Abstract
The domestication process in grapevines has facilitated the fixation of desired traits. Nowadays, vegetative propagation through cuttings enables easier preservation of these genotypes compared to sexual reproduction. Nonetheless, even with vegetative propagation, various phenotypes are often present within the same vineyard due to the accumulation of somatic mutations. These mutations are not the sole factors influencing phenotype. Alongside somatic variations, epigenetic variation has been proposed as a pivotal player in regulating phenotypic variability acquired during domestication. The emergence of these epialleles might have significantly influenced grapevine domestication over time. This study aims to investigate the impact of domestication on methylation patterns in cultivated grapevines. Reduced-representation bisulfite sequencing was conducted on 18 cultivated and wild accessions. Results revealed that cultivated grapevines exhibited higher methylation levels than their wild counterparts. Differential Methylation Analysis between wild and cultivated grapevines identified a total of 9955 differentially methylated cytosines, of which 78% were hypermethylated in cultivated grapevines. Functional analysis shows that core methylated genes (consistently methylated in both wild and cultivated accessions) are associated with stress response and terpenoid/isoprenoid metabolic processes. Meanwhile, genes with differential methylation are linked to protein targeting to the peroxisome, ethylene regulation, histone modifications, and defense response. Collectively, our results highlight the significant roles that epialleles may have played throughout the domestication history of grapevines.
Similar content being viewed by others
Background
Domestication syndrome is a phenomenon observed in crops. This results in a suite of traits that distinguish cultivated genotypes from their wild progenitors, including changes in morphology, physiology, and phenology that make them more amenable to cultivation. Thanks to the abundance of archaeobotanical, ecological and genetic information available for a handful of economically important seed propagated crops, the domestication syndrome has been well-documented in these species [1, 2]. However, less is known about the domestication trajectories of vegetatively propagated crops [2]. One of the main advantages of vegetative propagation is that it allows for the preservation of desirable traits from one generation to the next. This is because when a plant is propagated vegetatively, the offspring is genetically identical to the parent plant [2,3,4]. This means that desirable traits such as disease resistance, yield, and flavor can be maintained over many generations. This contrasts with sexual reproduction, where traits can be lost or diluted through the process of genetic recombination. The type of propagation used during domestication can result in diametrically opposed domestication syndromes. For example, while the use of vegetative propagation has been shown to negatively affect the capacity for sexual reproduction via the accumulation of mutations in genes associated to flower development, self-fertilization, and seed development, which lead to the production of self-fertilized fruits, flowering asynchrony, and lower seed viability [2, 5, 6]; crops domesticated by sexual reproduction, tend to present larger seeds, synchronic flowering and pollinator dependent fertilization [2].
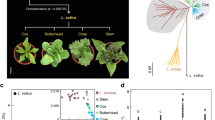
Vitis vinifera is a perennial woody liana belonging to the Vitaceae family. The species is divided into two different forms principally based on their reproductive system and whether it is a cultivated or a wild form. Wild grapevines (V. vinifera ssp. sylvestris), are commonly dioecious plants [7], and are naturally distributed across Asia and Europe. Cultivated grapevines (V. vinifera ssp. vinifera) mainly produce hermaphrodites flowers, and are broadly cultivated across the world, both for grape production to be consumed as a fruit, and for winemaking, grape juice or other derived products [8].
Although viticulture started at the Paleolithic age as a food source in Europe from wild accessions [9], there is evidence that the use of grapes by humans to produce wine started near to the seventh millennium BC [10]. This significantly influenced the domestication of grapevines by selecting varieties that produce a particular fruit quality and larger berries [7, 8]. It is believed that such selection occurred using vegetative propagation by cuttings to enhance the preservation of phenotypes of interest [7, 8, 11], which in turn had a negative effect on the crop tolerance to biotic and abiotic stresses. For example, populations of wild grapevines in North Africa and coastal regions of Northern Spain shown better adaptation to salt stress than cultivated grapevines [12, 13], while wild accessions from Germany, Iran and Georgia show higher resistance to mildew infections [14,15,16,17]. Moreover, despite the use of vegetative reproduction to maintain a desired genotype, the use of asexual reproduction in grapevine has resulted in novel phenotypes appearing within the same variety [5] and same vineyard [18]. Such phenotypic variants are frequently found in vegetatively propagated crops and often make up a significant portion of the cultivated varieties. Although a genetic basis is often presumed to be the reason for the noticeable differences in traits observed, epigenetic modifications have also been proposed to play an important role [19,20,36] to our data. Firstly, demultiplexing was performed in order to ensure the structure of the adapters to identify the samples [35], and a fastq-filter was performed using Stacks v2.55 [37]. The demultiplexed sequences from the triplicates from each accession were pooled to form a unique sample. Paired-end sequences were merged using PEAR v0.9.6 [38]. Alignment and methylation calling were performed with Bismark v0.23.0 [39] using the reference genome of Vitis vinifera L. PN40024 v4.1 [40]. Sequencing depth, coverage, and methylation differences between wild and cultivated accessions were visualized using ChromoMap R v1.0.0 [41].
Global differences in DNA methylation were visualized using hierarchical clustering and principal component analysis (PCA) performed using MethylKit R Package v1.16.1 [42] on the calculated percentage of methylation in all methylated cytosines present in at least four of the accessions. The percentage of total methylation was compared between cultivated and wild accessions in each context (CG, CHG and CHH (where H = A, T,C)) using T-test, after testing for normality in the data using Kolmogorov-Smirnoff Test, considering significant differences when p-value < 0.01. Finally, differentially methylated cytosines (DMCs) were identified using the methylKit R package v1.16.1 [42]. Cytosines were considered differentially methylated between wild and cultivated accessions when the observed difference in methylation was more than 25% and p-value < 0.01. To reduce the effect of genetic mutations on differential methylation data, for a genomic location to be included in the differential methylation analysis, such location must have a cytosine in a minimum of four samples per group and the location must have been sequenced to a minimum coverage of 10X. Additionally, a second more stringent filtering was implemented by identifying all genomic locations containing a SNP using the epiDiverse - SNP pipeline (available at “https://github.com/EpiDiverse/SNP”). Then, all epialleles located in genomic locations containing a SNP were removed from the analysis and hierarchical clustering was performed using all remaining epialleles.
To determine if DNA methylation patterns associated to the geographic origin of wild accessions were present, we performed a comparative analysis following the premises of De Andrés et al., (2012) [43]. For this, the methylation information gathered from wild accessions was filtered for epialleles associated to single nucleotide polymorphism as described above. Both the remaining epialleles and the SNPs identified using epiDiverse - SNP pipeline were used for hierarchical clustering analysis.
Protein coding genes presenting at least one methylated cytosine within 1000 bp of the transcription start site were deemed methylated. The annotated genome PN40024 v4.1 was used to determine the genic location (promoter, intro, exon) of methylated cytosines identified within genes. Then methylated genes were divided into 6 groups based on the type of methylation observed: 1. Core methylated genes, i.e., genes presenting unchanged methylated cytosines both in wild and cultivated accessions (CMCs); 2. Genes presenting CMCs and hypermethylated differentially methylated cytosines (DMCs) in cultivated compared to wild accessions; 3. genes presenting CMCs and hypomethylated DMCs in cultivated compared to wild accessions; 4. genes presenting CMCs and both hypomethylated and hypermethylated DMCs; 5. genes presenting hypermethylated DMCs in cultivated compared to wild accessions; and 6. genes presenting hypomethylated DMCs in cultivated compared to wild accessions. As above, DMCs associate with a SNP were removed from the analysis using the epiDiverse - SNP pipeline. Gene Ontology (GO) analysis was implemented with GOstats [44] and rrvgo [45] package in R, for each of these groups using all genes sequenced (i.e., presenting at least one read overlap** with a window of 1000 bp before and after the 5’ and 3’ UTRs respectively) as the gene universe. QuickGO Browser [46] (GO version 2023-09-20) was used to generate the ancestor charts for the main GO terms in each group.
Results
Differences in global levels of DNA methylation between wild and domesticated grapevine genotypes
EpiGBS2 libraries yielded a total of 44.5 million reads with an average of 2.5 million reads per sample (ranging from 1,106,659 to 8,249,031 reads). Bisulfite conversion efficiency showed on average 90% unmethylated cytosines converted to uracils. The mean percentage of mappable reads per sample after de-multiplexing was 49%, ranging from 37 to 60%. This resulted in an overall genome coverage of 1.5% (ranging between 0.7% and 2.6% (Supplementary Table S1)), with reads distributed evenly across the whole genome (See Fig. 1A for read distribution across chromosome 17 and Supplementary File 1 for read distribution across all chromosomes).
Methylation calling identified a total of 222,647 genomic locations containing methylated cytosines. The CG context presented the highest level of cytosine methylation, followed by CHG and CHH context (Fig. 1B). Cultivated varieties presented consistent significantly higher (T-test, p-value < 0.01) levels of DNA methylation than wild accessions in all sequence contexts (Fig. 1B). PCA plots built using the percentage of methylation for all sequenced cytosines as variables, show that wild and cultivated form two different clusters separated mainly by PC1 in all sequence contexts (Fig. 1C-E). Such observed separation between wild and cultivated accessions is particularly evident for the CHH context (Fig. 1E and Supplementary Fig. 3).
Analysis of differences in global levels of DNA methylation in cultivated andwild V. vinifera accessions. A Visualization of genomic and epigenomic information for chromosome 17 of Vitis vinifera using 100,000 bp windows. Vertical bars in panels (a) and (b) show the number of protein coding genes and transposable elements respectively per genomic window. Bars in panel (c) shows average sequencing depth per genomic window (Log 10 of calculated depth for sequenced bases). Panel (d) shows the average fold change in methylation in given window (blue and red bars indicate an average hypermethylated or hypomethylated window in cultivated vs. wild accessions. Panel containing chromosome number (i.e., chr17 here) shows average fold change in methylation in each window (hypomethylation (orange) hypermethylation (yellow). To visualize an interactive version of the figure containing all DMCs per window in all chromosomes see Supplementary File 1 (Follow instructions available in Supplementary File 2). Panels generated using ChromoMap R [41]. B Bars show the average percentage of methylation per sequence context (CG, CHG, CHH, and unknown) in cultivated (V. vinifera ssp. vinifera (n = 10); black bars), and wild type (V. vinifera ssp. sylvestris (n = 8); white bars) accessions. Error bars indicate the calculated Standard Deviation. ** T-test, p-value < 0.01. C-E Multivariate analysis of percentage of methylation for all individual cytosine sequenced in cultivated and wild V. vinifera accessions. Principal Component analysis plots show results for methylation analysis results in the CG (C), CHG (D), and CHH (E) contexts. Blue and red circles represent cultivated and wild accessions respectively. PCs 1 to 3 represent 53, 64, 69% of the total measured variability in CG, CHG, and CHH contexts respectively
Analysis of global methylation levels in wild and cultivated accessions at genomic feature level (i.e., intergenic and genic regions) showed that intergenic regions presented similar levels of DNA methylation to those observed genome-wide in all sequence contexts, with the exemption of CHGs, which showed higher levels of DNA methylation (Supplementary Fig. 1). Conversely, genic regions showed consistently lower levels of DNA methylation in all sequence contexts than those observed genome-wide (Supplementary Fig. 2). Finally, cultivated accessions presented significantly higher levels of DNA methylation (T-test, p-value < 0.01) than wild accessions in all sequence contexts and genomic features (Supplementary Figs. 1 and 2).
Identification of differentially methylated cytosines associated to domestication
Differential Methylation analysis identified a total of 9955 DMCs between wild and cultivated accessions evenly distributed across the genome (Fig. 1A and Supplementary File 1). Of those, 7793 DMCs were hypermethylated and 2162 DMCs were hypomethylated in cultivated vines compared to wild accessions. The majority of both hyper and hypomethylated DMCs were found in the CHH context (77 and 69% respectively) (Fig. 2A). From a gene feature context, DMCs were mainly found in intergenic regions (Fig. 2B). This is particularly evident in the CHH context, where 56 and 60% of hypermethylated and hypomethylated DMCs, respectively, were found in intergenic regions. The second most abundant genic feature presenting DMCs were introns, with percentages varying between 24 and 35% in hypermethylated DMCs, and 28 and 32% in hypomethylated DMCs, depending on the sequence context (Fig. 2B).
Identification of DMCs associated to grapevine’s domestication. Pie charts show (A) the total number and percentage of hypermethylated (top pie chart) and hypomethylated DMCs identified in cultivated vines compared to wild accessions in each sequence context (CG, CHG and CHH); and (B) the percentage of DMCs identified per genic feature and sequence context, in cultivated compared to wild type accessions
Effect of genetic differences to epigenetic differentiation between wild and cultivated accessions
The EpiDiverse-SNP pipeline identified 57,489 SNPs in the 222,711 genomic locations containing methylated cytosines. Of the remaining 165,189 genomic locations containing methylated cytosines, 5869 DMCs were hypermethylated and 1575 DMCs were hypomethylated in cultivated vines compared to wild accessions (i.e., 25% of the original DMCs were associated to a SNP). Hierarchical clustering analysis using all epialleles and only those not associated to SNPs showed no significant clustering differences (Supplementary Fig. 3).
Analysis of (epi)genetic signals of provenance in wild type accessions
We then compared wild accessions to determine if a genetic and or epigenetic signal associated to the location from where they were originally collected exist. Hierarchical cluster analysis showed no clear epigenetic signal irrespective of the use of all epialleles sequenced or after removing epialleles associated to a SNP (Fig. 3a). However, when only genetic information was used (i.e., clustering samples using the SNPs identified by the EpiDiverse-SNP pipeline, two separate clusters of wild accessions grouped by their provenance. One cluster contained all three accessions originally collected in the North of the Iberian Peninsula, in oceanic, continental and mountain climatic zones, while the second cluster contained all accession collected from the South of the Iberian Peninsula (Mediterranean climatic zone) (Fig. 3b) (see in Supplementary TableS1 for metadata associated to each accession).
Effect of region of origin on the methylome of Iberian Vitis vinifera ssp. sylvestris. Analysis of genetic (a) and epigenetic (b) differences among wild grapevine accessions originally collected from different regions of the Iberian Peninsula and grown in a common garden. Epigenetic analysis was performed using epialleles not associated to SNPs the epiDiverse-SNP pipeline to remove the effect of underlying genetic variation between wild grapevine populations. Samples highlighted in red, the represented branches correspond to wild accessions belonging to the South of Spain, and in blue, they correspond to wild accessions coming from the North of Spain, placed approximately to the map of the Spanish Climate Zones
Analysis of domestication associated DMCs within genic features
Collectively epiGBS2 results generated reads overlap** with a total of 7174 genes. Of those, a total of 2854 (40%) genes were identified as genes that contained at least one methylated cytosine (Supplementary Table S2A). Methylated cytosines were mainly found in introns (66–80%), followed by exons (15–20%), and promoters (4–14%) (Fig. 4) (Supplementary Table S2B). Genes containing methylated cytosines could be further divided into six groups, in order of abundance, (1) Genes presenting methylated cytosines both in wild and cultivated accessions (1883 genes) (core methylated genes (CMCs) hereafter); (2) genes presenting CMCs and hypermethylated DMCs in cultivated compared to wild accessions (564 genes); (3) genes presenting CMCs and hypomethylated DMCs in cultivated compared to wild accessions (252 genes); (4) genes presenting CMCs and both hypomethylated and hypermethylated DMCs (116 genes); (5) Genes presenting hypermethylated DMCs in cultivated compared to wild accessions (28 genes); and (6) genes presenting hypomethylated DMCs in cultivated compared to wild accessions (11 genes). Functional analysis of the genes identified within each group revealed that CMCs are significantly associated with the regulation of cellular response to stress and isoprenoid/terpenoid processes. Cultivated grapevines hypermethylated genes were associated mainly to processes associated to protein targeting to peroxisomes and histone lysine demethylation, while hypermethylated genes in wild grapevines related to ethylene regulation processes and response to ozone. The remaining group (i.e., genes both presenting hyper and hypomethylated cytosines between in both types of accessions) presented GO terms related to defense response (Fig. 4) (See Supplementary Table S2C for a complete list of GO terms in each group).
Schematic representation of methylated gene types in wild and cultivated grapevines. Boxes within gene models show the percentage of the total methylated cytosines in each gene group found in each genic context. Arrow heads color and size indicate the type of methylated cytosine found in each gene type (Core methylated cytosines (CMCs); Hypomethylated and hypermethylated cytosines in cultivated vs. wild grapevine accessions) and the abundance of that type of methylation within that gene type, respectively. Right panel shows the number of identified genes for each group and their correspondent most representative GO terms
Discussion
While significant strides have been made in understanding the genetic underpinnings of crop domestication, there is still a relative paucity of knowledge regarding the role of epigenetic mechanisms in this process. Epigenetics has emerged as a crucial regulator of various biological processes in both plants and animals. Recent studies have begun to hint at the potential involvement of epigenetic changes in the adaptation and phenotypic diversification of domesticated crops. However, a comprehensive understanding of how these epigenetic modifications may have been harnessed—or inadvertently altered—during the domestication process is still in its infancy. Studying the contribution of epigenetic mechanisms to domestication [47,48,49] will provide novel insights into the early stages of domestication and the selective pressures faced by ancestral agriculturists. At the same time, such studies will lay the foundation for the development of comprehensive models integrating plant adaptation to the environment through epigenetics mechanisms, facilitating their use for the development of novel cultivars more resilient to stress [33].
Epigenetic signal of domestication is independent of genetic variation
Previous studies have shown that DNA methylation variability in plants can be attributed to three main factors: genetic (sequence) differences, environmental induction, and stochasticity (see Konate et al., 2020 for a recent example) [50]. Moreover, ** the evolutionary and developmental trajectories of domesticated species, influencing in the crop’s plasticity and uniformity. Nevertheless, since this study only included hermaphrodite flower producing cultivated accessions, further studies including dioecious cultivated accessions are required to determine if the epigenetic differences identified here are really associated with domestication or to the sexual strategy of the studied plants. Additionally, future studies should analyze complete methylomes and focus on the consequences of methylation changes on gene expression to gain a comprehensive understanding of the role of DNA methylation in grapevine domestication.
Availability of data and materials
The datasets generated and analyzed during the current study are available in the European Nucleotide Archive (ENA), accession number PRJEB55284.
References
Meyer RS, DuVal AE, Jensen HR. Patterns and processes in crop domestication: an historical review and quantitative analysis of 203 global food crops. New Phytol. 2012;196(1):29–48.
Denham T, Barton H, Castillo C, Crowther A, Dotte-Sarout E, Florin SA, et al. The domestication syndrome in vegetatively propagated field crops. Ann Bot. 2020;125(4):581–97.
Ku YS, Contador CA, Ng MS, Yu J, Chung G, Lam HM. The effects of domestication on secondary metabolite composition in legumes. Front Genet [Internet]. 2020;11 https://www.frontiersin.org/articles/10.3389/fgene.2020.581357.
Raj S, Bräutigam K, Hamanishi ET, Wilkins O, Thomas BR, Schroeder W, et al. Clone history shapes Populus drought responses. Proc Natl Acad Sci USA. 2011;108(30):12521–6.
McKey D, Elias M, Pujol B, Duputié A. The evolutionary ecology of clonally propagated domesticated plants. New Phytol. 2010;186(2):318–32.
Tesfamicael KG, Gebre E, March TJ, Sznajder B, Mather DE, Rodríguez López CM. Accumulation of mutations in genes associated with sexual reproduction contributed to the domestication of a vegetatively propagated staple crop, enset. Hortic Res. 2020;7(1):185.
Arroyo-Garcia R, Ruíz-García L, Bolling L, Ocete R, López MA, Arnold C, et al. Multiple origins of cultivated grapevine (Vitis vinifera L. ssp. sativa) based on chloroplast DNA polymorphisms. Mol Ecol. 2006;15:3707–14.
Terral JF, Tabard E, Bouby L, Ivorra S, Pastor T, Figueiral I, et al. Evolution and history of grapevine (Vitis vinifera) under domestication: new morphometric perspectives to understand seed domestication syndrome and reveal origins of ancient European cultivars. Ann Bot. 2010;105(3):443–55.
Barker G. The origins and spread of agriculture and pastoralism in Eurasia. Antiquity. 1997;71(272):480–2.
McGovern PE, Glusker DL, Exner LJ, Voigt MM. Neolithic resinated wine. Nature. 1996;381(6582):480–1.
Levadoux L. Les populations sauvages et cultivées de Vitis vinifera L. Annales de l’Amélioration des Plantes. 1956;6:59–117.
Askri H, Daldoul S, Ammar AB, Rejeb S, Jardak R, Rejeb MN, et al. Short-term response of wild grapevines (Vitis vinifera L. ssp. Sylvestris) to NaCl salinity exposure: changes of some physiological and molecular characteristics. Acta Physiol Plant. 2012;34(3):957–68.
Carrasco D, Zhou-Tsang A, Rodriguez-Izquierdo A, Ocete R, Revilla MA, Arroyo-García R. Coastal Wild Grapevine Accession (Vitis vinifera L. ssp. Sylvestris) shows distinct late and early transcriptome changes under salt stress in comparison to Commercial Rootstock Richter 110. Plants. 2022;11(20):2688.
Bitsadze N, Aznarashvili M, Vercesi A, Chipashvili R, Failla O, Maghradze D. Screening of Georgian grapevine germplasm for susceptibility to downy mildew. Vitis. 2015;54:193–6.
Duan D, Halter D, Baltenweck R, Tisch C, Tröster V, Kortekamp A, et al. Genetic diversity of stilbene metabolism in Vitis Sylvestris. J Exp Bot. 2015;66(11):3243–57.
Riaz S, Boursiquot JM, Dangl GS, Lacombe T, Laucou V, Tenscher AC, et al. Identification of mildew resistance in wild and cultivated central Asian grape germplasm. BMC Plant Biol. 2013;13(1):149.
Toffolatti SL, Maddalena G, Salomoni D, Maghradze D, Bianco PA, Failla O. Evidence of resistance to the downy mildew agent Plasmopara Viticola in the Georgian Vitis vinifera germplasm. Vitis. 2016;55(3):121–8.
This P, Lacombe T, Thomas M. Historical origins and genetic diversity of wine grapes. Trends Genet. 2006;22(9):511–9.
Rodríguez López CM, Wilkinson MJ. Epi-fingerprinting and epi-interventions for improved crop production and food quality. Front Plant Sci [Internet]. 2015 Jun 5 [cited 2021 Mar 16];6. http://journal.frontiersin.org/Article/https://doi.org/10.3389/fpls.2015.00397/abstract.
Fortes AM, Gallusci P. Plant Stress Responses and Phenotypic Plasticity in the Epigenomics Era: Perspectives on the Grapevine Scenario, a Model for Perennial Crop Plants. Front Plant Sci [Internet]. 2017. http://journal.frontiersin.org/article/10.3389/fpls.2017.00082/full.
**e H, Konate M, Sai N, Tesfamicael KG, Cavagnaro T, Gilliham M, et al. Global DNA methylation patterns can play a role in defining Terroir in Grapevine (Vitis vinifera Cv. Shiraz). Front Plant Sci. 2017;8:1860.
Noshay JM, Springer NM. Stories that can’t be told by SNPs; DNA methylation variation in plant populations. Curr Opin Plant Biol. 2021;61:101989.
Wu Ct, Morris JR. Genes, Genetics, and epigenetics: a correspondence. Science. 2001;293(5532):1103–5.
Kumar S, Mohapatra T. Dynamics of DNA methylation and its functions in Plant Growth and Development. Front Plant Sci. 2021;12:596236.
Yao N, Zhang Z, Yu L, Hazarika R, Yu C, Jang H, et al. An evolutionary epigenetic clock in plants. Science. 2023;381(6665):1440–5.
Jackson SA. Epigenomics: dissecting hybridization and polyploidization. Genome Biol. 2017;18(1):117.
Yuan J, Jiao W, Liu Y, Ye W, Wang X, Liu B, et al. Dynamic and reversible DNA methylation changes induced by genome separation and merger of polyploid wheat. BMC Biol. 2020;18(1):171.
Song Q, Zhang T, Stelly DM, Chen ZJ. Epigenomic and functional analyses reveal roles of epialleles in the loss of photoperiod sensitivity during domestication of allotetraploid cottons. Genome Biol. 2017;18(1):99.
Cao S, Chen K, Lu K, Chen S, Zhang X, Shen C, et al. Asymmetric variation in DNA methylation during domestication and de-domestication of rice. Plant Cell. 2023;35(9):3429–43.
Guo H, Cao P, Wang C, Lai J, Deng Y, Li C, et al. Population analysis reveals the roles of DNA methylation in tomato domestication and metabolic diversity. Sci China Life Sci. 2023;66(8):1888–902.
Shen Y, Zhang J, Liu Y, Liu S, Liu Z, Duan Z, et al. DNA methylation footprints during soybean domestication and improvement. Genome Biol. 2018;19(1):128.
Xu G, Lyu J, Li Q, Liu H, Wang D, Zhang M, et al. Evolutionary and functional genomics of DNA methylation in maize domestication and improvement. Nat Commun. 2020;11(1):5539.
Tan JW, Rodríguez López CM. Epigenomics: a new tool for the generation of climate resilient grapevines. Front Hortic. 2023;2:1116866.
Gawehns F, Postuma M, van Antro M, Nunn A, Sepers B, Fatma S, et al. epiGBS2: improvements and evaluation of highly multiplexed, epiGBS-based reduced representation bisulfite sequencing. Mol Ecol Resour. 2022;22(5):2087–104.
van Gurp TP, Wagemaker NCAM, Wouters B, Vergeer P, Ouborg JNJ, Verhoeven KJF. epiGBS: reference-free reduced representation bisulfite sequencing. Nat Methods. 2016;13(4):322–4.
Werner O, Prudencio ÁS, de la Cruz-Martínez E, Nieto-Lugilde M, Martínez-Gómez P, Ros RM. A cost reduced variant of epi-genoty** by sequencing for studying DNA methylation in non-model organisms. Front Plant Sci. 2020;11:694.
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA. Stacks: an analysis tool set for population genomics. Mol Ecol. 2013;22(11):3124–40.
Zhang J, Kobert K, Flouri T, Stamatakis A. PEAR: a fast and accurate Illumina paired-end reAd mergeR. Bioinformatics. 2014;30(5):614–20.
Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27(11):1571–2.
Velt A, Frommer B, Blanc S, Holtgräwe D, Duchêne É, Dumas V, et al. An improved reference of the grapevine genome reasserts the origin of the PN40024 highly homozygous genotype. Whiteman N, editor. G3 Genes|Genomes|Genetics. 2023;13(5):jkad067
Anand L, Rodriguez Lopez CM. ChromoMap: an R package for interactive visualization of multi-omics data and annotation of chromosomes. BMC Bioinform. 2022;23(1):33.
Akalin A, Kormaksson M, Li S, Garrett-Bakelman FE, Figueroa ME, Melnick A, et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 2012;13(10):R87.
De Andrés MT, Benito A, Pérez-Rivera G, Ocete R, Lopez MA, Gaforio L, et al. Genetic diversity of wild grapevine populations in Spain and their genetic relationships with cultivated grapevines: GENETIC DIVERSITY OF WILD GRAPEVINE POPULATIONS FROM SPAIN. Mol Ecol. 2012;21(4):800–16.
Falcon S, Gentleman R. Using GOstats to test gene lists for GO term association. Bioinformatics. 2007;23(2):257–8.
Sayols S. rrvgo: a Bioconductor package to reduce and visualize Gene Ontology terms [Internet]. 2020. https://ssayols.github.io/rrvgo.
Binns D, Dimmer E, Huntley R, Barrell D, O’Donovan C, Apweiler R. QuickGO: a web-based tool for gene ontology searching. Bioinformatics. 2009;25(22):3045–6.
Consuegra S, Rodríguez López CM. Epigenetic-induced alterations in sex‐ratios in response to climate change: an epigenetic trap? BioEssays. 2016;38(10):950–8.
Bräutigam K, Cronk Q. DNA methylation and the evolution of developmental complexity in plants. Front Plant Sci. 2018;9:1447.
Róis AS, Rodríguez López CM, Cortinhas A, Erben M, Espírito-Santo D, Wilkinson MJ, et al. Epigenetic rather than genetic factors may explain phenotypic divergence between coastal populations of diploid and tetraploid Limonium spp. (Plumbaginaceae) in Portugal. BMC Plant Biol. 2013;13(1):205.
Konate M, Wilkinson M, Mayne B, Pederson S, Scott E, Berger B, et al. Salt stress induces Non-CG methylation in coding regions of Barley Seedlings (Hordeum vulgare). Epigenomes. 2018;2(2):12.
Angers B, Perez M, Menicucci T, Leung C. Sources of epigenetic variation and their applications in natural populations. Evol Appl. 2020;13(6):1262–78.
Arroyo-García R, Lefort F, de Andrés MT, Ibáñez J, Borrego J, Jouve N, et al. Chloroplast microsatellite polymorphisms in Vitis species. Genome. 2002;45(6):1142–9.
Daldoul S, Boubakri H, Gargouri M, Mliki A. Recent advances in biotechnological studies on wild grapevines as valuable resistance sources for smart viticulture. Mol Biol Rep. 2020;47(4):3141–53.
Guarino F, Cicatelli A, Brundu G, Heinze B, Castiglione S. Epigenetic diversity of clonal white poplar (Populus alba L.) populations: could methylation support the success of vegetative reproduction strategy? Labra M. Editor PLoS ONE. 2015;10(7):e0131480.
Vondras AM, Minio A, Blanco-Ulate B, Figueroa-Balderas R, Penn MA, Zhou Y, et al. The genomic diversification of grapevine clones. BMC Genom. 2019;20(1):972.
Chekanova JA. Long non-coding RNAs and their functions in plants. Curr Opin Plant Biol. 2015;27:207–16.
Zheng XM, Chen J, Pang HB, Liu S, Gao Q, Wang JR, et al. Genome-wide analyses reveal the role of noncoding variation in complex traits during rice domestication. Sci Adv. 2019;5(12):eaax3619.
Pinky, Jain R, Yadav A, Sharma R, Dhaka N. Emerging roles of long non-coding RNAs in regulating agriculturally important seed traits. Plant Physiol Biochem. 2023;203:108019.
He H, Zhou YF, Yang YW, Zhang Z, Lei MQ, Feng YZ, et al. Genome-wide analysis identified a set of conserved lncRNAs Associated with Domestication-related traits in Rice. IJMS. 2021;22(9):4742.
Wang G, Li H, Meng S, Yang J, Ye N, Zhang J. Analysis of global methylome and gene expression during Carbon Reserve Mobilization in Stems under Soil Drying. Plant Physiol. 2020;183(4):1809–24.
Zhang H, Lang Z, Zhu JK. Dynamics and function of DNA methylation in plants. Nat Rev Mol Cell Biol. 2018;19(8):489–506.
Wang X, Hu L, Wang X, Li N, Xu C, Gong L, et al. DNA methylation affects gene alternative splicing in plants: an Example from Rice. Mol Plant. 2016;9(2):305–7.
Maillot P, Velt A, Rustenholz C, Butterlin G, Merdinoglu D, Duchêne E. Alternative splicing regulation appears to play a crucial role in grape berry development and is also potentially involved in adaptation responses to the environment. BMC Plant Biol. 2021;21(1):487.
Zhou E, Wang G, Weng L, Li M, **ao H. Comparative analysis of environment-responsive alternative splicing in the inflorescences of cultivated and wild tomato species. IJMS. 2022;23(19):11585.
Wang X, Chen X, Luo S, Ma W, Li N, Zhang W, et al. Discovery of a DFR gene that controls anthocyanin accumulation in the spiny Solanum group: roles of a natural promoter variant and alternative splicing. Plant J. 2022;111(4):1096–109.
Li M, Wang R, Song B, Cao B, Sun M, Liu Y, et al. Alternative splicing analysis provides insight into the domestication and improvement of pear (Pyrus pyrifolia) fruit. Sci Hort. 2022;297:110976.
Yu K, Feng M, Yang G, Sun L, Qin Z, et al. Changes in alternative splicing in response to domestication and polyploidization in wheat. Plant Physiol. 2020;184(4):1955–68.
Sims RJ, Nishioka K, Reinberg D. Histone lysine methylation: a signature for chromatin function. Trends Genet. 2003;19(11):629–39.
Chen H, Bullock DA, Alonso JM, Stepanova AN. To fight or to grow: the balancing role of ethylene in plant abiotic stress responses. Plants. 2021;11(1):33.
Shi Z, Halaly-Basha T, Zheng C, Weissberg M, Ophir R, Galbraith DW, et al. Transient induction of a subset of ethylene biosynthesis genes is potentially involved in regulation of grapevine bud dormancy release. Plant Mol Biol. 2018;98(6):507–23.
Shi Z, Halaly-Basha T, Zheng C, Sharabi-Schwager M, Wang C, Galbraith DW, et al. Identification of potential post-ethylene events in the signaling cascade induced by stimuli of bud dormancy release in grapevine. Plant J. 2020;104(5):1251–68.
Böttcher C, Burbidge CA, Boss PK, Davies C. Interactions between ethylene and auxin are crucial to the control of grape (Vitis vinifera L.) berry ripening. BMC Plant Biol. 2013;13(1):222.
López ME, Roquis D, Becker C, Denoyes B, Bucher E. DNA methylation dynamics during stress response in woodland strawberry (Fragaria vesca). Hortic Res. 2022;9:uhac174.
Pan W, Liang J, Sui J, Li J, Liu C, **n Y, et al. ABA and Bud Dormancy in perennials: current knowledge and future perspective. Genes. 2021;12(10):1635.
Acknowledgements
Special recognition to Dr. Rosa Adela Arroyo García, one of the principal researchers of the project, that tragically deceased by a long illness. We miss you, Rosa.
Funding
This study was supported by the National Institute of Food and Agriculture, AFRI Competitive Grant Program Accession number 1018617, the National Institute of Food and Agriculture, United States Department of Agriculture, Hatch Program accession number 1020852, COST Action CA 17111 INTEGRAPE, supported by COST (European Cooperation in Science and Technology) and the Research and Science Ministry of Spain (project RTI2018-094470-R-C21).
Author information
Authors and Affiliations
Contributions
ARI, RAG, and CMRL conceived the study. ARI, DC, RAG, and CMRL developed the experimental design for the study. RAG provided the plant accessions and metadata. ARI and RM performed the laboratory work. ARI, DC, and PC, performed the data analysis. ARI, DC, and LA performed data visualization. ARI and CMRL wrote the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Rodriguez-Izquierdo, A., Carrasco, D., Anand, L. et al. Epigenetic differences between wild and cultivated grapevines highlight the contribution of DNA methylation during crop domestication. BMC Plant Biol 24, 504 (2024). https://doi.org/10.1186/s12870-024-05197-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-024-05197-z