Abstract
Background
Esophageal squamous cell carcinoma (ESCC) is a highly heterogeneous cancer that lacks comprehensive understanding and effective treatment. Although multi-omics study has revealed features and underlying drivers of advanced ESCC, research on molecular characteristics of the early stage ESCC is quite limited.
Materials and methods
We presented characteristics of genomics and transcriptomics in 10 matched pairs of tumor and normal tissues of early ESCC patients in the China region.
Results
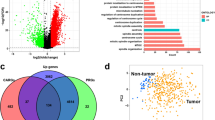
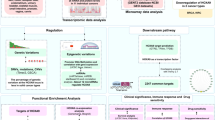
We identified the specific patterns of cancer gene mutations and copy number variations. We also found a dramatic change in the transcriptome, with more than 4,000 genes upregulated in cancer. Among them, more than one-third of HOX family genes were specifically and highly expressed in early ESCC samples of China and validated by RT-qPCR. Gene regulation network analysis indicated that alteration of Hox family genes promoted the proliferation and metabolism remodeling of early ESCC.
Conclusions
We characterized the genomic and transcriptomic landscape of 10 paired normal adjacent and early ESCC tissues in the China region, and provided a new perspective to understand the development of ESCC and insight into potential prevention and diagnostic targets for the management of early ESCC in China.
Similar content being viewed by others
Background
Esophageal cancer (EC), one of the most lethal cancers, is the seventh common cancer type and the sixth leading cause of cancer-related death in the world [1]. In China, there will be approximately 346,600 people newly diagnosed and 323,600 people dying from EC in 2022 [ Raw data can be accessed form GSE213565. Esophageal squamous cell carcinoma Early Esophageal squamous cell carcinoma Whole-exome sequencing Tumor mutational burden Microsatellite instability Copy number variation Gene regulation network Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71(3):209–49. https://doi.org/10.3322/caac.21660. **a C, Dong X, Li H, Cao M, Sun D, He S, et al. Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin Med J (Engl). 2022;135(5):584–90. https://doi.org/10.1097/CM9.0000000000002108. . Adato O, Orenstein Y, Kopolovic J, Juven-Gershon T, Unger R. Quantitative Analysis of Differential Expression of HOX Genes in Multiple Cancers. Cancers (Basel). 2020;12(6) https://doi.org/10.3390/cancers12061572 Rubenstein JH, Shaheen NJ. Epidemiology, Diagnosis, and Management of Esophageal Adenocarcinoma. Gastroenterology. 2015;149(2):302-17 e1. https://doi.org/10.1053/j.gastro.2015.04.053. Wang GQ, Abnet CC, Shen Q, Lewin KJ, Sun XD, Roth MJ, et al. Histological precursors of oesophageal squamous cell carcinoma: results from a 13 year prospective follow up study in a high risk population. Gut. 2005;54(2):187–92. https://doi.org/10.1136/gut.2004.046631. Wei WQ, Chen ZF, He YT, Feng H, Hou J, Lin DM, et al. Long-Term Follow-Up of a Community Assignment, One-Time Endoscopic Screening Study of Esophageal Cancer in China. J Clin Oncol. 2015;33(17):1951–7. https://doi.org/10.1200/JCO.2014.58.0423. Lin DC, Wang MR, Koeffler HP. Genomic and Epigenomic Aberrations in Esophageal Squamous Cell Carcinoma and Implications for Patients. Gastroenterology. 2018;154(2):374–89. https://doi.org/10.1053/j.gastro.2017.06.066. Song Y, Li L, Ou Y, Gao Z, Li E, Li X, et al. Identification of genomic alterations in oesophageal squamous cell cancer. Nature. 2014;509(7498):91–5. https://doi.org/10.1038/nature13176. Abnet CC, Arnold M, Wei WQ. Epidemiology of Esophageal Squamous Cell Carcinoma. Gastroenterology. 2018;154(2):360–73. https://doi.org/10.1053/j.gastro.2017.08.023. Li M, Zhang Z, Wang Q, Yi Y, Li B. Integrated cohort of esophageal squamous cell cancer reveals genomic features underlying clinical characteristics. Nat Commun. 2022;13(1):5268. https://doi.org/10.1038/s41467-022-32962-1. Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–60. https://doi.org/10.1093/bioinformatics/btp324. Repana D, Nulsen J, Dressler L, Bortolomeazzi M, Venkata SK, Tourna A, et al. The Network of Cancer Genes (NCG): a comprehensive catalogue of known and candidate cancer genes from cancer sequencing screens. Genome Biol. 2019;20(1):1. https://doi.org/10.1186/s13059-018-1612-0. Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018;28(11):1747–56. https://doi.org/10.1101/gr.239244.118. Talevich E, Shain AH, Botton T, Bastian BC. CNVkit: Genome-Wide Copy Number Detection and Visualization from Targeted DNA Sequencing. PLoS Comput Biol. 2016;12(4):e1004873. https://doi.org/10.1371/journal.pcbi.1004873. Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011;12(4):R41. https://doi.org/10.1186/gb-2011-12-4-r41. Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29(1):15–21. https://doi.org/10.1093/bioinformatics/bts635. Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30(7):923–30. https://doi.org/10.1093/bioinformatics/btt656. Kanehisa M, Furumichi M, Sato Y, Kawashima M, Ishiguro-Watanabe M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 2023;51(D1):D587–92. https://doi.org/10.1093/nar/gkac963. . Sherman BT, Hao M, Qiu J, Jiao X, Baseler MW, Lane HC, et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022https://doi.org/10.1093/nar/gkac194 . Huynh-Thu VA, Irrthum A, Wehenkel L, Geurts P. Inferring regulatory networks from expression data using tree-based methods. PLoS One. 2010;5(9) https://doi.org/10.1371/journal.pone.0012776 Aran D, Hu Z, Butte AJ. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017;18(1):220. https://doi.org/10.1186/s13059-017-1349-1. Kang SY, Kim DG, Ahn S, Ha SY, Jang KT, Kim KM. Comparative analysis of microsatellite instability by next-generation sequencing, MSI PCR and MMR immunohistochemistry in 1942 solid cancers. Pathol Res Pract. 2022;233:153874. https://doi.org/10.1016/j.prp.2022.153874. Roberts SA, Lawrence MS, Klimczak LJ, Grimm SA, Fargo D, Stojanov P, et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat Genet. 2013;45(9):970–6. https://doi.org/10.1038/ng.2702. Cooper DN, Mort M, Stenson PD, Ball EV, Chuzhanova NA. Methylation-mediated deamination of 5-methylcytosine appears to give rise to mutations causing human inherited disease in CpNpG trinucleotides, as well as in CpG dinucleotides. Hum Genomics. 2010;4(6):406–10. https://doi.org/10.1186/1479-7364-4-6-406. Voutsadakis IA. Amplification of 8p11.23 in cancers and the role of amplicon genes. Life Sci. 2021;264:118729. https://doi.org/10.1016/j.lfs.2020.118729. Cancer Genome Atlas Research N, Analysis Working Group: Asan U, Agency BCC, Brigham, Women’s H, Broad I, et al. Integrated genomic characterization of oesophageal carcinoma. Nature. 2017;541(7636):169–75. https://doi.org/10.1038/nature20805. Suo D, Wang Z, Li L, Chen Q, Zeng T, Liu R, et al. HOXC10 upregulation confers resistance to chemoradiotherapy in ESCC tumor cells and predicts poor prognosis. Oncogene. 2020;39(32):5441–54. https://doi.org/10.1038/s41388-020-1375-4. Yin J, Guo Y. HOXD13 promotes the malignant progression of colon cancer by upregulating PTPRN2. Cancer Med. 2021;10(16):5524–33. https://doi.org/10.1002/cam4.4078. . Amendola PG, Reuten R, Erler JT. Interplay Between LOX Enzymes and Integrins in the Tumor Microenvironment. Cancers (Basel). 2019;11(5) https://doi.org/10.3390/cancers11050729 Pez F, Dayan F, Durivault J, Kaniewski B, Aimond G, Le Provost GS, et al. The HIF-1-inducible lysyl oxidase activates HIF-1 via the Akt pathway in a positive regulation loop and synergizes with HIF-1 in promoting tumor cell growth. Cancer Res. 2011;71(5):1647–57. https://doi.org/10.1158/0008-5472.CAN-10-1516. Goodman FR, Bacchelli C, Brady AF, Brueton LA, Fryns JP, Mortlock DP, et al. Novel HOXA13 mutations and the phenotypic spectrum of hand-foot-genital syndrome. Am J Hum Genet. 2000;67(1):197–202. https://doi.org/10.1086/302961. Jorgensen EM, Ruman JI, Doherty L, Taylor HS. A novel mutation of HOXA13 in a family with hand-foot-genital syndrome and the role of polyalanine expansions in the spectrum of Mullerian fusion anomalies. Fertil Steril. 2010;94(4):1235–8. https://doi.org/10.1016/j.fertnstert.2009.05.057. Hung YC, Ueda M, Terai Y, Kumagai K, Ueki K, Kanda K, et al. Homeobox gene expression and mutation in cervical carcinoma cells. Cancer Sci. 2003;94(5):437–41. https://doi.org/10.1111/j.1349-7006.2003.tb01461.x. Leonard WJ, Lin JX, O’Shea JJ. The gammac Family of Cytokines: Basic Biology to Therapeutic Ramifications. Immunity. 2019;50(4):832–50. https://doi.org/10.1016/j.immuni.2019.03.028. . Rizzo A, Ricci AD, Brandi G. PD-L1, TMB, MSI, and Other Predictors of Response to Immune Checkpoint Inhibitors in Biliary Tract Cancer. Cancers (Basel). 2021;13(3) https://doi.org/10.3390/cancers13030558 Liu X, Zhang M, Ying S, Zhang C, Lin R, Zheng J, et al. Genetic Alterations in Esophageal Tissues From Squamous Dysplasia to Carcinoma. Gastroenterology. 2017;153(1):166–77. https://doi.org/10.1053/j.gastro.2017.03.033. Farshidfar F, Rhrissorrakrai K, Levovitz C, Peng C, Knight J, Bacchiocchi A, et al. Integrative molecular and clinical profiling of acral melanoma links focal amplification of 22q11.21 to metastasis. Nat Commun. 2022;13(1):898. https://doi.org/10.1038/s41467-022-28566-4. Zhang M, Liu ZZ, Aoshima K, Cai WL, Sun H, Xu T, et al. CECR2 drives breast cancer metastasis by promoting NF-kappaB signaling and macrophage-mediated immune suppression. Sci Transl Med. 2022;14(630):eabf5473. https://doi.org/10.1126/scitranslmed.abf5473. Shah N, Sukumar S. The Hox genes and their roles in oncogenesis. Nat Rev Cancer. 2010;10(5):361–71. https://doi.org/10.1038/nrc2826. Lv J, Cao XF, Ji L, Zhu B, Wang DD, Tao L, et al. Association of beta-catenin, Wnt1, Smad4, Hoxa9, and Bmi-1 with the prognosis of esophageal squamous cell carcinoma. Med Oncol. 2012;29(1):151–60. https://doi.org/10.1007/s12032-010-9816-5. Xu C, Li B, Zhao S, ** B, Jia R, Ge J, et al. MicroRNA-186-5p Inhibits Proliferation And Metastasis Of Esophageal Cancer By Mediating HOXA9. Onco Targets Ther. 2019;12:8905–14. https://doi.org/10.2147/OTT.S227920. Chen KN, Gu ZD, Ke Y, Li JY, Shi XT, Xu GW. Expression of 11 HOX genes is deregulated in esophageal squamous cell carcinoma. Clin Cancer Res. 2005;11(3):1044–9 (https://www.ncbi.nlm.nih.gov/pubmed/15709170). Tang L, Cao Y, Song X, Wang X, Li Y, Yu M, et al. HOXC6 promotes migration, invasion and proliferation of esophageal squamous cell carcinoma cells via modulating expression of genes involved in malignant phenotypes. PeerJ. 2019;7:e6607. https://doi.org/10.7717/peerj.6607. Zhang E, Han L, Yin D, He X, Hong L, Si X, et al. H3K27 acetylation activated-long non-coding RNA CCAT1 affects cell proliferation and migration by regulating SPRY4 and HOXB13 expression in esophageal squamous cell carcinoma. Nucleic Acids Res. 2017;45(6):3086–101. https://doi.org/10.1093/nar/gkw1247. Zhang X, Peng L, Luo Y, Zhang S, Pu Y, Chen Y, et al. Dissecting esophageal squamous-cell carcinoma ecosystem by single-cell transcriptomic analysis. Nat Commun. 2021;12(1):5291. https://doi.org/10.1038/s41467-021-25539-x. Zhang J, Wang H, Wu H, Qiang G. The Functionalities and Clinical Significance of Tumor-Infiltrating Immune Cells in Esophageal Squamous Cell Carcinoma. Biomed Res Int. 2021;2021:8635381. https://doi.org/10.1155/2021/8635381. Yuan J, Weng Z, Tan Z, Luo K, Zhong J, **e X, et al. Th1-involved immune infiltrates improve neoadjuvant chemoradiotherapy response of esophageal squamous cell carcinoma. Cancer Lett. 2023;553:215959. https://doi.org/10.1016/j.canlet.2022.215959. Not applicable. This research was funded by the Natural Science Foundation of Fujian Province of China, grant number 2019J01170 and the Natural Science Foundation of Gansu Province of China, grant number 21JR1RA021. Cai** Li, Jianjun Wang, Zhixiang Hu and Shenglin Huang designed the study; Cai** Li collected the data and clinical information; **g**g Zhao, Hena Zhang performed the bioinformatics analysis; Qiaojuan Li conducted the experiments; **g**g Zhao, **ya Jia and Zhixiang Hu wrote and revised the manuscript. All authors have read and agreed to the published version of the manuscript. The study was implemented in accordance with Declaration of Helsinki and relevant guidelines by the institutional ethics committee. The study was approved by Ethics Committee of Second Affiliated Hospital of Fujian Medical University (Approval-Number: 2020-246, 2020-05-22). All methods were carried out in accordance with relevant guidelines and regulations under the ethical approval. All patients gave their informed consents. Not applicable. The authors declare no competing interests. Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Figure S1. The 8p11.23 amplification impacted the transcriptome. Table S1. A summary of variations of early ESCC by WES. Table S2. The DEGs between tumor and normal. TableS3. Clinical data of the patients in this study. Table S4. TCGA samples used in the study. Table S5. Primers and RNA sequences used in this study. Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data. Zhao, J., Jia, X., Li, Q. et al. Genomic and transcriptional characterization of early esophageal squamous cell carcinoma.
BMC Med Genomics 16, 153 (2023). https://doi.org/10.1186/s12920-023-01588-7 Received: Accepted: Published: DOI: https://doi.org/10.1186/s12920-023-01588-7Availability of data and materials
Abbreviations
References
Acknowledgements
Funding
Author information
Contributions
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Consent for publication
Competing interests
Additional information
Publisher’s Note
Supplementary Information
Additional file 1:
Additional file 2:
Additional file 3:
Additional file 4:
Rights and permissions
About this article
Cite this article
Keywords