Abstract
Background
Ovarian folliculogenesis is a tightly regulated process leading to the formation of functional oocytes and involving successive quality control mechanisms that monitor chromosomal DNA integrity and meiotic recombination. A number of factors and mechanisms have been suggested to be involved in folliculogenesis and associated with premature ovarian insufficiency, including abnormal alternative splicing (AS) of pre-mRNAs. Serine/arginine-rich splicing factor 1 (SRSF1; previously SF2/ASF) is a pivotal posttranscriptional regulator of gene expression in various biological processes. However, the physiological roles and mechanism of SRSF1 action in mouse early-stage oocytes remain elusive. Here, we show that SRSF1 is essential for primordial follicle formation and number determination during meiotic prophase I.
Results
The conditional knockout (cKO) of Srsf1 in mouse oocytes impairs primordial follicle formation and leads to primary ovarian insufficiency (POI). Oocyte-specific genes that regulate primordial follicle formation (e.g., Lhx8, Nobox, Sohlh1, Sohlh2, Figla, Kit, Jag1, and Rac1) are suppressed in newborn Stra8-GFPCre Srsf1Fl/Fl mouse ovaries. However, meiotic defects are the leading cause of abnormal primordial follicle formation. Immunofluorescence analyses suggest that failed synapsis and an inability to undergo recombination result in fewer homologous DNA crossovers (COs) in the Srsf1 cKO mouse ovaries. Moreover, SRSF1 directly binds and regulates the expression of the POI-related genes Six6os1 and Msh5 via AS to implement the meiotic prophase I program.
Conclusions
Altogether, our data reveal the critical role of an SRSF1-mediated posttranscriptional regulatory mechanism in the mouse oocyte meiotic prophase I program, providing a framework to elucidate the molecular mechanisms of the posttranscriptional network underlying primordial follicle formation.
Similar content being viewed by others
Background
High-quality gametes are critical for successful reproduction [1]. Functional oocytes are derived from successful folliculogenesis in the ovaries, which includes primordial follicle formation; recruitment into the growing pool to form primary, secondary, and tertiary follicles; ovulation; and subsequent corpus luteum formation [2]. The size and quality of the surviving pool of primordial follicles are essential determinants of female fecundity and reproductive lifespan [3]. To maintain quality, diverse organisms have evolved three quality control mechanisms for eliminating meiocytes with defects in meiotic recombination or SPO11-linked DNA double-strand break (DSB) accumulation, namely, meiotic silencing, the synapsis checkpoint, and the DNA damage checkpoint [4]. During meiotic prophase I, defective meiosis results in POI due to early exhaustion of the follicle pool.
With the widespread application of next-generation sequencing (NGS), the genetic spectrum of POI has been expanded, especially by the recent identification of novel meiosis-related genes [5,6,7]. The cohesin complex regulates sister chromatid cohesion and synaptonemal complex (SC) formation and is composed of the meiosis-specific subunits STAG3, RAD21L, and SMC1β and the nonspecific subunits SMC3 and REC8 [8]. STAG3 aberrations have been identified to cause a rare monogenic type of POI [9,10,11,12,13,14,15,16,Full size image
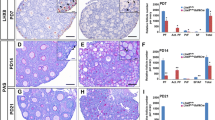
The adult cKO mice were normal in size (Fig. 2h, i), but the size of their ovaries was significantly reduced (Fig. 2j, k). To further confirm the effect of the loss of Srsf1 on female fertility, a breeding experiment was performed, and the data indicated that the absence of Srsf1 led to complete infertility in cKO females (Fig. 2l). Histological examination of cKO ovary sections revealed that folliculogenesis was arrested (Fig. 2m − p). Immunostaining of VASA indicated the absence or asymmetric deviation of oocytes in cKO ovaries (Fig. 2q − t). Follicle count data based on VASA immunohistochemistry analysis showed that the follicles in cKO ovaries were primarily arrested at the primary follicle stage (Fig. 2u, v). Together, these results indicate that SRSF1 is critical for follicle development.
SRSF1 is required for mouse primordial follicle formation
To confirm the effect of the loss of Srsf1 on POI, we examined mouse ovaries at 5 days post-partum (dpp), a timepoint at which the primordial follicle pool is successfully established [34]. Tissue analyses revealed that the cKO mice exhibited a reduced ovary size (Fig. 3a, b). Oocyte count data based on VASA immunohistochemistry analysis showed that the primordial follicle pool was consistently reduced in cKO ovaries (Fig. 3e). We verified that the numbers of naked oocytes or oocytes in the cysts were increased, but the numbers of primordial and primary follicles were reduced, in cKO ovaries (Fig. 3c, d, f). Together, these results suggest dysfunctional primordial follicle formation and number determination in cKO ovaries.
SRSF1 is required for mouse primordial follicle formation. a, b Ovarian atrophy in 5 dpp cKO mice. c, d Immunostaining of VASA in 5 dpp ovaries. PF, primary follicle (cʹ, dʹ), PmF, primordial follicle (cʹʹ), cyst oocytes (dʹʹ), naked oocyte (dʹʹʹ). e Quantification of total oocytes per 5 dpp ovary. f Quantification of primordial follicles, primary follicles, and oocytes in cysts or naked per 5 dpp ovary. n = 3. g, h Ovarian atrophy in newborn cKO mice. i–l Immunostaining of VASA in newborn ovaries. kʹ, lʹ The primordial follicle and oocytes in cysts are shown in magnified views. Arrowheads, PmF; arrows, cyst oocytes. m The mRNA level of Srsf1 decreased in newborn cKO ovaries. n The mRNA level of Vasa decreased in newborn cKO ovaries. o Quantification of total oocytes per newborn ovary. Ctrl, n = 3; cKO, n = 4. p Percentages of primordial follicles and oocytes in cysts per newborn ovary. Ctrl, n = 3; cKO, n = 4. q Co-immunostaining of GM130 and VASA in newborn ovaries. r Percentage of double-positive cells in newborn ovaries. n = 2. s Srsf1 deficiency downregulated expression of various genes (e.g., Kit, Rac1, Figla, Nobox). t Co-immunostaining of KIT and VASA in newborn ovaries. u Percentage of double-positive cells in newborn cKO ovaries. n = 3. v Co-immunostaining of JAG1 and VASA in newborn ovaries. w Percentage of double-positive cells is shown in newborn cKO ovaries. n = 3. Real-time qPCR data were normalized to Gapdh (n, m) or Vasa (s). n = 3. Significance was determined by unpaired Student’s t test; detailed P value P ≥ 0.05, *P < 0.05, ***P < 0.001, ****P < 0.0001. The error bar represents the mean ± SEM. Scale bar, 20 μm (q right, t magnified, v magnified left), 50 μm (kʹ, lʹ), 100 μm (c, d), 200 μm (a, d, i–l, q left, t left, v left), 400 μm (g, h)
The specific involvement of SRSF1 in primordial follicle formation and number determination was further verified by the observation of mild ovarian atrophy in newborn cKO mice, in which the primordial follicle pool is being established (Fig. 3g, h). RT–qPCR showed that the expression of Srsf1 was significantly decreased in cKO ovaries (Fig. 3m), and co-immunofluorescence analysis with SRSF1 and VASA antibodies showed an absence of the SRSF1 protein in all oocytes (Fig. 2d − g). Interestingly, VASA immunostaining revealed more oocytes in cysts in cKO ovaries (Fig. 3i − l). RT–qPCR results showed that the expression of Vasa was significantly decreased in the cKO ovaries, suggesting that the total number of oocytes was reduced (Fig. 3n). Additionally, count data showed that the numbers of both total oocytes and primary follicles were reduced in cKO ovaries (Fig. 3o, p). However, the number of oocytes in cysts was significantly increased (Fig. 3p). Abnormal primordial follicular pool formation was further confirmed by the observation of GM130 immunofluorescence in the ovaries of newborn cKO mice [37]. Our results revealed that the number of double-positive cells was reduced in newborn cKO ovaries, suggesting that SRSF1 is essential for cyst breakdown and primordial follicle formation (Fig. 3q, r). To understand the molecular mechanisms underlying these phenotypes, we examined vital genes that regulate primordial follicle formation by RT–qPCR and found that their expression was suppressed (Fig. 3s). Co-immunostaining of VASA and KIT or JAG1 showed that the number of double-positive cells was reduced in the ovaries of newborn cKO mice (Fig. 3t − w). These data suggest that vital genes that regulate primordial follicle formation are suppressed in the ovaries of newborn cKO mice.
Loss of SRSF1 impairs meiotic progression in oocytes
To investigate the molecular consequences of SRSF1 depletion in primordial follicle formation, we isolated mRNA from control (Ctrl) and cKO ovaries at 16.5 dpc and performed RNA sequencing. RNA-seq analyses identified 566 downregulated and 705 upregulated genes in cKO ovaries at 16.5 dpc (Fig. 4a, b; Additional file 2: Table 1). Surprisingly, Gene Ontology (GO) term enrichment indicated abnormal meiosis in cKO mouse oocytes (Fig. 4c). Next, we validated the abnormal expression of meiosis-related genes (downregulated: Rec8, Six6os1, Psmc3ip, Tdrd9, and Plk1; upregulated: Hormad2 and Syce1) by RT–qPCR (Fig. 4d). To assess meiotic progression in cKO ovaries, MSY2 immunostaining was employed as a marker of the diplotene stage in newborn mouse ovaries (Fig. 4e) [38]. Double-positive cell counting results showed that the number of diplotene oocytes was reduced in cKO ovaries (Fig. 4f). To further evaluate this phenotype, we performed SYCP3 immunofluorescence analysis in 5 dpp mouse ovaries (Fig. 4g). Normal meiotic arrest in the dictyate stage is crucial for primordial follicle formation. It can be identified by the presence of two to four visible nucleolus signals after staining with an SYCP3 antibody [39]. Dictyate oocyte count data revealed that SRSF1 deficiency impaired meiotic progression to the diplotene stage in oocytes (Fig. 4h). These results suggest that the few oocytes in cKO ovaries that reached the diplotene stage could not develop further to the dictyate stage.
SRSF1 is required for meiotic progression. a Volcano map displaying the distribution of differentially expressed genes from RNA-seq data. The abscissa in the figure represents the gene fold change in cKO and Ctrl mouse ovaries. |log2FoldChange|≥ 0. The ordinate indicates the significance of gene expression differences between cKO and Ctrl mouse ovaries. P ≤ 0.05. Upregulated genes are shown in red dots, and downregulated genes are shown in green dots. b Cluster heatmap of differentially expressed genes. The abscissa is the genotype, and the ordinate is the normalized FPKM (fragments per kilobase million) value of the differentially expressed gene. Red indicates a higher expression level, while green indicates a lower expression level. c Scatter plot of GO enrichment analysis of the 1271 differentially expressed genes. The 12 most effective terms were selected to draw a scatter diagram for display. d The misregulation of meiosis-related genes in 16.5 dpc cKO ovaries. Real-time qPCR data were normalized to Gapdh. The value in newborn Ctrl ovaries was set as 1.0, and the relative value in the ovaries of newborn cKO mice is indicated. n = 4. e Co-immunostaining of MSY2 and VASA in newborn Ctrl and cKO ovaries. DNA was stained with DAPI. Scale bar, 200 μm. f Percentage of double-positive cells is shown as the mean ± SEM. n = 3. g Immunostaining of SYCP3 in 5 dpp Ctrl and cKO ovaries. DNA was stained with DAPI. The dictyotene oocytes of Ctrl ovaries are shown in magnified views. Abnormal oocytes of cKO ovaries are shown in the magnified views. Scale bar, 50 μm. h Percentage of dictyotene oocytes is shown as the mean ± SEM in 5 dpp ovaries. n = 3. Significance was determined by unpaired Student’s t test; detailed P value P ≥ 0.05, *P < 0.05, ***P < 0.001, ****P < 0.0001. The error bar represents the mean ± SEM
SRSF1 deficiency leads to defects in synapsis and crossover recombination in cKO ovaries
The GO term enrichment results showed abnormal homologous chromosome segregation in cKO mouse oocytes (Fig. 4c). The COs cause the exchange of homologous DNA and establish the physical connections between homologues required for proper chromosome segregation [40]. MutL homologue 1 (MLH1) has been recognized as a classic marker of COs [41,42,43]. Therefore, the distribution of MLH1 foci was evaluated by MLH1 and SYCP3 immunostaining in newborn mouse oocyte surface spreads (Fig. 5a). The counts of MLH1 foci showed that the formation of COs was reduced in diplotene oocytes (Fig. 5b). The final and significant purpose of meiotic recombination is the formation of COs [44, 45]. To probe the nature of meiotic recombination in cKO ovaries, we examined chromosomal synapsis through oocyte surface spread analysis in newborn mouse ovaries. Aberrant synapsis was frequently observed in cKO oocytes based on the localization of synaptonemal complex protein 1 (SYCP1), SYCP3, and CREST (Fig. 5c). Surprisingly, SYCP1 could not be loaded onto chromosomes in cKO mouse oocytes (Fig. 5c). The provided schematic diagram shows the failure of synapsis in cKO oocytes (Fig. 5d, e). These data suggest that the failure of synapsis and the inability to undergo homologous recombination (HR) resulted in fewer COs in cKO oocytes.
SRSF1 deficiency impairs chromosome synapsis and the formation of COs at meiotic prophase I. a Co-immunostaining of MLH1 and SYCP3 in newborn Ctrl and cKO oocytes. DNA was stained with DAPI. Scale bar, 5 μm. b The number of COs, marked by MLH1 foci (red), was significantly reduced in cKO diplotene-like oocytes compared with Ctrl oocytes. Twenty-five Ctrl oocytes and thirty-three cKO oocytes were obtained from 3 animals. Significance was determined by unpaired Student’s t test; ****P < 0.0001. The error bar represents the mean ± SEM. c Localization of SYCP1, SYCP3, and CREST in newborn Ctrl and cKO oocytes. DNA was stained with DAPI. Scale bar, 5 μm. Diagrams illustrate aberrant synapsis in Ctrl (d) and cKO (e) oocytes
SRSF1 is essential for the resection of SPO11-linked DNA double-strand breaks (DSBs)
DSBs formed during meiosis recruit phosphorylated histone H2AX (γH2AX) [46]. Therefore, we monitored the progression of meiotic recombination in 17.5 dpc ovaries by the co-immunostaining with γH2AX and SYCP3. The immunostaining results revealed distinct γH2AX signals in cKO oocytes, whereas few oocytes in Ctrl ovaries showed positive γH2AX staining (Fig. 6a). The double-positive cell counting data confirmed that some DSBs were not repaired correctly in cKO oocytes (Fig. 6b). Replication protein A (RPA) is a single-strand DNA-binding heterotrimeric complex composed of RPA1, RPA2, and RPA3 that is essential for meiotic recombination [47]. We examined the localization of RPA1 as a representative of the RPA complex by nuclear spread analysis in newborn mouse ovaries. The co-immunostaining of RPA1 and SYCP3 showed that many RPA complexes remained uncleared in the diplotene stage in cKO oocytes (Fig. 6c, d). RecA-like proteins DMC1 and RAD51 form a nucleoprotein filament on ssDNA within DSBs to aid in the search and invasion of the homologous partner for successful recombination and synapsis. We further explored the distribution of proteins involved in recombination and DSB repair. Interestingly, the localization of DMC1, γH2AX, and SYCP3 showed that significant DMC1 foci were present on chromosomes in the pachytene stage in cKO oocytes (Fig. 6e). The DMC1 focus count data revealed an increase in recombinase on chromosomes in the pachytene stage in cKO oocytes (Fig. 6f). However, the recombinase had lost its ability to effect on DNA repair. These observations suggest that SRSF1 is essential for the resection of SPO11-linked DSBs in meiotic prophase.
SRSF1 is necessary for the resection of DSBs. a Co-immunostaining of γH2AX and SYCP3 in 18.5 dpc Ctrl and cKO ovaries. DNA was stained with DAPI. Scale bar of the top panel, 50 μm. Scale bar of the rest panel, 10 μm. b Percentage of double-positive cells is shown as the mean ± SEM in 18.5 dpc ovaries. Ctrl, n = 6; cKO, n = 5. c Co-immunostaining of RPA1 and SYCP3 in newborn Ctrl and cKO oocytes. DNA was stained with DAPI. Scale bar, 5 μm. d In cKO diplotene-like oocytes, the number of RPA1 foci (red) was maintained at a relatively high level. Twenty-five Ctrl oocytes and thirty-three cKO oocytes were obtained from 3 animals. e Localization of DMC1, γH2AX, and SYCP3 in 18.5 dpc Ctrl and cKO oocytes. DNA was stained with DAPI. Scale bar, 10 μm. f In cKO pachytene oocytes, the number of DMC1 foci (white) was maintained at a relatively high level. Sixty Ctrl oocytes and fifty-six cKO oocytes were obtained from 4 animals. Significance was determined by unpaired Student’s t test; detailed P value P ≥ 0.05, *P < 0.05, ***P < 0.001, ****P < 0.0001. The error bar represents the mean ± SEM
SRSF1 directly regulates the splicing of the POI-related genes Msh5 and Six6os1
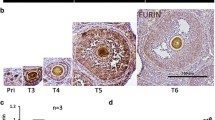
RNA-seq analyses showed 191 AS events that were identified as significantly affected (FDR < 0.05) in 16.5 dpc cKO ovaries (Additional file 3: Table 2). Among the 191 affected AS events, most (141) were classified as skipped exons (SEs). In addition, ten AS events were categorized as alternative 5′ splice sites (A5SSs), 10 as alternative 3′ splice sites (A3SSs), 15 as mutually exclusive exons (MXEs), and 15 as retained introns (RIs) (Fig. 7a). The GO enrichment analysis of the alternatively spliced genes revealed that eleven meiosis-related genes showed alterations in AS forms (Fig. 7b). At least two (Six6os1 and Msh5) of these genes have been associated with POI [19, 20, 48]. We then visualized the different types of AS based on RNA-seq data by using Integrative Genomics Viewer (IGV, 2.10.2) software (Fig. 7c). RT–PCR results showed that the pre-mRNAs of Six6os1 and Msh5 in cKO mouse oocytes exhibited abnormal AS (Fig. 7d). The results of RIP–qPCR showed that SRSF1 could bind to the pre-mRNAs of Msh5 and Six6os1 (Fig. 7e, f). Interestingly, Western blotting and immunostaining revealed that the protein levels of MSH5 and SIX6OS1 were significantly suppressed (Fig. 7g − k; Additional file 4: Fig. S2a, b). Additionally, abnormal AS changed the mRNA decay rates of Six6os1 (Fig. 7l) but not Msh5 (Additional file 4: Fig. S2c) in cKO mouse ovaries.
SRSF1 directly regulates the splicing of Msh5 and Six6os1. a Five AS events were significantly affected by SRSF1-deficient oocytes. rMATS (3.2.5) software was used to analyse AS events (FDR < 0.05). FDR, false discovery rate calculated from the P value. b Scatter plot of the GO enrichment analysis of the 191 affected AS events. c A schematic of the regulation of the splicing of Msh5 and Six6os1. Integrative Genomics Viewer (IGV, version 2.10.2) was used to visualize and confirm AS events in RNA-seq data. E, exon. d The ectopic splicing of Msh5 and Six6os1 in cKO ovaries was analysed by RT–PCR (n = 4 per group). The scheme and cumulative data on the percentage of the indicated fragment are shown accordingly. e, f SRSF1 directly regulated the expression of Msh5 and Six6os1 by RIP–qPCR in 16.5 dpc mouse ovaries. n = 3. g, h Western blotting of MSH5 and SIX6OS1 expression in 17.5 dpc Ctrl and cKO ovaries. GAPDH (g) or ACTB (h) served as a loading control. n = 4. i Localization of MSH5 and SYCP3 in 17.5 dpc Ctrl and cKO oocytes. DNA was stained with DAPI. Scale bar, 10 μm. j The number of MSH5 foci (green) was significantly reduced in cKO pachytene oocytes compared with Ctrl oocytes. Sixty Ctrl oocytes and sixty cKO oocytes were obtained from 4 animals. k Co-immunostaining of SIX6OS1 and SYCP3 in 17.5 dpc Ctrl and cKO ovaries. DNA was stained with DAPI. Zyg, zygotene; ePac, early pachytene; Pac, pachytene. Scale bar, 10 μm. l The expression of Six6os1 in 16.5 dpc cKO ovaries after ActD treatment at different times. n = 4. Significance was determined by unpaired Student’s t test; detailed P value P ≥ 0.05, *P < 0.05, ***P < 0.001, ****P < 0.0001. The error bar represents the mean ± SEM