Abstract
Background
Centrosome amplification is a well-recognized oncogenic driver of tumor initiation and progression across a variety of malignancies and has been linked with tumor aggressiveness, metastasis, and adverse prognosis. Nevertheless, the significance of centrosome amplification in HCC is not well understood.
Methods
The TCGA dataset was downloaded for centrosome amplification-related signature construction using the LASSO-penalized Cox regression algorithm, while the ICGC dataset was obtained for signature validation. Single-cell RNA sequencing from GSE149614 was analyzed to profile gene expression and the liver tumor niche.
Results
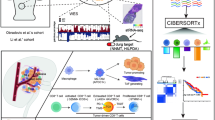
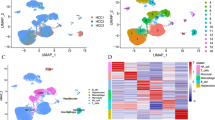
A total of 134 centrosome amplification-related prognostic genes in HCC were detected and 6 key prognostic genes (SSX2IP, SPAG4, SAC3D1, NPM1, CSNK1D, and CEP55) among them were screened out to construct a signature with both high sensitivity and specificity in diagnosis and prognosis of HCC patients. The signature, as an independent factor, was associated with frequent recurrences, high mortality rates, advanced clinicopathologic features, and high vascular invasions. Moreover, the signature was intimately associated with cell cycle-related pathways and TP53 mutation profile, suggesting its underlying role in accelerating cell cycle progression and leading to liver cancer development. Meanwhile, the signature was also closely correlated with immunosuppressive cell infiltration and immune checkpoint expression, making it a vital immunosuppressive factor in the tumor microenvironment. Upon single-cell RNA sequencing, SSX2IP and SAC3D1 were found to be specially expressed in liver cancer stem-like cells, where they promoted cell cycle progression and hypoxia.
Conclusions
This study provided a direct molecular link of centrosome amplification with clinical characteristics, tumor microenvironment, and clinical drug-response, highlighting the critical role of centrosome amplification in liver cancer development and therapy resistance, thereby providing valuable insights into prognostic prediction and therapeutic response of HCC.









Similar content being viewed by others
Data availability statement
The datasets obtained and explored in this study are available in the TCGA (https://portal.gdc.cancer.gov/) and ICGC (https://icgc.org/). The original contributions presented in the study are included in the article/supplementary material. Further inquiries can be directed to the corresponding author.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–249
Hilmi M, Neuzillet C, Calderaro J, Lafdil F, Pawlotsky JM, Rousseau B. Angiogenesis and immune checkpoint inhibitors as therapies for hepatocellular carcinoma: current knowledge and future research directions. J Immunother Cancer. 2019;7(1):333
Chaudhary K, Poirion OB, Lu L, Huang S, Ching T, Garmire LX. Multimodal meta-analysis of 1494 hepatocellular carcinoma samples reveals significant impact of consensus driver genes on phenotypes. Clin Cancer Res. 2019;25(2):463–472
Zhu XD, Tang ZY, Sun HC. Targeting angiogenesis for liver cancer: past, present, and future. Genes Dis. 2020;7(3):328–335
Donisi C, Puzzoni M, Ziranu P, Lai E, Mariani S, Saba G, et al. Immune checkpoint inhibitors in the treatment of HCC. Front Oncol. 2020;10: 601240
Ahn JC, Teng PC, Chen PJ, Posadas E, Tseng HR, Lu SC, et al. Detection of circulating tumor cells and their implications as a biomarker for diagnosis, prognostication, and therapeutic monitoring in hepatocellular carcinoma. Hepatology. 2021;73(1):422–436
Johnson P, Zhou Q, Dao DY, Lo YMD. Circulating biomarkers in the diagnosis and management of hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol. 2022;19(10):670–681
Erstad DJ, Tanabe KK. Prognostic and therapeutic implications of microvascular invasion in hepatocellular carcinoma. Ann Surg Oncol. 2019;26(5):1474–1493
Singal AG, Lampertico P, Nahon P. Epidemiology and surveillance for hepatocellular carcinoma: new trends. J Hepatol. 2020;72(2):250–261
LoMastro GM, Holland AJ. The emerging link between centrosome aberrations and metastasis. Dev Cell. 2019;49(3):325–331
So C, Seres KB, Steyer AM, Mönnich E, Clift D, Pejkovska A, et al. A liquid-like spindle domain promotes acentrosomal spindle assembly in mammalian oocytes. Science. 2019;364(6447):e9557
Nigg EA, Holland AJ. Once and only once: mechanisms of centriole duplication and their deregulation in disease. Nat Rev Mol Cell Biol. 2018;19(5):297–312
Morris EJ, Kawamura E, Gillespie JA, Balgi A, Kannan N, Muller WJ, et al. Stat3 regulates centrosome clustering in cancer cells via Stathmin/PLK1. Nat Commun. 2017;8:15289
Bao J, Yu Y, Chen J, He Y, Chen X, Ren Z, et al. MiR-126 negatively regulates PLK-4 to impact the development of hepatocellular carcinoma via ATR/CHEK1 pathway. Cell Death Dis. 2018;9(10):1045
Chan CY, Yuen VW, Chiu DK, Goh CC, Thu KL, Cescon DW, et al. Polo-like kinase 4 inhibitor CFI-400945 suppresses liver cancer through cell cycle perturbation and eliciting antitumor immunity. Hepatology. 2022. https://doi.org/10.1002/hep.32461
Liu Y, Liang W, Chang Y, He Z, Wu M, Zheng H, et al. CEP192 is a novel prognostic marker and correlates with the immune microenvironment in hepatocellular carcinoma. Front Immunol. 2022;13: 950884
Geeleher P, Cox N, Huang RS. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE. 2014;9(9): e107468
Ham SW, Jeon HY, ** X, Kim EJ, Kim JK, Shin YJ, et al. TP53 gain-of-function mutation promotes inflammation in glioblastoma. Cell Death Differ. 2019;26(3):409–425
Han H, Jain AD, Truica MI, Izquierdo-Ferrer J, Anker JF, Lysy B, et al. Small-molecule MYC inhibitors suppress tumor growth and enhance immunotherapy. Cancer Cell. 2019;36(5):483–97.e15
Finn RS, Qin S, Ikeda M, Galle PR, Ducreux M, Kim TY, et al. Atezolizumab plus bevacizumab in unresectable hepatocellular carcinoma. N Engl J Med. 2020;382(20):1894–1905
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24(10):1550–1558
Xu F, ** T, Zhu Y, Dai C. Immune checkpoint therapy in liver cancer. J Exp Clin Cancer Res. 2018;37(1):110
Lee TK, Guan XY, Ma S. Cancer stem cells in hepatocellular carcinoma—from origin to clinical implications. Nat Rev Gastroenterol Hepatol. 2022;19(1):26–44
Raoul JL, Forner A, Bolondi L, Cheung TT, Kloeckner R, de Baere T. Updated use of TACE for hepatocellular carcinoma treatment: how and when to use it based on clinical evidence. Cancer Treat Rev. 2019;72:28–36
Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, Blanc JF, et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359(4):378–390
Guinn BA, Bland EA, Lodi U, Liggins AP, Tobal K, Petters S, et al. Humoral detection of leukaemia-associated antigens in presentation acute myeloid leukaemia. Biochem Biophys Res Commun. 2005;335(4):1293–1304
Denniss FA, Breslin A, Ingram W, Hardwick NR, Mufti GJ, Guinn BA. The leukaemia-associated antigen, SSX2IP, is expressed during mitosis on the surface of myeloid leukaemia cells. Br J Haematol. 2007;138(5):668–669
Guinn B, Greiner J, Schmitt M, Mills KI. Elevated expression of the leukemia-associated antigen SSX2IP predicts survival in acute myeloid leukemia patients who lack detectable cytogenetic rearrangements. Blood. 2009;113(5):1203–1204
Li P, Lin Y, Zhang Y, Zhu Z, Huo K. SSX2IP promotes metastasis and chemotherapeutic resistance of hepatocellular carcinoma. J Transl Med. 2013;11:52
Han ME, Kim JY, Kim GH, Park SY, Kim YH, Oh SO. SAC3D1: a novel prognostic marker in hepatocellular carcinoma. Sci Rep. 2018;8(1):15608
Wang H, Shi X. SAC3D1 activates Wnt/β-catenin signalling in hepatocellular carcinoma. Mol Med Rep. 2022. https://doi.org/10.3892/mmr.2022.12833
Godinho SA, Picone R, Burute M, Dagher R, Su Y, Leung CT, et al. Oncogene-like induction of cellular invasion from centrosome amplification. Nature. 2014;510(7503):167–171
Liu T, Yu J, Ge C, Zhao F, Chen J, Miao C, et al. Sperm associated antigen 4 promotes SREBP1-mediated de novo lipogenesis via interaction with lamin A/C and contributes to tumor progression in hepatocellular carcinoma. Cancer Lett. 2022;536: 215642
Zhang D, Wu F, Song J, Meng M, Fan X, Lu C, et al. A role for the NPM1/PTPN14/YAP axis in mediating hypoxia-induced chemoresistance to sorafenib in hepatocellular carcinoma. Cancer Cell Int. 2022;22(1):65
Zhu M, Zhang J, Bian S, Zhang X, Shen Y, Ni Z, et al. Circadian gene CSNK1D promoted the progression of hepatocellular carcinoma by activating Wnt/β-catenin pathway via stabilizing dishevelled segment polarity protein 3. Biol Proced Online. 2022;24(1):21
Li L, Lei Q, Zhang S, Kong L, Qin B. Screening and identification of key biomarkers in hepatocellular carcinoma: evidence from bioinformatic analysis. Oncol Rep. 2017;38(5):2607–2618
Chen CH, Lu PJ, Chen YC, Fu SL, Wu KJ, Tsou AP, et al. FLJ10540-elicited cell transformation is through the activation of PI3-kinase/AKT pathway. Oncogene. 2007;26(29):4272–4283
Huang XY, Huang ZL, Zhang PB, Huang XY, Huang J, Wang HC, et al. CircRNA-100338 is associated with mTOR signaling pathway and poor prognosis in hepatocellular carcinoma. Front Oncol. 2019;9:392
Chevrier S, Levine JH, Zanotelli VRT, Silina K, Schulz D, Bacac M, et al. An immune atlas of clear cell renal cell carcinoma. Cell. 2017;169(4):736–49.e18
O’Leary B, Cutts RJ, Liu Y, Hrebien S, Huang X, Fenwick K, et al. The genetic landscape and clonal evolution of breast cancer resistance to palbociclib plus fulvestrant in the PALOMA-3 trial. Cancer Discov. 2018;8(11):1390–1403
Fukasawa K. Centrosome amplification, chromosome instability and cancer development. Cancer Lett. 2005;230(1):6–19
Adon AM, Zeng X, Harrison MK, Sannem S, Kiyokawa H, Kaldis P, et al. Cdk2 and Cdk4 regulate the centrosome cycle and are critical mediators of centrosome amplification in p53-null cells. Mol Cell Biol. 2010;30(3):694–710
Marteil G, Guerrero A, Vieira AF, de Almeida BP, Machado P, Mendonça S, et al. Over-elongation of centrioles in cancer promotes centriole amplification and chromosome missegregation. Nat Commun. 2018;9(1):1258
Zhang X, Wei C, Liang H, Han L. Polo-like kinase 4’s critical role in cancer development and strategies for Plk4-targeted therapy. Front Oncol. 2021;11: 587554
Levine MS, Bakker B, Boeckx B, Moyett J, Lu J, Vitre B, et al. Centrosome amplification is sufficient to promote spontaneous tumorigenesis in mammals. Dev Cell. 2017;40(3):313–22.e5
Otto T, Sicinski P. Cell cycle proteins as promising targets in cancer therapy. Nat Rev Cancer. 2017;17(2):93–115
Arnandis T, Monteiro P, Adams SD, Bridgeman VL, Rajeeve V, Gadaleta E, et al. Oxidative stress in cells with extra centrosomes drives non-cell-autonomous invasion. Dev Cell. 2018;47(4):409–24.e9
Lingle WL, Barrett SL, Negron VC, D’Assoro AB, Boeneman K, Liu W, et al. Centrosome amplification drives chromosomal instability in breast tumor development. Proc Natl Acad Sci U S A. 2002;99(4):1978–1983
Zhu J, Tsai HJ, Gordon MR, Li R. Cellular stress associated with aneuploidy. Dev Cell. 2018;44(4):420–431
Tripathi R, Modur V, Senovilla L, Kroemer G, Komurov K. Suppression of tumor antigen presentation during aneuploid tumor evolution contributes to immune evasion. Oncoimmunology. 2019;8(11):1657374
Nagarsheth N, Wicha MS, Zou W. Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat Rev Immunol. 2017;17(9):559–572
Suthen S, Lim CJ, Nguyen PHD, Dutertre CA, Lai HLH, Wasser M, et al. Hypoxia-driven immunosuppression by Treg and type-2 conventional dendritic cells in HCC. Hepatology. 2022;76(5):1329–1344
He Q, Liu M, Huang W, Chen X, Zhang B, Zhang T, et al. IL-1β-induced elevation of solute carrier family 7 member 11 promotes hepatocellular carcinoma metastasis through up-regulating programmed death ligand 1 and colony-stimulating factor 1. Hepatology. 2021;74(6):3174–3193
Acknowledgements
We appreciate each and every individual relevant to development and maintenance of the R packages such as seurat, limma, glmnet, and survival, which will hasten the retrospective statistical analysis of high-throughput sequencing data for cancer.
Funding
This work was supported by grants from National Natural Science Foundation of China, the Natural Science Foundation of China (81172537, 81272900, 81772828), Scientific and Technological Project of Guangdong Province (2016A020216028), the China Postdoctoral Science Fund (2018M643045), the Basic and Applied Basic Research Project of Guangzhou Science and Technology Bureau (202201011003), and the Ph.D. Start-up Fund of Second Affiliated Hospital of Guangzhou Medical University.
Author information
Authors and Affiliations
Contributions
Conception and design: YLL and JQC. Foundation support: YLL and JQC. Acquisition and analysis of data: YLL, MH, XRK, YTC, JZ, and ZQT. Interpretation of data: YLL, MH, XRK, YTC, JZ, and ZQT. Drafting the manuscript and revising for submission quality: YLL and JQC. Study supervision: JQC.
Corresponding author
Ethics declarations
Conflict of interest
Yanli Liu, Min He, **nrong Ke, Yuting Chen, Jie Zhu, Ziqing Tan and **gqi Chen declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Y., He, M., Ke, X. et al. Centrosome amplification-related signature correlated with immune microenvironment and treatment response predicts prognosis and improves diagnosis of hepatocellular carcinoma by integrating machine learning and single-cell analyses. Hepatol Int 18, 108–130 (2024). https://doi.org/10.1007/s12072-023-10538-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12072-023-10538-5