Abstract
Background
Cryptosporidium is a gastrointestinal protozoan that widely exists in nature, it is an established zoonotic pathogen. Infected cattle are considered to be associated with cryptosporidiosis outbreaks in humans. In the present study, we aimed to assess the prevalence and species distribution of Cryptosporidium in dairy cattle in Central Inner Mongolia.
Methods
We focused on the small subunit ribosomal RNA gene (SSU rRNA) of Cryptosporidium and 60-kDa glycoprotein gene (gp60) of Cryptosporidium parvum. We collected 505 dairy cattle manure samples from 6 sampling sites in Inner Mongolia in 2021; the samples were divided into 4 groups based on age. DNA extraction, polymerase chain reaction (PCR), sequence analysis, and restriction fragment length polymorphism (RFLP) using SspI and MboII restriction endonucleases were performed. RFLP analysis was performed to determine the prevalence and species distribution of Cryptosporidium.
Results
SSU rRNA PCR revealed that the overall prevalence of Cryptosporidium infection was 29.90% (151/505), with a prevalence of 37.67% (55/146) and 26.74% (96/359) in diarrheal and nondiarrheal samples, respectively; these differences were significant. The overall prevalence of Cryptosporidium infection at the 6 sampling sites ranged from 0 to 47.06% and that among the 4 age groups ranged from 18.50 to 43.81%. SSU rRNA sequence analysis and RFLP analysis revealed the presence of 4 Cryptosporidium species, namely, C. bovis (44.37%), C. andersoni (35.10%), C. ryanae (21.85%), and C. parvum (11.92%), along with a mixed infection involving two or three Cryptosporidium species. Cryptosporidium bovis or C. andersoni was the most common cause of infection in the four age groups. The subtype of C. parvum was successfully identified as IIdA via gp60 analysis; all isolates were identified as the subtype IIdA19G1.
Conclusions
To the best of our knowledge, this is the first report of dairy cattle infected with four Cryptosporidium species in Inner Mongolia, China, along with a mixed infection involving two or three Cryptosporidium species, with C. bovis and C. andersoni as the dominant species. Moreover, this is the first study to identify C. parvum subtype IIdA19G1 in cattle in Inner Mongolia. Our study findings provide detailed information on molecular epidemiological investigation of bovine cryptosporidiosis in Inner Mongolia, suggesting that dairy cattle in this region are at risk of transmitting cryptosporidiosis to humans.
Similar content being viewed by others
Background
Cryptosporidium is an important protozoan pathogen [15]; moreover, it may be lethal to immunosuppressed individuals [6]. In farm animals, cryptosporidiosis is the main cause of diarrhea in neonatal livestock and remains one of the most important diseases affecting neonatal calves [34]. In China, the average prevalence of Cryptosporidium infection in humans was 2.97% in 27 provinces between 1987, when it was first reported, and 2018 [17]. Between 1984 and 2016, 18.9% of common livestock (cattle, goats, sheep, horses, pigs, and buffaloes) were infected with Cryptosporidium spp. globally; moreover, domestic hoofed animals (camels, yaks, donkeys, alpacas, and llamas) exhibited a Cryptosporidium infection prevalence of 13.6%. Conventional microscopy (CM) and polymerase chain reaction (PCR) revealed that 23.4% of common livestock were positive for Cryptosporidium spp. infection. The pooled prevalence of Cryptosporidium infection in cattle was 22.5% (CM) or 29.1% (PCR). The prevalence of Cryptosporidium infection in livestock in different regions is mostly in the range of 5–30%. The highest and lowest prevalence of Cryptosporidium infection have been reported in America (26%) and Africa (14%), respectively; its highest prevalence observed in New Zealand is lower than that in other regions. Among 53 countries, livestock in Canada (60%) exhibited the highest infection rate, whereas those in China, Thailand, and Germany (8%) had the lowest infection rates [4]. In 1986, the first report of bovine Cryptosporidium infection in China was published in Lanzhou, Gansu Province [35]. Until 2016, Cryptosporidium species were distributed in 19 provinces in China, with an overall infection rate of 11.9% and average infection rate of 10.44% in dairy cattle [2]. During the same period, the overall infection rate of bovine Cryptosporidium in China was 14.50% and the prevalence in dairy cattle was 13.98% [13]. The pooled prevalence of Cryptosporidium infection in dairy cattle in 23 provinces in China was 17.0% during 2008–2018; this prevalence of varied among different provinces in China, with the highest and lowest prevalence observed in Heilongjiang (35.6%) and Tian** (4.3%), respectively [9]. Inner Mongolia is located on the northern border of China, spanning 28°52′ longitude from east to west, with a linear distance of > 2400 km, and 15°59′ latitude from north to south, with a linear distance of 1700 km. Currently, only two studies in Chinese in Inner Mongolia have reported the prevalence of Cryptosporidium infection in dairy cattle to be 24.56% (14/57) [36] and 14.92% (44/295) [37] using CM and PCR, respectively, and only C. andersoni was identified in the latter. In the present study, we aimed to investigate the prevalence and species distribution of Cryptosporidium in dairy cattle in Central Inner Mongolia.
Methods
Study areas and sample collection
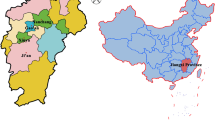
From March to September 2021, 505 fresh fecal samples were randomly collected from 4 intensive dairy farms and 2 free-ranging dairy farms in the vicinity of Tumed Left Banner, Horinger County, Togtoh County, Dalad Banner, and Hanggin Rear Banner (113°34′E–118°28′E, 24°29′N–30°04′N) in Central Inner Mongolia. The fecal samples were collected via rectal sampling from dairy cattle or from the inner top layer of the fresh feces. These samples were obtained from 103 preweaned calves (aged 0–60 days), 105 postweaned calves (aged 61–180 days), 124 young cattle (aged 181–360 days), and 173 adult cattle (aged > 361 days). Information regarding whether the animals experienced diseases such as diarrhea was recorded during sampling; the samples were transferred to the laboratory and stored at 4 °C until later use.
DNA extraction and PCR amplification
DNA was extracted from 505 fecal samples in a biosafety cabinet using E.Z.N.A® Stool DNA Kit (Omega Biotek, Norcross, GA, USA) according to the manufacturer’s instructions and was stored at − 20 °C for subsequent experiments.
The extracted DNA was used as a template and the small subunit ribosomal RNA gene (SSU rRNA) of Cryptosporidium [38] was amplified via nested PCR (annealing temperatures of 55 and 58 °C) using Premix Taq™ (TaKaRa Taq™ Version 2.0 plus dye) (TaKaRa, Bei**g, China). Positive PCR products were sent to a commercial company (Sangon Biotech, Shanghai, China) for sequencing. Simultaneously, SSU rRNA positive amplification products were subjected to restriction fragment length polymorphism (RFLP) analysis using the restriction enzymes (SspI and MboII (TaKaRa) [39]. The results of RFLP and SSU rRNA gene bidirectional sequencing analyses were used to analyze the extracted DNA of C. parvum and perform nested PCR (annealing temperatures of 52 °C and 50 °C) of gp60 [38]. The sequencing results of gp60 were used to identify the subtype of C. parvum [40].
Sequence analysis
The sequences were aligned with reference sequences downloaded from GenBank (http://www.ncbi.nlm.nih.gov) using the MEGA 5.0 software (http://www.megasoftware.net/). The BLAST online platform was used to analyze the sequencing results. Phylogenetic analyses were performed using the concatenated dataset of gp60 sequences. Using the NeighborJoining (NJ) algorithm, phylogenetic trees were constructed based on a matrix of evolutionary distances calculated via the Kimura 2-parameter model of the MEGA 7.0 software. Bootstrap analysis was performed using 1000 replicates to assess the robustness of clusters.
Statistical analysis
Chi-square test was performed and 95% confidence interval (CI) was determined using SPSS Statistics 21.0 (IBM Corp., New York, NY, USA) to compare Cryptosporidium infection rates among different sampling sites and age groups as well as between the diarrheal and nondiarrheal groups. A two-tailed p-value of < 0.05 was considered to indicate statistical significance.
Results
Cryptosporidium infection status
For the SSU rRNA of Cryptosporidium, the PCR amplification of 505 samples yielded positive results in 151 samples, with the overall prevalence of Cryptosporidium infection being 29.90% (151/505). The overall prevalence in diarrheal and nondiarrheal samples was 37.67% (55/146) and 26.74% (96/359), respectively (Table 1); this difference was significant, with an odds ratio (OR) of 1.656 (95% CI: 1.101–2.491, p = 0.015).
The overall prevalence of Cryptosporidium infection in all samples at the 6 sampling sites was 39.29% (54/140), 24.55% (27/110), 22.50% (27/120), 31.82% (35/100), 47.06% (8/17), and 0% (0/8). A significant difference was observed between Tumed Left Banner 1 and Tumed Left Banner 2, with an OR of 1.930 (95% CI: 1.112–3.351, p = 0.019). Moreover, there was a highly significant difference between Tumed Left Banner 1 and Horinger County, with an OR of 2.136 (95% CI: 1.251–3.738, p = 0.005). Further, no significant differences were observed between the other two farms (p > 0.05). The prevalence of Cryptosporidium infection in diarrheal samples at the 6 sampling sites was 45.45% (25/55), 50% (6/12), 27.50% (11/40), 33.33% (13/39), 0% (0/0), and 0% (0/0; Table 1); only Tumed Left Banner 2 farm showed significant difference in prevalence between diarrheal and nondiarrheal samples, with an OR of 3.667 (95% CI: 1.072–12.547, p = 0.030; Table 1).
The overall prevalence of Cryptosporidium infection in all samples was 27.18% (27/103), 43.81% (46/105), 37.10% (46/124), and 18.50% (32/173) in preweaned calves, postweaned calves, young cattle, and adult cattle, respectively. A highly significant difference was observed in the prevalence between pre- and postweaned calves [OR of 0.456 (95% CI: 0.254–0.817, p = 0.008)], between postweaned calves and adult cattle [OR of 3.435 (95% CI: 1.994–5.919, p = 0.000)], and between young and adult cattle [OR of 2.599 (95% CI: 1.531–4.411, p = 0.000)]. The differences in prevalence between the remaining two age groups were not significant (p > 0.05). The prevalence of Cryptosporidium infection in diarrheal samples was 30% (9/30), 45.59% (31/68), 50% (10/20), and 17.86% (5/28) in preweaned calves, postweaned calves, young cattle, and adult cattle, respectively, with no significant difference being observed between prevalence in diarrheal and nondiarrheal samples within each age group (Table 1).
RFLP and sequence analysis
Overall, 151 PCR amplification products of SSU rRNA gene were analyzed via RFLP, and the results were combined with those of sequencing analysis, four Cryptosporidium species were identified, namely, C. bovis (44.37%, 67/151), C. andersoni (35.10%, 53/151), C. ryanae (21.85%, 33/151), and C. parvum (11.92%, 18/151) along with the presence of mixed infections involving two or three Cryptosporidium species (Table 1). Three intensive dairy farms were infected with four Cryptosporidium species, one intensive dairy farm was infected with three Cryptosporidium species, and one free-ranging dairy farm was infected with two Cryptosporidium species.
Preweaned calves were frequently infected with C. bovis (15/27), followed by C. parvum (14/27), whereas postweaned calves were often infected with C. bovis (29/46), followed by C. ryanae (14/27). Young cattle were mostly infected with C. andersoni (28/46), followed by C. bovis (14/46), whereas adult cattle were often infected with C. andersoni (21/32), followed by C. bovis (9/32), but not with C. parvum. Infection with C. parvum alone occurred only in preweaned calves, whereas infections with the other three Cryptosporidium spp. alone were observed in all four age groups. Mixed infections and four Cryptosporidium species were identified in all age groups except adult cattle (Table 1).
The abovementioned four Cryptosporidium spp. were identified in both diarrheal and nondiarrheal samples; C. bovis (33/55) was the most frequently detected species in diarrheal samples, followed by C. ryanae (16/55), whereas C. andersoni (43/96) was the most frequently detected species in nondiarrheal samples, followed by C. bovis (34/96) (Table 2).
Identification ofC. parvumsubtype.
In total, 15 gp60 sequences were analyzed in this study; phylogenetic analysis of gp60 sequences based on C. parvum showed that gp60 obtained in the present study belonged to the same branch as the reference subtype IId (Fig. 1) and were successfully identified as C. parvum subtype family IIdA19G1 (Table 1).
A phylogenetic tree of Cryptosporidium parvum based on gp60 sequences. The phylogenetic tree was constructed via a NeighborJoining analysis of genetic distances calculated using the Kimura 2-parameter model. Percent bootstrap values of > 50% from 1000 replicates are shown to the left of nodes. The isolates indicated as black triangles (▲) represent the subtype IId, which was identified in cattle in this study
Discussion
To date, several studies worldwide have reported Cryptosporidium infection in cattle [43,44,45,46,47]. The overall prevalence of Cryptosporidium infection in dairy cattle was found to be 29.90% (151/505), which was close to the global pooled prevalence of 29.1% for bovine cryptosporidiosis [56,59,60,61,62,63,64,65,66,67,68,69]; moreover, it was higher than the prevalence reported in only two surveys on Cryptosporidium in Inner Mongolia [36, 37].
In the current study, the overall prevalence of Cryptosporidium infection was between 22.50% and 47.06% at the five sampling sites. The difference in prevalence between Tumed Left Banner 1 and Tumed Left Banner 2 was significant, and that between Tumed Left Banner 1 and Horinger County was highly significant. The maximum prevalence in other provinces in China also differed significantly, from 2.6% (Hebei/Tian**) [65] to 100% (Heilongjiang) [49]. However, it is difficult to compare the prevalence data as they are influenced by various factors, including geographic conditions, climate, sanitation conditions, rearing conditions, total number of samples, sampling season, age of animals, and diagnostic methods [2, 68]; however, there are several reports showing high prevalence in preweaned calves [50,51,52, 54, 55, 60, 63, 65]. Indeed, some studies have reported inconsistencies in the time interval between preweaned and postweaned cows or there was a lack of accurate information regarding the age of sampled cows. If calves aged < 3 months are classified as preweaned calves, it was observed that some postweaned calves should have been classified as preweaned calves. As mentioned above, several factors affect Cryptosporidium infection, including prevalence, age distribution, and the presence or absence of diarrhea. In addition, it is related to the nonspecific immunity acquired through factors such as breast milk, immature immune defenses [10], different feeding patterns [59, 61], unlike the finding in industrialized countries where C. parvum occurs almost exclusively [2, 80]; calves are considered to be the most important contributor to zoonotic cryptosporidiosis [5]. The prevalence of C. parvum infection in dairy cattle in China has dramatically increased in recent years with an increase in their populations [16]. The C. parvum IIa and IId subtypes are zoonotically transmitted [1, 5, 9, 12, 31], and IIa and IId subtypes have been detected in Chinese patients [16]. Although the IIa subtype has not yet been detected in cattle in China, it has been observed in various grazing animals in several provinces, including Inner Mongolia, and is prevalent in neighboring countries of China [16]. With the development of animal husbandry, the prevalence of cryptosporidiosis in China may follow the footsteps of that in industrialized countries and become a rampant zoonotic disease in China. In addition, human infections with C. andersoni and C. bovis have been reported [1, 9]. In summary, the results of this study suggest that there is a risk of Cryptosporidium infection in humans caused via dairy cattle in Inner Mongolia; and biosecurity measures are urgently required to delay the spread of local C. parvum IId subtype and imported C. parvum IIa subtype and other Cryptosporidium species.
Conclusions
To the best of our knowledge, this is the first study to report that dairy cattle in Inner Mongolia were infected with four species of Cryptosporidium and had mixed infections involving of two or three species. Cryptosporidium bovis and C. andersoni were identified to be dominant species infecting dairy cattle in Inner Mongolia. Further, the subtype of C. parvum in dairy cows was confirmed to be IIdA19G1, thereby providing a detailed information on the molecular epidemiological investigation of bovine cryptosporidiosis in this region. Further, studies on cryptosporidiosis in other animals in several regions are warranted to help in identifying and elucidating the zoonotic potential and distribution patterns of Cryptosporidium.
Data availability
All the sequences obtained in our laboratory have been uploaded to the GenBank database under the accession numbers OQ029566 to OQ029580, OP861556 to OP861567, OP861717 to OP861773, OP861775 to OP861799, OP861801 to OP861850. Reference sequence accession numbers: MH049734, KT235713, FJ897787, KC885904, JN867335, AF402285, KU852718, AB237137, KU852719, AF164491, AY873781, AY262034, DQ192501, AY873780, AM937006, KU670813, AY738188, AY873782, AY700401, AY382675, and KU852720.
Abbreviations
- gp60:
-
60 kDa glycoprotein
- SSU rRNA:
-
small subunit ribosomal RNA
- RFLP:
-
restriction fragmentlength polymorphism
- PCR:
-
polymerase chain reaction
- CI:
-
confidence interval
- OR:
-
odds ratio
References
Ryan UM, Feng Y, Fayer R, **ao L. Taxonomy and molecular epidemiology of Cryptosporidium and Giardia – a 50 year perspective (1971–2021). Int J Parasitol. 2021;51(13–14):1099–119.
Gong C, Cao X, Deng L, Li W, Huang X, Lan J, et al. Epidemiology of Cryptosporidium infection in cattle in China: a review. Parasite. 2017;24:1.
Tao W, Ni H, Du H, Jiang J, Zhang X. Molecular detection of Cryptosporidium and Enterocytozoon bieneusi in dairy calves and sika deer in four provinces in Northern China. Parasitol Res. 2020;119(1):105.
Hatam-Nahavandi K, Ahmadpour E, Carmena D, Spotin A, Bangoura B, **ao LH. Cryptosporidium infections in terrestrial ungulates with focus on livestock: a systematic review and meta-analysis. Parasite Vector. 2019;12(1):453.
Li N, Zhao W, Song S, Ye H, Chu W, Guo Y, et al. Diarrhoea outbreak caused by coinfections of Cryptosporidium parvum subtype IIdA20G1 and rotavirus in pre-weaned dairy calves. Transbound Emerg Dis. 2022;69:1606–17.
Kza B, Yin F, Jia B, Lza B. Public health and ecological significance of rodents in Cryptosporidium infections. One Health. 2021;14:100364.
Naguib D, Roellig DM, Arafat N, **ao LH. Genetic characterization of Cryptosporidium cuniculus from rabbits in Egypt. Pathogens. 2021;10:775.
FAO/WHO. Multicriteria-based ranking for risk management of food-borne parasites. Microbiological Risk Assessment Series No. 23. Rome: Food and Agriculture Organization of the United Nations/World Health Organization; 2014.
Ryan U, Zahedi A, Feng Y, **ao L. An update on zoonotic Cryptosporidium species and genotypes in humans. Animals-Basel. 2021;11(11):3307.
Cai Y, Zhang N, Gong Q, Zhao Q, Zhang XX. Prevalence of Cryptosporidium in dairy cattle in China during 2008–2018: a systematic review and meta-analysis. Microb Pathog. 2019;132:193–200.
Ma JY, Li MY, Qi ZZ, Fu M, Sun TF, Elsheikha HM, et al. Waterborne protozoan outbreaks: an update on the global, regional, and national prevalence from 2017 to 2020 and sources of contamination. Sci Total Environ. 2022;806:150562.
Feng Y, Ryan U, **ao L. Genetic diversity and population structure of Cryptosporidium. Trends Parasitol. 2018;34:997–1011.
Wang R, Zhao G, Gong Y, Zhang L. Advances and perspectives on the epidemiology of bovine Cryptosporidium in China in the past 30 years. Front Microbiol. 2017;8:1823.
Kumar D, Panda C, Yun S, Mahapatra RK. An update on Cryptosporidium biology and therapeutic avenues. J Parasit Dis. 2022;46(3):923–39.
Kotloff KL, Nataro JP, Blackwelder WC, Nasrin D, Farag TH, Panchalingam S, et al. Burden and aetiology of diarrhoeal disease in infants and young children in develo** countries (the Global Enteric Multicenter Study, GEMS): a prospective, case-control study. Lancet. 2013;382(9888):209–22.
Guo Y, Ryan U, Feng Y, **ao L. Emergence of zoonotic Cryptosporidium parvum in China. Trends Parasitol. 2022;4:38.
Liu A, Gong B, Liu X, Shen Y, Wu Y, Zhang W, et al. A retrospective epidemiological analysis of human Cryptosporidium infection in China during the past three decades (1987–2018). PLoS Negl Trop Dis. 2020;14(3):e0008146.
Zahedi A, Ryan U. Cryptosporidium – an update with an emphasis on foodborne and waterborne transmission. Res Vet Sci. 2020;132:500–12.
Santin M. Cryptosporidium and Giardia in ruminants. Vet Clin N Am Food Anim Pract. 2020;36(1):223–38.
Zhang Z, Su D, Meng X, Liang R, Wang W, Li N, et al. Cryptosporidiosis outbreak caused by Cryptosporidium parvum subtype IIdA20G1 in neonatal calves. Transbound Emerg Dis. 2021;00:1–8.
Feng Y, **ao L. Molecular epidemiology of Cryptosporidiosis in China. Front Microbiol. 2017;8:1701.
Qi M, Zhang K, Huang M, Wang S, Li J. Longitudinal detection of Cryptosporidium spp. in 1–10-week-old dairy calves on a farm in **njiang, China. Parasitol Res. 2020;119:3839–44.
Li N, Wang R, Cai M, Jiang W, Feng Y, **ao L. Outbreak of cryptosporidiosis due to Cryptosporidium parvum subtype IIdA19G1 in neonatal calves on a dairy farm in China - ScienceDirect. Int J Parasitol. 2019;49(7):569–77.
Azami M, Moghaddam DD, Salehi R, Salehi M. The identification of Cryptosporidium species (Protozoa) in Ifsahan, Iran by PCR-RFLP analysis of the 18S rRNA gene. Mol Biol. 2007;41(5):851–6.
Cai M, Guo Y, Pan B, Li N, Wang X, Tang C, et al. Longitudinal monitoring of Cryptosporidium species in pre-weaned dairy calves on five farms in Shanghai, China. Vet Parasitol. 2017;241:14–9.
Wang TP, Gao YQ, Roellig DM, Li N, Santín M, et al. Sympatric recombination in zoonotic Cryptosporidium leads to emergence of populations with modified host preference. Mol Biol Evol. 2022;39(7):msac150.
**ao L, Feng Y. Molecular epidemiologic tools for waterborne pathogens Cryptosporidium spp. and Giardia duodenalis. Food Waterborne Parasitol. 2017;8–9:14–32.
Mravcová K, Štrkolcová G, Mucha R, Barbušinová E, Goldová M, Kačírová J, et al. Cryptosporidium parvum – zoonotic subtype IIdA15G1 in a slovakian patient. Ann Agric Environ Med. 2020;27(3):485–8.
Yang X, Guo Y, **ao L, Feng Y. Molecular epidemiology of human cryptosporidiosis in low- and middle-income countries. Clin Microbiol Rev. 2021;34(2):e00087–19.
King P, Tyler KM, Hunter PR. Anthroponotic transmission of Cryptosporidium parvum predominates in countries with poorer sanitation: a systematic review and meta-analysis. Parasite Vector. 2019;12(1):16.
Jia R, Huang W, Huang N, Yu Z, Li N, **ao L, et al. High infectivity and unique genomic sequence characteristics of Cryptosporidium parvum in China. PLoS Negl Trop Dis. 2022;16(8):e0010714.
Guo Y, Li N, Ryan U, Feng Y, **ao L. Small ruminants and zoonotic cryptosporidiosis. Parasitol Res. 2021;120:4189–98.
Adamu H, Petros B, Zhang G, Kassa H, Amer S, Ye J, et al. Distribution and clinical manifestations of Cryptosporidium species and subtypes in HIV/AIDS patients in ethiopia. PLoS Negl Trop Dis. 2014;8(4):e2831.
Dong S, Yang Y, Wang J, Yang J, Yang Y, Shi Y, et al. Prevalence of Cryptosporidium infection in the global population:a systematic review and meta-analysis. Acta Parasitol. 2020;65(4):882–9.
Chen Y, Li G, Calf. Cryptosporidiosis in Lanzhou.Chin Vet Sci. 1986;12(12):41–2. (in Chinese).
Yang X, Liu Z, Yang L, Ma Z, Yan Z. A preliminary investigation of Cryptosporidium infection in cattle in Hohhot, Inner Mongolia. J Inn Mong Agric Univ (Nat Sci Ed). 2004;(01):40–2. (in Chinese).
Yin X, Nie F, Guo S, Qian Y, Chen X, Xu Y et al. Detection of Cryptosporidium infection and identification of Cryptosporidium species in dairy cattle in parts of Inner Mongolia by nested PCR. Heilongjiang Anim Sci Vet Med. 2018;(16):92–6. (in Chinese).
Alves M, **ao LH, Sulaiman I, Lal AA, Matos O, Antunes F. Subgenotype Analysis of Cryptosporidium isolates from humans, cattle, and zoo ruminants in Portugal. J Clin Microbiol. 2003;41(6):2744–7.
Feng Y, Ortega Y, He G, Das P, **ao L. Wide geographic distribution of Cryptosporidium bovis and the deer-like genotype in bovines. Vet Parasitol. 2007;144(1–2):1–9.
Sulaiman IM, Hira PR, Zhou L, Al-Ali FM, Al-Shelahi FA, Shweiki HM, et al. Unique endemicity of Cryptosporidiosis in children in Kuwait. J Clin Microbiol. 2005;43(6):2805–9.
Jian F, Wang A, Zhang R, Qi S, Zhao M, Shi W, et al. Common occurrence of Cryptosporidium hominis in horses and donkeys. Infect Genet Evol. 2016;43:261–6.
Li F, Su J, Ba C, Guo Q, Wang T, Yu Z, et al. Different distribution of Cryptosporidium species between horses and donkeys. Infect Genet Evol. 2019;75:103954.
Mi RS, Wang XJ, Huang Y, Mu GD, Zhang YH, Jia HY et al. Prevalence and genoty** of Cryptosporidium species in sheep in China. Appl Environ Microbiol. 2018;00868 – 18.
Fu Y, Dong H, Bian X, Qin Z, Han H, Lang J, et al. Molecular characterizations of Giardia duodenalis based on multilocus genoty** in sheep, goats, and beef cattle in Southwest Inner Mongolia, China. Parasite. 2022;29:33.
Feng S, Chang H, Wang Y, Huang C, He H. Molecular characterization of Cryptosporidium spp. in Brandt’s vole in China. Front Vet Sci. 2020;7:300.
Ni H, SunY, Qin S, Wang Y, Zhao Q, Sun Z, et al. Molecular detection of Cryptosporidium spp. and Enterocytozoon bieneusi infection in wild rodents from six provinces in China. Front Cell Infect Microbiol. 2021;11:783508.
Lang J, Han H, Dong H, Qin Z, Fu Y, Qin H et al. Molecular characterization and prevalence of Cryptosporidium spp. in sheep and goats in western Inner Mongolia, China. Parasitol Res. 2022;36526925.
Zhang K, Wu Y, Wei Z, Zhang Y, Zhang L. Genetic diversity of Cryptosporidium parvum in neonatal calves in **njiang. China Pathogens. 2019;9(9):692.
Zhang W, Wang R, Yang F, Zhang L, Shen Y. Distribution and genetic characterizations of Cryptosporidium spp. in pre-weaned dairy calves in northeastern China’s Heilongjiang Province. PLoS ONE. 2013;8(1):e54857.
Watanabe Y, Yang CH, Ooi HK. Cryptosporidium infection in livestock and first identification of Cryptosporidium parvum genotype in cattle feces in Taiwan. Parasitol Res. 2005;97(3):238–41.
Ma J, Li P, Zhao X, Xu H, Wu W, Wang Y, et al. Occurrence and molecular characterization of Cryptosporidium spp. and Enterocytozoon bieneusi in dairycattle, beef cattle and water buffaloes in China. Vet Parasitol. 2015;207:220–7.
Liang N, Wu Y, Sun M, Chang Y, Zhang L. Molecular epidemiology of Cryptosporidium spp. in dairy cattle in Guangdong Province, South China. Parasitology. 2018;146(1):1–5.
Feng Y, Gong X, Zhu K, Li N, **ao L, et al. Prevalence and genotypic identification of Cryptosporidium spp., Giardia duodenalis and Enterocytozoon bieneusi in pre-weaned dairy calves in Guangdong, China. Parasite Vector. 2019;12:41.
Huang J, Yue D, Qi M, Wang R, Zhao J, Li J, et al. Prevalence and molecular characterization of Cryptosporidium spp. and Giardia duodenalisin dairy cattle in Ningxia, northwestern China. BMC Vet Res. 2014;10(1):292.
Zhang X, Tan Q, Zhou D, Ni X, Liu G, Yang Y, et al. Prevalence and molecular characterization of Cryptosporidium spp. in dairy cattle, northwest China. Parasitol Res. 2015;114(7):2781–7.
Li S, Zou Y, Wang P, Qu M, Zhu X. Prevalence and multilocus genoty** of Cryptosporidium spp. in cattle in Jiangxi Province, southeastern China. Parasitol Res. 2021;120(4):1281–9.
Qi M, Wang H, **g B, Wang D, Wang R, Zhang L. Occurrence and molecular identification of Cryptosporidium spp. in dairy calves in **njiang, Northwestern China. Vet Parasitol. 2015;212:404–7.
Fan Y, Wang T, Koehler AV, Hu M, Gasser RB. Molecular investigation of Cryptosporidium and Giardia in pre- and post-weaned calves in Hubei Province, China. Parasite Vector. 2017;10(1):519.
Zhong Z, Dan J, Yan G, Tu R, Tian Y, Cao S, et al. Occurrence and genoty** of Giardia duodenalis and Cryptosporidium in pre-weaned dairy calves in central Sichuan province, China. Parasite. 2018;25:45.
Wang Y, Cao J, Chang Y, Yu F, Zhang S, Wang R, et al. Prevalence and molecular characterization of Cryptosporidium spp. and Giardia duodenalis in dairy cattle in Gansu, northwest China. Parasite. 2020;27:62.
Wang R, Wang H, Sun Y, Zhang L, Jian F, Qi M, et al. Characteristics of Cryptosporidium transmission in preweaned dairy cattle in Henan, China. J Clin Microbiol. 2010;49(3):1077–82.
Wang R, Ma G, Zhao J, Lu Q, Wang H, Zhang L, et al. Cryptosporidium andersoni is the predominant species in post-weaned and adult dairy cattle in China. Parasitol Int. 2011;60(1):1–4.
Tao W, Li Y, Yang H, Song M, Lu Y, Li W. Widespread occurrence of zoonotic Cryptosporidium species and subtypes in dairy cattle from northeast China: public health concerns. J Parasitol. 2018;104(1):10–7.
Liu A, Wang R, Li Y, Zhang L, Shu J, Zhang W, et al. Prevalence and distribution of Cryptosporidium spp. in dairy cattle in Heilongjiang Province, China. Parasitol Res. 2009;105(3):797–802.
Hu S, Liu Z, Yan F, Zhang Z, Zhang G, Zhang L et al. Zoonotic and host-adapted genotypes of Cryptosporidium spp., Giardia duodenalis and Enterocytozoon bieneusi in dairy cattle in Hebei and Tian**, China. Vet Parasitol. 2017;248:68–73.
Chen F, Huang K. Prevalence and molecular characterization of Cryptosporidium spp. in dairy cattle from farms in China. J Vet Sci. 2012;13(1):15–22.
Zhao G, Du S, Zhang L, Wang R, Guo Y, Qi MZ, et al. Molecular characterization of Cryptosporidium spp. in pre-weaned calves in Shaanxi Province, north-western China. J Med Microbiol. 2015;64:111–6.
Zhao G, Ren W, Gao M, Bian Q, Hu B, Cong M, et al. Genoty** Cryptosporidium andersoni in cattle in Shaanxi Province, Northwestern China. PLoS ONE. 2013;8(4):e60112.
Li F, Hu S, Wang H, Wang J, Wang R, Ming C, et al. Prevalence and molecular characterization of Cryptosporidium spp. and Giardia duodenalis in dairy cattle in Bei**g, China. Vet Parasitol. 2016;219:61–5.
de Graaf DC, Vanopdenbosch E, Ortega-Mora LM, Abbassi H, Peeters JE. A review of the importance of cryptosporidiosis in farm animals. Int J Parasitol. 1999;29(8):1269–87.
Ichikawa-Seki M, Aita J, Masatani T, Suzuki M, Nitta Y, Tamayose G, et al. Molecular characterization of Cryptosporidium parvum from two different japanese prefectures, Okinawa and Hokkaido. Parasitol Int. 2015;64(2):161–6.
Helmy YA, Krücken J, Nöckler K, von Samson-Himmelstjerna G, Zessin K-H. Molecular epidemiology of Cryptosporidium in livestock animals and humans in the Ismailia province of Egypt. Vet Parasitol. 2013;193(1):15–24.
Silverlås C, Bosaeus-Reineck H, Näslund K, Björkman C. Is there a need for improved Cryptosporidium diagnostics in swedish calves? Int J Parasitol. 2013;43(2):155–61.
Ibrahim MA, Abdel-Ghany AE, Abdel-Latef GK, Abdel-Aziz SA, Aboelhadid SM. Epidemiology and public health significance of Cryptosporidium isolated from cattle, buffaloes, and humans in Egypt. Parasitol Res. 2016;115(6):2439–48.
Garcia -RJCP, Anthony B, Velathanthiri, NilukaFrench NPHayman, David TS. Species and genotypes causing human cryptosporidiosis in New Zealand. Parasitol Res. 2020;119:2317–26.
Lebbad M, Winiecka-Krusnell J, Stensvold CR, Beser J. High diversity of Cryptosporidium species and subtypes identified in Cryptosporidiosis acquired in Sweden and abroad. Pathogens. 2021;10(5):523.
Costa D, Razakandrainibe R, Valot S, Vannier M, Sautour M, Basmaciyan L, et al. Epidemiology of Cryptosporidiosis in France from 2017 to 2019. Microorganisms. 2020;8(9):1358.
Cacciò SM, Chalmers RM. Human cryptosporidiosis in Europe. Clin Microbiol Infect. 2016;22(6):471–80.
O’ Leary JK, Blake L, Corcoran GD, Sleator RD, Lucey B. Increased diversity and novel subtypes among clinical Cryptosporidium parvum and Cryptosporidium hominis isolates in Southern Ireland. Exp Parasitol. 2020;218:107967.
Lochlainn NLM, Sane, Jussi, Schimmer B, et al. Risk factors for sporadic cryptosporidiosis in the Netherlands: analysis of a 3-year population based case-control study coupled with genoty**, 2013–2016. J Infect Dis. 2019;219:1121–9.
Acknowledgements
Not applicable.
Funding
This study was funded by the National Natural Science Foundation of China (32260887), National Natural Science Foundation of Inner Mongolia (2022MS03023), Inner Mongolia Agricultural University High-level Talents Research Initiation Fund Project (NDYB2019-3), and Supported by State Key Laboratory of Veterinary Biotechnology Foundation (SKLVBF202204).
Author information
Authors and Affiliations
Contributions
LZ, HLC, MYW and YHL conceived and designed the study and critically revised the manuscript. LZ, HLC, ZSZ, WXH, BY, and YHL performed the samples collection. HLC and YHL prepared Fig. 1. HLC, ZSZ, MYW, YW, SZ, WHZ, YMM, YJZ, LFW, YLD, JLW and LZ conducted the laboratory experiments. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was carried out in strict accordance with international standards as published in the “Guide to the feeding, management and use of experimental animals” (8th Edition) and follows the “Regulations on the management of experimental animals” and other relevant laws and regulations. The biomedical research ethics committee of Inner Mongolia Agricultural University specifically approved this study (No. 2020[081]). In addition, permission was obtained from the farm owners before the specimens were collected, and all efforts were made to minimize suffering.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Zhao, L., Chai, HL., Wang, MY. et al. Prevalence and molecular characterization of Cryptosporidium spp. in dairy cattle in Central Inner Mongolia, Northern China. BMC Vet Res 19, 134 (2023). https://doi.org/10.1186/s12917-023-03696-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12917-023-03696-z