Abstract
Background
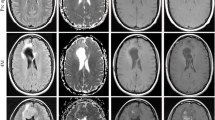
We created discriminative models of different regions of interest (ROIs) using radiomic texture features of neurite orientation dispersion and density imaging (NODDI) and evaluated the feasibility of each model in differentiating glioblastoma multiforme (GBM) from solitary brain metastasis (SBM).
Methods
We conducted a retrospective study of 204 patients with GBM (n = 146) or SBM (n = 58). Radiomic texture features were extracted from five ROIs based on three metric maps (intracellular volume fraction, orientation dispersion index, and isotropic volume fraction of NODDI), including necrosis, solid tumors, peritumoral edema, tumor bulk volume (TBV), and abnormal bulk volume. Four feature selection methods and eight classifiers were used for the radiomic texture feature selection and model construction. Receiver operating characteristic (ROC) curve analysis was used to evaluate the diagnostic performance of the models. Routine magnetic resonance imaging (MRI) radiomic texture feature models generated in the same manner were used for the horizontal comparison.
Results
NODDI-radiomic texture analysis based on TBV subregions exhibited the highest accuracy (although nonsignificant) in differentiating GBM from SBM, with area under the ROC curve (AUC) values of 0.918 and 0.882 in the training and test datasets, respectively, compared to necrosis (AUCtraining:0.845, AUCtest:0.714), solid tumor (AUCtraining:0.852, AUCtest:0.821), peritumoral edema (AUCtraining:0.817, AUCtest:0.762), and ABV (AUCtraining:0.834, AUCtest:0.779). The performance of the five ROI radiomic texture models in routine MRI was inferior to that of the NODDI-radiomic texture model.
Conclusion
Preoperative NODDI-radiomic texture analysis based on TBV subregions shows great potential for distinguishing GBM from SBM.
Key points
• NODDI-radiomic texture analysis can distinguish glioblastoma multiforme from solitary brain metastasis.
• ROI division affects efficiency, and tumor bulk volume outperformed other ROIs.
• NODDI-based outperforms routine magnetic resonance imaging-based radiomic texture analysis.
Similar content being viewed by others
Background
Glioblastoma multiforme (GBM) and solitary brain metastases (SBM) are the most common malignant brain tumors, and their correct identification is key for further diagnosis and treatment [1,2,3]. Although magnetic resonance imaging (MRI) is the main tool for differentiating between the two types of tumors, both GBM and SBM may show marked peritumoral edema and similar contrast-enhancement patterns on routine MRI, leading to great challenges in identification [4,5,6].
Previous studies reported that radiomics combined with routine MRI showed significant advantages in distinguishing GBM from SBM and suggested that specific imaging features are helpful in distinguishing between the two types of tumors [7, 8]. Currently, the acquisition of specific image features can be summarized into two trends: applying special MRI modalities or focusing on specific image feature types [9, 10].
Diffusion-weighted imaging (DWI) can provide a class of microscopic features related to the movement of water molecules in tissues, such as the current advanced diffusion imaging model and neurite orientation dispersion and density imaging (NODDI) [11, 12]. NODDI is a multi-spherical shell diffusion model based on the difference in the diffusion of water molecules inside and outside the cell and is more often used to characterize the difference in water diffusion between tumor infiltration and vasogenic edema [Full size image
Radiomic texture extraction and model construction
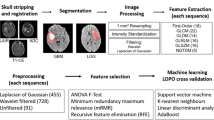
Feature extraction, feature selection, and model building were performed using the open-source software FeAture Explorer (FAE, version 0.5.2) [23]. Based on the automatically segmented ROIs, radiomic texture features were extracted using first-order statistical functions, gray-level co-occurrence matrix (GLCM) functions, and gray-level run-length matrix (GLRLM) functions on the original NODDI parametric maps, as well as eight sub-bands of its wavelet transformation. As controls, the features of routine MR images (T2WI, T2-dark-fluid, T1WI, and CE-T1 MPRAGE) were extracted in the same manner. Radiomic texture analysis for each ROI was based on a combination of three parametric map features from the NODDI or a combination of four routine MRI features. Finally, 234 features were extracted from each parameter map of NODDI (or each type of routine MRI). Details of the extracted features are presented in Supplementary Appendix E2.
Because of the imbalanced GBM-to-SBM sample ratio (2.5:1), we applied upsampling to the training dataset. After feature extraction, all radiomic texture feature values were normalized using the min-max or Z-score method. Four feature selection methods—Pearson’s correlation coefficient (PCC), analysis of variance, recursive feature elimination, and the Kruskal-Wallis test—as well as eight classifiers—support vector machine, linear discriminant analysis, auto-encoder, random forest, logistic regression, logistic regression via Lasso, ada-boost, and decision tree—were utilized to construct texture feature prediction models for each ROI. When the PCC value of a feature pair was greater than 0.90, only one of the features was randomly retained. Five-fold cross-validation was used to determine the hyperparameters of each model. After determining the hyperparameters, all training data were retrained for the final models. The maximum number of features included in the radiomic texture analysis model construction was four. Details of the sample size and feature number estimates are displayed in Supplementary Appendix E3. The final models were determined based on the highest area under the receiver operating characteristic (ROC) curve (AUC) value in the cross-validation, and a time-independent test dataset was used to evaluate the performance of the final model. The performance of the test dataset was determined through ROC curve analysis and evaluations of accuracy, AUC, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV).
Statistical analysis
Statistical analyses were performed using SPSS (version 21.0) and MedCalc (version 20.015) software. Differences in clinical characteristics between GBM and SBM were assessed using chi-square tests (or the Mann–Whitney U test, depending on the results of normality and homoscedasticity tests) and independent t-tests, as appropriate. DeLong’s test was used to assess differences in AUC values between models. Statistical significance was set at a two-sided p value < 0.05.