Abstract
Background
Quorum sensing (QS) is a cell density-based intercellular communication system that controls virulence gene expression and biofilm formation. In Pseudomonas aeruginosa (P. aeruginosa), the LasR system sits at the top of the QS hierarchy and coordinates the expression of a series of important traits. However, the role of lasR in phage infection remains unclear. This study aims to investigate the role of lasR QS in phage infection.
Methods
The P. aeruginosa phage was isolated from sewage, and its biological characteristics and whole genome were analyzed. The adsorption receptor was identified via a phage adsorption assay. Following lasR gene knockout, the adsorption rate and bactericidal activity of phage were analyzed. Finally, real-time quantitative polymerase chain reaction (RT-qPCR) was conducted to explore how lasR promoting phage infection.
Results
The lytic phage vB_Pae_PLY was isolated and lipopolysaccharide (LPS) was identified as its adsorption receptor. The adsorption rate and bactericidal activity of vB_Pae_PLY were reduced after lasR knockout. RT-qPCR results showed that the expression of galU, a key gene involved in LPS synthesis, was down-regulated, and several genes related to type IV pili (T4P) were also down-regulated in the lasR mutant PaΔlasR.
Conclusions
The study showed that QS lasR may promote phage vB_Pae_PLY infection by involving in the synthesis of LPS and T4P. This study provides an example of QS in promoting phage infection and deepens the understanding of phage-bacteria interactions.
Similar content being viewed by others
Background
Pseudomonas aeruginosa (P. aeruginosa) is an opportunistic pathogen that uses quorum sensing (QS) signaling molecules to regulate the expression of virulence genes and biofilm [1]. P. aeruginosa has a particularly complex QS signaling network, which primarily consists of four interconnected systems (LasI/R, RhlI/R, PQS, and IQS) [2]. In the Las and Rhl systems, the lasI and rhll genes are responsible for the synthesis of N -(3-oxo-dodecanoyl)- L -homoserine lactone (3- oxo -C12-HSL) and N -(butanoyl)- L -homoserine lactone (C4-HSL), respectively. Then, 3- oxo -C12-HSL and C4-HSL interact with LasR, respectively, to form LasR- 3- oxo -C12-HSL and RhlR- C4-HSL complex to regulate target genes [3]. The PQS system enables the biosynthesis of QS signal molecules 2-heptyl-4-hydroxyquinoline (HHQ) and 2-heptyl-3-hydroxy-4(1 H)-quinolone (PQS, Pseudomonas quinolone signal) [4]. IQS is the fourth and recently discovered QS system, which synthesizes the signaling molecule 2-(2-hydroxyphenyl)-thiazole-4-carbaldehyde (IQS) [1]. Among all the QS systems, the Las tops the signal network. The LasR- 3- oxo -C12-HSL complex positively regulates the expression of receptor and synthase genes in the downstream QS system, thus establishing a regulatory feedforward loop [5]. QS directly or indirectly regulates more than 10% of its genome and about 20% of its bacterial proteome [6, 7], and controls the expression of a series of important traits in P. aeruginosa [8, 9].
Phages are viruses that specifically infect bacteria and are abundant in environments associated with their host bacteria. Phages bind to the surfaces of bacterial cells by recognizing adsorption receptors [50].
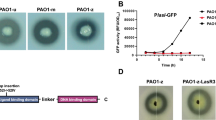
RT-qPCR analysis in the present study revealed that galU and T4P-related genes (pilA, pilB, pilC and pilD, pilQ and pilV, pilW and pilY) were significantly decreased in the PaΔlasR (Fig. 8). LPS is composed of O antigen, core oligosaccharide, and lipid A. galU encodes UDP-glucose pyrophosphorylase (GalU), and was found to be involved in the synthesis of the core region of P. aeruginosa LPS [43, 51]. The current study demonstrates that the phage vB_Pae_PLY utilizes surface LPS as the receptor for adsorption (Fig. 6). Thus, the reduced phage sensitivity in PaΔlasR can be largely attributed to the impaired LPS synthesis. In addition, pili also play an important role in phage infection; therefore, downregulation of T4P may also be another reason for reduced phage sensitivity. LPS and T4P are important virulence factors for the opportunistic pathogen P. aeruginosa. On the other hand, serving as phage receptors, LPS and T4P facilitate phage infection with their synthesis being positively regulated by QS. Therefore, lasR QS inhibition will affect the synthesis of bacterial LPS and T4P, resulting in reduced phage adsorption with LPS and T4P as receptors, and weaken the bactericidal effect of phages to a certain extent.
Conclusions
In conclusion, this study demonstrates that lasR may promote phage infection by positively regulating the biosynthesis of LPS and T4P. Disrupting lasR expression leads to decreased phage sensitivity. Given the complex and multifaceted role of QS in host-phage interactions, the future research is needed to uncover the various mechanisms by which QS participates in phage infection.
Data availability
The complete genome sequence of phage vB_Pae_PLY presented in this study is openly available in [GenBank, OR689712].
Abbreviations
- Pseudomonas aeruginosa :
-
P. aeruginosa
- QS:
-
Quorum sensing
- RT-qPCR:
-
Real-Time quantitative polymerase chain reaction
- LPS:
-
Lipopolysaccharide
- T4P:
-
Type IV pili-related
- AHL:
-
N-acylhomoserine lactones
- 3-oxo-C12-HSL:
-
N -(3-oxo-dodecanoyl)- L -homoserine lactone
- C4-HSL:
-
N -(butanoyl)- L -homoserine lactone
- HHQ:
-
2-heptyl-4-hydroxyquinoline
- PQS:
-
2-heptyl-3-hydroxy-4(1 H)-quinolone
- IQS:
-
2-(2-hydroxyphenyl)-thiazole-4-carbaldehyde
- LB:
-
Luria-Bertani
- Car:
-
Carbenicillin
- Gen:
-
Gentamicin
- PFU:
-
Plaque-forming unit
- CFU:
-
Colony-forming unit
- VFDB:
-
Virulence Factor Database
- CARD:
-
Comprehensive Antibiotic Resistance Database
- PCR:
-
Polymerase chain reaction
- PBS:
-
Phosphate-buffered saline
- MOI:
-
Multiplicity of infection
- TEM:
-
Transmission electron microscopy
- CDS:
-
Coding DNA sequences
- ICTV:
-
International Committee on Taxonomy of Viruses
- OmpK:
-
Outer membrane protein K
- GalU:
-
UDP-glucose pyrophosphorylase
References
Wang J, Wang C, Yu HB, Dela Ahator S, Wu X, Lv S, et al. Bacterial quorum-sensing signal IQS induces host cell apoptosis by targeting POT1-p53 signalling pathway. Cell Microbiol. 2019;21(10):e13076. https://doi.org/10.1111/cmi.13076.
Montagut EJ, Marco MP. Biological and clinical significance of quorum sensing alkylquinolones: current analytical and bioanalytical methods for their quantification. Anal Bioanal Chem. 2021;413(18):4599–618. https://doi.org/10.1007/s00216-021-03356-x.
Rather MA, Saha D, Bhuyan S, Jha AN, Mandal M. Quorum quenching: a Drug Discovery Approach against Pseudomonas aeruginosa. Microbiol Res. 2022;264:127173. https://doi.org/10.1016/j.micres.2022.127173.
Giallonardi G, Letizia M, Mellini M, Frangipani E, Halliday N, Heeb S, et al. Alkyl-Quinolone-dependent quorum sensing controls prophage-mediated autolysis in Pseudomonas aeruginosa colony biofilms. Front Cell Infect Microbiol. 2023;13:1183681. https://doi.org/10.3389/fcimb.2023.1183681.
Lee J, Zhang L. The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell. 2015;6(1):26–41. https://doi.org/10.1007/s13238-014-0100-x.
Chadha J, Harjai K, Chhibber S. Revisiting the virulence hallmarks of Pseudomonas aeruginosa: a chronicle through the perspective of quorum sensing. Environ Microbiol. 2022;24(6):2630–56. https://doi.org/10.1111/1462-2920.15784.
Schroven K, Putzeys L, Swinnen AL, Hendrix H, Paeshuyse J, Lavigne R. The phage-encoded protein PIT2 impacts Pseudomonas aeruginosa quorum sensing by direct interaction with LasR. iScience. 2023;26(10):107745. https://doi.org/10.1016/j.isci.2023.107745.
Wang Y, Dai J, Wang X, Wang Y, Tang F. Mechanisms of interactions between bacteria and bacteriophage mediate by quorum sensing systems. Appl Microbiol Biotechnol. 2022;106(7):2299–310. https://doi.org/10.1007/s00253-022-11866-6.
Rutherford ST, Bassler BL. Bacterial quorum sensing: its role in virulence and possibilities for its control. Cold Spring Harb Perspect Med. 2012;2(11). https://doi.org/10.1101/cshperspect.a012427.
Li P, Lin H, Mi Z, **ng S, Tong Y, Wang J. Screening of Polyvalent phage-resistant Escherichia coli strains based on phage receptor analysis. Front Microbiol. 2019;10:850. https://doi.org/10.3389/fmicb.2019.00850.
Labrie SJ, Samson JE, Moineau S. Bacteriophage resistance mechanisms. Nat Rev Microbiol. 2010;8(5):317–27. https://doi.org/10.1038/nrmicro2315.
Ahator SD, Sagar S, Zhu M, Wang J, Zhang LH. Nutrient availability and phage exposure alter the Quorum-Sensing and CRISPR-Cas-Controlled Population dynamics of Pseudomonas aeruginosa. mSystems. 2022;7(4):e0009222. https://doi.org/10.1128/msystems.00092-22.
De Smet J, Hendrix H, Blasdel BG, Danis-Wlodarczyk K, Lavigne R. Pseudomonas predators: understanding and exploiting phage-host interactions. Nat Rev Microbiol. 2017;15(9):517–30. https://doi.org/10.1038/nrmicro.2017.61.
Høyland-Kroghsbo NM, Paczkowski J, Mukherjee S, Broniewski J, Westra E, Bondy-Denomy J, et al. Quorum sensing controls the Pseudomonas aeruginosa CRISPR-Cas adaptive immune system. Proc Natl Acad Sci U S A. 2017;114(1):131–5. https://doi.org/10.1073/pnas.1617415113.
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315(5819):1709–12. https://doi.org/10.1126/science.1138140.
Høyland-Kroghsbo NM, Maerkedahl RB, Svenningsen SL. A quorum-sensing-induced bacteriophage defense mechanism. mBio. 2013;4(1):e00362–12. https://doi.org/10.1128/mBio.00362-12.
Tan D, Svenningsen SL, Middelboe M. Quorum sensing determines the choice of Antiphage Defense Strategy in Vibrio anguillarum. mBio. 2015;6(3):e00627. https://doi.org/10.1128/mBio.00627-15.
Xuan G, Lin H, Tan L, Zhao G, Wang J. Quorum sensing promotes phage infection in Pseudomonas aeruginosa PAO1. mBio. 2022;13(1):e0317421. https://doi.org/10.1128/mbio.03174-21.
Liu Y, Zhao Y, Qian C, Huang Z, Feng L, Chen L, et al. Study of combined effect of bacteriophage vB3530 and chlorhexidine on the inactivation of Pseudomonas aeruginosa. BMC Microbiol. 2023;23(1):256. https://doi.org/10.1186/s12866-023-02976-w.
Fang Q, Feng Y, McNally A, Zong Z. Characterization of phage resistance and phages capable of intestinal decolonization of carbapenem-resistant Klebsiella pneumoniae in mice. Commun Biol. 2022;5(1):48. https://doi.org/10.1038/s42003-022-03001-y.
Xuan G, Kong J, Wang Y, Lin H, Wang J. Characterization of the newly isolated Pseudomonas phage vB_Pae_LC3I3. Virus Res. 2023;323:198978. https://doi.org/10.1016/j.virusres.2022.198978.
Grant JR, Enns E, Marinier E, Mandal A, Herman EK, Chen CY, et al. Proksee: in-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 2023;51(W1):W484–92. https://doi.org/10.1093/nar/gkad326.
Li E, Yin Z, Ma Y, Li H, Lin W, Wei X, et al. Identification and molecular characterization of bacteriophage phiAxp-2 of Achromobacter xylosoxidans. Sci Rep. 2016;6:34300. https://doi.org/10.1038/srep34300.
Kiljunen S, Datta N, Dentovskaya SV, Anisimov AP, Knirel YA, Bengoechea JA, et al. Identification of the lipopolysaccharide core of Yersinia pestis and Yersinia pseudotuberculosis as the receptor for bacteriophage φA1122. J Bacteriol. 2011;193(18):4963–72. https://doi.org/10.1128/jb.00339-11.
Zhang Y, Wang L, Chen L, Zhu P, Huang N, Chen T, et al. Novel insight of transcription factor PtrA on pathogenicity and Carbapenems Resistance in Pseudomonas aeruginosa. Infect Drug Resist. 2022;15:4213–27. https://doi.org/10.2147/idr.S371597.
Jeon J, D’Souza R, Pinto N, Ryu CM, Park J, Yong D, et al. Characterization and complete genome sequence analysis of two myoviral bacteriophages infecting clinical carbapenem-resistant Acinetobacter baumannii isolates. J Appl Microbiol. 2016;121(1):68–77. https://doi.org/10.1111/jam.13134.
Li J, Yan B, He B, Li L, Zhou X, Wu N, et al. Development of phage resistance in multidrug-resistant Klebsiella pneumoniae is associated with reduced virulence: a case report of a personalised phage therapy. Clin Microbiol Infect. 2023. https://doi.org/10.1016/j.cmi.2023.08.022.
Liu Y, Wang Y, Kong J, Jiang X, Han Y, Feng L, et al. An effective antimicrobial strategy of colistin combined with the Chinese herbal medicine shikonin against colistin-resistant Escherichia coli. Microbiol Spectr. 2023;e0145923. https://doi.org/10.1128/spectrum.01459-23.
Adriaenssens EM, Sullivan MB, Knezevic P, van Zyl LJ, Sarkar BL, Dutilh BE, et al. Taxonomy of prokaryotic viruses: 2018–2019 update from the ICTV bacterial and archaeal viruses Subcommittee. Arch Virol. 2020;165(5):1253–60. https://doi.org/10.1007/s00705-020-04577-8.
Hoque MM, Naser IB, Bari SM, Zhu J, Mekalanos JJ, Faruque SM. Quorum Regulated Resistance of Vibrio cholerae against Environmental bacteriophages. Sci Rep. 2016;6:37956. https://doi.org/10.1038/srep37956.
Ding Y, Zhang D, Zhao X, Tan W, Zheng X, Zhang Q, et al. Autoinducer-2-mediated quorum-sensing system resists T4 phage infection in Escherichia coli. J Basic Microbiol. 2021;61(12):1113–23. https://doi.org/10.1002/jobm.202100344.
Xuan G, Tan L, Yang Y, Kong J, Lin H, Wang J. Quorum sensing autoinducers AHLs protect Shewanella baltica against phage infection. Int J Food Microbiol. 2023;403:110304. https://doi.org/10.1016/j.ijfoodmicro.2023.110304.
Broniewski JM, Chisnall MAW, Høyland-Kroghsbo NM, Buckling A, Westra ER. The effect of Quorum sensing inhibitors on the evolution of CRISPR-based phage immunity in Pseudomonas aeruginosa. Isme j. 2021;15(8):2465–73. https://doi.org/10.1038/s41396-021-00946-6.
Nobrega FL, Vlot M, de Jonge PA, Dreesens LL, Beaumont HJE, Lavigne R, et al. Targeting mechanisms of tailed bacteriophages. Nat Rev Microbiol. 2018;16(12):760–73. https://doi.org/10.1038/s41579-018-0070-8.
Dunne M, Hupfeld M, Klumpp J, Loessner MJ. Molecular basis of bacterial host interactions by Gram-positive targeting bacteriophages. Viruses. 2018;10(8). https://doi.org/10.3390/v10080397.
Donlan RM. Preventing biofilms of clinically relevant organisms using bacteriophage. Trends Microbiol. 2009;17(2):66–72. https://doi.org/10.1016/j.tim.2008.11.002.
Dowah ASA, Clokie MRJ. Review of the nature, diversity and structure of bacteriophage receptor binding proteins that target Gram-positive bacteria. Biophys Rev. 2018;10(2):535–42. https://doi.org/10.1007/s12551-017-0382-3.
Bertozzi Silva J, Storms Z, Sauvageau D. Host receptors for bacteriophage adsorption. FEMS Microbiol Lett. 2016;363(4). https://doi.org/10.1093/femsle/fnw002.
Harvey H, Bondy-Denomy J, Marquis H, Sztanko KM, Davidson AR, Burrows LL. Pseudomonas aeruginosa defends against phages through type IV pilus glycosylation. Nat Microbiol. 2018;3(1):47–52. https://doi.org/10.1038/s41564-017-0061-y.
McCutcheon JG, Peters DL, Dennis JJ. Identification and characterization of type IV Pili as the Cellular receptor of broad host Range Stenotrophomonas maltophilia bacteriophages DLP1 and DLP2. Viruses. 2018;10(6). https://doi.org/10.3390/v10060338.
Xuan G, Lin H, Wang J. Expression of a phage-encoded Gp21 protein protects Pseudomonas aeruginosa against phage infection. J Virol. 2022;96(5):e0176921. https://doi.org/10.1128/jvi.01769-21.
Testa S, Berger S, Piccardi P, Oechslin F, Resch G, Mitri S. Spatial structure affects phage efficacy in infecting dual-strain biofilms of Pseudomonas aeruginosa. Commun Biol. 2019;2:405. https://doi.org/10.1038/s42003-019-0633-x.
Dean CR, Goldberg JB. Pseudomonas aeruginosa galU is required for a complete lipopolysaccharide core and repairs a secondary mutation in a PA103 (serogroup O11) wbpM mutant. FEMS Microbiol Lett. 2002;210(2):277–83. https://doi.org/10.1111/j.1574-6968.2002.tb11193.x.
Li L, Pan X, Cui X, Sun Q, Yang X, Yang H. Characterization of Pseudomonas aeruginosa phage K5 genome and identification of its receptor related genes. J Basic Microbiol. 2016;56(12):1344–53. https://doi.org/10.1002/jobm.201600116.
Oechslin F, Piccardi P, Mancini S, Gabard J, Moreillon P, Entenza JM, et al. Synergistic Interaction between Phage Therapy and Antibiotics clears Pseudomonas Aeruginosa infection in Endocarditis and reduces virulence. J Infect Dis. 2017;215(5):703–12. https://doi.org/10.1093/infdis/jiw632.
Pires DP, Dötsch A, Anderson EM, Hao Y, Khursigara CM, Lam JS, et al. A Genotypic Analysis of Five P. aeruginosa strains after Biofilm infection by phages targeting different cell surface receptors. Front Microbiol. 2017;8:1229. https://doi.org/10.3389/fmicb.2017.01229.
Li N, Zeng Y, Wang M, Bao R, Chen Y, Li X, et al. Characterization of Phage Resistance and their impacts on bacterial fitness in Pseudomonas aeruginosa. Microbiol Spectr. 2022;10(5):e0207222. https://doi.org/10.1128/spectrum.02072-22.
Piepenbrink KH, Sundberg EJ. Motility and adhesion through type IV pili in Gram-positive bacteria. Biochem Soc Trans. 2016;44(6):1659–66. https://doi.org/10.1042/bst20160221.
Craig L, Forest KT, Maier B, Type. IV pili: dynamics, biophysics and functional consequences. Nat Rev Microbiol. 2019;17(7):429–40. https://doi.org/10.1038/s41579-019-0195-4.
Xuan G, Dou Q, Kong J, Lin H, Wang J. Pseudomonas aeruginosa resists phage infection via eavesdrop** on Indole Signaling. Microbiol Spectr. 2023;11(1):e0391122. https://doi.org/10.1128/spectrum.03911-22.
Jiang SS, Lin TY, Wang WB, Liu MC, Hsueh PR, Liaw SJ. Characterization of UDP-glucose dehydrogenase and UDP-glucose pyrophosphorylase mutants of Proteus mirabilis: defectiveness in polymyxin B resistance, swarming, and virulence. Antimicrob Agents Chemother. 2010;54(5):2000–9. https://doi.org/10.1128/aac.01384-09.
Funding
This work was supported by the Health Department of Zhejiang Province of the People’s Republic of China, No. 2022KY895; the research grants from the National Natural Science Foundation of China, No. 82172328; and the Key Laboratory of Clinical Laboratory Diagnosis and Translational Research of Zhejiang Province, No. 2022E10022.
Author information
Authors and Affiliations
Contributions
YL and ZCY performed the experiments, generated figures, analyzed the data, and drafted the manuscript under the supervision of QW and TLZ. ZZM, MRT, HCC, CRQ, and WLZ conducted the data analysis. The research plan for this project was conceived based on several rounds of discussions among all co-authors. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All the experimental methods were performed in accordance with approved guidelines and regulations. The First Affiliated Hospital of Wenzhou Medical University’s ethics committee authorized each investigative protocol used in this study. Informed consent was obtained from all participants.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Liu, Y., Yao, Z., Mao, Z. et al. Quorum sensing gene lasR promotes phage vB_Pae_PLY infection in Pseudomonas aeruginosa. BMC Microbiol 24, 207 (2024). https://doi.org/10.1186/s12866-024-03349-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12866-024-03349-7