weiss MJ, Cameron JL, He J, Goggins M, Rasheed ZA, Wolfgang CL, Wood LD. Circulating Tumor Cells Expressing Markers of Tumor-Initiating Cells Predict Poor Survival and Cancer Recurrence in Patients with Pancreatic Ductal Adenocarcinoma. Clin Cancer Res. 2017;23:2681–90.
Poruk KE, Valero V 3rd, Saunders T, Blackford AL, Griffin JF, Poling J, Hruban RH, Anders RA, Herman J, Zheng L, et al. Circulating Tumor Cell Phenotype Predicts Recurrence and Survival in Pancreatic Adenocarcinoma. Ann Surg. 2016;264:1073–81.
Park Y, Jun HR, Choi HW, Hwang DW, Lee JH, Song KB, Lee W, Kwon J, Ha SH, Jun E, Kim SC. Circulating tumour cells as an indicator of early and systemic recurrence after surgical resection in pancreatic ductal adenocarcinoma. Sci Rep. 2021;11:1644.
Wei T, Zhang X, Zhang Q, Yang J, Chen Q, Wang J, Li X, Chen J, Ma T, Li G, et al. Vimentin-positive circulating tumor cells as a biomarker for diagnosis and treatment monitoring in patients with pancreatic cancer. Cancer Lett. 2019;452:237–43.
Gemenetzis G, Groot VP, Yu J, Ding D, Teinor JA, Javed AA, Wood LD, Burkhart RA, Cameron JL, Makary MA, et al. Circulating Tumor Cells Dynamics in Pancreatic Adenocarcinoma Correlate With Disease Status: Results of the Prospective CLUSTER Study. Ann Surg. 2018;268:408–20.
Dhayat SA, Yang Z. Impact of circulating tumor DNA in hepatocellular and pancreatic carcinomas. J Cancer Res Clin Oncol. 2020;146:1625–45.
Yong E. Cancer biomarkers: Written in blood. Nature. 2014;511:524–6.
Li YZ, Kong SN, Liu YP, Yang Y, Zhang HM. Can Liquid Biopsy Based on ctDNA/cfDNA Replace Tissue Biopsy for the Precision Treatment of EGFR-Mutated NSCLC? J Clin Med. 2023;12(4):1438.
Campos-Carrillo A, Weitzel JN, Sahoo P, Rockne R, Mokhnatkin JV, Murtaza M, Gray SW, Goetz L, Goel A, Schork N, Slavin TP. Circulating tumor DNA as an early cancer detection tool. Pharmacol Ther. 2020;207:107458.
Sausen M, Parpart S, Diaz LA Jr. Circulating tumor DNA moves further into the spotlight. Genome Med. 2014;6:35.
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, LeBleu VS, Mittendorf EA, Weitz J, Rahbari N, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523:177–82.
Lee VH, Kwong DL, Leung TW, Choi CW, Lai V, Ng L, Lam KO, Ng SC, Sze CK, Tong CC, et al. Prognostication of serial post-intensity-modulated radiation therapy undetectable plasma EBV DNA for nasopharyngeal carcinoma. Oncotarget. 2017;8:5292–308.
Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, Thornton K, Agrawal N, Sokoll L, Szabo SA, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985–90.
Grønbaek K, Hother C, Jones PA. Epigenetic changes in cancer. APMIS. 2007;115:1039–59.
Wen X, Pu H, Liu Q, Guo Z, Luo D. Circulating Tumor DNA-A Novel Biomarker of Tumor Progression and Its Favorable Detection Techniques. Cancers (Basel). 2022;14(24):6025.
Raufi AG, May MS, Hadfield MJ, Seyhan AA, El-Deiry WS. Advances in Liquid Biopsy Technology and Implications for Pancreatic Cancer. Int J Mol Sci. 2023;24(4):4238.
Heredia-Soto V, Rodríguez-Salas N, Feliu J. Liquid Biopsy in Pancreatic Cancer: Are We Ready to Apply It in the Clinical Practice?. Cancers (Basel). 2021;13(8):1986.
Kinde I, Wu J, Papadopoulos N, Kinzler KW, Vogelstein B. Detection and quantification of rare mutations with massively parallel sequencing. Proc Natl Acad Sci U S A. 2011;108:9530–5.
Hon GC, Hawkins RD, Caballero OL, Lo C, Lister R, Pelizzola M, Valsesia A, Ye Z, Kuan S, Edsall LE, et al. Global DNA hypomethylation coupled to repressive chromatin domain formation and gene silencing in breast cancer. Genome Res. 2012;22:246–58.
Castells A, Puig P, Móra J, Boadas J, Boix L, Urgell E, Solé M, Capellà G, Lluís F, Fernández-Cruz L, et al. K-ras mutations in DNA extracted from the plasma of patients with pancreatic carcinoma: diagnostic utility and prognostic significance. J Clin Oncol. 1999;17:578–84.
Razavi P, Li BT, Brown DN, Jung B, Hubbell E, Shen R, Abida W, Juluru K, De Bruijn I, Hou C, et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat Med. 2019;25:1928–37.
Wang R, Zhao Y, Wang Y, Zhao Z, Chen Q, Duan Y, **ong S, Luan Z, Wang J, Cheng B. Diagnostic and Prognostic Values of KRAS Mutations on EUS-FNA Specimens and Circulating Tumor DNA in Patients With Pancreatic Cancer. Clin Transl Gastroenterol. 2022;13:e00487.
Eissa MAL, Lerner L, Abdelfatah E, Shankar N, Canner JK, Hasan NM, Yaghoobi V, Huang B, Kerner Z, Takaesu F, et al. Promoter methylation of ADAMTS1 and BNC1 as potential biomarkers for early detection of pancreatic cancer in blood. Clin Epigenetics. 2019;11:59.
Henriksen SD, Madsen PH, Larsen AC, Johansen MB, Drewes AM, Pedersen IS, Krarup H, Thorlacius-Ussing O. Cell-free DNA promoter hypermethylation in plasma as a diagnostic marker for pancreatic adenocarcinoma. Clin Epigenetics. 2016;8:117.
Nitschke C, Markmann B, Walter P, Badbaran A, Tölle M, Kropidlowski J, Belloum Y, Goetz MR, Bardenhagen J, Stern L, et al. Peripheral and Portal Venous KRAS ctDNA Detection as Independent Prognostic Markers of Early Tumor Recurrence in Pancreatic Ductal Adenocarcinoma. Clin Chem. 2023;69:295–307.
Watanabe K, Nakamura T, Kimura Y, Motoya M, Kojima S, Kuraya T, Murakami T, Kaneko T, Shinohara Y, Kitayama Y, et al. Tumor-Informed Approach Improved ctDNA Detection Rate in Resected Pancreatic Cancer. Int J Mol Sci. 2022;23(19):11521.
Bernard V, Kim DU, San Lucas FA, Castillo J, Allenson K, Mulu FC, Stephens BM, Huang J, Semaan A, Guerrero PA, et al. Circulating Nucleic Acids Are Associated With Outcomes of Patients With Pancreatic Cancer. Gastroenterology. 2019;156:108-118.e104.
Kirchweger P, Kupferthaler A, Burghofer J, Webersinke G, Jukic E, Schwendinger S, Wundsam H, Biebl M, Petzer A, Rumpold H. Prediction of response to systemic treatment by kinetics of circulating tumor DNA in metastatic pancreatic cancer. Front Oncol. 2022;12:902177.
Sugimori M, Sugimori K, Tsuchiya H, Suzuki Y, Tsuyuki S, Kaneta Y, Hirotani A, Sanga K, Tozuka Y, Komiyama S, et al. Quantitative monitoring of circulating tumor DNA in patients with advanced pancreatic cancer undergoing chemotherapy. Cancer Sci. 2020;111:266–78.
Lee B, Lipton L, Cohen J, Tie J, Javed AA, Li L, Goldstein D, Burge M, Cooray P, Nagrial A, et al. Circulating tumor DNA as a potential marker of adjuvant chemotherapy benefit following surgery for localized pancreatic cancer. Ann Oncol. 2019;30:1472–8.
Takahashi K, Taniue K, Ono Y, Fujiya M, Mizukami Y, Okumura T. Long Non-Coding RNAs in Epithelial-Mesenchymal Transition of Pancreatic Cancer. Front Mol Biosci. 2021;8:717890.
Donati S, Aurilia C, Palmini G, Falsetti I, Iantomasi T, Brandi ML. Autophagy-Related ncRNAs in Pancreatic Cancer. Pharmaceuticals (Basel). 2022;15(12):1547.
Zhang M, Dang P, Liu Y, Qiao B, Sun Z. Noncoding RNAs in pyroptosis and cancer progression: Effect, mechanism, and clinical application. Front Immunol. 2022;13:982040.
Huynh NP, Anderson BA, Guilak F, McAlinden A. Emerging roles for long noncoding RNAs in skeletal biology and disease. Connect Tissue Res. 2017;58:116–41.
Aurilia C, Donati S, Palmini G, Miglietta F, Falsetti I, Iantomasi T, Brandi ML. Are Non-Coding RNAs Useful Biomarkers in Parathyroid Tumorigenesis? Int J Mol Sci. 2021;22(19):10465.
Smolkova B, Kataki A, Earl J, Ruz-Caracuel I, Cihova M, Urbanova M, Buocikova V, Tamargo S, Rovite V, Niedra H, et al. Liquid biopsy and preclinical tools for advancing diagnosis and treatment of patients with pancreatic neuroendocrine neoplasms. Crit Rev Oncol Hematol. 2022;180:103865.
Zen K, Zhang CY. Circulating microRNAs: a novel class of biomarkers to diagnose and monitor human cancers. Med Res Rev. 2012;32:326–48.
Galoș D, Gorzo A, Balacescu O, Sur D. Clinical Applications of Liquid Biopsy in Colorectal Cancer Screening: Current Challenges and Future Perspectives. Cells. 2022;11(21):3493.
Zandieh MA, Farahani MH, Rajabi R, Avval ST, Karimi K, Rahmanian P, Razzazan M, Javanshir S, Mirzaei S, Paskeh MDA, et al. Epigenetic regulation of autophagy by non-coding RNAs in gastrointestinal tumors: Biological functions and therapeutic perspectives. Pharmacol Res. 2023;187:106582.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–54.
Brockdorff N, Ashworth A, Kay GF, McCabe VM, Norris DP, Cooper PJ, Swift S, Rastan S. The product of the mouse **st gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell. 1992;71:515–26.
Mi Z, Zhongqiang C, Caiyun J, Yanan L, Jianhua W, Liang L. Circular RNA detection methods: A minireview. Talanta. 2022;238:123066.
Deng R, Zhang K, Li J. Isothermal Amplification for MicroRNA Detection: From the Test Tube to the Cell. Acc Chem Res. 2017;50:1059–68.
Zhao Y, Chen F, Li Q, Wang L, Fan C. Isothermal Amplification of Nucleic Acids. Chem Rev. 2015;115:12491–545.
Masud MK, Umer M, Hossain MSA, Yamauchi Y, Nguyen NT, Shiddiky MJA. Nanoarchitecture Frameworks for Electrochemical miRNA Detection. Trends Biochem Sci. 2019;44:433–52.
Zhao G, Yan X, Zhang Y, Deng J, Liang X. Sensitive detection of MiRNA and CircRNA through DSN enzyme cooperating NEase assisted dual signal amplification. Anal Biochem. 2022;654:114744.
Dittmar RL, Liu S, Tai MC, Rajapakshe K, Huang Y, Longton G, DeCapite C, Hurd MW, Paris PL, Kirkwood KS, et al. Plasma miRNA Biomarkers in Limited Volume Samples for Detection of Early-stage Pancreatic Cancer. Cancer Prev Res (Phila). 2021;14:729–40.
Lou UK, Wong CH, Chen Y. A simple and rapid colorimetric detection of serum lncRNA biomarkers for diagnosis of pancreatic cancer. RSC Adv. 2020;10:8087–92.
Nicoletti A, Negri M, Paratore M, Vitale F, Ainora ME, Nista EC, Gasbarrini A, Zocco MA. Zileri Dal Verme L: Diagnostic and Prognostic Role of Extracellular Vesicles in Pancreatic Cancer: Current Evidence and Future Perspectives. Int J Mol Sci. 2023;24(1):885.
Li SR, Man QW, Gao X, Lin H, Wang J, Su FC, Wang HQ, Bu LL, Liu B, Chen G. Tissue-derived extracellular vesicles in cancers and non-cancer diseases: Present and future. J Extracell Vesicles. 2021;10:e12175.
Doyle LM, Wang MZ. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells. 2019;8(7):727.
Qian F, Huang Z, Zhong H, Lei Q, Ai Y, **e Z, Zhang T, Jiang B, Zhu W, Sheng Y, et al. Analysis and Biomedical Applications of Functional Cargo in Extracellular Vesicles. ACS Nano. 2022;16:19980–20001.
Théry C, Witwer KW, Aikawa E, Alcaraz MJ, Anderson JD, Andriantsitohaina R, Antoniou A, Arab T, Archer F, Atkin-Smith GK, et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles. 2018;7:1535750.
Dai J, Su Y, Zhong S, Cong L, Liu B, Yang J, Tao Y, He Z, Chen C, Jiang Y. Exosomes: key players in cancer and potential therapeutic strategy. Signal Transduct Target Ther. 2020;5:145.
Johnstone RM, Adam M, Hammond JR, Orr L, Turbide C. Vesicle formation during reticulocyte maturation. Association of plasma membrane activities with released vesicles (exosomes). J Biol Chem. 1987;262:9412–20.
Zhou B, Xu K, Zheng X, Chen T, Wang J, Song Y, Shao Y, Zheng S. Application of exosomes as liquid biopsy in clinical diagnosis. Signal Transduct Target Ther. 2020;5:144.
Yu W, Hurley J, Roberts D, Chakrabortty SK, Enderle D, Noerholm M, Breakefield XO, Skog JK. Exosome-based liquid biopsies in cancer: opportunities and challenges. Ann Oncol. 2021;32:466–77.
Keller S, Ridinger J, Rupp AK, Janssen JW, Altevogt P. Body fluid derived exosomes as a novel template for clinical diagnostics. J Transl Med. 2011;9:86.
Yu J, Ostowari A, Gonda A, Mashayekhi K, Dayyani F, Hughes CCW, Senthil M. Exosomes as a Source of Biomarkers for Gastrointestinal Cancers. Cancers (Basel). 2023;15(4):1263.
Mathieu M, Martin-Jaular L, Lavieu G, Théry C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat Cell Biol. 2019;21:9–17.
Irmer B, Chandrabalan S, Maas L, Bleckmann A, Menck K. Extracellular Vesicles in Liquid Biopsies as Biomarkers for Solid Tumors. Cancers (Basel). 2023;15(4):1307.
Zeringer E, Barta T, Li M, Vlassov AV. Strategies for isolation of exosomes. Cold Spring Harb Protoc. 2015;2015:319–23.
Yang F, Liao X, Tian Y, Li G. Exosome separation using microfluidic systems: size-based, immunoaffinity-based and dynamic methodologies. Biotechnol J. 2017;12(4). https://doi.org/10.1002/biot.201600699.
Liu J, Chen Y, Pei F, Zeng C, Yao Y, Liao W, Zhao Z. Extracellular Vesicles in Liquid Biopsies: Potential for Disease Diagnosis. Biomed Res Int. 2021;2021:6611244.
Zheng L, Wang H, Zuo P, Liu Y, Xu H, Ye BC. Rapid On-Chip Isolation of Cancer-Associated Exosomes and Combined Analysis of Exosomes and Exosomal Proteins. Anal Chem. 2022;94:7703–12.
Armakolas A, Kotsari M, Koskinas J. Liquid Biopsies, Novel Approaches and Future Directions. Cancers (Basel). 2023;15(5):1579.
Lee M, Kim J, Jang M, Park C, Lee JH, Lee T. Introduction of Nanomaterials to Biosensors for Exosome Detection: Case Study for Cancer Analysis. Biosensors (Basel). 2022;12(8):648.
Machida T, Tomofuji T, Maruyama T, Yoneda T, Ekuni D, Azuma T, Miyai H, Mizuno H, Kato H, Tsutsumi K, et al. miR-1246 and miR-4644 in salivary exosome as potential biomarkers for pancreatobiliary tract cancer. Oncol Rep. 2016;36:2375–81.
Goto T, Fujiya M, Konishi H, Sasajima J, Fujibayashi S, Hayashi A, Utsumi T, Sato H, Iwama T, Ijiri M, et al. An elevated expression of serum exosomal microRNA-191, - 21, -451a of pancreatic neoplasm is considered to be efficient diagnostic marker. BMC Cancer. 2018;18:116.
Nakamura S, Sadakari Y, Ohtsuka T, Okayama T, Nakashima Y, Gotoh Y, Saeki K, Mori Y, Nakata K, Miyasaka Y, et al. Pancreatic Juice Exosomal MicroRNAs as Biomarkers for Detection of Pancreatic Ductal Adenocarcinoma. Ann Surg Oncol. 2019;26:2104–11.
Xavier CPR, Castro I, Caires HR, Ferreira D, Cavadas B, Pereira L, Santos LL, Oliveira MJ, Vasconcelos MH. Chitinase 3-like-1 and fibronectin in the cargo of extracellular vesicles shed by human macrophages influence pancreatic cancer cellular response to gemcitabine. Cancer Lett. 2021;501:210–23.
Lu H, Lu S, Yang D, Zhang L, Ye J, Li M, Hu W. MiR-20a-5p regulates gemcitabine chemosensitivity by targeting RRM2 in pancreatic cancer cells and serves as a predictor for gemcitabine-based chemotherapy. Biosci Rep. 2019;39(5):BSR20181374.
Takahasi K, Iinuma H, Wada K, Minezaki S, Kawamura S, Kainuma M, Ikeda Y, Shibuya M, Miura F, Sano K. Usefulness of exosome-encapsulated microRNA-451a as a minimally invasive biomarker for prediction of recurrence and prognosis in pancreatic ductal adenocarcinoma. J Hepatobiliary Pancreat Sci. 2018;25:155–61.
Kawamura S, Iinuma H, Wada K, Takahashi K, Minezaki S, Kainuma M, Shibuya M, Miura F, Sano K. Exosome-encapsulated microRNA-4525, microRNA-451a and microRNA-21 in portal vein blood is a high-sensitive liquid biomarker for the selection of high-risk pancreatic ductal adenocarcinoma patients. J Hepatobiliary Pancreat Sci. 2019;26:63–72.
Sun H, Shi K, Qi K, Kong H, Zhang J, Dai S, Ye W, Deng T, He Q, Zhou M. Natural Killer Cell-Derived Exosomal miR-3607-3p Inhibits Pancreatic Cancer Progression by Targeting IL-26. Front Immunol. 2019;10:2819.
Li J, Li Z, Jiang P, Peng M, Zhang X, Chen K, Liu H, Bi H, Liu X, Li X. Circular RNA IARS (circ-IARS) secreted by pancreatic cancer cells and located within exosomes regulates endothelial monolayer permeability to promote tumor metastasis. J Exp Clin Cancer Res. 2018;37:177.
Zhang X, Zheng S, Hu C, Li G, Lin H, **a R, Ye Y, He R, Li Z, Lin Q, et al. Cancer-associated fibroblast-induced lncRNA UPK1A-AS1 confers platinum resistance in pancreatic cancer via efficient double-strand break repair. Oncogene. 2022;41:2372–89.
Chari ST. Detecting early pancreatic cancer: problems and prospects. Semin Oncol. 2007;34:284–94.
Yachida S, Jones S, Bozic I, Antal T, Leary R, Fu B, Kamiyama M, Hruban RH, Eshleman JR, Nowak MA, et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature. 2010;467:1114–7.
Choi JI, Jang SI, Hong J, Kim CH, Kwon SS, Park JS, Lim JB. Cancer-initiating cells in human pancreatic cancer organoids are maintained by interactions with endothelial cells. Cancer Lett. 2021;498:42–53.
Rhim AD, Mirek ET, Aiello NM, Maitra A, Bailey JM, McAllister F, Reichert M, Beatty GL, Rustgi AK, Vonderheide RH, et al. EMT and dissemination precede pancreatic tumor formation. Cell. 2012;148:349–61.
Rhim AD, Thege FI, Santana SM, Lannin TB, Saha TN, Tsai S, Maggs LR, Kochman ML, Ginsberg GG, Lieb JG, et al. Detection of circulating pancreas epithelial cells in patients with pancreatic cystic lesions. Gastroenterology. 2014;146:647–51.
Cauley CE, Pitman MB, Zhou J, Perkins J, Kuleman B, Liss AS, Fernandez-Del Castillo C, Warshaw AL, Lillemoe KD, Thayer SP. Circulating Epithelial Cells in Patients with Pancreatic Lesions: Clinical and Pathologic Findings. J Am Coll Surg. 2015;221:699–707.
Kuvendjiska J, Müller F, Bronsert P, Timme-Bronsert S, Fichtner-Feigl S, Kulemann B. Circulating Epithelial Cells in Patients with Intraductal Papillary Mucinous Neoplasm of the Pancreas. Life (Basel). 2023;13(7):1570.
Roa-Colomo A, López Garrido MÁ, Molina-Vallejo P, Rojas A, Sanchez MG, Aranda-García V, Salmeron J, Romero-Gomez M, Muntane J, Padillo J, et al. Hepatocellular carcinoma risk-stratification based on ASGR1 in circulating epithelial cells for cancer interception. Front Mol Biosci. 2022;9:1074277.
Lin HC, Liou MJ, Hsu HL, Hsieh JC, Chen YA, Tseng CP, Lin JD. Combined analysis of circulating epithelial cells and serum thyroglobulin for distinguishing disease status of the patients with papillary thyroid carcinoma. Oncotarget. 2016;7:17242–53.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, Kinzler KW. Cancer genome landscapes. Science. 2013;339:1546–58.
Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, Bartlett BR, Wang H, Luber B, Alani RM, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014;6(224):224ra24.
Cohen JD, Javed AA, Thoburn C, Wong F, Tie J, Gibbs P, Schmidt CM, Yip-Schneider MT, Allen PJ, Schattner M, et al. Combined circulating tumor DNA and protein biomarker-based liquid biopsy for the earlier detection of pancreatic cancers. Proc Natl Acad Sci U S A. 2017;114:10202–7.
Liu G, Shao C, Li A, Zhang X, Guo X, Li J. Diagnostic Value of Plasma miR-181b, miR-196a, and miR-210 Combination in Pancreatic Cancer. Gastroenterol Res Pract. 2020;2020:6073150.
Bartsch DK, Gercke N, Strauch K, Wieboldt R, Matthäi E, Wagner V, Rospleszcz S, Schäfer A, Franke FS, Mintziras I, et al. The Combination of MiRNA-196b, LCN2, and TIMP1 is a Potential Set of Circulating Biomarkers for Screening Individuals at Risk for Familial Pancreatic Cancer. J Clin Med. 2018;7(10):295.
Franklin O, Jonsson P, Billing O, Lundberg E, Öhlund D, Nyström H, Lundin C, Antti H, Sund M. Plasma Micro-RNA Alterations Appear Late in Pancreatic Cancer. Ann Surg. 2018;267:775–81.
Zhou X, Lu Z, Wang T, Huang Z, Zhu W, Miao Y. Plasma miRNAs in diagnosis and prognosis of pancreatic cancer: A miRNA expression analysis. Gene. 2018;673:181–93.
Vieira NF, Serafini LN, Novais PC, Neto FSL, Cirino MLA, Kemp R, Ardengh JC, Saggioro FP, Gaspar AF, Sankarankutty AK, et al. The role of circulating miRNAs and CA19-9 in pancreatic cancer diagnosis. Oncotarget. 2021;12:1638–50.
Yuan W, Tang W, **e Y, Wang S, Chen Y, Qi J, Qiao Y, Ma J. New combined microRNA and protein plasmatic biomarker panel for pancreatic cancer. Oncotarget. 2016;7:80033–45.
Wnuk J, Strzelczyk JK, Gisterek I. Clinical Value of Circulating miRNA in Diagnosis, Prognosis, Screening and Monitoring Therapy of Pancreatic Ductal Adenocarcinoma-A Review of the Literature. Int J Mol Sci. 2023;24(6):5113.
Sharma GG, Okada Y, Von Hoff D, Goel A. Non-coding RNA biomarkers in pancreatic ductal adenocarcinoma. Semin Cancer Biol. 2021;75:153–68.
Xu G, Ji Y, Wang L, Xu H, Shen C, Ye H, Yang X. M6A-Related Long Non-Coding RNA Displays Utility in Predicting Prognosis, Portraying the Tumor Immune Microenvironment and Guiding Immunotherapy in Pancreatic Ductal Adenocarcinoma. Vaccines (Basel). 2023;11(3):499.
Wu Q, Chen L, Miao D, ** Y, Zhu Z. Prognostic signature based on m6A-related lncRNAs to predict overall survival in pancreatic ductal adenocarcinoma. Sci Rep. 2022;12:3079.
Wu H, Wang B, Wang L, Xue Y. circular RNAs 0000515 and 0011385 as potential biomarkers for disease monitoring and determining prognosis in pancreatic ductal adenocarcinoma. Oncol Lett. 2022;23:56.
Xu Y, Yao Y, Gao P, Cui Y. Upregulated circular RNA circ_0030235 predicts unfavorable prognosis in pancreatic ductal adenocarcinoma and facilitates cell progression by sponging miR-1253 and miR-1294. Biochem Biophys Res Commun. 2019;509:138–42.
Zhang W, Campbell DH, Walsh BJ, Packer NH, Liu D, Wang Y. Cancer-derived small extracellular vesicles: emerging biomarkers and therapies for pancreatic ductal adenocarcinoma diagnosis/prognosis and treatment. J Nanobiotechnology. 2022;20:446.
Yu S, Li Y, Liao Z, Wang Z, Wang Z, Li Y, Qian L, Zhao J, Zong H, Kang B, et al. Plasma extracellular vesicle long RNA profiling identifies a diagnostic signature for the detection of pancreatic ductal adenocarcinoma. Gut. 2020;69:540–50.
Hsu SK, Jadhao M, Liao WT, Chang WT, Lin IL, Chiu CC. The Role of Exosomes in Pancreatic Ductal Adenocarcinoma Progression and Their Potential as Biomarkers. Cancers (Basel). 2023;15(6):1776.
Zeng S, Pöttler M, Lan B, Grützmann R, Pilarsky C, Yang H. Chemoresistance in Pancreatic Cancer. Int J Mol Sci. 2019;20(18):4504.
Kim H, Heo CM, Oh J, Chung HH, Lee EM, Park J, Lee SH, Lee KH, Lee KT, Lee JK, et al. Clinical significance of circulating tumor cells after chemotherapy in unresectable pancreatic ductal adenocarcinoma. Transl Oncol. 2022;16:101321.
Ju S, Chen C, Zhang J, Xu L, Zhang X, Li Z, Chen Y, Zhou J, Ji F, Wang L. Detection of circulating tumor cells: opportunities and challenges. Biomark Res. 2022;10:58.
Pan X, Zhang X. Utility of circulating tumor cells and DNA in the management of advanced colorectal cancer. Future Oncol. 2020;16:1289–99.
Beije N, Onstenk W, Kraan J, Sieuwerts AM, Hamberg P, Dirix LY, Brouwer A, de Jongh FE, Jager A, Seynaeve CM, et al. Prognostic Impact of HER2 and ER Status of Circulating Tumor Cells in Metastatic Breast Cancer Patients with a HER2-Negative Primary Tumor. Neoplasia. 2016;18:647–53.
Garrido-Navas MC, García-Díaz A, Molina-Vallejo MP, González-Martínez C, Alcaide Lucena M, Cañas-García I, Bayarri C, Delgado JR, González E, Lorente JA, Serrano MJ. The Polemic Diagnostic Role of TP53 Mutations in Liquid Biopsies from Breast, Colon and Lung Cancers. Cancers (Basel). 2020;12(11):3343.
Nicolazzo C, Raimondi C, Mancini M, Caponnetto S, Gradilone A, Gandini O, Mastromartino M, Del Bene G, Prete A, Longo F, et al. Monitoring PD-L1 positive circulating tumor cells in non-small cell lung cancer patients treated with the PD-1 inhibitor Nivolumab. Sci Rep. 2016;6:31726.
Fusi A, Liu Z, Kümmerlen V, Nonnemacher A, Jeske J, Keilholz U. Expression of chemokine receptors on circulating tumor cells in patients with solid tumors. J Transl Med. 2012;10:52.
Gardner KP, Tsai S, Aldakkak M, Gironda S, Adams DL. CXCR4 expression in tumor associated cells in blood is prognostic for progression and survival in pancreatic cancer. PLoS ONE. 2022;17:e0264763.
Wu YH, Hung YP, Chiu NC, Lee RC, Li CP, Chao Y, Shyr YM, Wang SE, Chen SC, Lin SH, et al. Correlation between drug sensitivity profiles of circulating tumour cell-derived organoids and clinical treatment response in patients with pancreatic ductal adenocarcinoma. Eur J Cancer. 2022;166:208–18.
Starlinger P, Brugger P, Reiter C, Schauer D, Sommerfeldt S, Tamandl D, Kuehrer I, Schoppmann SF, Gnant M, Brostjan C. Discrimination between circulating endothelial cells and blood cell populations with overlap** phenotype reveals distinct regulation and predictive potential in cancer therapy. Neoplasia. 2011;13:980–90.
Sabbaghian MS, Rothberger G, Alongi AP, Gagner JP, Goldberg JD, Rolnitzky L, Chiriboga L, Hajdu CH, Zagzag D, Basch R, Shamamian P. Levels of elevated circulating endothelial cell decline after tumor resection in patients with pancreatic ductal adenocarcinoma. Anticancer Res. 2010;30:2911–7.
Dall’Olio FG, Marabelle A, Caramella C, Garcia C, Aldea M, Chaput N, Robert C, Besse B. Tumour burden and efficacy of immune-checkpoint inhibitors. Nat Rev Clin Oncol. 2022;19:75–90.
Wei T, Zhang Q, Li X, Su W, Li G, Ma T, Gao S, Lou J, Que R, Zheng L, et al. Monitoring Tumor Burden in Response to FOLFIRINOX Chemotherapy Via Profiling Circulating Cell-Free DNA in Pancreatic Cancer. Mol Cancer Ther. 2019;18:196–203.
Groot VP, Mosier S, Javed AA, Teinor JA, Gemenetzis G, Ding D, Haley LM, Yu J, Burkhart RA, Hasanain A, et al. Circulating Tumor DNA as a Clinical Test in Resected Pancreatic Cancer. Clin Cancer Res. 2019;25:4973–84.
van der Sijde F, Homs MYV, van Bekkum ML, van den Bosch TPP, Bosscha K, Besselink MG, Bonsing BA, de Groot JWB, Karsten TM, Groot Koerkamp B, et al. Serum miR-373–3p and miR-194–5p Are Associated with Early Tumor Progression during FOLFIRINOX Treatment in Pancreatic Cancer Patients: A Prospective Multicenter Study. Int J Mol Sci. 2021;22(20):10902.
Mikamori M, Yamada D, Eguchi H, Hasegawa S, Kishimoto T, Tomimaru Y, Asaoka T, Noda T, Wada H, Kawamoto K, et al. MicroRNA-155 Controls Exosome Synthesis and Promotes Gemcitabine Resistance in Pancreatic Ductal Adenocarcinoma. Sci Rep. 2017;7:42339.
Amaral MJ, Oliveira RC, Donato P, Tralhão JG. Pancreatic Cancer Biomarkers: Oncogenic Mutations, Tissue and Liquid Biopsies, and Radiomics-A Review. Dig Dis Sci. 2023;68:2811–23.
Hugenschmidt H, Labori KJ, Borgen E, Brunborg C, Schirmer CB, Seeberg LT, Naume B, Wiedswang G. Preoperative CTC-Detection by Cell Search(®) Is Associated with Early Distant Metastasis and Impaired Survival in Resected Pancreatic Cancer. Cancers (Basel). 2021;13(3):485.
Sun Y, Wu G, Cheng KS, Chen A, Neoh KH, Chen S, Tang Z, Lee PF, Dai M, Han RPS. CTC phenoty** for a preoperative assessment of tumor metastasis and overall survival of pancreatic ductal adenocarcinoma patients. EBioMedicine. 2019;46:133–49.
Guo S, Shi X, Shen J, Gao S, Wang H, Shen S, Pan Y, Li B, Xu X, Shao Z, ** G. Preoperative detection of KRAS G12D mutation in ctDNA is a powerful predictor for early recurrence of resectable PDAC patients. Br J Cancer. 2020;122:857–67.
Kirchweger P, Kupferthaler A, Burghofer J, Webersinke G, Jukic E, Schwendinger S, Weitzendorfer M, Petzer A, Függer R, Rumpold H, Wundsam H. Circulating tumor DNA correlates with tumor burden and predicts outcome in pancreatic cancer irrespective of tumor stage. Eur J Surg Oncol. 2022;48:1046–53.
DiPardo BJ, Winograd P, Court CM, Tomlinson JS. Pancreatic cancer circulating tumor cells: applications for personalized oncology. Expert Rev Mol Diagn. 2018;18:809–20.
Kamyabi N, Huang J, Lee JJ, Bernard V, Semaan A, Stephens B, Hurd MW, Vanapalli SA, Maitra A, Guerrero PA. A microfluidic device for label-free isolation of tumor cell clusters from unprocessed blood samples. Biomicrofluidics. 2019;13:044111.
Pan Y, Li D, Yang J, Wang N, **ao E, Tao L, Ding X, Sun P, Li D. Portal Venous Circulating Tumor Cells Undergoing Epithelial-Mesenchymal Transition Exhibit Distinct Clinical Significance in Pancreatic Ductal Adenocarcinoma. Front Oncol. 2021;11:757307.
Song BG, Kwon W, Kim H, Lee EM, Han YM, Kim H, Byun Y, Lee KB, Lee KH, Lee KT, et al. Detection of Circulating Tumor Cells in Resectable Pancreatic Ductal Adenocarcinoma: A Prospective Evaluation as a Prognostic Marker. Front Oncol. 2020;10:616440.
Nitschke C, Markmann B, Tölle M, Kropidlowski J, Belloum Y, Goetz MR, Schlüter H, Kwiatkowski M, Sinn M, Izbicki J, et al. Characterization of RARRES1 Expression on Circulating Tumor Cells as Unfavorable Prognostic Marker in Resected Pancreatic Ductal Adenocarcinoma Patients. Cancers (Basel). 2022;14(18):4405.
Mancuso P, Colleoni M, Calleri A, Orlando L, Maisonneuve P, Pruneri G, Agliano A, Goldhirsch A, Shaked Y, Kerbel RS, Bertolini F. Circulating endothelial-cell kinetics and viability predict survival in breast cancer patients receiving metronomic chemotherapy. Blood. 2006;108:452–9.
Bidard FC, Mathiot C, Degeorges A, Etienne-Grimaldi MC, Delva R, Pivot X, Veyret C, Bergougnoux L, de Cremoux P, Milano G, Pierga JY. Clinical value of circulating endothelial cells and circulating tumor cells in metastatic breast cancer patients treated first line with bevacizumab and chemotherapy. Ann Oncol. 2010;21:1765–71.
Strijbos MH, Gratama JW, Schmitz PI, Rao C, Onstenk W, Doyle GV, Miller MC, de Wit R, Terstappen LW, Sleijfer S. Circulating endothelial cells, circulating tumour cells, tissue factor, endothelin-1 and overall survival in prostate cancer patients treated with docetaxel. Eur J Cancer. 2010;46:2027–35.
Miglio U, Oldani A, Mezzapelle R, Veggiani C, Paganotti A, Garavoglia M, Boldorini R. KRAS mutational analysis in ductal adenocarcinoma of the pancreas and its clinical significance. Pathol Res Pract. 2014;210:307–11.
Ako S, Kato H, Nouso K, Kinugasa H, Terasawa H, Matushita H, Takada S, Saragai Y, Mizukawa S, Muro S, et al. Plasma KRAS mutations predict the early recurrence after surgical resection of pancreatic cancer. Cancer Biol Ther. 2021;22:564–70.
Hadano N, Murakami Y, Uemura K, Hashimoto Y, Kondo N, Nakagawa N, Sueda T, Hiyama E. Prognostic value of circulating tumour DNA in patients undergoing curative resection for pancreatic cancer. Br J Cancer. 2016;115:59–65.
Wang Y, Ji Y, Xu Q, Huang W. Prognostic N6-methyladenosine (m6A)-related lncRNA patterns to aid therapy in pancreatic ductal adenocarcinoma. Front Genet. 2022;13:866340.
Yang H, Zhang W, Ding J, Hu J, Sun Y, Peng W, Chu Y, **e L, Mei Z, Shao Z, **ao Y. A novel genomic instability-derived lncRNA signature to predict prognosis and immune characteristics of pancreatic ductal adenocarcinoma. Front Immunol. 2022;13:970588.
Kandimalla R, Shimura T, Mallik S, Sonohara F, Tsai S, Evans DB, Kim SC, Baba H, Kodera Y, Von Hoff D, et al. Identification of Serum miRNA Signature and Establishment of a Nomogram for Risk Stratification in Patients With Pancreatic Ductal Adenocarcinoma. Ann Surg. 2022;275:e229–37.
Reese M, Flammang I, Yang Z, Dhayat SA. Potential of Exosomal microRNA-200b as Liquid Biopsy Marker in Pancreatic Ductal Adenocarcinoma. Cancers (Basel). 2020;12(1):197.
Park SJ, Park JY, Shin K, Hong TH, Lee M, Kim Y, Kim IH. Clinical significance of serum-derived exosomal PD-L1 expression in patients with advanced pancreatic cancer. BMC Cancer. 2023;23:389.
Freed-Pastor WA, Lambert LJ, Ely ZA, Pattada NB, Bhutkar A, Eng G, Mercer KL, Garcia AP, Lin L, Rideout WM 3rd, et al. The CD155/TIGIT axis promotes and maintains immune evasion in neoantigen-expressing pancreatic cancer. Cancer Cell. 2021;39:1342-1360.e1314.
Murakami D, Matsuda K, Iwamoto H, Mitani Y, Mizumoto Y, Nakamura Y, Matsuzaki I, Iwamoto R, Takahashi Y, Kojima F, et al. Prognostic value of CD155/TIGIT expression in patients with colorectal cancer. PLoS ONE. 2022;17:e0265908.
Wang D, Gu Y, Yan X, Huo C, Wang G, Zhao Y, Teng M, Li Y. Role of CD155/TIGIT in Digestive Cancers: Promising Cancer Target for Immunotherapy. Front Oncol. 2022;12:844260.
Chang WH, Nguyen TT, Hsu CH, Bryant KL, Kim HJ, Ying H, Erickson JW, Der CJ, Cerione RA, Antonyak MA. KRAS-dependent cancer cells promote survival by producing exosomes enriched in Survivin. Cancer Lett. 2021;517:66–77.
Yang Z, LaRiviere MJ, Ko J, Till JE, Christensen T, Yee SS, Black TA, Tien K, Lin A, Shen H, et al. A Multianalyte Panel Consisting of Extracellular Vesicle miRNAs and mRNAs, cfDNA, and CA19-9 Shows Utility for Diagnosis and Staging of Pancreatic Ductal Adenocarcinoma. Clin Cancer Res. 2020;26:3248–58.
Satoh K. Molecular Approaches Using Body Fluid for the Early Detection of Pancreatic Cancer. Diagnostics (Basel). 2021;11(2):375.
Malara N, Kovacs G, Bussu F, Ferrazzo T, Garo V, Raso C, Cornacchione P, Iezzi R, Tagliaferri L. Liquid Biopsy-Guided Interventional Oncology: A Proof of Concept with a Special Focus on Radiotherapy and Radiology. Cancers (Basel). 2022;14(19):4676.
Vidal L, Pando E, Blanco L, Fabregat-Franco C, Castet F, Sierra A, Macarulla T, Balsells J, Charco R, Vivancos A. Liquid biopsy after resection of pancreatic adenocarcinoma and its relation to oncological outcomes. Systematic review and meta-analysis. Cancer Treat Rev. 2023;120:102604.
Acknowledgements
We would like to extend our sincere gratitude to Professor Ying Cheng and the reviewers for their invaluable and constructive comments provided during the manuscript revision process. The graphics in this article were supported and licensed by BioRender.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
ZB and PQ designed the study, WKC and ZB collected the related papers and drafted the manuscript, PQ and WX generated figures and critically reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Wang, K., Wang, X., Pan, Q. et al. Liquid biopsy techniques and pancreatic cancer: diagnosis, monitoring, and evaluation. Mol Cancer 22, 167 (2023). https://doi.org/10.1186/s12943-023-01870-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12943-023-01870-3