Abstract
Background
Solid pulmonary nodules are different from subsolid nodules and the diagnosis is much more challenging. We intended to evaluate the diagnostic and prognostic value of radiomics and deep learning technologies for solid pulmonary nodules.
Methods
Retrospectively enroll patients with pathologically-confirmed solid pulmonary nodules and collect clinical data. Obtain pre-treatment high-resolution thoracic CT and manually delineate the nodule in 3D. Then, all patients were randomly divided into training and testing sets at a ratio of 7:3, and convolutional neural networks (CNN) models and random forest (RF) models were established. Survival analyses were performed for patients with solid adenocarcinomas.
Results
Totally 720 solid pulmonary nodules were enrolled, 348 benign and 372 malignant. The CNN model with clinical features achieved the highest AUC [0.819, 95% confidence interval (CI): 0.760–0.877] with a sensitivity of 0.778, specificity of 0.788 and accuracy of 0.783. No significant differences were observed between the CNN and radiomics models. There were 295 solid adenocarcinomas in survival analysis. Different disease-free survival was observed between the low-risk and high-risk groups divided according to the radiomics Rad-score. However, the groups based on deep learning signatures showed similar survival. Cox regression analysis indicated that the radiomics Rad-score (hazard ratio: 5.08, 95% CI: 2.61–9.90) was an independent predictor of recurrence.
Conclusions
The radiomics and deep learning models can well predict the malignancy of solid pulmonary nodules. Radiomics signatures also demonstrate prognostic value in solid adenocarcinomas.
Similar content being viewed by others
Background
Lung cancer screening with low-dose computed tomography among high-risk individuals can reduce lung cancer modality [1, 2]. However, it’s challenging to manage pulmonary nodules detected on thoracic CT either during screening or routine clinical practice. Solid pulmonary nodules are usually distinct from subsolid nodules (SSNs) and therefore different recommendations were provided in guidelines [3,4,5,6]. Most SSNs exhibit indolent nature and grow slowly or stay stable over years [7], and often pathologically diagnosed as lung adenocarcinomas [8]. However, the solid nodules which could be caused by various respiratory diseases, can grow rapidly and are more prone to distant metastasis when they are malignant [9, 10]. Therefore, the risk prediction of solid pulmonary nodules should be important, as it can help clinicians make the right decision and save time for patients during medical care.
Recently, radiographic assessment of disease is being improved by advanced computational analyses. On the one hand, radiomics approach can digitally decode radiographic images into quantitative features (e.g., descriptors of shape, size and textural patterns), and therefore classify the medical image into a predefined group [11]. On the other hand, deep learning has made great strides in automatically characterizing radiographic images. It uses convolutional neural networks (CNN) to automatically learn feature representations from sample images, which could match and even surpass human performance in task-specific applications [12].
In previous studies, researchers have investigated the diagnostic performance of radiomics and deep learning technologies for solid pulmonary nodules [13,14,15,16,17,18,19]. However, some of the studies focused mainly on small solid pulmonary nodules, such as nodules less than 15 mm or 20 mm [14, 15, 17]. Some studies differentiated solid nodules between one specific benign lung disease and lung adenocarcinoma, like focal organizing pneumonia, solitary granulomatous nodules or tuberculosis [13, 16, 19]. Besides, none of the studies investigated prognostic values of radiomics and deep learning technologies for solid nodules.
Therefore, the current study intended to establish CNN and radiomics models for solid pulmonary nodules without restricting the nodule size and pathology. Furthermore, survival analyses were performed for patients with solid adenocarcinomas.
Methods
Patients and clinical variables
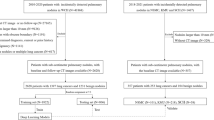
This retrospective study was approved by the institutional review board of the West China Hospital of Sichuan University. We collected possible cases by reviewing discharge records of patients in West China Hospital from January 2010 to July 2017. The following terms were used to extract the data: lung cancer, lung adenocarcinoma, lung squamous carcinoma, non-small cell lung cancer, small cell lung cancer; inflammatory lung nodule, benign lung nodule, benign lung tumor, lung hamartoma, lung sclerosing hemangioma, lung tuberculosis, lung granuloma. Then, the patient was enrolled based on the following criteria: (a) there was an untreated, pathologically confirmed, 5–30 mm noncalcified solid nodule detected on chest CT; (b) the slice thickness of CT was less than or equal to 1 mm. Otherwise, patients were excluded if (a) there were multiple pulmonary nodules, or pleural effusion, atelectasis, lymph node enlargement was observed; (b) it wasn’t a primary lung tumor.
Totally, the current study enrolled 720 patients with 720 nodules, 348 benign and 372 malignant. The pathology of benign nodules was confirmed by surgery (N = 315, 90.5%) and CT guided percutaneous lung biopsy (N = 33, 9.5%), while the malignant nodules was confirmed by surgery (N = 365, 98.1%), CT guided percutaneous lung biopsy (N = 4, 1.1%) and transbronchial lung biopsy (N = 3, 0.8%), respectively.
Following clinical characteristics were recorded, including age, sex, smoking status, history of malignancy, family history of malignancy, nodule diameter, location, pathology and clinical stage. As surgically resected adenocarcinomas were predominant among all malignant nodules, prognostic data were collected for survival analysis.
CT image acquisition and nodule segmentation
Thoracic CT before treatment was obtained for each patient. All images were acquired from GE, Siemens or Philips scanners, with tube voltage and current being 100 ~ 120 Kvp and 60 ~ 250 mAs. Reconstructions were performed using a standard convolution kernel. The detailed information on manufacturer, manufacturer’s model and slice thickness were summarized in Table S1 and Table S2.
All target nodules were first manually segmented in 3D by one author with 4 years of clinical experience in pulmonology, using the ITK-SNAP software. Then, randomly selecting 100 patients, both the same author and another author manually segmented the target nodules again to assess the consistency of the intra-rater and inter-rater segmentations by calculating Dice similarity coefficient. Both authors were blinded to pathological results of lesions.
CNN models
Patients were randomly divided into training and testing set at a ratio of 7:3 during model establishment. The overall framework of the CNN model is shown in Fig. 1. Here we used transfer learning from a pre-trained benign-malignant nodule classification model, in which 1715 pathologically-confirmed nodules and 14,735 unlabeled nodules were used [20]. In detail, there were one 3D convolution layer with a kernel size of 3 and stride of 1 as input block, four 3D convolution layers with a kernel size of 3 and stride of 2 as downsample block, and two fully connected layers as output block for the benign-malignant classification task. Besides, the class activation map** was used to guide the network focusing on the nodule region, where attention maps were generated by back-propagating weights of the fully-connected layer onto the convolutional feature maps [Full size image
Radiomics models
Firstly, radiomics features were extracted of segmented nodules, including 42 dedicated handcrafted features and 104 widely-used radiomics features. Details of handcrafted features were described in previous study [22]. The widely-used first-order image intensity statistics, shape and texture features were extracted using PyRadiomics [23]. Then, three RF models were established by using radiomics features, clinical features and both features, respectively. To avoid overfitting and obtain predictive features, least absolute shrinkage and selection operator (LASSO) was applied for radiomics feature selection where regression coefficients of irrelevant variables were shrunk to zero. To achieve the best performance, an optimization algorithm based on Bayesian was used to optimize the hyperparameters.
To compare the diagnostic performance of established models with manual visual assessment, two junior radiologists were invited to blindly classify the solid nodules in the testing set.
Statistical analysis
The continuous variables, age and nodule diameter, were presented with mean ± standard deviation and compared with Student’s t-test. The follow-up time was compared with Mann-Whitney U test. The other categorical data were described in number of cases (proportion) and compared with Chi-square test.
The classification performance of the models was evaluated on sensitivity, specificity, accuracy, receiver operating characteristic curves (ROC) and values of area under the ROC curve (AUC). Calibration curves were also plotted to evaluate the accuracy of risk estimate. Additionally, Brier scores were calculated that quantitatively measure the distance in the probability domain and a lower score means better prediction. Differences in the AUC values were assessed by Delong test [24].
For prognostic analysis, a Rad-score was computed for each patient by combining LASSO selected radiomics features. According to the Rad-score, patients were classified into low-risk or high-risk group split by X-tile (version 3.6.1, http://tissuearray.org/) [25]. The potential association of radiomics signature with disease-free survival (DFS) was evaluated by Kaplan-Meier survival analysis and multivariate Cox regression. Similarly, the prognostic value of malignancy-score derived from CNN model (with clinical features) was also evaluated. Differences in survival curves were assessed by log-rank test.
The LASSO analysis, ROC curves, calibration curves and Brier scores were implemented with an open source “Scikit-learn 1.1.2” in Python. The Kaplan-Meier survival analysis and multivariate Cox regression were performed with “survival 3.1-8, survminer 0.4.8” packages in R. The statistical tests were all two-sided and differences with P < 0.05 were considered statistically significant. All statistical analyses were conducted using R version 3.6.0 and Python version 3.7.0.