Abstract
TMEM16A (known as anoctamin 1) Ca2+-activated chloride channel is overexpressed in many tumors. TMEM16A overexpression can be caused by gene amplification in many tumors harboring 11q13 amplification. TMEM16A expression is also controlled in many cancer cells via transcriptional regulation, epigenetic regulation and microRNAs. In addition, TMEM16A activates different signaling pathways in different cancers, e.g. the EGFR and CAMKII signaling in breast cancer, the p38 and ERK1/2 signaling in hepatoma, the Ras-Raf-MEK-ERK1/2 signaling in head and neck squamous cell carcinoma and bladder cancer, and the NFκB signaling in glioma. Furthermore, TMEM16A overexpression has been reported to promote, inhibit, or produce no effects on cell proliferation and migration in different cancer cells. Since TMEM16A exerts different roles in different cancer cells via activation of distinct signaling pathways, we try to develop the idea that TMEM16A regulates cancer cell proliferation and migration in a cell-dependent mechanism. The cell-specific role of TMEM16A may depend on the cellular environment that is predetermined by TMEM16A overexpression mechanisms specific for a particular cancer type. TMEM16A may exert its cell-specific role via its associated protein networks, phosphorylation by different kinases, and involvement of different signaling pathways. In addition, we discuss the role of TMEM16A channel activity in cancer, and its clinical use as a prognostic and predictive marker in different cancers. This review highlights the cell-type specific mechanisms of TMEM16A in cancer, and envisions the promising use of TMEM16A inhibitors as a potential treatment for TMEM16A-overexpressing cancers.
Similar content being viewed by others
Background
TMEM16A (also known as anoctamin 1) was identified as a Ca2+-activated chloride channel (CaCC) in 2008 [1,2,3], and is the first member of the ten-member family of “Transmembrane protein 16” (abbreviated as TMEM16). Besides TMEM16A, TMEM16B and TMEM16F have been found to function as CaCCs [4,5,6]. TMEM16F can also function as a Ca2+-dependent phospholipid scramblase [7, 8], and Ca2+-activated nonselective cation channel [9]. However, controversies exist among other members of the TMEM16 family regarding whether they are CaCCs or Ca2+-dependent lipid scramblases [7, 10]. The Ca2+-dependent lipid scrambling function of TMEM16 family members have been implicated in the regulation of membrane trafficking, the release of extracellular vesicle, and the modulation of cell-cell communication [11].
As a CaCC, TMEM16A is activated by intracellular Ca2+, and the current is characterized by voltage-dependent activation, strong outward rectification, and a deactivating tail current on depolarization at low intracellular Ca2+ concentrations, and voltage-independent activation and linear current-voltage relationship at high [Ca2+]i [12]. Based on the crystal structure of a TMEM16 family member from the fungus Nectria haematococcaten (nhTMEM16), a conserved Ca2+-binding site located within the membrane has been identified [13]. Ca2+-dependent properties of TMEM16A such as rectification, activation and deactivation kinetics, and rundown can be well explained by the presence of Ca2+ binding site within the membrane [14]. However, because nhTMEM16 is a scramblase, not an ion channel, the location of the pore in the TMEM16A channel remains unclear. A “double-barrel” pore architecture of TMEM16A channel has been recently proposed [15, 16].
TMEM16A is widely expressed in many cells including secretory epithelia [1, 17,18,19], airway and vascular smooth muscle cells [18, 20,21,22], vascular endothelium [23, 24], interstitial cells of Cajal [25, 26], and nociceptive neurons [27,28,29]. TMEM16A regulates many cellular functions, such as fluid secretion in secretory epithelia, smooth muscle contraction, gut mobility, cell volume regulation, apoptosis, and pain (reviewed in [30,31,32,33]). In addition, TMEM16A dysfunction contributes to many diseases such as cancer, hypertension, gastrointestinal motility disorders, and cystic fibrosis [31, 34,35,36]. Recently, growing evidence has shown that TMEM16A is overexpressed in many tumors (Table 1). However, conflicting results exist regarding the role of TMEM16A in cell proliferation and migration in cancer cells. In addition, it remains unclear how TMEM16A overexpression contributes to tumorigenesis.
In this review, we examine recent findings in the study of TMEM16A in cancer, and focus on the role of TMEM16A in cancer cell proliferation and migration. We summarize the mechanisms of TMEM16A overexpression, the signaling pathways that are activated by TMEM16A, and potential clinical use of TMEM16A as a prognostic and predictive marker in cancer. Since TMEM16A plays different roles in different cancer cells, we try to develop the idea that TMEM16A regulates cancer cell proliferation and migration via a cell-specific mechanism.
TMEM16A Overexpression in cancer
Before it was identified as a CaCC, TMEM16A had been found to be amplified in oral cancer, head and neck squamous cell carcinoma (HNSCC), gastrointestinal stromal tumor (GIST), breast cancer, and esophageal squamous cell (ESCC) cancer under other names such as FLJ10261, TAOS1 (tumor amplified and overexpressed sequence 1) and DOG1 (discovered on GISTs protein 1) [37,38,39,40,41]. Recently, TMEM16A has been reported to be highly expressed in many human tumors including breast cancer [42, 43], HNSCC [44,45,46,47], colorectal cancer (CRC) [48, 49], ESCC [50], lung cancer [51], hepatocellular carcinoma [52], prostate cancer [53], gastric cancer [54, 55], and glioma [56] (Table 1).
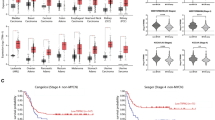
TMEM16A is located on chromosome 11q13, which is frequently amplified in many malignant tumors [57, 58]. Several studies have examined the copy number of TMEM16A in many tumors including breast cancer, HNSCC, and ESCC, and found that gene amplification commonly accounts for TMEM16A overexpression in these cancers (Table 1). To further confirm TMEM16A gene amplification in cancers, we performed bioinformatics analysis to detect TMEM16A gene alterations using the cBioPortal database (cBioPortal for Cancer Genomic). TMEM16A gene amplification accounts for the most alterations, and more frequently occurs in HNSCC, ESCC, breast cancer, and lung cancer than in other tumors (Fig. 1a). Interestingly, many tumors have missense mutations and deletions in the TMEM16A gene. A total of 165 missense mutations have been identified in TMEM16A, and the most frequent mutations are R561L/Q/W, R433Q, and R588G/Q (Fig. 1b). However, the role of these mutations has not been investigated in cancer.
The alterations of the TMEM16A gene in cBioPortal database. a TMEM16A gene was examined in 29 studies with >100 human cancer samples and >5% gene alterations. The copy number alteration (CNA) occurs more frequently in cancer. b TMEM16A missense mutations identified in cBioPortal database. A total of 165 missense mutations are shown. The most frequent mutations are R561L/Q/W, R433Q, and R588G/Q
Several studies have reported that 11q13 amplification is associated with poor prognosis in patients with malignant tumors [57, 58]. Consistent with the idea, Ruiz et al. found that 11q13 gene amplification correlated with TMEM16A expression in human HNSCC cancer, and TMEM16A overexpression was associated with poor overall survival in HNSCC patients [45]. In addition, Ayoub et al. reported that TMEM16A gene amplification and protein overexpression were associated with distant metastasis in patients with papillomavirus (HPV)-negative HNSCC [46]. Similarly, Bristschgi et al. reported that 11q13 amplification resulted in a higher TMEM16A expression in human breast cancer than in non-11q13-amplified tumors, and TMEM16A gene amplification and protein overexpression correlated with poor prognosis [42]. Shi et al. found that TMEM16A gene amplification and protein overexpression was associated with lymph node metastasis and advanced clinical stage in patients with ESCC [50].
Consistent with the results from the human tumor samples, TMEM16A has been found to be highly expressed in many cell lines with 11q13 amplification, including ZR75–1, HCC1954, and MDA-MB-415 breast cancer cell lines, UM-SCC1, BHY, and Te11 HNSCC cell lines, and FaDu, KYSE30 and KYSE510 ESCC cell lines [42, 44, 50] (Table 1). Knockdown of TMEM16A in cancer cells with 11q13 amplification results in a decrease in cell proliferation and an inhibition in xenograft tumor growth [42, 44, 50, 59]. These studies indicate that TMEM16A is critical for cell proliferation and tumor growth in 11q13-amplified tumors.
Although TMEM16A gene amplification is responsible for TMEM16A overexpression in many tumors, it is clearly not the only mechanism for TMEM16A expression. For example, in breast cancer, 11q13 amplification only occurs in approximately 15% of breast cancer patients, but TMEM16A overexpression occurs in >78% human breast cancer samples [42, 43]. Similarly, TMEM16A overexpression was more pervasive than gene amplification in human gastric cancer samples [54]. In contrast, in HNSCC, TMEM16A gene amplification was more frequently detected than protein expression [45, 47]. Therefore, other mechanisms that regulate TMEM16A expression must exist.
Multiple regulatory mechanisms of TMEM16A overexpression in cancer
In non-tumor cells, TMEM16A expression is regulated by many signaling pathways under physiological and pathological conditions. For example, in the airway epithelial cells, IL-4 induces TMEM16A upregulation, which is important for goblet cell differentiation [2, 60]. In human aortic smooth muscle cells, myocardin promotes TMEM16A expression by forming a complex with serum response factor (SRF) on the TMEM16A promoter, and angiotensin II inhibits TMEM16A expression via Kruppel-like factor 5, which competes with SRF to interact with myocardin [61]. In endothelial cells, angiotensin II increases TMEM16A expression [23]. In pulmonary arterial smooth muscle cells, chronic hypoxia increases TMEM16A expression [62]. In human lung epithelial A549 cells, TMEM16A expression is upregulated after lipopolysaccharide treatment [63]. Therefore, it appears that TMEM16A expression is controlled by various molecules and stimuli and the regulatory mechanisms varies in different cells. Here, we summarize the regulatory mechanisms of TMEM16A expression in cancer cells, and TMEM16A expression is controlled via transcriptional regulation, epigenetic regulation, and microRNAs (Fig. 2).
TMEM16A expression is upregulated via transcriptional regulation, epigenetic regulation and microRNAs in cancer. TMEM16A upregulation is induced by IL-4 and IL-13 [64, 65], which bind to their receptors and subsequently activate JAK/STAT6 signaling. STAT6 binds to the TMEM16A promoter and increases the transcription of the TMEM16A gene. Testosterone (T) induces TMEM16A upregulation by binding to the androgen receptor (AR), which subsequently increases the transcription of the TMEM16A gene [71]. Histone deacetylase (HDAC) inhibitors reduce TMEM16A expression in breast and prostate cell lines [77]. Promoter hypomethylation contributed to TMEM16A overexpression in HPV-negative HNSCC [75] and promoter hypermethylation results in decreased TMEM16A expression in metastatic lymph node tissues [74]. miR-132 and miR-381 binds to the 3′ UTR of TMEM16A mRNA, resulting in TMEM16A downregulation [49, 55]. Downregulation of miR-132 and miR-318 contributes to TMEM16A in patients with colorectal cancer [49] and gastric cancer [55]
Transcriptional regulation
Bioinformatics analyses show that the promoter region of the TMEM16A gene lacks TATAT box sequences, but contains many INRs (initiator elements) and/or TSSs (transcriptional start sites), suggesting that TMEM16A expression can be regulated by diverse transcription factors [64]. The TMEM16A promoter region contains a signal transducer and activator of transcription 6 (STAT6) binding site [64], which mediates TMEM16A upregulation induced by IL-4 and IL-13 [64, 65]. Zhang et al. reported that the expression of TMEM16A and MUC5AC was increased in nasal epithelial cells from patients with chronic rhinosinusitis [66]. IL-13 stimulated MUC5AC expression in human airway and nasal epithelial cells, and this effect was blocked by TMEM16A inhibitors, suggesting that TMEM16A might mediate IL-13-induced mucin secretion [65, 66]. These studies suggest that TMEM16A may play an important role in airway inflammation diseases. It is well known that IL-4 and IL-13 play an important role in cancer development [67,68,69,70]. To date, it remains unclear whether TMEM16A can be regulated by IL-4 and IL-13 in cancer cells. Future studies are required to demonstrate whether TMEM16A upregulation by IL-4 and IL-13 is involved in tumorigenesis.
The transcriptional regulation of TMEM16A expression has been demonstrated in testosterone-induced prostate hyperplasia by Cha et al. [71]. They found that the promoter region of the TMEM16A gene contains three putative binding sites for androgen response element (ARE), which mediates testosterone-induced TMEM16A upregulation in prostate epithelial cells. The testosterone-induced TMEM16A upregulation was blocked by small interfering RNAs (siRNAs) against the androgen receptor, which binds to the ARE region and subsequently promotes gene transcription. This study implies that TMEM16A upregulation induced by testosterone may contribute to the progression of prostate cancer.
Epigenetic regulation
DNA methylation of the target gene promoter plays an important role in the epigenetic regulation of genes that are essential for tumorigenesis [72, 73]. Promoter hypermethylation can repress gene expression, whereas promoter hypomehtylation can result in active transcription of the gene. TMEM16A promoter contains CpG islands, suggesting that DNA may be involved in the regulation of transcription of the TMEM16A gene [64, 74]. Indeed, Dixit et al. reported that TMEM16A expression was higher in HPV-negative HNSCC than in HPV-positive HNSCC, and promoter hypomethylation contributed to the higher expression of TMEM16A in HPV-negative HNSCC [75]. In addition, Shiwarski et al. found that compared with primary HNSCC tumors, methylation of the TMEM16A promoter region was increased in the metastatic lymph node tissue, thus resulting in decreased TMEM16A expression [74]. Promoter methylation-mediated inhibition of TMEM16A expression is believed to drive HNSCC cells from growth to metastic spread [74].
Histone deacetylase (HDAC) plays an important role in epigenetic regulation of gene expression by deacetylating the lysine residues in the histone, and dysregulation of HDACs has been implicated in the pathogenesis of cancer [76]. Matsuba et al. reported that HDAC inhibitors reduced TMEM16A expression and reduced cancer cell viability in breast and prostate cancer cell lines [77]. Wanitchakool et al. reported that HDAC inhibitors decreased TMEM16A expression and inhibited cell proliferation in HNSCC cells [59]. These studies further suggest that HDAC inhibitors may inhibited cell proliferation via downregulation of TMEM16A. However, the molecular mechanisms underlying the epigenetic regulation of TMEM16A transcription by HDAC have not been elucidated yet.
MicroRNAs
MicroRNAs are small, noncoding RNA molecules of ~22 nucleotides that inhibit gene expression by targeting the 3′ UTR of the target mRNAs. MicroRNAs regulate cell proliferation, apoptosis, angiogenesis and invasion, and contribute to tumorigenic processes in human cancers [78]. Recently, Mokutani et al. found that the 3′ UTR of TMEM16A mRNA contained a complementary site for miR-132, and the luciferase reporter assay showed that TMEM16A was the direct target of miR-132 [49]. In addition, TMEM16A overexpression was inversely associated with downregulation of miR-132 in human CRC, and correlated with poor clinical outcomes in patients with CRC [49]. Similarly, Cao et al. found that TMEM16A is the direct target of miR-381, and downregulation of miR-381 was inversely correlated with TMEM16A expression in human gastric cancer tissues [55]. These findings suggest that downregulation of microRNAs may contribute to TMEM16A overexpression in human cancers.
The signaling pathways activated by TMEM16A in cancer
As a CaCC, TMME16A overexpression can result in increased channel function, and opening of TMEM16A chloride channel can lead to changes in intracellular Cl− concentration ([Cl−]i) and membrane potential. This change in [Cl−]i and membrane potential may activate many signaling pathways that are involved in cancer cell proliferation and migration. In addition, as a membrane protein, TMEM16A interacts with several membrane proteins including SNARE proteins that control vesicle trafficking and the ezrin-radixin-moesin (ERM) complex that links membrane proteins with cytoskeleton [79]. It is possible that TMEM16A activates many signaling pathways via its interactome. Here, we summarize the signaling pathways that are activated by TMEM16A in cancer. TMEM16A activates many signaling pathways that participate in cell proliferation, migration, and invasion (Table 1, and Fig. 3).
The signaling pathways that are activated by TMEM16A in cancer. TMEM16A directly interacts with EGFR [81], and promotes EGFR phosphorylation, which activates the AKT/SRC/ERK1/2 signaling [42]. In addition, TMEM16A increases autocrine secretion of EGF in breast cancer cells [42]. TMEM16A directly interacts with IP3R, and increased Ca2+ release from the ER [85]. TMEM16A activates CaMKII by increasing intracellular Ca2+ concentrations, and CaMKII subsequently activates the AKT/SRC/ERK1/2 signaling [42]. TMEM16A also activates the Ras-Raf-Mek-ERK1/2 signaling pathway in UM-SCC1 HNSCC cells and T24 bladder cells [44]. In SMMC-7721 human hepatoma cells, TMEM16A activates the p38 signaling pathway [52]. TMEM16A activates the NFκB signaling pathway and promotes the gene transcription in glioma cells [56]. +, activates the signaling pathway.?, the mechanisms of how TMEM16A activates the signaling pathway are unknown
Epidermal growth factor receptor (EGFR) signaling
EGFR is a tyrosine kinase receptor that is overexpressed in many tumors such as HNSCC and breast cancer, and contributes to tumorigenesis [80]. Bill et al. reported that TMEM16A promoted EGFR phosphorylation, and increased the expression in a posttranslational and degradation-independent mechanism in HNSCC cells [81]. In addition, TMEM16A formed a complex with EGFR, and the complex regulated cancer proliferation in HNSCC cells [81]. Activation of EGFR signaling by TMEM16A is further demonstrated by Britschgi et al. showing that TMEM16A knockdown reduced EGFR phosphorylation and subsequently inhibited AKT, SRC, and ERK activation in breast cancer cell lines [42]. Furthermore, they demonstrated that TMEM16A knockdown reduced the autocrine secretion of EGFR ligands, EGF and TGF-α in breast cancer cells, suggesting that TMEM16A can activate EGFR signaling by increasing autocrine secretion of EGFR-ligands [42]. Therefore, TMEM16A activates the EGFR signaling pathway by increasing EGFR expression, phosphorylation, and autocrine EGFR-ligand secretion (Fig. 3).
Ca2+/Calmodulin-dependent protein kinase II (CAMKII) signaling
Britschgi et al. also found that EGFR inhibition only partially reduced TMEM16A overexpression-induced cell viability, and EGFR activation partially reversed the inhibitory effect of TMEM16A inhibitors on cell viability in breast cancer cells, suggesting that TMEM16A activates additional signaling pathways that are involved in cell viability [42]. Furthermore, they found that TMEM16A overexpression increased calcium/CAMKII phosphorylation, indicating that TMEM16A overexpression activates calcium-dependent CAMKII signaling. It has been reported that TMEM16A is located in the lipid raft of the plasma membrane in nociceptive neurons, where it is in close proximity to IP3R [82], and is believed to play a role in modulating intracellular Ca2+ levels [83]. In addition, TMEM16A inhibition has been found to reduce intracellular Ca2+ flux from both the plasma membrane and sarcoplasmic reticulum in airway smooth muscle [84]. Recently, Cabrita et al. found that TMEM16A directly interacted with the IP3R, and increased compartmentalized Ca2+ release from the ER store induced by ATP in HeLa cells [85]. TMEM16A inhibitors reduced ATP-induced increase in [Ca2+]i [85], suggesting that Cl− transport through TMEM16A channels may be important for Ca2+ release from the Ca2+ store in cancer cells. In addition, TMEM16A did not interacted with ORAI, and TMEM16A activation was not affected by ORAI inhibitors [85], suggesting that TMEM16A may not regulate ORAI-mediated Ca2+ entry in cancer cells.
Mitogen-activated protein kinase (MAPK) signaling
The MAPK signaling pathways regulate many cellular processes such as proliferation, apoptosis, migration, differentiation, and growth, and play an important role in the development and progression of cancer [86]. Duvvuri et al. found that TMEM16A overexpression activated the Ras-Raf-MEK-ERK1/2 signaling pathway in UM-SCC1 HNSCC cells and T24 bladder cells, and ERK1/2 inhibition reduced TMEM16A-induced cell growth [46], and correlates with poor prognosis in patients with breast cancer [42], gastric cancer [54], and HNSCC [45]. However, it has been reported that TMEM16A overexpression has no effects on cell migration when transfected into HEK293 cells [94]. TMEM16A inhibition reduces cell migration in HNSCC cells [14, 125]. TMEM16A couples to different Ca2+-permeable ion channels that are predominantly expressed in different cells. For example, TMEM16A has been found to be activated by Ca2+ influx via TRPV1 in mouse dorsal root ganglion neurons [28], TRPV4 in the choroid plexus [126], TRPV6 in epithelial principal cells of the rat epididymis [127], TRPC1 in salivary gland cells [128], TRPC2 in rat thyroid cells [129], TRPC6 in cerebral artery myocytes [130], Cav1.4 at the photoreceptor ribbon synapse [131], Cav1.2 in canine ventricular myocytes [132], and store-operated Ca2+ entry in eccrine sweat glands [133]. Therefore, TMEM16A is activated by Ca2+ via different Ca2+-permeable ion channels in a cell-specific manner. However, it remains unclear whether TMEM16A may couple to different Ca2+-permeable ion channels in different cancers. Recently, Cabrita et al. have reported that TMEM16A directly interacts with IP3R in HeLa cells, and is activated by Ca2+ release from the ER via the IP3R, but not via ORAI-mediated Ca2+ influx [85]. This finding suggests that TMEM16A may be primarily activated by IP3R-mediated Ca2+ release from the ER in cancer cells. ** et al. reported a similar finding in small neurons from dorsal root ganglia, showing activation of TMEM16A by IP3R-mediated Ca2+ release, but not by Ca2+ influx via voltage-gated Ca2+ channels [82]. Further studies are required to investigate whether IP3R-mediated Ca2+ release from the ER represents a general mechanism for TMEM16A activation in cancer.
TMEM16A overexpression can be used as a prognostic and predictive marker for clinical outcomes in cancer patients (Table 1). We have previously found that TMEM16A overexpression is associated with good prognosis in PR-positive or HER2-negative breast cancer patients following tamoxifen treatment [43]. Since tamoxifen inhibits TMEM16A currents [1], the beneficial effect of tamoxifen in breast cancer patients may be associated with its inhibition on TMEM16A channel function. In addition, TMEM16A inhibition by T16A-inhA01 and CaCC-inhA01 has been reported to increase responses to EGFR/HER2-targeted therapy in HNSCC cells [110]. Recently, several other TMEM16A inhibitors has been discovered, including MONNA [134], eugenol [135], dehydroandrographolide [136], 9-Phenanthrol [137], Ani9 [138], idebenone [139], and luteolin [111]. Although some TMEM16A inhibitors have been tested in certain cancer cell lines [111, 136, 139], it remains unclear whether these compounds can effectively inhibit cancer growth in vivo, since pharmacological sensitivity of TMEM16A channels may be affected by cellular environment [10, 140, 141]. Both animal and clinical studies are required to investigate the efficacy of a TMEM16A inhibitor on cancer cell growth and metastasis before it can be used for cancer therapy.
Abbreviations
- [Ca2+]i :
-
Intracellular Ca2+ concentration
- [Cl−]i :
-
Intracellular Cl− concentration
- ARE:
-
Androgen response element
- CaCC:
-
Ca2+-activated chloride channel
- CAMKII:
-
Calmodulin-dependent protein kinase II
- CRC:
-
Colorectal cancer
- CTCs:
-
Circulating tumor cells
- DOG1:
-
Discovered on GISTs protein 1
- EGFR:
-
Epidermal growth factor receptor
- ELA:
-
Ehrlich Lettre ascites
- EMT:
-
Epithelial-to-mesenchymal transition
- ER:
-
Endoplasmic reticulum
- ERM:
-
Ezrin-radixin-moesin
- ESCC:
-
Esophageal squamous cell
- GIST:
-
Gastrointestinal stromal tumor
- HDAC:
-
Histone deacetylase
- HNSCC:
-
Head and neck squamous cell carcinoma
- HPV:
-
Human papillomavirus
- INRs:
-
Initiator elements
- IκB:
-
the inhibitor of κB
- MAPK:
-
Mitogen-activated protein kinase
- NFκB:
-
Nuclear factor κB
- S970:
-
Serine 970
- siRNAs:
-
Small interfering RNAs
- SRF:
-
Serum response factor
- STAT6:
-
Signal transducer and activator of transcription 6
- T9:
-
Threonine 9
- TAOS1:
-
Tumor amplified and overexpressed sequence 1
- TMEM16:
-
Transmembrane protein 16
- TSSs:
-
Transcriptional start sites
References
Yang YD, Cho H, Koo JY, Tak MH, Cho Y, Shim WS, Park SP, Lee J, Lee B, Kim BM, Raouf R, Shin YK, Oh U. TMEM16A Confers receptor-activated calcium-dependent chloride conductance. Nature. 2008;455:1210–5.
Caputo A, Caci E, Ferrera L, Pedemonte N, Barsanti C, Sondo E, Pfeffer U, Ravazzolo R, Zegarra-Moran O, Galietta LJ. TMEM16A, A membrane protein associated with calcium-dependent chloride channel activity. Science. 2008;322:590–4.
Schroeder BC, Cheng T, Jan YN, Jan LY. Expression cloning of TMEM16A as a calcium-activated chloride channel subunit. Cell. 2008;134:1019–29.
Grubb S, Poulsen KA, Juul CA, Kyed T, Klausen TK, Larsen EH, Hoffmann EK. TMEM16F (Anoctamin 6), an anion channel of delayed ca(2+) activation. J Gen Physiol. 2013;141:585–600.
Schreiber R, Uliyakina I, Kongsuphol P, Warth R, Mirza M, Martins JR, Kunzelmann K. Expression and function of epithelial anoctamins. J Biol Chem. 2010;285:7838–45.
Pifferi S, Dibattista M, Menini A. TMEM16B Induces chloride currents activated by calcium in mammalian cells. Pflugers Arch. 2009;458:1023–38.
Suzuki J, Fujii T, Imao T, Ishihara K, Kuba H, Nagata S. Calcium-dependent phospholipid scramblase activity of TMEM16 protein family members. J Biol Chem. 2013;288:13305–16.
Suzuki J, Umeda M, Sims PJ, Nagata S. Calcium-dependent phospholipid scrambling by TMEM16F. Nature. 2010;468:834–8.
Yang H, Kim A, David T, Palmer D, ** T, Tien J, Huang F, Cheng T, Coughlin SR, Jan YN, Jan LY. TMEM16F Forms a Ca2+−activated cation channel required for lipid scrambling in platelets during blood coagulation. Cell. 2012;151:111–22.
Tian Y, Schreiber R, Kunzelmann K. Anoctamins are a family of Ca2+−activated Cl- channels. J Cell Sci. 2012;125:4991–8.
Whitlock JM, Hartzell HC. Anoctamins/TMEM16 proteins: chloride channels flirting with lipids and extracellular vesicles. Annu Rev Physiol. 2016;
**ao Q, Yu K, Perez-Cornejo P, Cui Y, Arreola J, Hartzell HC. Voltage- and calcium-dependent gating of TMEM16A/Ano1 chloride channels are physically coupled by the first intracellular loop. Proc Natl Acad Sci U S A. 2011;108:8891–6.
Brunner JD, Lim NK, Schenck S, Duerst A, Dutzler R. X-ray structure of a calcium-activated TMEM16 lipid scramblase. Nature. 2014;516:207–12.
Ma K, Wang H, Yu J, Wei M, **ao Q. New insights on the regulation of Ca2+ −activated Chloride Channel TMEM16A. J Cell Physiol. 2017;232:707–16.
Jeng G, Aggarwal M, Yu WP, Chen TY. Independent activation of distinct pores in dimeric TMEM16A channels. J Gen Physiol. 2016;148:393–404.
Lim NK, Lam AK, Dutzler R. Independent activation of ion conduction pores in the double-barreled calcium-activated chloride channel TMEM16A. J Gen Physiol. 2016;148:375–92.
Rock JR, O'Neal WK, Gabriel SE, Randell SH, Harfe BD, Boucher RC, Grubb BR. Transmembrane protein 16A (TMEM16A) is a Ca2+−regulated Cl- secretory channel in mouse airways. J Biol Chem. 2009;284:14875–80.
Huang F, Zhang H, Wu M, Yang H, Kudo M, Peters CJ, Woodruff PG, Solberg OD, Donne ML, Huang X, Sheppard D, Fahy JV, Wolters PJ, Hogan BL, Finkbeiner WE, Li M, Jan YN, Jan LY, Rock JR. Calcium-activated chloride channel TMEM16A modulates mucin secretion and airway smooth muscle contraction. Proc Natl Acad Sci U S A. 2012;109:16354–9.
Almaca J, Tian Y, Aldehni F, Ousingsawat J, Kongsuphol P, Rock JR, Harfe BD, Schreiber R, Kunzelmann K. TMEM16 Proteins produce volume-regulated chloride currents that are reduced in mice lacking TMEM16A. J Biol Chem. 2009;284:28571–8.
Wang M, Yang H, Zheng LY, Zhang Z, Tang YB, Wang GL, Du YH, Lv XF, Liu J, Zhou JG, Guan YY. Downregulation of TMEM16A calcium-activated chloride channel contributes to cerebrovascular remodeling during hypertension by promoting basilar smooth muscle cell proliferation. Circulation. 2012;125:697–707.
Manoury B, Tamuleviciute A, Tammaro P. TMEM16A/Anoctamin 1 protein mediates calcium-activated chloride currents in pulmonary arterial smooth muscle cells. J Physiol. 2010;588:2305–14.
Thomas-Gatewood C, Neeb ZP, Bulley S, Adebiyi A, Bannister JP, Leo MD, Jaggar JH. TMEM16A Channels generate ca(2)(+)-activated Cl(−) currents in cerebral artery smooth muscle cells. Am J Physiol Heart Circ Physiol. 2011;301:H1819–27.
Ma MM, Gao M, Guo KM, Wang M, Li XY, Zeng XL, Sun L, Lv XF, Du YH, Wang GL, Zhou JG, Guan YY. TMEM16A Contributes to endothelial dysfunction by facilitating Nox2 NADPH Oxidase-derived reactive oxygen species generation in hypertension. Hypertension. 2017;
Wu MM, Lou J, Song BL, Gong YF, Li YC, Yu CJ, Wang QS, Ma TX, Ma K, Hartzell HC, Duan DD, Zhao D, Zhang ZR. Hypoxia augments the calcium-activated chloride current carried by anoctamin-1 in cardiac vascular endothelial cells of neonatal mice. Br J Pharmacol. 2014;171:3680–92.
Hwang SJ, Blair PJ, Britton FC, O'Driscoll KE, Hennig G, Bayguinov YR, Rock JR, Harfe BD, Sanders KM, Ward SM. Expression of anoctamin 1/TMEM16A by interstitial cells of Cajal is fundamental for slow wave activity in gastrointestinal muscles. J Physiol. 2009;587:4887–904.
Huang F, Rock JR, Harfe BD, Cheng T, Huang X, Jan YN, Jan LY. Studies on expression and function of the TMEM16A calcium-activated chloride channel. Proc Natl Acad Sci U S A. 2009;106:21413–8.
Liu B, Linley JE, Du X, Zhang X, Ooi L, Zhang H, Gamper N. The acute nociceptive signals induced by bradykinin in rat sensory neurons are mediated by inhibition of M-type K+ channels and activation of Ca2+−activated Cl- channels. J Clin Invest. 2010;120:1240–52.
Takayama Y, Uta D, Furue H, Tominaga M. Pain-enhancing mechanism through interaction between TRPV1 and anoctamin 1 in sensory neurons. Proc Natl Acad Sci U S A. 2015;112:5213–8.
Cho H, Yang YD, Lee J, Lee B, Kim T, Jang Y, Back SK, Na HS, Harfe BD, Wang F, Raouf R, Wood JN, Oh U. The calcium-activated chloride channel anoctamin 1 acts as a heat sensor in nociceptive neurons. Nat Neurosci. 2012;15:1015–21.
Oh U, Jung J. Cellular functions of TMEM16/anoctamin. Pflugers Arch. 2016;468:443–53.
Pedemonte N, Galietta LJ. Structure and function of TMEM16 proteins (anoctamins). Physiol Rev. 2014;94:419–59.
Hartzell HC, Yu K, **ao Q, Chien LT, Qu Z. Anoctamin/TMEM16 family members are Ca2+−activated Cl- channels. J Physiol. 2009;587:2127–39.
Wanitchakool P, Ousingsawat J, Sirianant L, MacAulay N, Schreiber R, Kunzelmann K. Cl- channels in apoptosis. Eur Biophys J. 2016;
Mazzone A, Bernard CE, Strege PR, Beyder A, Galietta LJ, Pasricha PJ, Rae JL, Parkman HP, Linden DR, Szurszewski JH, Ordog T, Gibbons SJ, Farrugia G. Altered expression of Ano1 variants in human diabetic gastroparesis. J Biol Chem. 2011;286:13393–403.
Sondo E, Caci E, Galietta LJ. The TMEM16A chloride channel as an alternative therapeutic target in cystic fibrosis. Int J Biochem Cell Biol. 2014;52:73–6.
Wang B, Li C, Huai R, Qu Z. Overexpression of ANO1/TMEM16A, an arterial Ca2+−activated Cl- channel, contributes to spontaneous hypertension. J Mol Cell Cardiol. 2015;82:22–32.
Huang X, Gollin SM, Raja S, Godfrey TE. High-resolution map** of the 11q13 amplicon and identification of a gene, TAOS1, that is amplified and overexpressed in oral cancer cells. Proc Natl Acad Sci U S A. 2002;99:11369–74.
Carles A, Millon R, Cromer A, Ganguli G, Lemaire F, Young J, Wasylyk C, Muller D, Schultz I, Rabouel Y, Dembele D, Zhao C, Marchal P, Ducray C, Bracco L, Abecassis J, Poch O, Wasylyk B. Head and neck squamous cell carcinoma transcriptome analysis by comprehensive validated differential display. Oncogene. 2006;25:1821–31.
West RB, Corless CL, Chen X, Rubin BP, Subramanian S, Montgomery K, Zhu S, Ball CA, Nielsen TO, Patel R, Goldblum JR, Brown PO, Heinrich MC, van de Rijn M. The novel marker, DOG1, is expressed ubiquitously in gastrointestinal stromal tumors irrespective of KIT or PDGFRA mutation status. Am J Pathol. 2004;165:107–13.
Katoh M, Katoh M. FLJ10261 Gene, located within the CCND1-EMS1 locus on human chromosome 11q13, encodes the eight-transmembrane protein homologous to C12orf3, C11orf25 and FLJ34272 gene products. Int J Oncol. 2003;22:1375–81.
Carneiro A, Isinger A, Karlsson A, Johansson J, Jonsson G, Bendahl PO, Falkenback D, Halvarsson B, Nilbert M. Prognostic impact of array-based genomic profiles in esophageal squamous cell cancer. BMC Cancer. 2008;8:98.
Britschgi A, Bill A, Brinkhaus H, Rothwell C, Clay I, Duss S, Rebhan M, Raman P, Guy CT, Wetzel K, George E, Popa MO, Lilley S, Choudhury H, Gosling M, Wang L, Fitzgerald S, Borawski J, Baffoe J, Labow M, Gaither LA, Bentires-Alj M. Calcium-activated chloride channel ANO1 promotes breast cancer progression by activating EGFR and CAMK signaling. Proc Natl Acad Sci U S A. 2013;110:E1026–34.
Wu H, Guan S, Sun M, Yu Z, Zhao L, He M, Zhao H, Yao W, Wang E, ** F, **ao Q, Wei M. Ano1/TMEM16A Overexpression is associated with good prognosis in PR-positive or HER2-negative breast cancer patients following Tamoxifen treatment. PLoS One. 2015;10:e0126128.
Duvvuri U, Shiwarski DJ, **ao D, Bertrand C, Huang X, Edinger RS, Rock JR, Harfe BD, Henson BJ, Kunzelmann K, Schreiber R, Seethala RS, Egloff AM, Chen X, Lui VW, Grandis JR, Gollin SM. TMEM16A Induces MAPK and contributes directly to tumorigenesis and cancer progression. Cancer Res. 2012;72:3270–81.
Ruiz C, Martins JR, Rudin F, Schneider S, Dietsche T, Fischer CA, Tornillo L, Terracciano LM, Schreiber R, Bubendorf L, Kunzelmann K. Enhanced expression of ANO1 in head and neck squamous cell carcinoma causes cell migration and correlates with poor prognosis. PLoS One. 2012;7:e43265.
Ayoub C, Wasylyk C, Li Y, Thomas E, Marisa L, Robe A, Roux M, Abecassis J, de Reynies A, Wasylyk B. ANO1 Amplification and expression in HNSCC with a high propensity for future distant metastasis and its functions in HNSCC cell lines. Br J Cancer. 2010;103:715–26.
Rodrigo JP, Menendez ST, Hermida-Prado F, Alvarez-Teijeiro S, Villaronga MA, Alonso-Duran L, Vallina A, Martinez-Camblor P, Astudillo A, Suarez C, Maria G-PJ. Clinical significance of Anoctamin-1 gene at 11q13 in the development and progression of head and neck squamous cell carcinomas. Sci Rep. 2015;5:15698.
Sui Y, Sun M, Wu F, Yang L, Di W, Zhang G, Zhong L, Ma Z, Zheng J, Fang X, Ma T. Inhibition of TMEM16A expression suppresses growth and invasion in human colorectal cancer cells. PLoS One. 2014;9:e115443.
Mokutani Y, Uemura M, Munakata K, Okuzaki D, Haraguchi N, Takahashi H, Nishimura J, Hata T, Murata K, Takemasa I, Mizushima T, Doki Y, Mori M, Yamamoto H. Down-regulation of microRNA-132 is associated with poor prognosis of colorectal cancer. Ann Surg Oncol. 2016;
Shi ZZ, Shang L, Jiang YY, Hao JJ, Zhang Y, Zhang TT, Lin DC, Liu SG, Wang BS, Gong T, Zhan QM, Wang MR. Consistent and differential genetic aberrations between esophageal dysplasia and squamous cell carcinoma detected by array comparative genomic hybridization. Clin Cancer Res. 2013;19:5867–78.
Jia L, Liu W, Guan L, Lu M, Wang K. Inhibition of calcium-activated Chloride Channel ANO1/TMEM16A suppresses tumor growth and invasion in human lung cancer. PLoS One. 2015;10:e0136584.
Deng L, Yang J, Chen H, Ma B, Pan K, Su C, Xu F, Zhang J. Knockdown of TMEM16A suppressed MAPK and inhibited cell proliferation and migration in hepatocellular carcinoma. Onco Targets Ther. 2016;9:325–33.
Liu W, Lu M, Liu B, Huang Y, Wang K. Inhibition of ca(2+)-activated Cl(−) channel ANO1/TMEM16A expression suppresses tumor growth and invasiveness in human prostate carcinoma. Cancer Lett. 2012;326:41–51.
Liu F, Cao QH, Lu DJ, Luo B, Lu XF, Luo RC, Wang XG. TMEM16A Overexpression contributes to tumor invasion and poor prognosis of human gastric cancer through TGF-beta signaling. Oncotarget. 2015;6:11585–99.
Cao Q, Liu F, Ji K, Liu N, He Y, Zhang W, Wang L. MicroRNA-381 inhibits the metastasis of gastric cancer by targeting TMEM16A expression. J Exp Clin Cancer Res. 2017;36:29.
Liu J, Liu Y, Ren Y, Kang L, Zhang L. Transmembrane protein with unknown function 16A overexpression promotes glioma formation through the nuclear factor-kappaB signaling pathway. Mol Med Rep. 2014;9:1068–74.
Akervall JA, ** Y, Wennerberg JP, Zatterstrom UK, Kjellen E, Mertens F, Willen R, Mandahl N, Heim S, Mitelman F. Chromosomal abnormalities involving 11q13 are associated with poor prognosis in patients with squamous cell carcinoma of the head and neck. Cancer. 1995;76:853–9.
Ormandy CJ, Musgrove EA, Hui R, Daly RJ, Sutherland RL. Cyclin D1, EMS1 and 11q13 amplification in breast cancer. Breast Cancer Res Treat. 2003;78:323–35.
Wanitchakool P, Wolf L, Koehl GE, Sirianant L, Schreiber R, Kulkarni S, Duvvuri U, Kunzelmann K. Role of anoctamins in cancer and apoptosis. Philos Trans R Soc Lond Ser B Biol Sci. 2014;369:20130096.
Scudieri P, Caci E, Bruno S, Ferrera L, Schiavon M, Sondo E, Tomati V, Gianotti A, Zegarra-Moran O, Pedemonte N, Rea F, Ravazzolo R, Galietta LJ. Association of TMEM16A chloride channel overexpression with airway goblet cell metaplasia. J Physiol. 2012;590:6141–55.
Zhang XH, Zheng B, Yang Z, He M, Yue LY, Zhang RN, Zhang M, Zhang W, Zhang X, Wen JK. TMEM16A And Myocardin form a positive feedback loop that is disrupted by KLF5 during Ang II-induced vascular remodeling. Hypertension. 2015;66:412–21.
Sun H, **a Y, Paudel O, Yang XR, Sham JS. Chronic hypoxia-induced upregulation of Ca2+−activated Cl- channel in pulmonary arterial myocytes: a mechanism contributing to enhanced vasoreactivity. J Physiol. 2012;590:3507–21.
Zhang A, Yan X, Li H, Gu Z, Zhang P, Zhang H, Li Y, Yu H. TMEM16A Protein attenuates lipopolysaccharide-mediated inflammatory response of human lung epithelial cell line A549. Exp Lung Res. 2014;40:237–50.
Mazzone A, Gibbons SJ, Bernard CE, Nowsheen S, Middha S, Almada LL, Ordog T, Kendrick ML, Reid Lombardo KM, Shen KR, Galietta LJ, Fernandez-Zapico ME, Farrugia G. Identification and characterization of a novel promoter for the human ANO1 gene regulated by the transcription factor signal transducer and activator of transcription 6 (STAT6). FASEB J. 2015;29:152–63.
Qin Y, Jiang Y, Sheikh AS, Shen S, Liu J, Jiang D. Interleukin-13 stimulates MUC5AC expression via a STAT6-TMEM16A-ERK1/2 pathway in human airway epithelial cells. Int Immunopharmacol. 2016;40:106–14.
Zhang Y, Wang X, Wang H, Jiao J, Li Y, Fan E, Zhang L, Bachert C. TMEM16A-Mediated Mucin secretion in IL-13-induced nasal epithelial cells from chronic Rhinosinusitis patients. Allergy Asthma Immunol Res. 2015;7:367–75.
Dmitrieva OS, Shilovskiy IP, Khaitov MR, Grivennikov SI. Interleukins 1 and 6 as main mediators of inflammation and cancer. Biochemistry (Mosc). 2016;81:80–90.
Bharti R, Dey G, Mandal M. Cancer development, chemoresistance, epithelial to mesenchymal transition and stem cells: a snapshot of IL-6 mediated involvement. Cancer Lett. 2016;375:51–61.
May RD, Fung M. Strategies targeting the IL-4/IL-13 axes in disease. Cytokine. 2015;75:89–116.
Suzuki A, Leland P, Joshi BH, Puri RK. Targeting of IL-4 and IL-13 receptors for cancer therapy. Cytokine. 2015;75:79–88.
Cha JY, Wee J, Jung J, Jang Y, Lee B, Hong GS, Chang BC, Choi YL, Shin YK, Min HY, Lee HY, Na TY, Lee MO, Oh U. Anoctamin 1 (TMEM16A) is essential for testosterone-induced prostate hyperplasia. Proc Natl Acad Sci U S A. 2015;112:9722–7.
Jones PA, Issa JP, Baylin S. Targeting the cancer epigenome for therapy. Nat Rev Genet. 2016;17:630–41.
Biswas S, Rao CM. Epigenetics in cancer: fundamentals and beyond. Pharmacol Ther. 2017;
Shiwarski DJ, Shao C, Bill A, Kim J, **ao D, Bertrand CA, Seethala RS, Sano D, Myers JN, Ha P, Grandis J, Gaither LA, Puthenveedu MA, Duvvuri U. To "grow" or "go": TMEM16A expression as a switch between tumor growth and metastasis in SCCHN. Clin Cancer Res. 2014;20:4673–88.
Dixit R, Kemp C, Kulich S, Seethala R, Chiosea S, Ling S, Ha PK, Duvvuri U. TMEM16A/ANO1 Is differentially expressed in HPV-negative versus HPV-positive head and neck squamous cell carcinoma through promoter methylation. Sci Rep. 2015;5:16657.
Zhang H, Shang YP, Chen HY, Li J. Histone deacetylases function as novel potential therapeutic targets for cancer. Hepatol Res. 2016; 10.1111/hepr.12757.
Matsuba S, Niwa S, Muraki K, Kanatsuka S, Nakazono Y, Hatano N, Fujii M, Zhan P, Suzuki T, Ohya S. Downregulation of Ca2+−activated Cl- channel TMEM16A by the inhibition of histone deacetylase in TMEM16A-expressing cancer cells. J Pharmacol Exp Ther. 2014;351:510–8.
Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16:203–22.
Perez-Cornejo P, Gokhale A, Duran C, Cui Y, **ao Q, Hartzell HC, Faundez V. Anoctamin 1 (Tmem16A) Ca2+−activated chloride channel stoichiometrically interacts with an ezrin-radixin-moesin network. Proc Natl Acad Sci U S A. 2012;109:10376–81.
Steelman LS, Fitzgerald T, Lertpiriyapong K, Cocco L, Follo MY, Martelli AM, Neri LM, Marmiroli S, Libra M, Candido S, Nicoletti F, Scalisi A, Fenga C, Drobot L, Rakus D, Gizak A, Laidler P, Dulinska-Litewka J, Basecke J, Mijatovic S, Maksimovic-Ivanic D, Montalto G, Cervello M, Milella M, Tafuri A, Demidenko Z, Abrams SL, McCubrey JA. Critical roles of EGFR family members in breast cancer and breast cancer stem cells: targets for therapy. Curr Pharm Des. 2016;22:2358–88.
Bill A, Gutierrez A, Kulkarni S, Kemp C, Bonenfant D, Voshol H, Duvvuri U, Gaither LA. ANO1 Interacts with EGFR and correlates with sensitivity to EGFR-targeting therapy in head and neck cancer. Oncotarget. 2015;6:9173–88.
** X, Shah S, Liu Y, Zhang H, Lees M, Fu Z, Lippiat JD, Beech DJ, Sivaprasadarao A, Baldwin SA, Gamper N. Activation of the Cl- channel ANO1 by localized calcium signals in nociceptive sensory neurons requires coupling with the IP3 receptor. Sci Signal. 2013;6:ra73.
Kunzelmann K, Cabrita I, Wanitchakool P, Ousingsawat J, Sirianant L, Benedetto R, Schreiber R. Modulating ca signals: a common theme for TMEM16, Ist2, and TMC. Pflugers Arch. 2016;468:475–90.
Danielsson J, Perez-Zoghbi J, Bernstein K, Barajas MB, Zhang Y, Kumar S, Sharma PK, Gallos G, Emala CW. Antagonists of the TMEM16A calcium-activated Chloride Channel modulate airway smooth muscle tone and intracellular calcium. Anesthesiology. 2015;123:569–81.
Cabrita I, Benedetto R, Fonseca A, Wanitchakool P, Sirianant L, Skryabin BV, Schenk LK, Pavenstadt H, Schreiber R, Kunzelmann K. Differential effects of anoctamins on intracellular calcium signals. FASEB J. 2017;31:2123–34.
Dhillon AS, Hagan S, Rath O, Kolch W. MAP kinase signalling pathways in cancer. Oncogene. 2007;26:3279–90.
Hoesel B, Schmid JA. The complexity of NF-kappaB signaling in inflammation and cancer. Mol Cancer. 2013;12:86.
Alberti C, Pinciroli P, Valeri B, Ferri R, Ditto A, Umezawa K, Sensi M, Canevari S, Tomassetti A. Ligand-dependent EGFR activation induces the co-expression of IL-6 and PAI-1 via the NFkB pathway in advanced-stage epithelial ovarian cancer. Oncogene. 2012;31:4139–49.
Yang H, Huang LY, Zeng DY, Huang EW, Liang SJ, Tang YB, Su YX, Tao J, Shang F, Wu QQ, **ong LX, Lv XF, Liu J, Guan YY, Zhou JG. Decrease of intracellular chloride concentration promotes endothelial cell inflammation by activating nuclear factor-kappaB pathway. Hypertension. 2012;60:1287–93.
Buchholz B, Faria D, Schley G, Schreiber R, Eckardt KU, Kunzelmann K. Anoctamin 1 induces calcium-activated chloride secretion and proliferation of renal cyst-forming epithelial cells. Kidney Int. 2014;85:1058–67.
Stanich JE, Gibbons SJ, Eisenman ST, Bardsley MR, Rock JR, Harfe BD, Ordog T, Farrugia G. Ano1 As a regulator of proliferation. Am J Physiol Gastrointest Liver Physiol. 2011;301:G1044–51.
Sauter DR, Novak I, Pedersen SF, Larsen EH, Hoffmann EK. ANO1 (TMEM16A) in pancreatic ductal adenocarcinoma (PDAC). Pflugers Arch. 2015;467:1495–508.
Simon S, Grabellus F, Ferrera L, Galietta L, Schwindenhammer B, Muhlenberg T, Taeger G, Eilers G, Treckmann J, Breitenbuecher F, Schuler M, Taguchi T, Fletcher JA, Bauer S. DOG1 Regulates growth and IGFBP5 in gastrointestinal stromal tumors. Cancer Res. 2013;73:3661–70.
Ubby I, Bussani E, Colonna A, Stacul G, Locatelli M, Scudieri P, Galietta L, Pagani F. TMEM16A Alternative splicing coordination in breast cancer. Mol Cancer. 2013;12:75.
Wu H, Wang H, Guan S, Zhang J, Chen Q, Wang X, Ma K, Zhao P, Zhao H, Yao W, ** F, **ao Q, Wei M. Cell-specific regulation of proliferation by Ano1/TMEM16A in breast cancer with different ER, PR, and HER2 status. Oncotarget. 2017;
Lee YS, Lee JK, Bae Y, Lee BS, Kim E, Cho CH, Ryoo K, Yoo J, Kim CH, Yi GS, Lee SG, Lee CJ, Kang SS, Hwang EM, Park JY. Suppression of 14-3-3gamma-mediated surface expression of ANO1 inhibits cancer progression of glioblastoma cells. Sci Rep. 2016;6:26413.
Li Y, Zhang J, Hong S. ANO1 As a marker of oral squamous cell carcinoma and silencing ANO1 suppresses migration of human SCC-25 cells. Med Oral Patol Oral Cir Bucal. 2014;19:e313–9.
Ruffin M, Voland M, Marie S, Bonora M, Blanchard E, Blouquit-Laye S, Naline E, Puyo P, Le Rouzic P, Guillot L, Corvol H, Clement A, Tabary O. Anoctamin 1 dysregulation alters bronchial epithelial repair in cystic fibrosis. Biochim Biophys Acta. 1832;2013:2340–51.
Jacobsen KS, Zeeberg K, Sauter DR, Poulsen KA, Hoffmann EK, Schwab A. The role of TMEM16A (ANO1) and TMEM16F (ANO6) in cell migration. Pflugers Arch. 2013;465:1753–62.
Kunzelmann K, Tian Y, Martins JR, Faria D, Kongsuphol P, Ousingsawat J, Thevenod F, Roussa E, Rock J, Schreiber R. Anoctamins. Pflugers Arch. 2011;462:195–208.
Schwab A, Fabian A, Hanley PJ, Stock C. Role of ion channels and transporters in cell migration. Physiol Rev. 2012;92:1865–913.
Lee YS, Bae Y, Park N, Yoo JC, Cho CH, Ryoo K, Hwang EM, Park JY. Surface expression of the Anoctamin-1 (ANO1) channel is suppressed by protein-protein interactions with beta-COP. Biochem Biophys Res Commun. 2016;475:216–22.
Ha JR, Siegel PM, Ursini-Siegel J. The tyrosine Kinome dictates breast cancer heterogeneity and therapeutic responsiveness. J Cell Biochem. 2016;117:1971–90.
Fleuren ED, Zhang L, Wu J, Daly RJ. The kinome 'at large' in cancer. Nat Rev Cancer. 2016;16:83–98.
Ponuwei GA. A glimpse of the ERM proteins. J Biomed Sci. 2016;23:35.
Clucas J, Valderrama F. ERM proteins in cancer progression. J Cell Sci. 2015;128:1253.
Huang X, Godfrey TE, Gooding WE, McCarty KS Jr, Gollin SM. Comprehensive genome and transcriptome analysis of the 11q13 amplicon in human oral cancer and synteny to the 7F5 amplicon in murine oral carcinoma. Genes Chromosomes Cancer. 2006;45:1058–69.
Namkung W, Phuan PW, Verkman AS. TMEM16A Inhibitors reveal TMEM16A as a minor component of calcium-activated chloride channel conductance in airway and intestinal epithelial cells. J Biol Chem. 2011;286:2365–74.
De La Fuente R, Namkung W, Mills A, Verkman AS. Small-molecule screen identifies inhibitors of a human intestinal calcium-activated chloride channel. Mol Pharmacol. 2008;73:758–68.
Kulkarni S, Bill A, Godse NR, Khan NI, Kass JI, Steehler K, Kemp C, Davis K, Bertrand CA, Vyas AR, Holt DE, Grandis JR, Gaither LA, Duvvuri U. TMEM16A/ANO1 Suppression improves response to antibody mediated targeted therapy of EGFR and HER2/ERBB2. Genes Chromosomes Cancer. 2017;
Seo Y, Ryu K, Park J, Jeon DK, Jo S, Lee HK, Namkung W. Inhibition of ANO1 by luteolin and its cytotoxicity in human prostate cancer PC-3 cells. PLoS One. 2017;12:e0174935.
Bill A, Hall ML, Borawski J, Hodgson C, Jenkins J, Piechon P, Popa O, Rothwell C, Tranter P, Tria S, Wagner T, Whitehead L, Gaither LA. Small molecule-facilitated degradation of ANO1 protein: a new targeting approach for anticancer therapeutics. J Biol Chem. 2014;289:11029–41.
Gianotti A, Ferrera L, Philp AR, Caci E, Zegarra-Moran O, Galietta LJ, Flores CA. Pharmacological analysis of epithelial chloride secretion mechanisms in adult murine airways. Eur J Pharmacol. 2016;781:100–8.
Liu Y, Zhang H, Huang D, Qi J, Xu J, Gao H, Du X, Gamper N. Characterization of the effects of Cl(−) channel modulators on TMEM16A and bestrophin-1 ca(2)(+) activated Cl(−) channels. Pflugers Arch. 2015;467:1417–30.
Kucherenko YV, Wagner-Britz L, Bernhardt I, Lang F. Effect of chloride channel inhibitors on cytosolic Ca2+ levels and Ca2+−activated K+ (Gardos) channel activity in human red blood cells. J Membr Biol. 2013;246:315–26.
Kunzelmann K, Schreiber R, Kmit A, Jantarajit W, Martins JR, Faria D, Kongsuphol P, Ousingsawat J, Tian Y. Expression and function of epithelial anoctamins. Exp Physiol. 2012;97:184–92.
Hiraoka K, Miyazaki H, Niisato N, Iwasaki Y, Kawauchi A, Miki T, Marunaka Y. Chloride ion modulates cell proliferation of human androgen-independent prostatic cancer cell. Cell Physiol Biochem. 2010;25:379–88.
Habela CW, Ernest NJ, Swindall AF, Sontheimer H. Chloride accumulation drives volume dynamics underlying cell proliferation and migration. J Neurophysiol. 2009;101:750–7.
Duran C, Thompson CH, **ao Q, Hartzell HC. Chloride channels: often enigmatic, rarely predictable. Annu Rev Physiol. 2010;72:95–121.
Ohsawa R, Miyazaki H, Niisato N, Shiozaki A, Iwasaki Y, Otsuji E, Marunaka Y. Intracellular chloride regulates cell proliferation through the activation of stress-activated protein kinases in MKN28 human gastric cancer cells. J Cell Physiol. 2010;223:764–70.
Shang L, Hao JJ, Zhao XK, He JZ, Shi ZZ, Liu HJ, Wu LF, Jiang YY, Shi F, Yang H, Zhang Y, Liu YZ, Zhang TT, Xu X, Cai Y, Jia XM, Li M, Zhan QM, Li EM, Wang LD, Wei WQ, Wang MR. ANO1 Protein as a potential biomarker for esophageal cancer prognosis and precancerous lesion development prediction. Oncotarget. 2016;
Espinosa I, Lee CH, Kim MK, Rouse BT, Subramanian S, Montgomery K, Varma S, Corless CL, Heinrich MC, Smith KS, Wang Z, Rubin B, Nielsen TO, Seitz RS, Ross DT, West RB, Cleary ML, van de Rijn M. A novel monoclonal antibody against DOG1 is a sensitive and specific marker for gastrointestinal stromal tumors. Am J Surg Pathol. 2008;32:210–8.
Reddy RB, Bhat AR, James BL, Govindan SV, Mathew R, Ravindra DR, Hedne N, Illiayaraja J, Kekatpure V, Khora SS, Hicks W, Tata P, Kuriakose MA, Suresh A. Meta-analyses of microarray datasets identifies ANO1 and FADD as prognostic markers of head and neck cancer. PLoS One. 2016;11:e0147409.
Li Q, Zhi X, Zhou J, Tao R, Zhang J, Chen P, Roe OD, Sun L, Ma L. Circulating tumor cells as a prognostic and predictive marker in gastrointestinal stromal tumors: a prospective study. Oncotarget. 2016;
Ferrera L, Caputo A, Galietta LJ. TMEM16A Protein: a new identity for ca(2+)-dependent Cl(−) channels. Physiology (Bethesda). 2010;25:357–63.
Takayama Y, Shibasaki K, Suzuki Y, Yamanaka A, Tominaga M. Modulation of water efflux through functional interaction between TRPV4 and TMEM16A/anoctamin 1. FASEB J. 2014;28:2238–48.
Gao da Y, Zhang BL, Leung MC, Au SC, Wong PY, Shum WW. Coupling of TRPV6 and TMEM16A in epithelial principal cells of the rat epididymis. J Gen Physiol. 2016;148:161–82.
Sun Y, Birnbaumer L, Singh BB. TRPC1 Regulates calcium-activated chloride channels in salivary gland cells. J Cell Physiol. 2015;230:2848–56.
Viitanen TM, Sukumaran P, Lof C, Tornquist K. Functional coupling of TRPC2 cation channels and the calcium-activated anion channels in rat thyroid cells: implications for iodide homeostasis. J Cell Physiol. 2013;228:814–23.
Wang Q, Leo MD, Narayanan D, Kuruvilla KP, Jaggar JH. Local coupling of TRPC6 to ANO1/TMEM16A channels in smooth muscle cells amplifies vasoconstriction in cerebral arteries. Am J Physiol Cell Physiol. 2016; 10.1152/ajpcell.00092.2016.
Mercer AJ, Rabl K, Riccardi GE, Brecha NC, Stella SL Jr, Thoreson WB. Location of release sites and calcium-activated chloride channels relative to calcium channels at the photoreceptor ribbon synapse. J Neurophysiol. 2011;105:321–35.
Horvath B, Vaczi K, Hegyi B, Gonczi M, Dienes B, Kistamas K, Banyasz T, Magyar J, Baczko I, Varro A, Seprenyi G, Csernoch L, Nanasi PP, Szentandrassy N. Sarcolemmal Ca2+−entry through L-type Ca2+ channels controls the profile of Ca2+−activated Cl- current in canine ventricular myocytes. J Mol Cell Cardiol. 2016;97:125–39.
Concepcion AR, Vaeth M, Wagner LE 2nd, Eckstein M, Hecht L, Yang J, Crottes D, Seidl M, Shin HP, Weidinger C, Cameron S, Turvey SE, Issekutz T, Meyts I, Lacruz RS, Cuk M, Yule DI, Feske S. Store-operated Ca2+ entry regulates Ca2+−activated chloride channels and eccrine sweat gland function. J Clin Invest. 2016;126:4303–18.
Oh SJ, Hwang SJ, Jung J, Yu K, Kim J, Choi JY, Hartzell HC, Roh EJ, Lee CJ. MONNA, a potent and selective blocker for transmembrane protein with unknown function 16/anoctamin-1. Mol Pharmacol. 2013;84:726–35.
Yao Z, Namkung W, Ko EA, Park J, Tradtrantip L, Verkman AS. Fractionation of a herbal antidiarrheal medicine reveals eugenol as an inhibitor of Ca2+−activated Cl- channel TMEM16A. PLoS One. 2012;7:e38030.
Sui Y, Wu F, Lv J, Li H, Li X, Du Z, Sun M, Zheng Y, Yang L, Zhong L, Zhang X, Zhang G. Identification of the novel TMEM16A inhibitor Dehydroandrographolide and its anticancer activity on SW620 cells. PLoS One. 2015;10:e0144715.
Burris SK, Wang Q, Bulley S, Neeb ZP, Jaggar JH. 9-Phenanthrol inhibits recombinant and arterial myocyte TMEM16A channels. Br J Pharmacol. 2015;172:2459–68.
Seo Y, Lee HK, Park J, Jeon DK, Jo S, Jo M, Namkung W. Ani9, A novel potent small-molecule ANO1 inhibitor with negligible effect on ANO2. PLoS One. 2016;11:e0155771.
Seo Y, Park J, Kim M, Lee HK, Kim JH, Jeong JH, Namkung W. Inhibition of ANO1/TMEM16A Chloride Channel by Idebenone and its Cytotoxicity to cancer cell lines. PLoS One. 2015;10:e0133656.
Tian Y, Schreiber R, Wanitchakool P, Kongsuphol P, Sousa M, Uliyakina I, Palma M, Faria D, Traynor-Kaplan AE, Fragata JI, Amaral MD, Kunzelmann K. Control of TMEM16A by INO-4995 and other inositolphosphates. Br J Pharmacol. 2013;168:253–65.
Boedtkjer DM, Kim S, Jensen AB, Matchkov VM, Andersson KE. New selective inhibitors of calcium-activated chloride channels - T16Ainh -A01, CaCCinh -A01 and MONNA - what do they inhibit? Br J Pharmacol. 2015;172:4158–72.
Acknowledgements
Not applicable.
Funding
This work is supported by grants from the National Natural Science Foundation of China (No. 81572613 and No. 31371145 to Qinghuan **ao) and Lianoning Pandeng Scholar (to Qinghuan **ao).
Availability of data and materials
Not applicable.
Author information
Authors and Affiliations
Contributions
QX collected and interpreted studies, provided guidance throughout the preparation of this manuscript, and wrote and edited the manuscript. HW and LZ collected and interpreted studies, and drafted the manuscript. KM, JY, and HW drafted and revised the figures in the manuscript. MW reviewed and made a significant revision on the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Wang, H., Zou, L., Ma, K. et al. Cell-specific mechanisms of TMEM16A Ca2+-activated chloride channel in cancer. Mol Cancer 16, 152 (2017). https://doi.org/10.1186/s12943-017-0720-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12943-017-0720-x