Abstract
Background
Acute heat stress could induce high mortality and cause huge economic losses in the poultry industry. Although many studies have revealed heat stress-induced injuries of multiple tissues, the main target tissue and molecular mechanism of death under acute heat stress was largely unknown. This study systematically compared the transcriptome data of five main visceral tissues in chickens to reveal the response of multiple tissues to acute heat stress and determine the main target tissue of acute heat stress, further revealing the injuries of main target tissue and their potential mechanism by combing pathological section and qRT-PCR technologies.
Results
The transcriptome data of five visceral tissues revealed that acute heat stress broadly caused inflammatory response and damaged tissues metabolic homeostasis. Among the five tested visceral tissues, the number of differentially expressed genes in the lung was the highest, and their fold changes were the greatest, indicating that the lung was the main target tissue of acute heat stress. The results of pathological section revealed severe inflammation, emphysema and pulmonary hemorrhage in the lung under acute heat stress. Our study found that some pro-inflammatory genes, including CNTFR, FURIN, CCR6, LIFR and IL20RA, were significantly up-regulated both in the heat-stress and heat-death groups, and their fold changes in the heat-death group were significantly greater than that in the heat-stress group. We also found an anti-inflammatory gene, AvBD9, exhibiting an extremely high expression in the heat-stress group but a low expression in the heat-death group.
Conclusions
Our study found that acute heat stress caused multiple tissue injuries broadly and the lung was the main target tissue of acute heat stress in chicken. Acute heat stress caused a severe inflammatory response, emphysema, and pulmonary haemorrhage, The severe inflammatory response in the heat-death group was related to the up-regulation of pro-inflammatory genes and down-regulation of anti-inflammatory genes.
Similar content being viewed by others
Introduction
With global warming, heat stress caused huge economic losses in the poultry industry [1]. According to the statistics, heat stress has caused losses of 125–165 million dollars to the poultry industry in the United States [2]. Heat stress is the sum of non-specific responses of chickens when they are under an excessively high temperature exceeding their thermoregulatory capacity, and can be roughly divided into acute and chronic heat stress according to the environmental temperature and exposure time. Under a high temperature, acute heat stress in chickens will be induced in a short time, causing high levels of tissue-damage and mortality [3]. Due to the influence of global warming, the occurrence frequency of extreme heat weather in various regions is increasing, which is short-lived but can still cause a high level of mortality. For example, in the summer of 2012, a large number of deaths of laying hens were caused by experiencing three periods of temperatures exceeding 38℃ in a short time [4]. Some regions, especially the tropical desert areas, have extremely high temperatures causing several problems for local poultry industry. For example, the highest temperature in the Cairo area of Egypt exceeded 44℃ for five years in the past decade (https://rp5.ru). Additionally, in modern poultry industry, temperature is controlled at the cost of a lot of energy, in which unexpected events such as power outages can’t be completely avoided. When power outages occurred, the environmental temperature would rise sharply. A reported example was that the environmental temperature was as much as 55 °C after a power outage and the mortality of population experienced such an event was as high as 98% [5].
In fact, a series of studies have proved that heat stress could cause injuries to multiple tissues in chickens. For example, heat stress could cause mitochondrial damage and produce excess reactive oxygen species (ROS) [6], thus resulting in the overwhelmed liver buffering system and oxidative injuries to enzymes, cellular lipids and mitochondrial membranes [7]. Heat stress could cause myocardial fiber rupture [8] and up-regulation of multiple genes involved in cardiac contraction [9]. Additionally, heat stress caused reduced layers and structural abnormalities of spermatogenic cells in chicken testis [1); three genes were transcribed from intronic regions of HSPA8, including LOC112530276, LOC112530277 and LOC112530278; one gene was overlapped with HSPH (Table 1).
Comparative analysis of five visceral tissues between the control and heat-stress groups. (A) Summary of differentially expressed gene numbers in five visceral tissues between the control and heat-stress groups. (B) Distribution of fold changes of differentially expressed genes between the control and heat-stress groups. (C) Significant pathways enriched by differentially expressed genes between the control and heat-stress groups. (D) The number of differentially expressed genes from five visceral tissues involved in significant pathways
Then, all DEGs were used for KEGG pathway enrichment analysis. The results showed that a total of 23 pathways exhibited significance, and 13 of these pathways, including cytokine-cytokine receptor interaction, signal transduction, metabolic pathway, neuronal system, and calcium signalling pathway, were shared in all tissues (Fig. 1C). These results indicated that acute heat stress could cause inflammatory responses broadly in every visceral tissue, and signal transduction and the nervous system might play an important regulatory role in response to acute heat stress, and metabolic homeostasis might also be remodelled under acute heat stress.
Transcriptome data of multiple tissues reveals that the lung is the main target tissue under acute heat stress
As mentioned above, the DEG number in the lung was much higher than that in other tissues, and the fold changes of DEGs were significantly greater than that of other tissues. In the shared pathways, the number of DEGs involved in the GPCR signalling pathway from the lung was significantly higher than that of other tissues (Fig. 1D), indicating the signalling transduction in the lung under acute heat stress was more active. The number of DEGs enriched in neural ligand-receptor interaction in the lung was significantly higher than that in other tissues (Fig. 1D), indicating that the connection between lung and nervous system under acute heat stress was more intimate than that of other tissues. Besides, the number of DEGs in the lung related to the metabolic pathway was higher than that in other tissues (Fig. 1D), suggesting a greater effect on the lung metabolic homeostasis under acute heat stress. The number of DEGs in the heart and lung involved in cytokine-cytokine receptor interaction was higher than that in other tissues (Fig. 1D), indicating a more severe inflammatory response in the heart and lung under acute heat stress.
Pathological section reveals severe inflammation and emphysema in the lung under acute heat stress
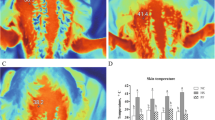
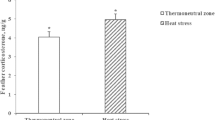
Additionally, we set heat-stress and heat-death groups to examine the lung injuries under acute heat stress. The heat-death time of these individuals ranged from 49 to 75 min and we randomly selected three individuals for further analysis with their heat-death time from 53 to 75 min (Table S3). We found the right lung index of the heat-death group was significantly higher than that of the control group (P < 0.01) (Fig. 3A), and the ratio between right lung weight after drying at 37℃ for 48 h and fresh weight of right lung in the heat-death group was lower than that of the control group (P = 0.17) (Fig. 3B). Compared to the control group (Fig. 3C, F, I), we detected excessive inflammatory cell infiltration in the lung epithelial mucosa of the heat-stress group (Fig. 3D) and heat-death group (Fig. 3E), mild emphysema in heat-stress group (Fig. 3G), severe emphysema and alveolar rupture in heat-death group (Fig. 3H), pulmonary haemorrhage in both groups (Fig. 3J and K). In short, our results revealed that acute heat stress could damage the normal structure of the lung, and cause severe inflammatory response, emphysema and pulmonary hemorrhage.
Pathological findings in the lung under acute heat stress. (A) the index of right lung weight divided by body weight between the control and heat-death groups. (B) the dry weight of right lung after drying at 37℃ for 48 h divided by its fresh weight. The different letters indicate significant difference between groups. (C-K) the representative photographs of HE-stained lungs, and their scale is 200 μm. (C) (F) and (I) show that pathological lung sections in the control group were normal. Severe inflammation was observed in the heat-stress group (D) and heat-death group (E). Emphysema was observed in the heat-stress group (G) and heat-death group (H), and alveolar ruptures were observed in the heat-death group (H). Pulmonary hemorrhage was observed in the heat-stress group (J) and heat-death group (K)
Evidence reveals the potential mechanism of inflammation in the lung under acute heat stress
A total of 10 genes involved in cytokine-cytokine receptor interaction were differentially expressed in the heat-stress group, including BMP10, IL20RA, GDF2, GDF15, THPO, CNTFR, INHBA, CCR6, LIFR and BMP3 (Figure S3). In these 10 genes, all genes were up-regulated significantly in the heat-stress group except CCR6, LIFR and BMP3. CCR6, LIFR, IL20RA and CNTFR were selected to examine their expression in the heat-stress and heat-death groups because they have been reported to be pro-inflammatory genes. Besides, the other 3 genes, including FURIN, AvBD9 and BAIAP9, were also targeted for validation because they were reported to be related to the innate immune system. In these 7 genes used for validation, the expression trends of five genes in transcriptome data were consistent with the results of qRT-PCR (Table S2). Transcriptome data revealed that CCR6 and LIFR were significantly down-regulated with low-fold changes in the heat-stress group (Table S2). The results of qRT-PCR revealed their up-regulation, suggesting that we should be particularly careful about DEGs with low fold change when using the transcriptome data. All these genes for validation exhibited up-regulations in the heat-stress group, including 5 pro-inflammatory genes and 2 anti-inflammatory genes (Fig. 4). The 4 pro-inflammatory genes, including CNTFR, CCR6, LIFR and FURIN, were up-regulated both in the heat-stress and heat-death groups, with higher fold changes in the heat-death group than in the heat-stress group. The pro-inflammatory gene IL20RA also exhibited up-regulation in both groups, but its fold change in the heat-death group was lower than that in the heat-stress group. The anti-inflammatory gene AvBD9 exhibited significant up-regulation in the heat-stress group but down-regulation in the heat-death group with potential significance. Another anti-inflammatory gene BAIAP2 exhibited insignificant up-regulation in the heat-stress and heat-death groups.
Relative expression level of several inflammatory genes by qRT-PCR. Among these genes, CNTFR, CCR6, LIFR, FURIN and IL20RA belong to pro-inflammatory genes, and AvBD9 and BAIAP2 belong to anti-inflammatory genes. Expression levels were normalized against that of the GAPDH gene. Values are means (n = 3) and error bars represent the standard deviation. Different letters above the bars indicate a significant difference in different groups
Discussion
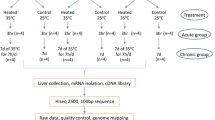
Chicken has no sweat glands and is covered with feathers, thus chicken has a higher temperature than mammals. Thus, chickens are more susceptible to heat stress than mammals when under high temperatures. Many previous studies have found that heat stress could induce multiple tissue injuries [6, 7, Total RNA extraction, RNA-Seq and DEGs identification To examine the tissue injuries under heat stress, 3 samples of the control and heat-stress groups were randomly selected to perform RNA-Seq. Total RNA was extracted by TRIzol reagent based on the manufacturer’s instruction, whose quality was examined by NanoDrop and Agilent 2100. The mRNA was isolated by Oligo (dT)-attached magnetic beads, and randomly fragmented in a fragmentation buffer. First-strand cDNA was synthesized with the fragmented mRNA as a template and random hexamers as primers, followed by second-strand synthesis with the addition of PCR buffer, dNTPs, RNase H and DNA polymerase I, and cDNA was purified with AMPure XP beads. Double-strand cDNA was subjected to the end repair, and a cDNA library was obtained by certain rounds of PCR. A total of 36 libraries were sequenced on Illumina HiSeq 2500 platform (Illumina Inc., San Diego, USA) according to Illumina’s RNA-Seq instructions to obtain 150-bp paired-end reads for further study. The low-quality reads and adapters were removed with Trimmomatic-0.39 (Bolger, et al., 2014). High-quality reads were aligned to the reference genome (GRCg6a) by hisat2 [34], and gene count was generated by HTSeq [35]. Differentially expressed genes (DEGs) were detected by DESeq2 package [36]. The genes with P-value < 0.05 and |Log2 (fold change)| >1 were defined as DEGs. DEGs were used to examine their enriched pathways with KOBAS 3.0 [37]. For the most important pathways, the raw images were obtained from KEGG [38,39,40], and the related significant gene information was highlighted using Pathview[41]. The samples fixed in 4% formaldehyde solution (National Pharmaceutical Group Chemical Reagent Co., Ltd., China) were paraffin-embedded, and 3 μm-thick sections of the paraffin embedding samples were obtained and used for hematoxylin-eosin (H&E) staining. We examined all the histological slides with an Olympus light microscope and digitized images with a Nikon DS-U3 camera (Olympus Corporation, Tokyo, Japan) control unit connected to a Nikon Eclipse CI upright optical microscope. We performed qRT-PCR technology with 3 samples of the control, heat-stress and heat-death groups to quantify the expressions of key genes and verify the reliability of RNA-Seq. Total RNA from every sample was converted into cDNA using HiScript Reverse Transcriptase (Vazyme Biotech Co., Ltd., Nan**g, China), and cDNA was used for qRT-PCR with GAPDH as the internal control gene. qRT-PCR was performed with SYBR Green I kit (Vazyme Biotech Co., Ltd., Nan**g, China), with 1.6ul cDNA and 0.2ul of forward and reversed primers in a final volume of 10ul. The samples were centrifuged briefly and run on the PCR machine using the program ( 95℃ for 5 min, 95℃ for 30s, 58℃ for 30s, 72℃ for 30s, 72℃ for 5 min and 25℃ for 1 min) in triplicate. The relative gene expression levels were determined using the 2–ΔΔCt method, the mean value of three technological replications was used to represent the expression of the individual and three biological replications of different groups was used to estimate the difference in different groups. We used LSD.test function to estimate the difference in gene expression in different groups. The primer sequences of genes for validation are represented in Table 2.Histopathological examination
Quantitative real-time PCR (qRT-PCR)
Data Availability
Sequencing data in this study are submitted to the Genome Sequence Archive in BIG Data Center under the BioProject: PRJCA010811 (https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA010811).
Abbreviations
- DEG:
-
Differentail expressed gene
- HE:
-
Hematoxylin-eosin staining
- qRT-PCR:
-
Quantitative real-time PCR
- PCA:
-
Principle component analysis
- HSP:
-
Heat shock protein
- DNAJA4:
-
DnaJ heat shock protein family (Hsp40) member A4
- HSPA8:
-
Heat shock protein family A (Hsp70) member 8
- HSPA2:
-
Heat shock protein family A (Hsp70) member 2
- BAG3:
-
BAG cochaperone 3
- HSP90AA1:
-
Heat shock protein 90 alpha family class A member 1
- HSPB9:
-
Heat shock protein family B (small) member 9
- IL20RA:
-
Interleukin 20 receptor subunit alpha
- GDF2:
-
Growth differentiation factor 2
- GDF15:
-
Growth differentiation factor 15
- THPO:
-
Thrombopoietin
- CNTFR:
-
Ciliary neurotrophic factor receptor
- INHBA:
-
Inhibin subunit beta A
- CCR6:
-
C-C motif chemokine receptor 6
- LIFR:
-
LIF receptor subunit alpha
- BMP3:
-
Bone morphogenetic protein 3
- FURIN:
-
Furin, paired basic amino acid cleaving enzyme
- AvBD9:
-
Avian beta-defensin 9
- BAIAP2:
-
BAR/IMD domain containing adaptor protein 2
- TGFβ:
-
Transforming growth factor beta family
References
Khan RU, Naz S, Ullah H, Ullah Q, Laudadio V, Qudratullah, Bozzo G, Tufarelli V. Physiological dynamics in broiler chickens under heat stress and possible mitigation strategies. Animal Biotechnol. 2023;34(2):438–47.
St-Pierre N, Cobanov B, Schnitkey G. Economic losses from heat stress by US livestock industries. J Dairy Sci. 2003;86:E52–E77.
Branton S, Reece F, Deaton J. Use of ammonium chloride and sodium bicarbonate in acute heat exposure of broilers. Poult Sci. 1986;65(9):1659–63.
Wolc A, Arango J, Settar P, Fulton J, O’Sullivan N, Dekkers J. Genome wide association study for heat stress induced mortality in a white egg layer line. Poult Sci. 2019;98(1):92–6.
Asadollahi H, Vaez Torshizi R, Ehsani A, Masoudi AA. An association of CEP78, MEF2C, VPS13A and ARRDC3 genes with survivability to heat stress in an F2 chicken population. J Anim Breed Genet 2022.
Mujahid A, Pumford NR, Bottje W, Nakagawa K, Miyazawa T, Akiba Y, Toyomizu M. Mitochondrial oxidative damage in chicken skeletal muscle induced by acute heat stress. J Poult Sci. 2007;44(4):439–45.
Emami NK, Jung U, Voy B, Dridi S. Radical response: effects of heat stress-induced oxidative stress on lipid metabolism in the avian liver. Antioxidants. 2020;10(1):35.
Tang S, Yin B, Xu J, Bao E. Rosemary reduces heat stress by inducing CRYAB and HSP70 expression in broiler chickens. Oxidative Medicine and Cellular Longevity 2018, 2018.
Zhang Q, Zhang B, Luo Y. Cardiac transcriptome study of the effect of heat stress in yellow-feather broilers. Italian J Anim Sci 2019.
**ong Y, Yin Q, Li J, He S. Oxidative stress and endoplasmic reticulum stress are involved in the protective effect of alpha lipoic acid against heat damage in chicken testes. Animals. 2020;10(3):384.
Wang S-H, Cheng C-Y, Tang P-C, Chen C-F, Chen H-H, Lee Y-P, Huang S-Y. Differential gene expressions in testes of L2 strain Taiwan country chicken in response to acute heat stress. Theriogenology. 2013;79(2):374–82. e377.
Gyawali I, Paudel R. The effect of heat stress on Meat Quality, Growth performance and antioxidant capacity of broiler chickens: a review. Poult Sci J. 2022;10(1):1–12.
Sun L, Lamont SJ, Cooksey AM, McCarthy F, Tudor CO, Vijay-Shanker K, DeRita RM, Rothschild M, Ashwell C, Persia ME. Transcriptome response to heat stress in a chicken hepatocellular carcinoma cell line. Cell Stress Chaperones. 2015;20(6):939–50.
Bowman MJ, Park W, Bauer PJ, Udall JA, Page JT, Raney J, Scheffler BE, Jones DC, Campbell BT. RNA-Seq transcriptome profiling of upland cotton (Gossypium hirsutum L.) root tissue under water-deficit stress. PLoS ONE. 2013;8(12):e82634.
Khatab AA, Li J, Hu L, Yang J, Fan C, Wang L, **e G. Global identification of quantitative trait loci and candidate genes for cold stress and chilling acclimation in rice through GWAS and RNA-seq. Planta. 2022;256(4):1–17.
Collier RJ, Baumgard LH, Zimbelman RB, **ao Y. Heat stress: physiology of acclimation and adaptation. Anim Front. 2019;9(1):12–9.
Adu-Asiamah P, Zhang Y, Amoah K, Leng Q, Zheng J, Yang H, Zhang W, Zhang L. Evaluation of physiological and molecular responses to acute heat stress in two chicken breeds. Animal. 2021;15(2):100106.
Hosseindoust A, Kang H, Kim J. Quantifying heat stress; the roles on metabolic status and intestinal integrity in poultry, a review. Domest Anim Endocrinol. 2022;81:106745.
Mohyuddin SG, Khan I, Zada A, Qamar A, Arbab AAI, Ma X-b, Yu Z-c, Liu X-X, Yong Y-H, Ju XH. Influence of heat stress on intestinal epithelial barrier function, tight junction protein, and immune and reproductive physiology. BioMed Research International 2022, 2022.
Jastrebski SF, Lamont SJ, Schmidt CJ. Chicken hepatic response to chronic heat stress using integrated transcriptome and metabolome analysis. PLoS ONE. 2017;12(7):e0181900.
Zhang Q, Luo YK, Zhang BH, Chan YZ, Huang LL, Wang Y, Liang JM, Zhang XQ. RNA-Seq study of hepatic response of yellow-feather chickens to acute heat stress. Annals of Animal Science. 2020;20(1):55–69.
Ebi KL, Capon A, Berry P, Broderick C, de Dear R, Havenith G, Honda Y, Kovats RS, Ma W, Malik A. Hot weather and heat extremes: health risks. The Lancet. 2021;398(10301):698–708.
Su BH, Tseng YL, Shieh GS, Chen YC, Wu P, Shiau AL, Wu CL. Over-expression of prothymosin‐α antagonizes TGFβ signalling to promote the development of emphysema. J Pathol. 2016;238(3):412–22.
Verhamme F, Seys L, De Smet E, Provoost S, Janssens W, Elewaut D, Joos G, Brusselle G, Bracke K. Elevated GDF-15 contributes to pulmonary inflammation upon cigarette smoke exposure. Mucosal Immunol. 2017;10(6):1400–11.
Soler Palacios B, Estrada-Capetillo L, Izquierdo E, Criado G, Nieto C, Municio C, González‐Alvaro I, Sánchez‐Mateos P, Pablos JL, Corbí AL. Macrophages from the synovium of active rheumatoid arthritis exhibit an activin A‐dependent pro‐inflammatory profile. J Pathol. 2015;235(3):515–26.
Koivisto L, Bi J, Häkkinen L, Larjava H. Integrin αvβ6: structure, function and role in health and disease. Int J Biochem Cell Biol. 2018;99:186–96.
Chovatiya R, Medzhitov R. Stress, inflammation, and defense of homeostasis. Mol Cell. 2014;54(2):281–8.
Kim JW, Marquez CP, Sperberg RAP, Wu J, Bae WG, Huang PS, Sweet-Cordero EA, Cochran JR. Engineering a potent receptor superagonist or antagonist from a novel IL-6 family cytokine ligand. Proc Natl Acad Sci. 2020;117(25):14110–8.
Cordova ZM, Grönholm A, Kytölä V, Taverniti V, Hämäläinen S, Aittomäki S, Niininen W, Junttila I, Ylipää A, Nykter M. Myeloid cell expressed proprotein convertase FURIN attenuates inflammation. Oncotarget. 2016;7(34):54392.
Lukacs NW, Prosser DM, Wiekowski M, Lira SA, Cook DN. Requirement for the chemokine receptor CCR6 in allergic pulmonary inflammation. J Exp Med. 2001;194(4):551–6.
Wei K, Nguyen HN, Brenner MB. Fibroblast pathology in inflammatory diseases. J Clin Investig 2021, 131(20).
Lamichhane S, Mo J-S, Sharma G, Choi T-Y, Chae S-C. MicroRNA 452 regulates IL20RA-mediated JAK1/STAT3 pathway in inflammatory colitis and colorectal cancer. Inflamm Res. 2021;70(8):903–14.
Tu J, Qi K, Xue T, Wei H, Zhang Y, Wu Y, Zhou X, Lv X. Construction of recombinant pichia pastoris carrying a constitutive AvBD9 gene and analysis of its activity. J Microbiol Biotechnol. 2015;25(12):2082–9.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12(4):357–60.
Anders S, Pyl PT, Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31(2):166–9.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):1–21.
Bu D, Luo H, Huo P, Wang Z, Zhang S, He Z, Wu Y, Zhao L, Liu J, Guo J. KOBAS-i: intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res. 2021;49(W1):W317–25.
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28(1):27–30.
Kanehisa M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 2019;28(11):1947–51.
Kanehisa M, Furumichi M, Sato Y, Kawashima M, Ishiguro-Watanabe M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 2023;51(D1):D587–92.
Luo W, Brouwer C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics. 2013;29(14):1830–1.
Acknowledgements
Thanks to all co-authors for their dedication to this article. We also appreciate for valued comments and suggestions from Professor Xu Wang, Huazhong Agricultural University, China.
Funding
This work was supported by National Key R&D Program of China (Grant No. 2021YFD1300100) ; Natural Science Foundation of China (No.32072707); and the grants from the Science and Technology Major Project of Hubei Province (2021ABA016).
Author information
Authors and Affiliations
Contributions
Shijun Li and Jiuhong Nan dessigned the experiments. Jiuhong Nan, Hongrui Yang, Li Rong Zijia Jia and Sendong Yang collected the samples, Jiuhong Nan analyzed the data, performed the experiments and wrote the draft. Shijun Li and Jiuhong Nan revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Animal experiments were approved by the Animal Welfare Committee of Huazhong Agricultural University (Hubei, China), and all experimental procedures strictly followed the related laboratory regulations and the relevant guidelines. The study adheres to the ARRIVE 2.0 guidelines for reporting animal research.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Nan, J., Yang, H., Rong, L. et al. Transcriptome analysis of multiple tissues reveals the potential mechanism of death under acute heat stress in chicken. BMC Genomics 24, 459 (2023). https://doi.org/10.1186/s12864-023-09564-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-023-09564-2