Abstract
Breast cancer is one of the most common malignancies that pose a serious threat to women's health. Reprogramming of energy metabolism is a major feature of the malignant transformation of breast cancer. Compared to normal cells, tumor cells reprogram metabolic processes more efficiently, converting nutrient supplies into glucose, amino acid and lipid required for malignant proliferation and progression. Non-coding RNAs(ncRNAs) are a class of functional RNA molecules that are not translated into proteins but regulate the expression of target genes. NcRNAs have been demonstrated to be involved in various aspects of energy metabolism, including glycolysis, glutaminolysis, and fatty acid synthesis. This review focuses on the metabolic regulatory mechanisms and clinical applications of metabolism-regulating ncRNAs involved in breast cancer. We summarize the vital roles played by metabolism-regulating ncRNAs for endocrine therapy, targeted therapy, chemotherapy, immunotherapy, and radiotherapy resistance in breast cancer, as well as their potential as therapeutic targets and biomarkers. Difficulties and perspectives of current targeted metabolism and non-coding RNA therapeutic strategies are discussed.
Similar content being viewed by others
Introduction
Breast cancer, the most prevalent cancer in women and a leading cause of cancer-related deaths, has surpassed lung cancer in prevalence among women according to 2020 Global Cancer Statistics [1]. Treatment decisions are significantly influenced by molecular ty** and histologic features. Molecular classifications include luminal A, luminal B, HER2-enriched, and triple-negative breast cancer. Histologically, invasive ductal carcinoma is the most common, followed by invasive lobular carcinoma. These classifications impact prognosis and treatment options [2]. Breast cancer management involves local and systemic therapies. Local treatment includes surgical removal and radiotherapy. Systemic therapy varies based on subtypes such as endocrine therapy, HER2-targeted therapy, chemotherapy, and immunotherapy.
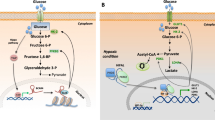
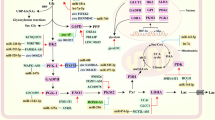
Tumors exhibit a distinct metabolic reprogramming, a hallmark characterized by the Warburg effect [3,4,5]. In the dynamic tumor microenvironment, cells adjust their metabolism to efficiently use glucose, lipids, and amino acids for rapid proliferation, survival, and metastasis. Despite lower ATP efficiency, the Warburg effect in cancer cells supports their high energy demands [6]. Increased glycolysis directs intermediates to biosynthetic pathways, promoting the synthesis of lipids, amino acids, and nucleosides for cell growth [7]. Tumor cells also display a 'lipogenic phenotype', enhancing fatty acid synthesis independently of exogenous sources [8]. While aerobic glycolysis predominates, some carbon is redirected to the tricarboxylic acid (TCA) cycle via glutamine metabolism, contributing to energy cycling and fatty acid synthesis [6, 9]. The intertwined reprogramming of these pathways collaborates to facilitate tumor growth and proliferation.
Non-coding RNAs (ncRNAs) are functional transcripts without protein-coding potential [10, 11]. They play key roles in developmental and pathological processes involving chromatin remodeling, transcription, post-transcriptional modifications and signal transduction [12]. Breast cancer exhibits a significant number of differentially expressed ncRNAs, some of which are linked to specific subtypes [13,14,15,163]. For miRNA targeting that requires overexpression, synthetic oligonucleotides consisting of miRNA duplexes (miRNA mimics) are used. In contrast, to achieve inhibition of oncogenic miRNAs, single-stranded antisense RNA were used (antagomiRs) [164]. Du et al. performed functional analysis of miR-210-3p using miRNA mimics and found that miR-210-3p promoted aerobic glycolysis by regulating glycolytic genes downstream of HIF-1α and p53. This activity conferred a growth advantage to TNBC and anti-apoptotic activity, suggesting that miR-210-3p may be a valuable target for the treatment of TNBC [165]. In the case of gene silencing for tsRNAs, a similar approach as for miRNAs is utilized. Zhu et al. constructed a tRFLys−CTT−010 knockdown model by small RNA inhibitors, which resulted in decreased G6PC protein levels and a significant reduction in cellular lactate production. Knockdown of tRFLys−CTT−010 inhibited the proliferation, migration and invasion of TNBC cells in vitro [80]. Therefore, targeting tRFLys−CTT−010, a regulator of G6PC, presents a promising approach for TNBC treatment.
RNA interference (RNAi) technology may be an effective method for treating breast cancer based on lncRNA and circRNA. In this strategy, exogenous or mimic double-stranded RNAs, such as short interfering RNAs (siRNAs) and shRNAs, are often used to specifically knock down target genes. For instance, Qin et al. demonstrated that silencing lnc030 expression through shRNA transfection led to a significant reduction in SQLE expression, resulting in decreased cellular cholesterol synthesis and inhibition of BCSC stemness maintenance. Animal experiments further validated the effectiveness of lnc030 or SQLE knockdown in reducing tumor-initiating ability and inhibiting tumor growth in vivo [112]. This highlights the potential of targeting Lnc030 and its downstream signaling as an effective therapeutic option. Wang et al. used shRNA knockdown of circSEPT9, which led to the inhibition of glutamine uptake and cell proliferation in BC cells. Subsequent mouse model experiments corroborated the in vitro anticancer activity of circSEPT9 silencing [119]. In addition, as described previously, LncRNAs HOXC-AS3 [57], DIO3OS [72], circKIF4A [68], and circZFR [97] can regulate metabolism by targeting PFK-1, LHDA, PKM2, and FABP, which in turn exert pro-cancer effects. Silencing these ncRNAs may be a target for potential therapeutic effects in breast cancer.
In conclusion, the combination of targeting ncRNAs and metabolic therapy presents a promising therapeutic strategy that may offer new insights and solutions for individualized tumor treatment. However, further basic research and clinical practice are necessary to validate its safety and efficacy.
Biomarkers
Early detection, diagnosis and treatment are key to improving the prognosis of breast cancer. Currently, there are still limited methods for predicting postoperative outcomes as well as testing for treatment efficacy. Therefore, the search for accurate biomarkers is crucial for early diagnosis and accurate prognosis of breast cancer. Numerous findings have shown that dysregulated ncRNAs expression is observed in breast cancer and ncRNAs are expected to serve as a diagnostic and prognostic biomarker.
There is growing evidence that metabolism-regulating ncRNAs play an important role as biomarkers in the diagnosis of BC, by being detected in breast cancer tissue or fluids. As mentioned previously, Li et al. found that LncRNA PlncRNA-1 inhibited breast cancer growth by down-regulating PHGDH, the enzyme that catalyzes the first step of the serine biosynthetic pathway. The authors used ROC curve analysis to evaluate the diagnostic value of PlncRNA-1 expression in breast tissue and serum for breast cancer. The area under the curve (AUC) of PlncRNA-1 expression in breast tissue in the diagnosis of breast cancer was 0.8994, and the AUC of serum PlncRNA-1 in the diagnosis of breast cancer was 0.8667. suggesting that PlncRNA-1 can accurately predict breast cancer in both tissue and serum [133]. Huang identified differentially expressed tRFs in normal and breast cancer cell lines, and the AUCs of tDR-7816, -5334, and -4733 were 0.859, 0.661, and 0.621, respectively, according to the ROC curve results. It can be inferred that tDR-7816, tDR-5334 and tDR-4733 may serve as potential candidates for non-TNBC breast cancer biomarkers. Functional analysis of target genes showed that the target genes of these three tRFs play a role in lipid metabolism processes such as glucuronic acid metabolism, steroid metabolism, and lipid biosynthesis [107]. Therefore, ncRNAs that regulate metabolism could serve as potential diagnostic markers for breast cancer.
Determining the prognostic value of metabolism-regulating ncRNAs is an essential field of BC research. The lncRNA breast cancer anti-estrogen resistance 4 (BCAR4) is required for YAP-dependent glycolysis. The expression levels of BCAR4 and YAP were positively correlated in tissue samples from breast cancer patients, where high expression of BCAR4 and YAP was associated with poor survival prognosis [90]. CircPDCD11 accelerated the rate of glucose uptake, lactate production and extracellular acidification in TNBC cells. Clinical results showed that high circPDCD11 expression was closely associated with poor prognosis and was an independent risk factor for TNBC prognosis [168,169,170]. Develo** different therapeutic strategies for the metabolic vulnerabilities of breast cancer presents both opportunities and challenges. Therefore, more advanced methods for assessing metabolic phenotypes, such as metabolomics, metabolic imaging, single-cell, and spatial assays, are needed. Tailoring personalized therapeutic strategies by analyzing a patient's unique tumor metabolic profile serves as a viable future direction.
Conclusion
This review summarizes the roles and mechanisms of non-coding RNAs that regulate metabolism in breast cancer. Non-coding RNAs that regulate metabolism can have an impact on resistance to existing treatment modalities in breast cancer. In addition to this, they can serve as therapeutic targets and biomarkers in their own right. Therefore, therapeutic approaches targeting non-coding RNAs and metabolism present both opportunities and challenges in the future treatment of breast cancer.
Availability of data and materials
Not applicable.
Abbreviations
- ncRNAs:
-
Non-coding RNAs
- TCA:
-
Tricarboxylic acid
- miRNA:
-
MicroRNA
- LncRNA:
-
Long non-coding RNA
- circRNA:
-
Circular RNA
- tsRNA:
-
TRNA-derived small RNA
- miRISC:
-
MiRNA-induced silencing complex
- ceRNAs:
-
Competitive endogenous RNAs
- tRFs:
-
TRNA-related small RNA fragments
- tiRNAs:
-
TRNA-derived stress-induced RNAs
- SLCs:
-
Solute carriers
- GLUT:
-
Glucose transporter
- ECAR:
-
Extracellular acidification rate
- EREG:
-
Epidermal regulator
- HK2:
-
Hexokinase 2
- PDK1:
-
Pyruvate dehydrogenase kinase 1
- PFK-1:
-
Phosphofructokinase-1
- PK:
-
Pyruvate kinase
- G6P:
-
Glucose-6-phosphate
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- F6P:
-
Fructose 6-phosphate
- F-1,6-BP:
-
Fructose 1,6-bisphosphate
- F-2,6-BP:
-
Fructose-2,6-bisphosphate
- PFKFB:
-
PFK-2/FBPase
- PEP:
-
Phosphoenolpyruvate
- PKM2:
-
Pyruvate kinase isoenzyme M2
- PHD3:
-
Prolyl hydroxylase 3
- hnRNPF:
-
Heterogeneous nuclear ribonucleoprotein F
- PGK1:
-
Phosphoglycerate kinase 1
- OCR:
-
Oxygen consumption rate
- LDHA:
-
Lactate dehydrogenase
- PDH:
-
Pyruvate dehydrogenase
- PTBP1:
-
Polypyrimidine tract binding protein 1
- PDHX:
-
Pyruvate dehydrogenase protein X
- GS:
-
Glycogen synthase
- GP:
-
Glycogen phosphorylase
- PGM:
-
Phosphate glucose metastases
- EMT:
-
Epithelial–mesenchymal transition
- G6PC:
-
Glucose-6-phosphatase catalytic
- TNBC:
-
Triple-negative breast cancer
- FABPs:
-
Fatty acid binding proteins
- ACC:
-
Acetyl CoA carboxylase
- FASN:
-
Fatty acid synthase
- FAO:
-
Fatty acid oxidation
- CPT-1:
-
Carnitine palmitoyltransferase I
- NEAT1:
-
Nuclear paraspeckle assembly transcript 1
- HMG-CoA:
-
3-Hydroxy-3-methylglutaryl CoA
- MVA:
-
Mevalonate
- HMGCR:
-
HMG-CoA reductase
- SQLE:
-
Squalene epoxidase
- MOS:
-
2,3(S)-monoxysqualene
- PCBP2:
-
Poly(rC)-binding protein 2
- BCSCs:
-
Breast cancer stem cells
- FFAs:
-
Free fatty acids
- PC:
-
Phosphatidylcholine
- PG:
-
Glycerophosphoglycerol
- SLC1A5:
-
Solute carrier family 1 neutral amino acid transporter member 5
- GPX4:
-
Glutathione peroxidase 4
- GSH:
-
Glutathione
- VGLUT2:
-
Vesicular glutamate transporter 2
- XBP1SBM:
-
XBP1s binding micropeptide
- SSP:
-
Serine synthetic pathway
- PHGDH:
-
Phosphoglycerate dehydrogenase
- PSAT:
-
Phosphoserine aminotransferase
- PSPH:
-
Phospho serine phosphatase
- 3-PPyr:
-
3-Phosphohydroxypyruvate
- AIs:
-
Aromatase inhibitors
- shRNAs:
-
Small hairpin RNA
- CTLA-4:
-
Cytotoxic T-lymphocyte-associated antigen-4
- PD-1:
-
Programmed cell death receptor-1
- PD-L1:
-
Programmed cell death ligand-1
- TME:
-
Tumor microenvironment
- ROS:
-
Reactive oxygen species
- LNA:
-
Locked nucleic acid
- siRNAs:
-
Short interfering RNAs
- AUC:
-
The area under the curve
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Waks AG, Winer EP. Breast cancer treatment: a review. JAMA. 2019;321:288–300.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Paul S, Ghosh S, Kumar S. Tumor glycolysis, an essential sweet tooth of tumor cells. Semin Cancer Biol. 2022;86:1216–30.
Hsu PP, Sabatini DM. Cancer cell metabolism: Warburg and beyond. Cell. 2008;134:703–7.
Li Z, Zhang H. Reprogramming of glucose, fatty acid and amino acid metabolism for cancer progression. Cell Mol Life Sci CMLS. 2016;73:377–92.
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324:1029–33.
Ligorio F, Pellegrini I, Castagnoli L, Vingiani A, Lobefaro R, Zattarin E, et al. Targeting lipid metabolism is an emerging strategy to enhance the efficacy of anti-HER2 therapies in HER2-positive breast cancer. Cancer Lett. 2021;511:77–87.
Altman BJ, Stine ZE, Dang CV. From Krebs to Clinic: Glutamine metabolism to cancer therapy. Nat Rev Cancer. 2016;16:619–34.
Lin X, Wu Z, Hu H, Luo M-L, Song E. Non-coding RNAs rewire cancer metabolism networks. Semin Cancer Biol. 2021;75:116–26.
Carninci P. Non-coding RNA transcription: turning on neighbours. Nat Cell Biol. 2008;10:1023–4.
Anastasiadou E, Jacob LS, Slack FJ. Non-coding RNA networks in cancer. Nat Rev Cancer. 2018;18:5–18.
Søkilde R, Persson H, Ehinger A, Pirona AC, Fernö M, Hegardt C, et al. Refinement of breast cancer molecular classification by miRNA expression profiles. BMC Genomics. 2019;20:503.
Kudela E, Samec M, Koklesova L, Liskova A, Kubatka P, Kozubik E, et al. MiRNA expression profiles in Luminal A breast cancer-implications in biology, prognosis, and prediction of response to hormonal treatment. Int J Mol Sci. 2020;21:7691.
Zhao Z, Guo Y, Liu Y, Sun L, Chen B, Wang C, et al. Individualized lncRNA differential expression profile reveals heterogeneity of breast cancer. Oncogene. 2021;40:4604–14.
Zhong Y, Pan S, Zhi S, Li Y, **u Z, Wei C, et al. Construction and investigation of circRNA-associated ceRNA regulatory network in molecular subtypes of breast cancer. Curr Comput Aided Drug Des. 2022;18:185–95.
Tan Y, Lin J, Li T, Li J, Xu R, Ju H. LncRNA-mediated posttranslational modifications and reprogramming of energy metabolism in cancer. Cancer Commun. 2020;41:109–20.
Yan H, Bu P. Non-coding RNA in cancer. Essays Biochem. 2021;65:625–39.
Tiwari A, Mukherjee B, Dixit M. MicroRNA key to angiogenesis regulation: miRNA biology and therapy. Curr Cancer Drug Targets. 2018;18:266–77.
Kim T, Croce CM. MicroRNA: trends in clinical trials of cancer diagnosis and therapy strategies. Exp Mol Med. 2023;55:1314–21.
Sempere LF, Azmi AS, Moore A. microRNA-based diagnostic and therapeutic applications in cancer medicine. Wiley Interdiscip Rev RNA. 2021;12: e1662.
Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010;11:597–610.
Hill M, Tran N. Global miRNA to miRNA Interactions: Impacts for miR-21. Trends Cell Biol. 2021;31:3–5.
Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. 2018;172:393–407.
Statello L, Guo C-J, Chen L-L, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22:96–118.
Chen YG, Satpathy AT, Chang HY. Gene regulation in the immune system by long noncoding RNAs. Nat Immunol. 2017;18:962–72.
Li X, Yang L, Chen L-L. The biogenesis, functions, and challenges of circular RNAs. Mol Cell. 2018;71:428–42.
Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20:675–91.
Chen L, Wang C, Sun H, Wang J, Liang Y, Wang Y, et al. The bioinformatics toolbox for circRNA discovery and analysis. Brief Bioinform. 2021;22:1706–28.
Balatti V, Nigita G, Veneziano D, Drusco A, Stein GS, Messier TL, et al. tsRNA signatures in cancer. Proc Natl Acad Sci USA. 2017;114:8071–6.
Wang J-H, Chen W-X, Mei S-Q, Yang Y-D, Yang J-H, Qu L-H, et al. tsRFun: a comprehensive platform for decoding human tsRNA expression, functions and prognostic value by high-throughput small RNA-Seq and CLIP-Seq data. Nucleic Acids Res. 2022;50:D421–31.
Di Fazio A, Gullerova M. An old friend with a new face: tRNA-derived small RNAs with big regulatory potential in cancer biology. Br J Cancer. 2023;128:1625–35.
Yamasaki S, Ivanov P, Hu G-F, Anderson P. Angiogenin cleaves tRNA and promotes stress-induced translational repression. J Cell Biol. 2009;185:35–42.
Saikia M, Krokowski D, Guan B-J, Ivanov P, Parisien M, Hu G, et al. Genome-wide identification and quantitative analysis of cleaved tRNA fragments induced by cellular stress. J Biol Chem. 2012;287:42708–25.
Akiyama Y, Kharel P, Abe T, Anderson P, Ivanov P. Isolation and initial structure-functional characterization of endogenous tRNA-derived stress-induced RNAs. RNA Biol. 2020;17:1116–24.
Li X, Liu X, Zhao D, Cui W, Wu Y, Zhang C, et al. tRNA-derived small RNAs: novel regulators of cancer hallmarks and targets of clinical application. Cell Death Discov. 2021;7:249.
Krishna S, Raghavan S, DasGupta R, Palakodeti D. tRNA-derived fragments (tRFs): establishing their turf in post-transcriptional gene regulation. Cell Mol Life Sci CMLS. 2021;78:2607–19.
Wen J-T, Huang Z-H, Li Q-H, Chen X, Qin H-L, Zhao Y. Research progress on the tsRNA classification, function, and application in gynecological malignant tumors. Cell Death Discov. 2021;7:388.
Yu M, Lu B, Zhang J, Ding J, Liu P, Lu Y. tRNA-derived RNA fragments in cancer: current status and future perspectives. J Hematol OncolJ Hematol Oncol. 2020;13:121.
Ancey P-B, Contat C, Meylan E. Glucose transporters in cancer – from tumor cells to the tumor microenvironment. FEBS J. 2018;285:2926–43.
**ao H, Wang J, Yan W, Cui Y, Chen Z, Gao X, et al. GLUT1 regulates cell glycolysis and proliferation in prostate cancer. Prostate. 2018;78:86–94.
Dai W, Xu Y, Mo S, Li Q, Yu J, Wang R, et al. GLUT3 induced by AMPK/CREB1 axis is key for withstanding energy stress and augments the efficacy of current colorectal cancer therapies. Signal Transduct Target Ther. 2020;5:177.
Kim S, Lee E, Jung J, Lee JW, Kim HJ, Kim J, et al. microRNA-155 positively regulates glucose metabolism via PIK3R1-FOXO3a-cMYC axis in breast cancer. Oncogene. 2018;37:2982–91.
Xu J, Li T, Zhang Y, Qiu D, Chen N, Chai X, et al. C-myc/TSPEAR-AS2 axis facilitates breast cancer growth and metastasis in a GLUT1-dependent glycolysis manner. BioMed Res Int. 2022;2022:4239500.
Cheng H, Kuang S, Tan L, Sun S. Circ_0001955 plays a carcinogenic role in breast cancer via positively regulating GLUT1 via decoying miR-1299. Thorac Cancer. 2022;13:913–24.
Qi C, Qin X, Zhou Z, Wang Y, Yang Q, Liao T. Circ_0072995 promotes cell carcinogenesis via up-regulating miR-149-5p-mediated SHMT2 in breast cancer. Cancer Manag Res. 2020;12:11169–81.
Wan L, Han Q, Zhu B, Kong Z, Feng E. Circ-TFF1 facilitates breast cancer development via regulation of miR-338-3p/FGFR1 axis. Biochem Genet. 2022;60:315–35.
Li Y, Li H, Wang W, Yu X, Xu Q. LINC00346 regulates glycolysis by modulation of glucose transporter 1 in breast cancer cells. Mol Cell Probes. 2020;54: 101667.
Poliaková M, Aebersold DM, Zimmer Y, Medová M. The relevance of tyrosine kinase inhibitors for global metabolic pathways in cancer. Mol Cancer. 2018;17:27.
Shelly M, Pinkas-Kramarski R, Guarino BC, Waterman H, Wang LM, Lyass L, et al. Epiregulin is a potent pan-ErbB ligand that preferentially activates heterodimeric receptor complexes. J Biol Chem. 1998;273:10496–505.
He M, ** Q, Chen C, Liu Y, Ye X, Jiang Y, et al. The miR-186-3p/EREG axis orchestrates tamoxifen resistance and aerobic glycolysis in breast cancer cells. Oncogene. 2019;38:5551–65.
Mikawa T, LLeonart ME, Takaori-Kondo A, Inagaki N, Yokode M, Kondoh H. Dysregulated glycolysis as an oncogenic event. Cell Mol Life Sci CMLS. 2015;72:1881–92.
Sundaram SM, Doughty LA, Sereda MW. Location matters: hexokinase 1 in glucose metabolism and inflammation. Trends Endocrinol Metab TEM. 2022;33:665–7.
Wiel C, Le Gal K, Ibrahim MX, Jahangir CA, Kashif M, Yao H, et al. BACH1 stabilization by antioxidants stimulates lung cancer metastasis. Cell. 2019;178:330-345.e22.
Huang S-L, Huang Z-C, Zhang C-J, **e J, Lei S-S, Wu Y-Q, et al. LncRNA SNHG5 promotes the glycolysis and proliferation of breast cancer cell through regulating BACH1 via targeting miR-299. Breast Cancer. 2022;29:65–76.
Webb BA, Forouhar F, Szu F-E, Seetharaman J, Tong L, Barber DL. Structures of human phosphofructokinase-1 and atomic basis of cancer-associated mutations. Nature. 2015;523:111–4.
Zhu W, Chen X, Guo X, Liu H, Ma R, Wang Y, et al. Low glucose–induced overexpression of HOXC-AS3 promotes metabolic reprogramming of breast cancer. Cancer Res. 2022;82:805–18.
Shi L, Pan H, Liu Z, **e J, Han W. Roles of PFKFB3 in cancer. Signal Transduct Target Ther. 2017;2:17044.
Pilkis SJ, Claus TH, Kurland IJ, Lange AJ. 6-Phosphofructo-2-kinase/fructose-2,6-bisphosphatase: a metabolic signaling enzyme. Annu Rev Biochem. 1995;64:799–835.
Yu H, Luo H, Liu X. Knockdown of circ_0102273 inhibits the proliferation, metastasis and glycolysis of breast cancer through miR-1236-3p/PFKFB3 axis. Anticancer Drugs. 2022;33:323–34.
Wang B, Li D, Ilnytskyy Y, Kovalchuk I, Kovalchuk O. A miR-34a-guided, tRNAiMet-derived, piR_019752-like fragment (tRiMetF31) suppresses migration and angiogenesis of breast cancer cells via targeting PFKFB3. Cell Death Discov. 2022;8:355.
Wong N, Ojo D, Yan J, Tang D. PKM2 contributes to cancer metabolism. Cancer Lett. 2015;356:184–91.
Christofk HR, Vander Heiden MG, Harris MH, Ramanathan A, Gerszten RE, Wei R, et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature. 2008;452:230–3.
Mazurek S, Boschek CB, Hugo F, Eigenbrodt E. Pyruvate kinase type M2 and its role in tumor growth and spreading. Semin Cancer Biol. 2005;15:300–8.
Marsboom G, Zhang G-F, Pohl-Avila N, Zhang Y, Yuan Y, Kang H, et al. Glutamine metabolism regulates the pluripotency transcription factor OCT4. Cell Rep. 2016;16:323–32.
Morfouace M, Lalier L, Oliver L, Cheray M, Pecqueur C, Cartron P-F, et al. Control of glioma cell death and differentiation by PKM2-Oct4 interaction. Cell Death Dis. 2014;5: e1036.
Shen Y, Xu J, Pan X, Zhang Y, Weng Y, Zhou D, et al. LncRNA KCNQ1OT1 sponges miR-34c-5p to promote osteosarcoma growth via ALDOA enhanced aerobic glycolysis. Cell Death Dis. 2020;11:278.
Huang J, Deng X, Chen X, Chang Z, Lu Q, Tang A, et al. Circular RNA KIF4A promotes liver metastasis of breast cancer by reprogramming glucose metabolism. J Oncol. 2022;2022:8035083.
Zheng F, Chen J, Zhang X, Wang Z, Chen J, Lin X, et al. The HIF-1α antisense long non-coding RNA drives a positive feedback loop of HIF-1α mediated transactivation and glycolysis. Nat Commun. 2021;12:1341.
Ye T, Liang Y, Zhang D, Zhang X. MicroRNA-16-1-3p represses breast tumor growth and metastasis by inhibiting PGK1-mediated Warburg effect. Front Cell Dev Biol. 2020;8: 615154.
Sharma D, Singh M, Rani R. Role of LDH in tumor glycolysis: regulation of LDHA by small molecules for cancer therapeutics. Semin Cancer Biol. 2022;87:184–95.
Chen X, Luo R, Zhang Y, Ye S, Zeng X, Liu J, et al. Long noncoding RNA DIO3OS induces glycolytic-dominant metabolic reprogramming to promote aromatase inhibitor resistance in breast cancer. Nat Commun. 2022;13:7160.
**ng Z, Wang R, Wang X, Liu J, Zhang M, Feng K, et al. CircRNA circ-PDCD11 promotes triple-negative breast cancer progression via enhancing aerobic glycolysis. Cell Death Discov. 2021;7:218.
Zan X, Li W, Wang G, Yuan J, Ai Y, Huang J, et al. Circ-CSNK1G1 promotes cell proliferation, migration, invasion and glycolysis metabolism during triple-negative breast cancer progression by modulating the miR-28-5p/LDHA pathway. Reprod Biol Endocrinol RBE. 2022;20:138.
Inoue J, Kishikawa M, Tsuda H, Nakajima Y, Asakage T, Inazawa J. Identification of PDHX as a metabolic target for esophageal squamous cell carcinoma. Cancer Sci. 2021;112:2792–802.
Eastlack SC, Dong S, Ivan C, Alahari SK. Suppression of PDHX by microRNA-27b deregulates cell metabolism and promotes growth in breast cancer. Mol Cancer. 2018;17:100.
Liu Q, Li J, Zhang W, **ao C, Zhang S, Nian C, et al. Glycogen accumulation and phase separation drives liver tumor initiation. Cell. 2021;184:5559-5576.e19.
Favaro E, Bensaad K, Chong MG, Tennant DA, Ferguson DJP, Snell C, et al. Glucose utilization via glycogen phosphorylase sustains proliferation and prevents premature senescence in cancer cells. Cell Metab. 2012;16:751–64.
Ran F, Zhang Y, Shi Y, Liu J, Li H, Ding L, et al. miR-1224-3p promotes breast cancer cell proliferation and migration through PGM5-mediated aerobic glycolysis. J Oncol. 2021;2021:5529770.
Zhu P, Lu J, Zhi X, Zhou Y, Wang X, Wang C, et al. tRNA-derived fragment tRFLys-CTT-010 promotes triple-negative breast cancer progression by regulating glucose metabolism via G6PC. Carcinogenesis. 2021;42:1196–207.
Fong MY, Zhou W, Liu L, Alontaga AY, Chandra M, Ashby J, et al. Breast cancer-secreted miR-122 reprograms glucose metabolism in pre-metastatic niche to promote metastasis. Nat Cell Biol. 2015;17:183–94.
Ren S, Liu J, Feng Y, Li Z, He L, Li L, et al. Knockdown of circDENND4C inhibits glycolysis, migration and invasion by up-regulating miR-200b/c in breast cancer under hypoxia. J Exp Clin Cancer Res CR. 2019;38:388.
Zhang X, Li J, Feng Q. CircRNA circYY1 (hsa_circ_0101187) modulates cell gycolysis and malignancy through regulating YY1 expression by sponging miR-769-3p in breast cancer. Cancer Manag Res. 2021;13:1145–58.
Dou D, Ren X, Han M, Xu X, Ge X, Gu Y, et al. Circ_0008039 supports breast cancer cell proliferation, migration, invasion, and glycolysis by regulating the miR-140-3p/SKA2 axis. Mol Oncol. 2021;15:697–709.
Chen Q, Yang Z, Ding H, Li H, Wang W, Pan Z. CircWHSC1 promotes breast cancer progression by regulating the FASN/AMPK/mTOR axis through sponging miR-195-5p. Front Oncol. 2022;11: 649242.
Sui C, Qu W, Lian Y, Feng C, Zhan Y. Hsa_circ_0069094 knockdown inhibits cell proliferation, migration, invasion and glycolysis, while induces cell apoptosis by miR-661/HMGA1 axis in breast cancer. Anticancer Drugs. 2021;32:829–41.
Liu J, Liu J. Circ_0000442 functions as a tumor repressor in breast cancer by impacting miR-1229-3p and upregulating ZBTB1. Mamm Genome. 2022;33:543–54.
Qiu Z, Wang L, Liu H. Hsa_circ_0001982 promotes the progression of breast cancer through miR-1287-5p/MUC19 axis under hypoxia. World J Surg Oncol. 2021;19:161.
Cao L, Wang M, Dong Y, Xu B, Chen J, Ding Y, et al. Circular RNA circRNF20 promotes breast cancer tumorigenesis and Warburg effect through miR-487a/HIF-1α/HK2. Cell Death Dis. 2020;11:145.
Zheng X, Han H, Liu G-P, Ma Y-X, Pan R-L, Sang L-J, et al. LncRNA wires up Hippo and Hedgehog signaling to reprogramme glucose metabolism. EMBO J. 2017;36:3325–35.
Liu Y, Ma L, Hua F, Min Z, Zhan Y, Zhang W, et al. Exosomal circCARM1 from spheroids reprograms cell metabolism by regulating PFKFB2 in breast cancer. Oncogene. 2022;41:2012–25.
Fang K, Xu Z-J, Jiang S-X, Tang D-S, Yan C-S, Deng Y-Y, et al. ncRNA FGD5-AS1 promotes breast cancer progression by regulating the hsa-miR-195-5p/NUAK2 axis. Mol Med Rep. 2021;23:460.
**ao X, Huang X, Ye F, Chen B, Song C, Wen J, et al. The miR-34a-LDHA axis regulates glucose metabolism and tumor growth in breast cancer. Sci Rep. 2016;6:21735.
Yi G, Wang D, Han J, Jia L, Liu X, He J. circKLHL24 blocks breast cancer development by regulating the miR-1204/ALX4 network. Cancer Biother Radiopharm. 2022;37:684–96.
Zhao Y, Zhong R, Deng C, Zhou Z. Circle RNA circABCB10 modulates PFN2 to promote breast cancer progression, as well as aggravate radioresistance through facilitating glycolytic metabolism via miR-223-3p. Cancer Biother Radiopharm. 2021;36:477–90.
Kagawa Y, Umaru BA, Ariful I, Shil SK, Miyazaki H, Yamamoto Y, et al. Role of FABP7 in tumor cell signaling. Adv Biol Regul. 2019;71:206–18.
Tian X, Yang H, Fang Q, Quan H, Lu H, Wang X. Circ_ZFR affects FABP7 expression to regulate breast cancer progression by acting as a sponge for miR-223-3p. Thorac Cancer. 2022;13:1369–80.
Wu H, Xu J, Gong G, Zhang Y, Wu S. CircARL8B contributes to the development of breast cancer via regulating miR-653-5p/HMGA2 axis. Biochem Genet. 2021;59:1648–65.
Schroeder B, Vander Steen T, Espinoza I, Venkatapoorna CMK, Hu Z, Silva FM, et al. Fatty acid synthase (FASN) regulates the mitochondrial priming of cancer cells. Cell Death Dis. 2021;12:977.
Menendez JA, Lupu R. Fatty acid synthase (FASN) as a therapeutic target in breast cancer. Expert Opin Ther Targets. 2017;21:1001–16.
Huang X, Tan W, Liu Z, Fu X, Li Z, Lai S, et al. EIF4A3-induced circZFAND6 promotes breast cancer proliferation and metastasis through the miR-647/FASN axis. Life Sci. 2023;324: 121745.
Han S, Wei R, Zhang X, Jiang N, Fan M, Huang JH, et al. CPT1A/2-mediated FAO enhancement—a metabolic target in radioresistant breast cancer. Front Oncol. 2019;9:1201.
Das M, Giannoudis A, Sharma V. The role of CPT1A as a biomarker of breast cancer progression: a bioinformatic approach. Sci Rep. 2022;12:16441.
**ong Y, Liu Z, Li Z, Wang S, Shen N, **n Y, et al. Long non-coding RNA nuclear paraspeckle assembly transcript 1 interacts with microRNA-107 to modulate breast cancer growth and metastasis by targeting carnitine palmitoyltransferase-1. Int J Oncol. 2019;55:1125–36.
Nazih H, Bard JM. Cholesterol, oxysterols and LXRs in breast cancer pathophysiology. Int J Mol Sci. 2020;21:1356.
Chen Y-Y, Ge J-Y, Zhu S-Y, Shao Z-M, Yu K-D. Copy number amplification of ENSA promotes the progression of triple-negative breast cancer via cholesterol biosynthesis. Nat Commun. 2022;13:791.
Huang Y, Ge H, Zheng M, Cui Y, Fu Z, Wu X, et al. Serum tRNA-derived fragments (tRFs) as potential candidates for diagnosis of nontriple negative breast cancer. J Cell Physiol. 2020;235:2809–24.
Singh R, Yadav V, Kumar S, Saini N. MicroRNA-195 inhibits proliferation, invasion and metastasis in breast cancer cells by targeting FASN, HMGCR, ACACA and CYP27B1. Sci Rep. 2015;5:17454.
Sun H, Li L, Li W, Yang F, Zhang Z, Liu Z, et al. p53 transcriptionally regulates SQLE to repress cholesterol synthesis and tumor growth. EMBO Rep. 2021;22: e52537.
Helms MW, Kemming D, Pospisil H, Vogt U, Buerger H, Korsching E, et al. Squalene epoxidase, located on chromosome 8q24.1, is upregulated in 8q+ breast cancer and indicates poor clinical outcome in stage I and II disease. Br J Cancer. 2008;99:774–80.
Stopsack KH, Gerke TA, Sinnott JA, Penney KL, Tyekucheva S, Sesso HD, et al. Cholesterol metabolism and prostate cancer lethality. Cancer Res. 2016;76:4785–90.
Qin Y, Hou Y, Liu S, Zhu P, Wan X, Zhao M, et al. A novel long non-coding RNA lnc030 maintains breast cancer stem cell stemness by stabilizing SQLE mRNA and increasing cholesterol synthesis. Adv Sci Weinh Baden-Wurtt Ger. 2021;8:2002232.
Duncan RE, Sarkadi-Nagy E, Jaworski K, Ahmadian M, Sul HS. Identification and functional characterization of adipose-specific phospholipase A2 (AdPLA). J Biol Chem. 2008;283:25428–36.
**ong S, Tu H, Kollareddy M, Pant V, Li Q, Zhang Y, et al. Pla2g16 phospholipase mediates gain-of-function activities of mutant p53. Proc Natl Acad Sci USA. 2014;111:11145–50.
Liu S, Sun Y, Hou Y, Yang L, Wan X, Qin Y, et al. A novel lncRNA ROPM-mediated lipid metabolism governs breast cancer stem cell properties. J Hematol OncolJ Hematol Oncol. 2021;14:178.
Wei X, Tao S, Mao H, Zhu H, Mao L, Pei W, et al. Exosomal lncRNA NEAT1 induces paclitaxel resistance in breast cancer cells and promotes cell migration by targeting miR-133b. Gene. 2023;860: 147230.
Vettore L, Westbrook RL, Tennant DA. New aspects of amino acid metabolism in cancer. Br J Cancer. 2020;122:150–6.
Yuan M, Zhang J, He Y, Yi G, Rong L, Zheng L, et al. Circ_0062558 promotes growth, migration, and glutamine metabolism in triple-negative breast cancer by targeting the miR-876-3p/SLC1A5 axis. Arch Gynecol Obstet. 2022;306:1643–55.
Wang J, Yang K, Cao J, Li L. Knockdown of circular RNA septin 9 inhibits the malignant progression of breast cancer by reducing the expression of solute carrier family 1 member 5 in a microRNA-149-5p-dependent manner. Bioengineered. 2021;12:10624–37.
Li J, Cao F, Yin H-L, Huang Z-J, Lin Z-T, Mao N, et al. Ferroptosis: past, present and future. Cell Death Dis. 2020;11:88.
Koppula P, Zhuang L, Gan B. Cystine transporter SLC7A11/xCT in cancer: ferroptosis, nutrient dependency, and cancer therapy. Protein Cell. 2021;12:599–620.
Yadav P, Sharma P, Sundaram S, Venkatraman G, Bera AK, Karunagaran D. SLC7A11/ xCT is a target of miR-5096 and its restoration partially rescues miR-5096-mediated ferroptosis and anti-tumor effects in human breast cancer cells. Cancer Lett. 2021;522:211–24.
Wang S, Wang Y, Li Q, Li X, Feng X. A novel circular RNA confers trastuzumab resistance in human epidermal growth factor receptor 2-positive breast cancer through regulating ferroptosis. Environ Toxicol. 2022;37:1597–607.
Martineau M, Guzman RE, Fahlke C, Klingauf J. VGLUT1 functions as a glutamate/proton exchanger with chloride channel activity in hippocampal glutamatergic synapses. Nat Commun. 2017;8:2279.
Santos MS, Foss SM, Park CK, Voglmaier SM. Protein interactions of the vesicular glutamate transporter VGLUT1. PLoS ONE. 2014;9: e109824.
Yin J, Tu G, Peng M, Zeng H, Wan X, Qiao Y, et al. GPER-regulated lncRNA-Glu promotes glutamate secretion to enhance cellular invasion and metastasis in triple-negative breast cancer. FASEB J. 2020;34:4557–72.
Yoo HC, Yu YC, Sung Y, Han JM. Glutamine reliance in cell metabolism. Exp Mol Med. 2020;52:1496–516.
Wise DR, Thompson CB. Glutamine addiction: a new therapeutic target in cancer. Trends Biochem Sci. 2010;35:427–33.
Wu Y-Z, Chen Y-H, Cheng C-T, Ann DK, Kuo C-Y. Amino acid restriction induces a long non-coding RNA UBA6-AS1 to regulate GCN2-mediated integrated stress response in breast cancer. FASEB J. 2022;36: e22201.
Choi S-W, Kim H-W, Nam J-W. The small peptide world in long noncoding RNAs. Brief Bioinform. 2019;20:1853–64.
Wu S, Guo B, Zhang L, Zhu X, Zhao P, Deng J, et al. A micropeptide XBP1SBM encoded by lncRNA promotes angiogenesis and metastasis of TNBC via XBP1s pathway. Oncogene. 2022;41:2163–72.
Possemato R, Marks KM, Shaul YD, Pacold ME, Kim D, Birsoy K, et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature. 2011;476:346–50.
Li Q, Gao H, Zhou S, Liao Y. LncRNA PlncRNA-1 overexpression inhibits the growth of breast cancer by upregulating TGF-β1 and downregulating PHGDH. Breast Cancer. 2018;25:619–25.
Luo M-Y, Zhou Y, Gu W-M, Wang C, Shen N-X, Dong J-K, et al. Metabolic and nonmetabolic functions of PSAT1 coordinate signaling cascades to confer EGFR inhibitor resistance and drive progression in lung adenocarcinoma. Cancer Res. 2022;82:3516–31.
Wang H, Fang Q, You S, Wu Y, Zhang C. miRNA-195-5p/PSAT1 feedback loop in human triple-negative breast cancer cells. Genes Genomics. 2023;45:39–47.
Petri BJ, Piell KM, Wilt AE, Howser AD, Winkler L, Whitworth MR, et al. MicroRNA regulation of the serine synthesis pathway in endocrine-resistant breast cancer cells. Endocr Relat Cancer. 2023;30: e230148.
Mishra A, Srivastava A, Pateriya A, Tomar MS, Mishra AK, Shrivastava A. Metabolic reprograming confers tamoxifen resistance in breast cancer. Chem Biol Interact. 2021;347: 109602.
Walter W, Thomalla J, Bruhn J, Fagan DH, Zehowski C, Yee D, et al. Altered regulation of PDK4 expression promotes antiestrogen resistance in human breast cancer cells. Springerplus. 2015;4:689.
Liu X, Miao W, Huang M, Li L, Dai X, Wang Y. Elevated hexokinase II expression confers acquired resistance to 4-Hydroxytamoxifen in breast cancer cells. Mol Cell Proteomics MCP. 2019;18:2273–84.
Sun M, Zhao S, Duan Y, Ma Y, Wang Y, Ji H, et al. GLUT1 participates in tamoxifen resistance in breast cancer cells through autophagy regulation. Naunyn Schmiedebergs Arch Pharmacol. 2021;394:205–16.
Early Breast Cancer Trialists’ Collaborative Group (EBCTCG). Aromatase inhibitors versus tamoxifen in premenopausal women with oestrogen receptor-positive early-stage breast cancer treated with ovarian suppression: a patient-level meta-analysis of 7030 women from four randomised trials. Lancet Oncol. 2022;23:382–92.
Early Breast Cancer Trialists’ Collaborative Group (EBCTCG). Aromatase inhibitors versus tamoxifen in early breast cancer: patient-level meta-analysis of the randomised trials. Lancet Lond Engl. 2015;386:1341–52.
Chakraborty B, Byemerwa J, Krebs T, Lim F, Chang C-Y, McDonnell DP. Estrogen receptor signaling in the immune system. Endocr Rev. 2023;44:117–41.
Pernas S, Tolaney SM. HER2-positive breast cancer: new therapeutic frontiers and overcoming resistance. Ther Adv Med Oncol. 2019;11:1758835919833519.
Gandhi N, Das GM. Metabolic reprogramming in breast cancer and its therapeutic implications. Cells. 2019;8:89.
Zhao Y, Liu H, Liu Z, Ding Y, Ledoux SP, Wilson GL, et al. Overcoming trastuzumab resistance in breast cancer by targeting dysregulated glucose metabolism. Cancer Res. 2011;71:4585–97.
Huang W-H, Yang Q, Zhang C. eIF4A3-induced circWAC promotes breast cancer progression through mediating miR-599/E2F3 axis. Kaohsiung J Med Sci. 2022;38:321–35.
Wang L, Zhou Y, Jiang L, Lu L, Dai T, Li A, et al. CircWAC induces chemotherapeutic resistance in triple-negative breast cancer by targeting miR-142, upregulating WWP1 and activating the PI3K/AKT pathway. Mol Cancer. 2021;20:43.
Park MK, Zhang L, Min K-W, Cho J-H, Yeh C-C, Moon H, et al. NEAT1 is essential for metabolic changes that promote breast cancer growth and metastasis. Cell Metab. 2021;33:2380-2397.e9.
Adams S, Gatti-Mays ME, Kalinsky K, Korde LA, Sharon E, Amiri-Kordestani L, et al. Current landscape of immunotherapy in breast cancer: a review. JAMA Oncol. 2019;5:1205–14.
Zappasodi R, Serganova I, Cohen IJ, Maeda M, Shindo M, Senbabaoglu Y, et al. CTLA-4 blockade drives loss of Treg stability in glycolysis-low tumours. Nature. 2021;591:652–8.
Liu Z, Zheng N, Li J, Li C, Zheng D, Jiang X, et al. N6-methyladenosine-modified circular RNA QSOX1 promotes colorectal cancer resistance to anti-CTLA-4 therapy through induction of intratumoral regulatory T cells. Drug Resist Updat Rev Comment Antimicrob Anticancer Chemother. 2022;65: 100886.
Qin A, Wen Z, Zhou Y, Li Y, Li Y, Luo J, et al. MicroRNA-126 regulates the induction and function of CD4(+) Foxp3(+) regulatory T cells through PI3K/AKT pathway. J Cell Mol Med. 2013;17:252–64.
Ma G, Li C, Zhang Z, Liang Y, Liang Z, Chen Y, et al. Targeted glucose or glutamine metabolic therapy combined with PD-1/PD-L1 checkpoint blockade immunotherapy for the treatment of tumors—mechanisms and strategies. Front Oncol. 2021;11: 697894.
Guo D, Tong Y, Jiang X, Meng Y, Jiang H, Du L, et al. Aerobic glycolysis promotes tumor immune evasion by hexokinase2-mediated phosphorylation of IκBα. Cell Metab. 2022;34:1312-1324.e6.
Huang X, **e X, Wang H, **ao X, Yang L, Tian Z, et al. PDL1 And LDHA act as ceRNAs in triple negative breast cancer by regulating miR-34a. J Exp Clin Cancer Res CR. 2017;36:129.
Boyages J. Radiation therapy and early breast cancer: current controversies. Med J Aust. 2017;207:216–22.
Pitroda SP, Wakim BT, Sood RF, Beveridge MG, Beckett MA, MacDermed DM, et al. STAT1-dependent expression of energy metabolic pathways links tumour growth and radioresistance to the Warburg effect. BMC Med. 2009;7:68.
Li X, Lu P, Li B, Yang R, Chu Y, Zhang Z, et al. Sensitization of hepatocellular carcinoma cells to irradiation by miR-34a through targeting lactate dehydrogenase-A. Mol Med Rep. 2016;13:3661–7.
Stankevicins L, Almeida da Silva AP, Ventura Dos Passos F, Dos Santos Ferreira E, Menks Ribeiro MC, David GM, et al. MiR-34a is up-regulated in response to low dose, low energy X-ray induced DNA damage in breast cells. Radiat Oncol Lond Engl. 2013;8:231.
Wang B, Zheng J, Li R, Tian Y, Lin J, Liang Y, et al. Long noncoding RNA LINC02582 acts downstream of miR-200c to promote radioresistance through CHK1 in breast cancer cells. Cell Death Dis. 2019;10:764.
**ao Y, Yu T-J, Xu Y, Ding R, Wang Y-P, Jiang Y-Z, et al. Emerging therapies in cancer metabolism. Cell Metab. 2023;35:1283–303.
Slack FJ, Chinnaiyan AM. The role of non-coding RNAs in oncology. Cell. 2019;179:1033–55.
Winkle M, El-Daly SM, Fabbri M, Calin GA. Noncoding RNA therapeutics - challenges and potential solutions. Nat Rev Drug Discov. 2021;20:629–51.
Du Y, Wei N, Ma R, Jiang S, Song D. A miR-210-3p regulon that controls the Warburg effect by modulating HIF-1α and p53 activity in triple-negative breast cancer. Cell Death Dis. 2020;11:731.
**ao M, Lou C, **ao H, Yang Y, Cai X, Li C, et al. MiR-128 regulation of glucose metabolism and cell proliferation in triple-negative breast cancer. Br J Surg. 2018;105:75–85.
Faubert B, Solmonson A, DeBerardinis RJ. Metabolic reprogramming and cancer progression. Science. 2020;368:e473.
Gong Y, Ji P, Yang Y-S, ** of triple-negative breast cancer reveals potential therapeutic targets. Cell Metab. 2021;33:51-64.e9.
El Ansari R, McIntyre A, Craze ML, Ellis IO, Rakha EA, Green AR. Altered glutamine metabolism in breast cancer; subtype dependencies and alternative adaptations. Histopathology. 2018;72:183–90.
Cho ES, Kim NH, Yun JS, Cho SB, Kim HS, Yook JI. Breast cancer subtypes underlying EMT-mediated catabolic metabolism. Cells. 2020;9:E2064.
Acknowledgements
Not applicable.
Funding
This study was supported by the National Natural Science Foundation of China (Grant No. 81702078, No. 81901599); the Natural Science Foundation of Jiangsu Province (Grant No. BK20170356); Suzhou Science, Education and Health Youth Science and Technology Project (Grant No. KJXW2020019); Gusu Talent Program; Support for the project of nuclear technology medical application supported by discipline construction (Grant No. XKTJ-HRC2021001); Science and Technology Program of Suzhou (Grant No. SKY2021045, SKJY2021097); Maternal and Child Health Association Project of Jiangsu province (Grant No. FYX202123); “National Tutor System” Training Program for Health Youth Key Talents in Suzhou (Qngg2023008).
Author information
Authors and Affiliations
Contributions
JL and HY designed the review. SLX and LXW collected the literature and wrote the manuscript. YXZ, TM and BW revised the manuscript. All authors read, reviewed, and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Xu, S., Wang, L., Zhao, Y. et al. Metabolism-regulating non-coding RNAs in breast cancer: roles, mechanisms and clinical applications. J Biomed Sci 31, 25 (2024). https://doi.org/10.1186/s12929-024-01013-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12929-024-01013-w