Abstract
Background
Epidermal patterning factor / -like (EPF/EPFL) gene family encodes a class of cysteine-rich secretory peptides, which are widelyfound in terrestrial plants.Multiple studies has indicated that EPF/EPFLs might play significant roles in coordinating plant development and growth, especially as the morphogenesis processes of stoma, awn, stamen, and fruit skin. However, few research on EPF/EPFL gene family was reported in Gossypium.
Results
We separately identified 20 G. raimondii, 24 G. arboreum, 44 G. hirsutum, and 44 G. barbadense EPF/EPFL genes in the 4 representative cotton species, which were divided into four clades together with 11 Arabidopsis thaliana, 13 Oryza sativa, and 17 Selaginella moellendorffii ones based on their evolutionary relationships. The similar gene structure and common motifs indicated the high conservation among the EPF/EPFL members, while the uneven distribution in chromosomes implied the variability during the long-term evolutionary process. Hundreds of collinearity relationships were identified from the pairwise comparisons of intraspecifc and interspecific genomes, which illustrated gene duplication might contribute to the expansion of cotton EPF/EPFL gene family. A total of 15 kinds of cis-regulatory elements were predicted in the promoter regions, and divided into three major categories relevant to the biological processes of development and growth, plant hormone response, and abiotic stress response. Having performing the expression pattern analyses with the basic of the published RNA-seq data, we found most of GhEPF/EPFL and GbEPF/EPFL genes presented the relatively low expression levels among the 9 tissues or organs, while showed more dramatically different responses to high/low temperature and salt or drought stresses. Combined with transcriptome data of develo** ovules and fibers and quantitative Real-time PCR results (qRT-PCR) of 15 highly expressed GhEPF/EPFL genes, it could be deduced that the cotton EPF/EPFL genes were closely related with fiber development. Additionally, the networks of protein–protein interacting among EPF/EPFLs concentrated on the cores of GhEPF1 and GhEPF7, and thosefunctional enrichment analyses indicated that most of EPF/EPFLs participate in the GO (Gene Ontology) terms of stomatal development and plant epidermis development, and the KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways of DNA or base excision repair.
Conclusion
Totally, 132 EPF/EPFL genes were identified for the first time in cotton, whose bioinformatic analyses of cis-regulatory elements and expression patterns combined with qRT-PCR experiments to prove the potential functions in the biological processes of plant growth and responding to abiotic stresses, specifically in the fiber development. These results not only provide comprehensive and valuable information for cotton EPF/EPFL gene family, but also lay solid foundation for screening candidate EPF/EPFL genes in further cotton breeding.
Similar content being viewed by others
Background
In the process of plant growth and development, information must be transmitted between cells to ensure the basic life activities, among which protein secretion is one of the important ways of cell–cell communication. There are two common pathways of protein secretion, and the classical ER-Golgi pathway aims at the N-terminal of proteins with signal peptide sequences, meanwhile depends on endoplasmic (ER) and Golgi apparatus. The other non-classical pathway aims at the same N-terminal of proteins while without signal peptide sequences, nevertheless is independent of ER and Golgi apparatus [1]. The proteolytic enzymes, also namely as protease in plants, are generally consisted of these secreted proteins. They could break substrates into small peptide segments by hydrolyzing peptide bonds, which are divided into cystein protease, serine protease, aspartate protease, and metalloprptease based on their different catalytic sites [2]. Among the total 500–800 kinds of plant proteases encoded by Arabidopsis thaliana genome, cystein proteases (CPs) could contribute approximately 140 kinds, and are consisted of five protein families, such as papain proteinases (family C1), vacuolar processing enzyme (family VPE), caspases (family C14), calcium-dependent proteinases (family C2), and other CP family [3]. Given that CPs presented the different expression patterns in diverse tissues and organs, the previous reports have proved that CPs could play important roles in affecting the biological processes, such as seed germination, root growth, leaf senescence, and programmed cell death (PCD) of tracheary elements in stems and of tapetum cells in anthers [4,5,6,7,8]. These CPs makes significant differences in the specific positions through protein degradation for providing nutrients and components for the development of new cells, ensuring that the biological processes involved in plant growth proceed smoothly.
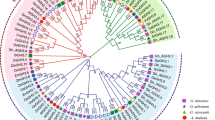
Epidermal patterning factor / -like (EPF/EPFL) proteins are one of the cysteine-rich plant-specific secreted peptides, which normally have 6 or 8 conserved cysteine residues in the C-terminal of peptide chain. These cysteine residues could form inter-molecular disulfide bond so as to affect the folding of peptide chain and protein activity [9]. Besides the C-terminal signal peptide, there is 1 alpha helix, 2 reverse beta folds, and 1 irregular ring region connecting with 2 beta folds, forming the core skeleton of EPF/EPFL peptides. Among the peptides, the 2 beta folds constitute the active region of the molecular, while the irregular ring region takes the responsibility for molecular specificity [10]. EPF/EPFL genes are ubiquitous in terrestrial plants, such as Physcomotrella patens, Selaginella moellendorffii, Picea glauca, Sorghum bicolor, Populus trichocarpa, Medicago truncatula, Carica papaya, while their genome-wide family analyses were only reported on A. thaliana, Oryza sativa, and Malus domestica [11,12,13]. During the specific stages of plant growth and development, EPF1 and EPF2 were found to combine with 3 LRR-RLKs (Leucine-rich repeat receptor kinases), namely as ERECTA (ER), ERECTA-LIKE1 (ERL1), and ERL2, and to interact with 1 LRR-receptor-like protein (LRR-RLP), namely as TOO MANY MOUTHS (TMM), collectively transmitting extracellular specific stomatal development signals [14,15,16,17]. In addition, the homologous EPFL1 was reported to induce the awn elongation in rice [18]. Meanwhile, the ectopic expression of wheat EPFL1 in A. thaliana resulted in the shorter filaments and peduncles, implying its correlation with flower development [19]. Gene editing experiments conducted in Kasalath (one rice cultivar) to knockout 11 OsEPF/EPFL genes by CRISPR/Cas9 (Clustered regularly interspaced short palindromic repeats) indicated that OsEPFL2 participated in the regulation of awn development [ The software TBtools was chosen in this study to extract the position information of EPF/EPFL genes in the chromosomes according to the genome sequences and annotation files of the four representative cotton species. Thesoftware MapChart (https://help.salesforce.com/s/articleView?id=sf.bi_chart_intro_ map.htm&type = 5) was utilized to draw and visualize the physical locations in the cotton chromosomes. Firstly, the annotation files of cotton EPF/EPFL genes were entered into Gene Structure Display Server (GSDS, http://gsds.cbi.pku.edu.cn) so as to analyze their exon–intron structures [46]. Subsequently, the on-line website MEME SUITE (http://meme-suite.org/memei) was utilized for identifying the conserved motifs in cotton EPF/EPFL proteins [47]. Finally, the software TBtools was used to perform the visual-merge map** on the phylogenetic tree, gene structure, and conserved protein motif of all the cotton EPF/EPFL family members. The gene sequences of cotton EPF/EPFL family was subjected to collinearity analysis by the MCScanX software [48], whose visualization was displayed by the software TBtools. The collinearity analyses were composed of intraspecific and interspecific BLAST, which were separately conducted on the diploid genomes of A2 (G. arboretum) and D5 (G. raimondii) and the allotetraploid genomes of AD1 (G. hirsutum) and AD2 (G. barbadense). Subsequently, the duplicated gene pairs were identified from the intraspecific collinearity as Ga-Ga, Gr-Gr, Gh-Gh, and Gb-Gb, which were also identified from the interspecific collinearity as Ga-Gr, Ga-Gh, Ga-Gb, Gr-Gh, Gr-Gb, and Gh-Gb. The obtained duplication events were finally presented as collinearity relationships with the intraspecific and interspecific covariance circles. The DNA sequences of 2000 bp upstream of initiation codon (ATG) of all the 132 cotton EPF/EPFL genes were downloaded as their promoter regions from CottonFGD (http://cottonfgd.net/) database, and on-line tool PlantCARE (http://bioinformatics.psb. ugent.be/webtools/plantcare/html/) [49] was chosen to perform prediction analysis of cis-regulatory elements. The visualization of the predicted cis-regulatory elements were shown by TBtools, and the colorful rectangles presented the different cis-regulatory elements with the same clades of evolution relationships. The transcriptome data of G. hirsutum TM-1 and G. barbadense Hai7124 on the different tissues/organs (root, stem, leaf, petal, torus, sepal, epicalyx, anther, and pistill) were downloaded from the SRA (Sequence Read Archive) database of NCBI website (http://www.ncbi.nlm.nih.gov/, and the accession number was PRJNA490626). Meanwhile, the transcriptome data of G. hirsutum TM-1 in response to multiple abiotic stresses (low temperature at 4℃, high temperature at 37℃, salt treatment of 0.4 M NaCl, and drought treatment of 200 g/liter PEG6000) were also obtained from the SRA database under the accession number PRJNA248163 [32]. The filtering treatment was firstly carried out on the published RNA-seq data by Trimmomatic software [50], and the obtained clean data were subsequently subjected to map** on the reference genome databases built by the HISAT software [51]. The software Cufflinks was chosen to calculate the expression levels of cotton EPF/EPFL genes with the presentation of FPKM (fragments per kilobase of transcript per million fragments) values [52], which were utilized to show whether to express or not, and to show high or low expression levels of all the GhEPF/EPFL and GbEPF/EPFL genes in the different tissues/organs. The FPKM values of all the GhEPF/EPFL genes under the different adversity stresses were subjected to uniformization treatment by the Z-score algorithm, which was performed in order to investigate their up-regulated and down-regulated expression patterns along with the processes of stress occurrence and proceeding [53]. The heat-map of expression levels and patterns of cotton EPF/EPFL genes was finally drawn by the softwareTBtools. The transcriptome data of G. hirsutum TM-1 and G. barbadense Hai7124 on the develo** ovules (0, 1, 3, 5, 10, and 20 days after anthesis, DPA) and fibers (10, 20, and 25 DPA) were downloaded from the SRA database under the accession number of PRJNA490626. These data were subjected to the same treatments in turn as the aforesaid descriptions, including filtering the low-quality data, map** to the reference genome, calculating the expression level, and Z-score uniformization. The positive and negative values represented the up-regulated and down-regulated expression patterns during the development of ovules and fibers, respectively, whose heat-map was also drawn by the software TBtools. In addition, two cultivated species, namely G. hirsutum CCRI36 and G. barbadense Hai1, were chosen in this study to perform quantitative Real-time PCR (qRT-PCR) experiment. The CCRI36 harbored the merits of high yield and wide adaptability, while subjected to normal fiber quality and low resistance to Verticillium wilt, and the Hai1 had the characteristics of superior fiber quality, high VW-resistance, low yield, and poor adaptability [54, 56]. In consideration of the fact that there was no protein data of cotton species recorded into the STRING database (http://string-db.org), homologous alignment was firstly performed between the 44 GhEPF/EPFLs and 11 AtEPF/EPFLs, of which the protein sequences of the latter ones were inputted as the protein models into STRING database for constructing the protein–protein interacting network [57]. Meanwhile, the 44 GhEPF/EPFL genes were also subjected to functional enrichment by GO (Gene Ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) databases, of which the former was generally divided into three categories, namely biological process, cellular component, and molecular function [58], and the latter was composed of cellular processes, environmental information processing, genetic information processing, human disease, metabolism, and organismal systems [59]. The enrichment analyses of GO and KEGG were accomplished by the on-line platform OmicShare (https://www.omicshare.com/), and the detailed procedures were as followed: the ID numbers of GO and KEGG of all the cotton genes were firstly extracted from the released annotation information of G. hirsutum TM-1 [32], which were separately inputted into the OmicShare tools of GO Enrichment Analysis Advanced and Pathway Enrichment Analysis Advanced as the background files together with the 44 GhEPF/EPFL ids as the queries, finally generating the classification results. Twenty-four GaEPF/EPFL, 20 GrEPF/EPFL, 44 GhEPF/EPFL, and 44 GbEPF/EPFL genes were separately identified from the genomes of G. arboreum (A2), G. raimondii (D5), G. hirsutum (AD1) and G. barbadense (AD2) [33, 36, 37], reaching the total number of 132 members (Additional file 1: Table S2). Meanwhile, the analyses of physico-chemical properties of cotton EPF/EPFL family showed that the lengths of amino acid sequences of EPF/EPFL proteins ranged from 87 aa (GrEPF20 and GhEPF35) to 161 aa (GhEPF36 and GbEPF35), and those molecular weight ranges were from 9760.13 kDa (GrEPF20) to 18,049.99 kDa (GhEPF36 and GbEPF35). The isoelectric points of cotton EPF/EPFLs ranged from 5.53 (GhEPF15 and GbEPF17) to 9.91 (GhEPF44), whose average value was 8.84. The ranges of instability index of cotton EPF/EPFLs were from 30.62 (GhEPF21) to 84.76 (GhEPF5), including 12 possibly stable members (≦ 40) and 120 possibly unstable members (> 40). The results of subcellular localization prediction showed that most of cotton EPF/EPFL genes were located in chloroplast (59 members) and extracellular (51 members), and the minority ones were located in mitochondria (9 members), endoplasmic reticulum (4 members), nucleus (4 members), vacuole (4 members), and plasmalemma (1 member). The amino acid alignment of EPF/EPFLs were firstly conducted among the four cotton species, A. thaliana, O. sativa, and S. moellendorffii (Additional file 1: Figure S1), and the results showed the 6-cysteine residues conservatively located in the C-terminal mature peptide region for the both two clades. There were another 2-cysteine residues conserved in the loop region of EPF1-EPF2-EPFL7 clade, implying their diversity with EPFL9/Stomagen clade. Subsequently, 11 AtEPF/EPFL proteins, 13 OsEPF/EPFL proteins, and 17 SmEPF/EPFL proteins were utilized in this study to construct evolutionary tree together with 132 cotton EPF/EPFL proteins, resulting in four clades by virtue of neighbor-joining method (Fig. 1). The largest number of cotton EPF/EPFL proteins (71 ones, 53.8% of the total 132 ones) were classified into one clade with AtEPFL1-3 of the plant model species, therefore this clade was named as EPFL1-3. The most EPF/EPFL proteins of A. thaliana, namely as AtEPFL4, AtEPFL5, AtEPFL6, and AtEPFL8, and the second largest number of cotton EPF/EPFL proteins (24 ones, 18.2%) were divided into the same clade named as EPFL4-6-EPFL8. Eighteen EPF/EPFL proteins in cotton (13.6%) and 3 AtEPF/EPFL proteins, namely as AtEPF1, AtEPF2, and AtEPFL7, were classified into EPF1-EPFL2-EPFL7 clade, and 19 cotton EPF/EPFL proteins (14.4%) and AtEPFL9/Stomagen were divided into EPFL9/Stomagen clade. Phylogenetic analysis of EPF/EPFL proteins from Gossypium, Arabidopsis thaliana, Oryza sativa, and Selaginella moellendorffii. The blue checkmark presents the AtEPF/EPFL proteins, and the purple square presents the OsEPF/EPFL proteins. The red check mark presents the SmEPF/EPFL proteins, and the green triangle presents the GaEPF/EPFL proteins. The red square presents the GrEPF/EPFL proteins, and the yellow star and blue circle present the GhEPF/EPFL and GbEPF/EPFL proteins, respectively As shown in Fig. 2, 24 GaEPF/EPFL genes were unevenly distributed in 10 chromosomes and 1 scaffold except for 4th, 8th, and 13th chromosome in A2 genome, of which the numerous members were found in 5th (5 GaEPF/EPFL genes) and 11th (4 GaEPF/EPFL genes) chromosome. The similar results of 20 GrEPF/EPFL genes also occurred in D5 gnome, and there was no GrEPF/EPFL genes in 12th chromosome besides 4th, 8th, and 13th chromosome, while the different fact was that 9th chromosom chromosome harbored the most number (4 GrEPF/EPFL genes). As for the two AADD genomes, we noticed the same EPF/EPFL members and chromosome distributions between G. hirsutum and G. barbadense. The main difference was found that AD1 genome had two more GhEPF/EPFL genes separately located in D05 and D07 chromosomes, while AD2 genome had two more GbEPF/EPFL genes separately located in A10 and D06 chromosomes. With the comparison between the allotetraploid and diploid genomes, there were 21 GhEPF/EPFL genes and 22 GbEPF/EPFL genes in A subgenomes, and the numbers were less than 24 GaEPF/EPFL genes. On the contrary, 24 GhEPF/EPFL genes and 22 GbEPF/EPFL genes were found in D subgenomes that were more than 20 GrEPF/EPFL genes, despite their total number of EPF/EPFL genes were the same as 44. Besides, the distributed chromosomes between A2 and AD1 or AD2 were one-to-one corresponding, and we found one GhEPF/EPFL gene located in D12 chromosome in AD1 genome while not in D5 and AD2 genomes. These similarities and differences derived from the above results indicated EPF/EPFL gene family showed the conservatism and variability during the long-term evolutionary process. The evolutionary relationships were also investigated among the cotton EPF/EPFLs based on their protein sequences, and the clustering result of 4 clades (Fig. 3a) maintained the consistency with the increased Arabidopsis EPF/EPFL proteins (Fig. 1). The prediction analysis of conserved motifs on the cotton EPF/EPFL proteins (Fig. 3b) indicated that a total of 8 conserved motifs were identified and named as Motif 1 to Motif 8 in turn (Additional file 2: Figure S1). The number of conserved motifs of cotton EPF/EPFLs ranged from 3 to 5, of which the most ones had 4 conserved motifs. Only Motif 1 was commonly observed in all the EPF/EPFL proteins, therefore it was deemed as the most conserved motif with the second lowest E-value. Motif 2 with the minimum E-value and the longest width was also commonly identified among the EPF/EPFL proteins of EPFL1-3, EPF1-EPF2-EPFL7, and EPFL4-6-EPFL8 clades, and Motif 3 with the third lowest E-value and the shortest width was also found among the PF/EPFL proteins of EPFL1-3, EPFL4-6-EPFL8, and EPFL9/Stomagen clades. Besides, we noticed that Motif 5 uniquely while commonly appeared in EPFL9/Stomagen clade, implying its potential significance. The results of gene structure of cotton EPF/EPFL genes showed that the exon number of cotton EPFL/EPFL genes ranged from 1 to 4 (Fig. 3c). The main gene structure of cotton EPF/EPFL genes contained 3 exons and 2 introns (60/132), followed by the gene structures with 2 exons and 1 intron, with 4 exons and 3 introns, and with 1 exon and 0 intron. Gene duplication has been deemed as the main force to extend the number of gene family, which was generally consisted with tandem duplication, fragment duplication, and whole genome duplication [ The expression patterns and qRT-PCR verification of cotton EPF/EPFL genes during fiber development. A presented the analyses of expression patterns of GhEPF/EPFL and GbEPF/EPFL genes on the develo** ovule (0, 1, 3, 5, 10, and 20 DPA) and fiber (10, 20, and 25 DPA), and B presented the qRT-PCR verification of 15 highly expressed GhEPF/EPFL genes on the high-yield and wide-adaptability CCRI36 and superior fiber-quality and high VW-resistance Hai1 at 10, 20, and 25 DPA, respectively During the initial period (0 to 3 DPA) of ovule development, 8 of 44 GhEPF/EPFL genes showed the highest up-regulated fold changes at 0DAP, namely as GhEPF2, GhEPF4, GhEPF14, GhEPF15, GhEPF23, GhEPF24, GhEPF26, and GhEPF36, which were observed at 1 DPA as GhEPF25, GhEPF35, GhEPF37, and GhEPF39, while at 3 DPA as GhEPF9, GhEPF18, and GhEPF28. As for the elongation period (5 to 20 DPA) of ovule development, we noticed that GhEPF6, GhEPF36, GhEPF40, and GhEPF43 at 5 DPA, GhEPF3, GhEPF12, GhEPF13, GhEPF18, GhEPF19, GhEPF24, GhEPF27, and GhEPF42 at 10 DPA, and GhEPF7, GhEPF8, GhEPF22, GhEPF30, and GhEPF31 at 20 DPA, showed the highest up-regulated fold changes. During the elongation and secondary wall thickening periods (10 to 25 DPA) of fiber development, GhEPF20 and GhEPF43 at 10 DPA, GhEPF10, GhEPF16, GhEPF17, GhEPF29, GhEPF39, and GhEPF41 at 20 DPA, and GhEPF1, GhEPF33, and GhEPF41 at 25 DPA harbored the highest up-regulated fold changes. Similarly in the ovule and fiber development of Hai7124, we identified the highest up-regulated fold changes as GbEPF1, GbEPF17, GbEPF30, and GbEPF42 in the ovule at 0 DPA, GbEPF10 and GbEPF21 in the ovule at 1 DPA, GbEPF6-8, GbEPF14, and GbEPF43 in the ovule at 5 DPA, GbEPF12, GbEPF13, GbEPF19, GbEPF21, GbEPF29, GbEPF33, GbEPF34, and GbEPF41 in the ovule at 10 DPA, GbEPF16 and GbEPF24 in the ovule at 20 DPA, GbEPF18 and GbEPF40 in the fiber at 10 DPA, GbEPF2, GbEPF15, and GbEPF38 in the fiber at 20 DPA, and GbEPF3, GbEPF4, GbEPF9, GbEPF25, GbEPF26, GbEPF28, GbEPF32, and GbEPF39 in the fiber at 25 DPA. These data indicated that cotton EPF/EPFL genes might play important roles affecting the development and growth of ovules and fibers. After conducting the relative quantization on the 13 high-expressed GhEPF genes during the key development periods of cotton fiber (10, 20, and 25 DPA), the largest number of GhEPF genes showed the dramatic changes on the expression levels either between the two different varieties or among the three develo** periods, while little changes were mainly observed in GhEPF20, GhEPF26, and GhEPF43 between CCRI36 and Hai1 at 20 and 25 DPA. As for the fiber-elongation period (10 DPA), we noticed GhEPF10, GhEPF26, and GhEPF43 presented higher expression levels in Hai1 than those in CCRI36. During the secondary-wall thickening periods, GhEPF13, GhEPF28, GhEPF29, GhEPF39, GhEPF40, and GhEPF43 were more highly expressed in Hai1 than those in CCRI36 at 20 DPA, and GhEPF13, GhEPF14, GhEPF23, GhEPF25, GhEPF26, GhEPF28, and GhEPF29 showed higher expression levels in Hai1 than those in CCRI36 at 25 DPA. To sum up, despite there were no GhEPF genes harboring the steadily higher expression levels in Hai1 under the key developmental periods of fiber elongation and secondary-wall thickening periods, GhEPF26 and GhEPF43 were separately observed to highly expressed in sea island cotton than those in upland cotton between 10 DAP and 25 DPA, and between 10 and 20 DPA, while GhEPF13, GhEPF28, and GhEPF29 were found with higher expression levels in sea island cotton than those in upland cotton between 20 and 25 DPA. These highly expressed GhEPF genes might be the candidate genes determining why the sea island cotton had the superior fiber quality than upland cotton, which definitely require the further experiments of genetic transformation to verify their potential functions in fiber development.Map** EPF/EPFL genes in cotton chromosomes
Analyses of gene structure and conserved protein motif of cotton EPF/EPFL genes
Collinearity analysis of cotton EPF/EPFL genes
Analyses of cis-regulatory elementsof cotton EPF/EPFL genes
Analyses of expression patterns and quantitative Real-time PCR verification
Analyses of protein–protein interaction and functional enrichment of GhEPF/EPFL genes
Results
Identification and physiochemical characteristic analyses of cotton EPF/EPFL genes
Phylogenetic analysis of EPF/EPFL proteins in Gossypium and Arabidopsis
Chromosomal location and gene duplication of cotton EPF/EPFLs
Gene structure and conserved motif prediction of cotton EPF/EPFLs
Collinearity analysis of EPF/EPFL genes in cotton
Analyses of protein–protein interaction and functional enrichment of GhEPF/EPFL genes
For screening the candidate EPF/EPFL genes that could be utilized to cultivate the high-yield, superior fiber-quality, and multiple-resistance varieties in cotton by over-expression and gene-editing technology, the analyses of protein–protein interaction and functional enrichment were conducted on the cotton EPF/EPFL genes, respectively. Due to the fact that there was no uploaded protein data of cotton species in the STRING database (http://string-db.org/), homologous alignment was firstly carried out between the EPF/EPFL proteins of A. thaliana and G. hirsutum, which were subsequently utilized to draw the interaction networks together with their interacting proteins (Fig. 8A). The results indicated that the core interaction networks concentrated on the eight EPF/EPFL proteins of A. thaliana, namely EPFL8 (homologous proteins as GhEPF8, GhEPF28, and GhEPF38), EPFL2 (homologous proteins as GhEPF10, GhEPF17, GhEPF18, GhEPF21, GhEPF29, GhEPF41, and GhEPF42), EPFL5 (homologous protein as GhEPF39), EPFL4 (homologous proteins as GhEPF5, GhEPF14, GhEPF16, GhEPF23, and GhEPF36), EPFL6 (homologous proteins as GhEPF4 and GhEPF26), EPF1 (homologous protein as GhEPF1), EPF2 (homologous proteins as GhEPF3, GhEPF7, GhEPF24, and GhEPF31), and EPFL3(homologous proteins as GhEPF6, GhEPF9, GhEPF11, GhEPF15, GhEPF27, GhEPF30, GhEPF32, and GhEPF37) ordered by the interacting extents. In order to further investigate the potential functions of these GhEPF/EPFL proteins, enrichment analyses of GO and KEGG pathways were separately accomplished as shown in Fig. 8B and 8C. As for the GO categories, the most significant enriched terms were stomatal complex development (GO:0010374), regulation of stomatal complex development (GO:2,000,038), and plant epidermis development (GO:0090558) in biological processes, membrane (GO:0016020), integral component of membrane (GO:0016021), and membrane part (GO:0044425) in cellular component, and hydrolase activity (GO:0016787), hydrolase activity (GO:0016787), and nucleotide binding (GO:0000166) in molecular function. Among the annotated EPF/EPFL genes in GO terms, GhEPF1, GhEPF7, GhEPF13, GhEPF20, GhEPF22, GhEPF24, GhEPF31, GhEPF33, GhEPF34, and GhEPF43 were the most enriched ones that participate in the top 10 biological processes. While, only DNA replication (ko03030), base excision repair (ko03410), nucleotide excision repair (ko03420), and nucleotide excision repair (ko03440) were the enriched KEGG pathways by the same 5 EPF/EPFL genes, namely GhEPF1, GhEPF7, GhEPF22, GhEPF24, and GhEPF31. The comprehensive analyses of protein–protein interaction and functional enrichment on the GhEPF/EPFL genes indicated the two comment enriched genes were found namely as GhEPF1 and GhEPF7, which might deserve more attentions for further functional verification and interacting-mechanism research.
Discussion
Cotton is one of the most widely cultivated cash crops all around the world, whose natural fiber, edible oil, vegetable protein, and other products have greatly contributed to the national economy and social development [61]. However, the cotton production is suffering the increasingly serious disputes on arable lands between grain and cotton. The deteriorating ecological environment by salinization phenomenon and extreme weather (high/low temperature or flood and drought), the ever-expanding diseases (Verticillium wilt and Fusarium wilt) and pets (cotton bollworms and aphids), while the growing population of human beings and the rising living standards and consumption levels, resulting in that how to simultaneously improve the fiber yield and quality so as to cultivate and promote new varieties with high production, superior quality, and multiple resistance has become a different issue to be solved urgently at present [62,63,64]. Therefore, it is of necessity and significance to concentrate on the functional genes relevant to fiber development and growth since the specific biological process could determine the final fiber yield and quality. Coincidentally, EPF/EPFL gene family was annotated as the epidermal patterning factor [65], and cotton fiber was a single-cell seed trichome differentiated from the epidermal cell of ovule [66], implying the close relationships between the two above-mentioned ones. Despite there was few reports on EPF/EPFL genes in cotton, EPF/EPFL gene family has been confirmed in multiple plant species to participate in plant development and growth, especially as the morphogenesis processes of stoma, awn, stamen, and fruit skin [14,15,16,17,18,19,11,12,13], the same diploid cotton species had more numbers of EPF/EPFL genes, which might be relevant to their differences on genome complexity and evolutionary origins while not depend on the more or less of chromosome numbers and belonging phyla of herbaceous or ligneous plants. Interestingly, the GhEPF/EPFL or GbEPF/EPFL number was the sum number of GrEPF/EPFL and GaEPF/EPFL genes, while the tetraploid ones were not strictly twice of the diploid ones, which was proved by the non-absolute one to one correspondences of chromosome locations of EPF/EPFL gene between A-subgenome or D-subgeome of allotetraploid cotton species and A2 or D5 of diploid cotton species (Fig. 2). The fact of the incomplete equilibrium in cotton EPF/EPFL gene family indicated the conservatism and variability during the natural hybridization and chromosome reduplication. In the meantime, the on-line tools were utilized to predict the physical and chemical characteristics, and either amino acid length and molecular weight or isoelectir point and instability index widely ranged among all the cotton EPF/EPFL proteins. The largest number of cotton EPF/EPFL gene family were predicted to be located in chloroplast and extracellular, of which the distribution in former location might be related with ROS (reactive oxygen species) concentrations caused by some over-expressed EPF/EPFL genes against adversity stresses, such as MdEPF2 [13], while the latter location indicated these secretory polypeptides might combine with receptors on the cell membrane to transfer the extracellular signals and to play important roles in the process of signaling between cells [67, 68].
A total of 173 EPF/EPFL proteins were used to construct the phylogenetic tree, including 11 AtEPF/EPFL, 13 OsEPF/EPFL, 17 SmEPF/EPFL, and 132 cotton EPF/EPFL proteins, which were divided into four clades (Fig. 1). Each clade was composed of four kinds of EPF/EPFLs derived from four cotton species, and was separately named as EPF1-EPF2-EPFL7, EPFL1-3, EPFL4-6-EPFL8, and EPFL9/Stomagen according to the included AtEPF/EPFLs, which was consistent with the previous studies [11,12,13]. After removing the AtEPF/EPFL proteins, all the cotton EPF/EPFLs were also subjected to investigate their evolutionary relationships (Fig. 3a), and the similar clustering results in gene structure and conserved motif confirmed the phylogenetic categories. Besides, we noticed that the number of conserved motifs ranged from 3 to 5 in the cotton EPF/EPFL proteins, while only Motif 1 was commonly found in all the EPF/EPFLs (Fig. 3b). The numbers of exon and intron of cotton EPF/EPFL genes ranged from 1 to 4 and 0 to 3, respectively, of which the main gene structure was composed of 3 exons and 2 introns (Fig. 3c). Despite more EPF/EPFL genes were observed in cotton species, the evolutionary classification, conserved motif, and exon–intron structure were generally accorded with those in A. thaliana, O. sativa and M. domestica [11,12,13], indicating the high conservation during the natural evolution and species expansion.
The uneven distribution of cotton EPF/EPFL genes in the chromosomes between the tetraploid subgenome and the diploid genome promoted us to perform the collineary analysis for estimating the possible reasons. As we all know, gene duplication has been proved to provide the main contribution to the extension of gene family, generally including tandem duplication, fragment duplication, and whole genome duplication [39]. The intraspecific and interspecific results of collineary relationships among the four cotton species showed the total duplication genes were 740 pairs (Fig. 4), which provided strong evidences that gene duplication could make difference in expanding EPF/EPFL genes like other families in cotton [69, 70]. The increased size and complexity of plant genome were accomplished by the events of chromosome doubling and gene duplication, which also made it harder to accurately assembly and obtain the whole genome information even though combing the second-generation and third-generation sequencing technologies with High-through chromosome conformation capture (Hi-C) and Bionano optical maps [32, 38]. The inevitable mistakes and unassembled scaffolds in cotton genomes might also be cause for explaining the non-strict duplication phenomenon between the tetraploid and diploid species.
The gene functions were closely related with their coding regions, while their transcriptions did occur or not were up to the combination between the transcription factors and promoter regions [71]. Cis-regulatory elements of the cotton EPF/EPFL genes were analyzed to comprehend their potential functions in the multiple biological processes, and the 15 identified kinds of cis-regulatory elements were divided into three categories: plant development and growth, plant hormone response, and abiotic stresses (Fig. 5). The most researches of EPF/EPFL functions were concentrated on the stoma development [9, 10, 12] followed by the development of awn and stamen [18,19,7A. Compared with the expression patterns between ovule and fiber development, we noticed that more EPF/EPFL genes showed the down-regulated expression patterns in fiber development, while more patterns of firstly down-regulated and then up-regulated expression were observed in ovule development. These data indicated the cotton EPF/EPFL genes presented the more dramatic differences to affect the ovule development, which could determine the final fiber yield [76]. The various changes of EPF/EPFL genes during the development of fibers also implied their significant roles to control fiber quality, therefore qRT-PCR experiments were adopted to verify and screen the potential candidate genes in two tetraploid cotton varieties harboring the extreme difference between the cotton yield and fiber quality (Fig. 7B). Most of the selected 15 GhEPF/EPFL genes were differentially expressed either in the two materials or during the different periods of fiber development, of which GhEPF10, GhEPF26, and GhEPF43 presented higher expression levels in Hai1 than those in CCRI36 during the fiber-elongation period (10 DPA). As for the period of secondary cell-wall thickening, GhEPF13, GhEPF28, GhEPF29, GhEPF39, GhEPF40, and GhEPF43 were more highly expressed in Hai1 than those in CCRI36 at 20 DPA, while GhEPF13, GhEPF14, GhEPF23, GhEPF25, GhEPF26, GhEPF28, and GhEPF29 showed higher expression levels in Hai1 than those in CCRI36 at 25 DPA. On one hand, these differentially expresses GhEPF/EPFL genes could make contributions to fiber development, and on the other hand, the common high-expression GhEPF26 between 10 and 25 DPA, GhEPF43 between 10 and 20 DPA, and GhEPF13, GhEPF28, and GhEPF29 between 20 and 25 DPA in the superior fiber-quality cotton variety provided the abundant gene resource for further genetic transformation verification and molecular breeding in cotton production.
Protein interacting network, also namely as protein–protein interaction (PPI) analysis, is composed of individual proteins interacting with each other, afterwards to participate in the biological signal transmission, gene expression regulation, energy and material metabolism, and cell cycle control [77]. PPI analysis has been widely utilized in the gene family researches to construct the protein interacting network for screening the core candidate genes [78,79,80], and our results also identified the core proteins namely as EPFL8, EPFL2, EPFL5, EPFL4, EPFL6, EPF1, EPF2, and EPFL3 (Fig. 8A). However, more homologous proteins were found in cotton genomes, which required other database to perform conjoint analyses for narrowing the range of the key candidate genes. The enrichment analyses of GO and KEGG pathways were adopted to excavate the potential functional categories and significant signal transduction (Fig. 8B and 8C), of which the most significant enriched biological processes of GO terms were stomatal complex development (GO:0010374), regulation of stomatal complex development (GO:2,000,038), and plant epidermis development (GO:0090558), and DNA replication (ko03030), base excision repair (ko03410), nucleotide excision repair (ko03420), and nucleotide excision repair (ko03440) were identified as the enriched KEGG pathways. The close relationship of cotton EPF/EPFL genes with stoma development and epidermis development were consistent with the previous studies [11,12,13]. Having employing the comprehensive analyses of PPI, GO enrichment, and KEGG pathway on the GhEPF/EPFL genes, GhEPF1 and GhEPF7 were finally screened as the common enriched genes for further research by cultivating transgenetic varieties and investigating the interacting-mechanism among the signal pathways.
Conclusion
Genome identification of EPF/EPFL gene family was performed on the four representative cotton species, and 132 cotton EPF/EPFL proteins were divided into 4 clades based on their evolutionary relationships. The comprehensive analyses of chromosome location, gene structure, conserved motifs, and collineary relationship indicated the conservatism and variability during the natural hybridization and chromosome reduplication, and expression patterns in response to multiple abiotic stresses and during the development of ovules and fibers demonstrated that cotton EPF/EPFL genes participate in coordinating the adversity responsiveness and plant development. Finally, the crucial candidate EPF/EPFL genes were comprehensively screened by protein interacting network and enrichment analyses of GO and KEGG pathway, and these results not only provided useful information for further functional verification, but also established a solid foundation for molecular breeding in cotton production.
Availability of data and materials
All data generated or analyzed in this study are included in this published article.
References
Drakakaki G, Dandekar A. Protein secretion: how many secretory routes does a plant cell have? Plant Sci. 2013;203–204:74–8. https://doi.org/10.1016/j.plantsci.2012.12.017.
Grudkowska M, Zagdańska B. Multifunctional role of plant cysteine proteinases. Acta Biochim Pol. 2004;51(3):609–24.
Rawlings ND, Barrett AJ, Thomas PD, Huang X, Bateman A, Finn RD. The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res. 2018;46(D1):D624–32. https://doi.org/10.1093/nar/gkx1134.
Szewińska J, Simińska J, Bielawski W. The roles of cysteine proteases and phytocystatins in development and germination of cereal seeds. J Plant Physiol. 2016;207:10–21. https://doi.org/10.1016/j.jplph.2016.09.008.
Höwing T, Dann M, Müller B, Helm M, Scholz S, Schneitz K, Hammes UZ, Gietl C. The role of KDEL-tailed cysteine endopeptidases of Arabidopsis (AtCEP2 and AtCEP1) in root development. PLoS ONE. 2018;13(12): e0209407. https://doi.org/10.1371/journal.pone.0209407.
Liu H, Hu M, Wang Q, Cheng L, Zhang Z. Role of Papain-Like Cysteine Proteases in Plant Development. Front Plant Sci. 2018;9:1717. https://doi.org/10.3389/fpls.2018.01717.
Minami A, Fukuda H. Transient and specific expression of a cysteine endopeptidase associated with autolysis during differentiation of Zinnia mesophyll cells into tracheary elements. Plant Cell Physiol. 1995;36(8):1599–606.
Cheng Z, Guo X, Zhang J, Liu Y, Wang B, Li H, Lu H. βVPE is involved in tapetal degradation and pollen development by activating proprotease maturation in Arabidopsis thaliana. J Exp Bot. 2020;71(6):1943–55. https://doi.org/10.1093/jxb/erz560.
Hara K, Kajita R, Torii KU, Bergmann DC, Kakimoto T. The secretory peptide gene EPF1 enforces the stomatal one-cell-spacing rule. Genes Dev. 2007;21(14):1720–5. https://doi.org/10.1101/gad.1550707.
Ohki S, Takeuchi M, Mori M. The NMR structure of stomagen reveals the basis of stomatal density regulation by plant peptide hormones. Nat Commun. 2011;2:512. https://doi.org/10.1038/ncomms1520.
Takata N, Yokota K, Ohki S, Mori M, Taniguchi T, Kurita M. Evolutionary relationship and structural characterization of the EPF/EPFL gene family. PLoS ONE. 2013;8(6): e65183. https://doi.org/10.1371/journal.pone.0065183.
Lu J, He J, Zhou X, Zhong J, Li J, Liang YK. Homologous genes of epidermal patterning factor regulate stomatal development in rice. J Plant Physiol. 2019;234–235:18–27. https://doi.org/10.1016/j.jplph.2019.01.010.
Jiang Q, Yang J, Wang Q, Zhou K, Mao K, Ma F. Overexpression of MdEPF2 improves water use efficiency and reduces oxidative stress in tomato. Environ Exp Bot. 2019. https://doi.org/10.1016/j.envexpbot.2019.03.009.
Shpak ED, McAbee JM, Pillitteri LJ, Torii KU. Stomatal patterning and differentiation by synergistic interactions of receptor kinases. Science. 2005;309(5732):290–3. https://doi.org/10.1126/science.1109710.
Yang M, Sack FD. The too many mouths and four lips mutations affect stomatal production in Arabidopsis. Plant Cell. 1995;7(12):2227–39. https://doi.org/10.1105/tpc.7.12.2227.
Vatén A, Bergmann DC. Mechanisms of stomatal development: an evolutionary view. EvoDevo. 2012;3(1):11. https://doi.org/10.1186/2041-9139-3-11.
Jangra R, Brunetti SC, Wang X, Kaushik P, Gulick PJ, Foroud NA, Wang S, Lee JS. Duplicated antagonistic EPF peptides optimize grass stomatal initiation. Development. 2021, 148(16):dev199780. https://doi.org/10.1242/dev.199780.
Bessho-Uehara K, Wang DR, Furuta T, Minami A, Nagai K, Gamuyao R, Asano K, Angeles-Shim RB, Shimizu Y, Ayano M, Komeda N, Doi K, Miura K, Toda Y, Kinoshita T, Okuda S, Higashiyama T, Nomoto M, Tada Y, Shinohara H, Matsubayashi Y, Greenberg A, Wu J, Yasui H, Yoshimura A, Mori H, McCouch SR, Ashikari M. Loss of function at RAE2, a previously unidentified EPFL, is required for awnlessness in cultivated Asian rice. Proc Natl Acad Sci U S A. 2016, 113(32):8969–74. https://doi.org/10.1073/pnas.1604849113.
Sun Q, Qu J, Yu Y, Yang Z, Wei S, Wu Y, Yang J, Peng Z. TaEPFL1, an EPIDERMAL PATTERNING FACTOR-LIKE (EPFL) secreted peptide gene, is required for stamen development in wheat. Genetica. 2019;147(2):121–30. https://doi.org/10.1007/s10709-019-00061-7.
**ong L, Huang Y, Liu Z, Li C, Yu H, Shahid MQ, Lin Y, Qiao X, **ao J, Gray JE, ** J. Small EPIDERMAL PATTERNING FACTOR-LIKE2 peptides regulate awn development in rice. Plant Physiol. 2022;190(1):516–31. https://doi.org/10.1093/plphys/kiac278.
Tameshige T, Okamoto S, Lee JS, Aida M, Tasaka M, Torii KU, Uchida N. A Secreted Peptide and Its Receptors Shape the Auxin Response Pattern and Leaf Margin Morphogenesis. Curr Biol. 2016;26(18):2478–85. https://doi.org/10.1016/j.cub.2016.07.014.
Abrash EB, Davies KA, Bergmann DC. Generation of signaling specificity in Arabidopsis by spatially restricted buffering of ligand-receptor interactions. Plant Cell. 2011;23(8):2864–79. https://doi.org/10.1105/tpc.111.086637.
Uchida N, Lee JS, Horst RJ, Lai HH, Kajita R, Kakimoto T, Tasaka M, Torii KU. Regulation of inflorescence architecture by intertissue layer ligand-receptor communication between endodermis and phloem. Proc Natl Acad Sci U S A. 2012;109(16):6337–42. https://doi.org/10.1073/pnas.1117537109.
Uchida N, Tasaka M. Regulation of plant vascular stem cells by endodermis-derived EPFL-family peptide hormones and phloem-expressed ERECTA-family receptor kinases. J Exp Bot. 2013;64(17):5335–43. https://doi.org/10.1093/jxb/ert196.
Sugano SS, Shimada T, Imai Y, Okawa K, Tamai A, Mori M, Hara-Nishimura I. Stomagen positively regulates stomatal density in Arabidopsis. Nature. 2010;463(7278):241–4. https://doi.org/10.1038/nature08682.
Kondo T, Kajita R, Miyazaki A, Hokoyama M, Nakamura-Miura T, Mizuno S, Masuda Y, Irie K, Tanaka Y, Takada S, Kakimoto T, Sakagami Y. Stomatal density is controlled by a mesophyll-derived signaling molecule. Plant Cell Physiol. 2010;51(1):1–8. https://doi.org/10.1093/pcp/pcp180.
Wang C, Liu S, Dong Y, Zhao Y, Geng A, **a X, Yin W. PdEPF1 regulates water-use efficiency and drought tolerance by modulating stomatal density in poplar. Plant Biotechnol J. 2016;14(3):849–60. https://doi.org/10.1111/pbi.12434.
Hughes J, Hepworth C, Dutton C, Dunn JA, Hunt L, Stephens J, Waugh R, Cameron DD, Gray JE. Reducing Stomatal Density in Barley Improves Drought Tolerance without Impacting on Yield. Plant Physiol. 2017;174(2):776–87. https://doi.org/10.1104/pp.16.01844.
Caine RS, Yin X, Sloan J, Harrison EL, Mohammed U, Fulton T, Biswal AK, Dionora J, Chater CC, Coe RA, Bandyopadhyay A, Murchie EH, Swarup R, Quick WP, Gray JE. Rice with reduced stomatal density conserves water and has improved drought tolerance under future climate conditions. New Phytol. 2019;221(1):371–84. https://doi.org/10.1111/nph.15344.
Hepworth C, Doheny-Adams T, Hunt L, Cameron DD, Gray JE. Manipulating stomatal density enhances drought tolerance without deleterious effect on nutrient uptake. New Phytol. 2015;208(2):336–41. https://doi.org/10.1111/nph.13598.
Hepworth C, Turner C, Landim MG, Cameron D, Gray JE. Balancing Water Uptake and Loss through the Coordinated Regulation of Stomatal and Root Development. PLoS ONE. 2016;11(6): e0156930. https://doi.org/10.1371/journal.pone.0156930.
Hu Y, Chen J, Fang L, Zhang Z, Ma W, Niu Y, Ju L, Deng J, Zhao T, Lian J, Baruch K, Fang D, Liu X, Ruan YL, Rahman MU, Han J, Wang K, Wang Q, Wu H, Mei G, Zang Y, Han Z, Xu C, Shen W, Yang D, Si Z, Dai F, Zou L, Huang F, Bai Y, Zhang Y, Brodt A, Ben-Hamo H, Zhu X, Zhou B, Guan X, Zhu S, Chen X, Zhang T. Gossypium barbadense and Gossypium hirsutum genomes provide insights into the origin and evolution of allotetraploid cotton. Nat Genet. 2019;51(4):739–48. https://doi.org/10.1038/s41588-019-0371-5.
Chen ZJ, Sreedasyam A, Ando A, Song Q, De Santiago LM, Hulse-Kemp AM, Ding M, Ye W, Kirkbride RC, Jenkins J, Plott C, Lovell J, Lin YM, Vaughn R, Liu B, Simpson S, Scheffler BE, Wen L, Saski CA, Grover CE, Hu G, Conover JL, Carlson JW, Shu S, Boston LB, Williams M, Peterson DG, McGee K, Jones DC, Wendel JF, Stelly DM, Grimwood J, Schmutz J. Genomic diversifications of five Gossypium allopolyploid species and their impact on cotton improvement. Nat Genet. 2020;52(5):525–33. https://doi.org/10.1038/s41588-020-0614-5.
Fang L, Wang Q, Hu Y, Jia Y, Chen J, Liu B, Zhang Z, Guan X, Chen S, Zhou B, Mei G, Sun J, Pan Z, He S, **ao S, Shi W, Gong W, Liu J, Ma J, Cai C, Zhu X, Guo W, Du X, Zhang T. Genomic analyses in cotton identify signatures of selection and loci associated with fiber quality and yield traits. Nat Genet. 2017;49(7):1089–98. https://doi.org/10.1038/ng.3887.
Zhang D, Zhang Z, Unver T, Zhang B. CRISPR/Cas: A powerful tool for gene function study and crop improvement. J Adv Res. 2020;29:207–21. https://doi.org/10.1016/j.jare.2020.10.003.
Du X, Huang G, He S, Yang Z, Sun G, Ma X, Li N, Zhang X, Sun J, Liu M, Jia Y, Pan Z, Gong W, Liu Z, Zhu H, Ma L, Liu F, Yang D, Wang F, Fan W, Gong Q, Peng Z, Wang L, Wang X, Xu S, Shang H, Lu C, Zheng H, Huang S, Lin T, Zhu Y, Li F. Resequencing of 243 diploid cotton accessions based on an updated A genome identifies the genetic basis of key agronomic traits. Nat Genet. 2018;50(6):796–802. https://doi.org/10.1038/s41588-018-0116-x.
Udall JA, Long E, Hanson C, Yuan D, Ramaraj T, Conover JL, Gong L, Arick MA, Grover CE, Peterson DG, Wendel JF. De Novo Genome Sequence Assemblies of Gossypium raimondii and Gossypium turneri. G3 (Bethesda). 2019, 9(10):3079–3085. https://doi.org/10.1534/g3.119.400392.
Wang M, Tu L, Yuan D, Zhu D, Shen C, Li J, Liu F, Pei L, Wang P, Zhao G, Ye Z, Huang H, Yan F, Ma Y, Zhang L, Liu M, You J, Yang Y, Liu Z, Huang F, Li B, Qiu P, Zhang Q, Zhu L, ** S, Yang X, Min L, Li G, Chen LL, Zheng H, Lindsey K, Lin Z, Udall JA, Zhang X. Reference genome sequences of two cultivated allotetraploid cottons, Gossypium hirsutum and Gossypium barbadense. Nat Genet. 2019;51(2):224–9. https://doi.org/10.1038/s41588-018-0282-x.
Peng R, Xu Y, Tian S, Unver T, Liu Z, Zhou Z, Cai X, Wang K, Wei Y, Liu Y, Wang H, Hu G, Zhang Z, Grover CE, Hou Y, Wang Y, Li P, Wang T, Lu Q, Wang Y, Conover JL, Ghazal H, Wang Q, Zhang B, Van Montagu M, Van de Peer Y, Wendel JF, Liu F. Evolutionary divergence of duplicated genomes in newly described allotetraploid cottons. Proc Natl Acad Sci USA. 2022;119(39):e2208496119. https://doi.org/10.1073/pnas.2208496119.
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, **a R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol Plant. 2020;13(8):1194–202. https://doi.org/10.1016/j.molp.2020.06.009.
Hao XD, Liu Y, Li BW, Wu W, Zhao XW. Exome sequencing analysis identifies novel homozygous mutation in ABCA4 in a Chinese family with Stargardt disease. Int J Ophthalmol. 2020;13(4):671–6. https://doi.org/10.18240/ijo.2020.04.22.
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K. WoLF PSORT: protein localization predictor. Nucleic Acids Res. 2007, 35(Web Server issue):W585–7. https://doi.org/10.1093/nar/gkm259.
Thompson JD, Gibson TJ, Higgins DG. Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinformatics. 2002, Chapter 2:Unit 2.3. https://doi.org/10.1002/0471250953.bi0203s00. PMID: 18792934.
Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol. 2016;33(7):1870–4. https://doi.org/10.1093/molbev/msw054.
He Z, Zhang H, Gao S, Lercher MJ, Chen WH, Hu S. Evolview v2: an online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Res. 2016;44(W1):W236–41. https://doi.org/10.1093/nar/gkw370.
Hu B, ** J, Guo AY, Zhang H, Luo J, Gao G. GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics. 2015, 31(8):1296–7. https://doi.org/10.1093/bioinformatics/btu817.
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009, 37(Web Server issue):W202–8. https://doi.org/10.1093/nar/gkp335.
Wang Y, Tang H, Debarry JD, Tan X, Li J, Wang X, Lee TH, ** H, Marler B, Guo H, Kissinger JC, Paterson AH. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012;40(7):e49. https://doi.org/10.1093/nar/gkr1293.
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002;30(1):325–7. https://doi.org/10.1093/nar/30.1.325.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30(15):2114–20. https://doi.org/10.1093/bioinformatics/btu170.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12(4):357–60. https://doi.org/10.1038/nmeth.3317.
Ghosh S, Chan CK. Analysis of RNA-Seq Data Using TopHat and Cufflinks. Methods Mol Biol. 2016;1374:339–61. https://doi.org/10.1007/978-1-4939-3167-5_18.
Alkhateeb A, Rueda L. Zseq: An Approach for Preprocessing Next-Generation Sequencing Data. J Comput Biol. 2017;24(8):746–55. https://doi.org/10.1089/cmb.2017.0021.
Li PT, Wang M, Lu QW, Ge Q, Rashid MHO, Liu AY, Gong JW, Shang HH, Gong WK, Li JW, Song WW, Guo LX, Su W, Li SQ, Guo XP, Shi YZ, Yuan YL. Comparative transcriptome analysis of cotton fiber development of Upland cotton (Gossypium hirsutum) and Chromosome Segment Substitution Lines from G. hirsutum × G. barbadense. BMC Genomics. 2017;18(1):705. https://doi.org/10.1186/s12864-017-4077-8.
Li PT, Rashid MHO, Chen TT, Lu QW, Ge Q, Gong WK, Liu AY, Gong JW, Shang HH, Deng XY, Li JW, Li SQ, **ao XH, Liu RX, Zhang Q, Duan L, Zou XY, Zhang Z, Jiang X, Zhang Y, Peng RH, Shi YZ, Yuan YL. Transcriptomic and biochemical analysis of upland cotton (Gossypium hirsutum) and a chromosome segment substitution line from G. hirsutum × G. barbadense in response to Verticillium dahliae infection. BMC Plant Biol. 2019;19(1):19. https://doi.org/10.1186/s12870-018-1619-4.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–8. https://doi.org/10.1006/meth.2001.1262.
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021;49(D1):D605–12. https://doi.org/10.1093/nar/gkaa1074.
Gene Ontology Consortium. Gene Ontology Consortium: going forward. Nucleic Acids Res. 2015, 43(Database issue):D1049–56. https://doi.org/10.1093/nar/gku1179.
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016;44(D1):D457–62. https://doi.org/10.1093/nar/gkv1070.
Li S, Kong L, **ao X, Li P, Liu A, Li J, Gong J, Gong W, Ge Q, Shang H, Pan J, Chen H, Peng Y, Zhang Y, Lu Q, Shi Y, Yuan Y. Genome-wide artificial introgressions of Gossypium barbadense into G. hirsutum reveal superior loci for simultaneous improvement of cotton fiber quality and yield traits. J Adv Res. 2022, S2090–1232(22)00265-X. https://doi.org/10.1016/j.jare.2022.11.009.
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38(4):576–89. https://doi.org/10.1016/j.molcel.2010.05.004.
Peng R, Jones DC, Liu F, Zhang B. From Sequencing to Genome Editing for Cotton Improvement. Trends Biotechnol. 2021;39(3):221–4. https://doi.org/10.1016/j.tibtech.2020.09.001.
Yang Z, Qanmber G, Wang Z, Yang Z, Li F. Gossypium Genomics: Trends, Scope, and Utilization for Cotton Improvement. Trends Plant Sci. 2020;25(5):488–500. https://doi.org/10.1016/j.tplants.2019.12.011.
Yang Z, Gao C, Zhang Y, Yan Q, Hu W, Yang L, Wang Z, Li F. Recent progression and future perspectives in cotton genomic breeding. J Integr Plant Biol. 2023;65(2):548–69. https://doi.org/10.1111/jipb.13388.
Wang M, Yang K, Le J. Organ-specific effects of brassinosteroids on stomatal production coordinate with the action of Too Many Mouths. J Integr Plant Biol. 2015;57(3):247–55. https://doi.org/10.1111/jipb.12285.
Qin Y, Sun M, Li W, Xu M, Shao L, Liu Y, Zhao G, Liu Z, Xu Z, You J, Ye Z, Xu J, Yang X, Wang M, Lindsey K, Zhang X, Tu L. Single-cell RNA-seq reveals fate determination control of an individual fibre cell initiation in cotton (Gossypium hirsutum). Plant Biotechnol J. 2022;20(12):2372–88. https://doi.org/10.1111/pbi.13918.
Richardson LG, Torii KU. Take a deep breath: peptide signalling in stomatal patterning and differentiation. J Exp Bot. 2013;64(17):5243–51. https://doi.org/10.1093/jxb/ert246.
Shimada T, Sugano SS, Hara-Nishimura I. Positive and negative peptide signals control stomatal density. Cell Mol Life Sci. 2011;68(12):2081–8. https://doi.org/10.1007/s00018-011-0685-7.
Wei Y, Li Z, Wedegaertner TC, Jaconis S, Wan S, Zhao Z, Liu Z, Liu Y, Zheng J, Hake KD, Peng R, Zhang B. Conservation and Divergence of Phosphoenolpyruvate Carboxylase Gene Family in Cotton. Plants (Basel). 2022;11(11):1482. https://doi.org/10.3390/plants11111482.
Gu H, Zhao Z, Wei Y, Li P, Lu Q, Liu Y, Wang T, Hu N, Wan S, Zhang B, Hu S, Peng R. Genome-Wide Identification and Functional Analysis of RF2 Gene Family and the Critical Role of GhRF2-32 in Response to Drought Stress in Cotton. Plants (Basel). 2023;12(14):2613. https://doi.org/10.3390/plants12142613.
Haberle V, Stark A. Eukaryotic core promoters and the functional basis of transcription initiation. Nat Rev Mol Cell Biol. 2018;19(10):621–37. https://doi.org/10.1038/s41580-018-0028-8.
Liu S, Jia F, Jiao Z, Wang J, **a X, Yin W. Ectopic expression of secretory peptide PdEPF3 in Arabidopsis confers drought tolerance with reduced stomatal density. Acta Soc Bot Pol. 2019;88(2):3627. https://doi.org/10.5586/asbp.3627.
Hara K, Yokoo T, Kajita R, Onishi T, Yahata S, Peterson KM, Torii KU, Kakimoto T. Epidermal cell density is autoregulated via a secretory peptide, EPIDERMAL PATTERNING FACTOR 2 in Arabidopsis leaves. Plant Cell Physiol. 2009;50(6):1019–31. https://doi.org/10.1093/pcp/pcp068.
Buckley CR, Caine RS, Gray JE. Pores for Thought: Can Genetic Manipulation of Stomatal Density Protect Future Rice Yields? Front Plant Sci. 2020;10:1783. https://doi.org/10.3389/fpls.2019.01783.
Negoro S, Hirabayashi T, Iwasaki R, Torii KU, Uchida N. EPFL peptide signalling ensures robust self-pollination success under cool temperature stress by aligning the length of the stamen and pistil. Plant Cell Environ. 2023;46(2):451–63. https://doi.org/10.1111/pce.14498.
Tian Y, Zhang T. MIXTAs and phytohormones orchestrate cotton fiber development. Curr Opin Plant Biol. 2021;59:101975. https://doi.org/10.1016/j.pbi.2020.10.007.
Sun Z, Li S, Li F, **ang J. Bioinformatic prediction of WSSV-host protein-protein interaction. Biomed Res Int. 2014, 416543. https://doi.org/10.1155/2014/416543.
Hao Y, Wang X, Li X, Bassa C, Mila I, Audran C, Maza E, Li Z, Bouzayen M, van der Rest B, Zouine M. Genome-wide identification, phylogenetic analysis, expression profiling, and protein-protein interaction properties of TOPLESS gene family members in tomato. J Exp Bot. 2014;65(4):1013–23. https://doi.org/10.1093/jxb/ert440.
Lei P, Wei X, Gao R, Huo F, Nie X, Tong W, Song W. Genome-wide identification of PYL gene family in wheat: Evolution, expression and 3D structure analysis. Genomics. 2021;113(2):854–66. https://doi.org/10.1016/j.ygeno.2020.12.017.
Guan H, Huang B, Chen M, Wang X, Song S, Liu H, Chen R, Hao Y. Genome-wide identification, phylogeny analysis, expression profiling, and determination of protein-protein interactions of the LEUNIG gene family members in tomato. Gene. 2018;679:1–10. https://doi.org/10.1016/j.gene.2018.08.075.
Funding
This research was funded by the National Natural Science Foundation of China (32272179 and 31801404), the Regional Innovation Guidance Project of **njiang Production and Construction Corps (2021BB012), the Program for Innovative Research Team (in Science and Technology) in University of Henan Province (20IRTSTHN021), the Postgraduate Education Reform and Quality Improvement Project of Henan Province (YJS2022JD47), the National Key R&D Program of China (2021YFE0101200), Zhongyuan Scholars Workstation (224400510020), and Technology development Project of Henan Province (242102110262).
Author information
Authors and Affiliations
Contributions
PTL, QWL, and YLY conceived and designed the experiments; PTL, ZLZ, and WKW performed the experiments; TW, NH, YYW, ZHS, YFL, and QKL analyzed the data; SHY, JWG, XHX, YLL, and YZS contributed reagents, materials, and analysis tools; PTL, QWL, and YLY wrote the paper; RHP, QWL, and YLY revised the paper; All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
12870_2024_5262_MOESM1_ESM.docx
Additional file 1: Table S1. Primer information of 15 GhEPF/EPFL and Ubiquitin7 for qRT-PCR experiments. Table S2. Identification and physiochemical characteristic analysis of cotton EPF/EPFL genes.
12870_2024_5262_MOESM2_ESM.docx
Additional file 2: Figure S1. The conserved peptides among the EPF/EPFL proteins. Figure S2. The information of 8 conserved motifs.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Li, P., Zhao, Z., Wang, W. et al. Genome-wide analyses of member identification, expression pattern, and protein–protein interaction of EPF/EPFL gene family in Gossypium. BMC Plant Biol 24, 554 (2024). https://doi.org/10.1186/s12870-024-05262-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-024-05262-7