Abstract
Background
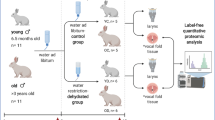
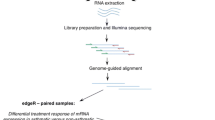
Voice disorders are a worldwide problem impacting human health, particularly for occupational voice users. Avoidance of surface dehydration is commonly prescribed as a protective factor against the development of dysphonia. The available literature inconclusively supports this practice and a biological mechanism for how surface dehydration of the laryngeal tissue affects voice has not been described. In this study, we used an in vivo male New Zealand white rabbit model to elucidate biological changes based on gene expression within the vocal folds from surface dehydration. Surface dehydration was induced by exposure to low humidity air (18.6% + 4.3%) for 8 h. Exposure to moderate humidity (43.0% + 4.3%) served as the control condition. Ilumina-based RNA sequencing was performed and used for transcriptome analysis with validation by RT-qPCR.
Results
There were 103 statistically significant differentially expressed genes identified through Cuffdiff with 61 genes meeting significance by both false discovery rate and fold change. Functional annotation enrichment and predicted protein interaction map** showed enrichment of various loci, including cellular stress and inflammatory response, ciliary function, and keratinocyte development. Eight genes were selected for RT-qPCR validation. Matrix metalloproteinase 12 (MMP12) and macrophage cationic peptide 1 (MCP1) were significantly upregulated and an epithelial chloride channel protein (ECCP) was significantly downregulated after surface dehydration by RNA-Seq and RT-qPCR. Suprabasin (SPBN) and zinc activated cationic channel (ZACN) were marginally, but non-significantly down- and upregulated as evidenced by RT-qPCR, respectively.
Conclusions
The data together support the notion that surface dehydration induces physiological changes in the vocal folds and justifies targeted analysis to further explore the underlying biology of compensatory fluid/ion flux and inflammatory mediators in response to airway surface dehydration.
Similar content being viewed by others
Background
Voice disorders are a prevalent communication disorder affecting human health worldwide [1,2,3,4,5,6]. In the United States general population, the prevalence of voice disorders has been estimated at 6.2% [7], and more recently, at 7.6% [8]. Data from the National Longitudinal Study of Adolescent to Adult Health shows the same 6% estimate among the adolescent population [9]. The development of voice disorders is identified as an occupational hazard, particularly among speakers who depend on a healthy voice for their livelihood. School teachers, entertainers, legal professionals are all at greater risk of dysphonia from voice disorders [3, 7, 10,11,12,13]. The economic impact of voice disorders is substantial. The average associated health care costs in the United States have been estimated at almost 200 million dollars [14], and a study of Brazilian teachers having to take time away from work due to dysphonia illustrates the potential impact of a loss of productivity in the workforce [15]. Taken together, the impact of voice disorders on society supports the need for a more comprehensive understanding of the development of voice disorders and therapies to address them.
Interventions for voice disorders exist along a continuum of non-invasive behavioral modifications to phonosurgery. The focus of this study is on the molecular biological responses to laryngeal surface dehydration as a means of substantiating the commonly prescribed prophylactic and therapeutic practice among speech-language pathologists [4, 16,17,18,19].
Dehydration, as it relates to voice, occurs under two paradigms: systemic dehydration and airway surface dehydration. Systemic dehydration, decreased total body water, has been shown to negatively impact phonatory effort in humans and acoustic measures in humans and ex vivo animal models [20,21,22,23]. Surface dehydration as related to voice is defined as loss of water from the luminal surface of the larynx and vocal folds. In everyday life, this may be caused by exposure to air of low humidity or increased respiratory rate from exercise. While there is evidence suggesting that surface dehydration within the larynx negatively impacts phonation with similar outcomes as systemic dehydration, recent studies in humans [24,25,26,27] do not always find a significant correlation between the two.
Unfortunately, rigorous in vivo analysis of the physiology of laryngeal surface dehydration is precluded by the invasive nature of data collection and the ethical implications of causing vocal injury in human subjects. Human studies are, therefore, generally limited to acoustic and aerodynamic measures or post-mortem evaluation. Conversely, animal models have largely allowed for ex vivo studies, which provide ample evidence that surface dehydration impacts vocal fold biomechanics and function [28,29,30], but the molecular pathobiology and resulting homeostatic compensatory mechanisms remain unclear. An attractive surrogate to the vocal folds is the airway distal to the larynx, which has been studied in the context of airway surface fluid homeostasis and response to luminal perturbations [31,32,33]. It has long been established that the humidity of inspired air can affect the magnitude of water lost to respiration [34] and that the resulting concentration of luminal electrolytes can cause dramatic physiological responses in the trachea, upper and lower airways [35, 36]. The vocal folds are covered by nonkeratinized stratified squamous epithelium, and the laryngeal lumen is predominately covered by respiratory epithelium. Therefore, the larynx may respond to perturbations similarly to the tracheal epithelium. This potential is supported in studies assessing vocal fold ion flux to altered composition of luminal surface fluid [37,38,39]. However, these were in vitro studies limiting the generalization of the data. Further studies are required to address questions of the specific underlying biology.
To probe for potential physiological responses to surface dehydration, we used an in vivo rabbit model. Anatomically, the rabbit larynx is grossly similar to the human larynx. Its size has been approximated to 8.6 × 5.5 mm at the level of the arytenoids [40, 41], consistent with the dimensions of the human newborn larynx [42]. Additionally, the literature demonstrates that rabbit larynges exhibit sufficient biological similarity to humans and have been used in molecular and histological studies of the vocal folds [43,44,45,46,47]. The rabbit larynx has also been used to characterize the physiological response to injury secondary to phonation [46, 47] or laryngeal and vocal fold surgery [48,49,50]. The common use of rabbits for laryngeal studies and the relatively small size for handling and housing makes this animal a suitable model for this study.
In this study, we sought to identify transcriptional-level changes in response to low humidity exposure that suggest a response to surface dehydration within the membranous vocal folds or the vocal fold lamina propria. We successfully addressed the following aims: [1] construction and evaluation of an environmental chamber capable of exposing rabbits to a consistent, physiologically-realistic low relative humidity environment and [2] investigation of the effects of 8 h of low humidity exposure on rabbit larynx by way of RNA sequencing (RNA-Seq). An 8-h exposure was selected as representative of a typical working day for human subjects. We used low humidity rather than desiccated air as the surface dehydration challenge to increase the ecological validity of the study. Rabbits exposed to moderate humidity served as the control condition.
Results
Humidity challenge and gross physical assessment
A total of eight rabbits were challenged with low humidity, and six rabbits were exposed to moderate humidity (control condition) in a specially fabricated environmental chamber (Fig. 1; see Methods section for details). Low humidity was 18.6 ± 4.3% (mean ± standard deviation) over the 8 h. The moderate humidity exposure was 43.0 ± 4.3% over the 8 h (Fig. 2). There was no observable behavioral differences or evidence of respiratory distress following exposure in either group. No gross evidence of inflammation or damage to the laryngeal mucosa was observed during visual examination under a dissecting microscope.
Environmental chamber used in this experiment. a Schematic design of the environmental chamber. Air output toward dehumidifier (a), air intake plenum from dehumidifier (b), latches for chamber doors that open longitudinally (c), mobile divider for separating challenge compartment into two sections (d, 1, 2), permanent divider separating challenge from control compartment (e, 3), and gated vent caps for titration of room air (f). b Picture of chamber showing actual materials and dimensions. Schematic design and photograph are both property of the authors
Packed cell volume (PCV)
The pre-experiment PCV (%) across all 14 rabbits was 46.7 ± 2.8 (mean ± SD). The % change in PCV from baseline to after the experiment did not differ significantly between the low and moderate humidity groups (p = 0.1692).
Sequence read map** and RNA-Seq
Approximately 69 to 112 million paired reads were obtained by RNA-Seq with an average of 70% quality reads map** to genes in the rabbit genome in each sample. In total, 23,669 annotations were obtained. Differential gene expression by Cuffdiff revealed 103 genes reaching an FDR < 0.05 with 61 meeting the additional fold change (|log2 FC| ≥ 1) filtering criterion. Of these, 48 genes were considered significantly downregulated and 13 genes were significantly upregulated. The 10 genes with the greatest up- and downregulated fold changes from this list of 61 are shown in Table 1. A complete list of all genes identified is provided within the Additional file 1: Table S1.
Rabbits were compared using principal component analysis based on FPKM obtained from Cuffdiff without low expression genes being removed (Fig. 3). The first two principal components explain 44% of the total variability. Although neither PC1 nor PC2 were able to distinguish low humidity rabbits from control rabbits, rabbits tended to cluster according to their treatment information based on PC1 and PC2 together. The rabbits with the most prominent deviations, LH26 and CH35, were not found to be consistent outliers within the qRT-PCR analyses discussed below.
Functional enrichment analysis
Functional enrichment analysis by DAVID and STRING provided similar but distinct sets with FDR < 0.05. DAVID identified 4 GO terms for biological process, 6 GO terms for cellular component, 2 GO terms for molecular function, and 7 processes by KEGG with FDR < 0.05. GO terms and KEGG processes included cardiac muscle function, calcium binding, chemical carcinogenesis, and ECM-receptor interaction. STRING provided a richer set with 7, 15, and 19 GO terms for biological process, cellular component, and molecular function, respectively, and 2 KEGG processes. GO terms included stress and inflammatory response, cytoskeleton, and ion binding.
For GSEA, 17 genes sets were significantly enriched in the moderate humidity group with an FDR < 0.25. There were 5, 6, 6 terms for biological process, cellular compartment, and molecular function, respectively. These include collagen, basement and plasma membrane, epidermis development, and epithelial cell differentiation. In the low humidity group 4 gene sets were significantly enriched with FDR < 0.25. There were 2, 1, and 1 terms for biological process, cellular compartment, and molecular function, respectively. These include olfactory receptor activity and cellular response to calcium. The full lists of terms, functions, associated genes, and statistics for the aforementioned DAVID and STRING analyses, and enrichment data in moderate and low humidity groups from GSEA are provided in Additional file 2: Table S2.
The predicted protein-interacting network generated by STRING is shown in Fig. 4. There were 8 clusters identified with between 2 to 10 gene products. Larger clusters contain members that are associated with cellular response to external stimuli and immune response (dark green, lavender), muscle function (red), keratinocyte development (light green), and ciliary function (aqua).
Protein interaction network was created using STRING. A 100 node network was obtained from an input set of 103 differentially expressed genes identified by Cuffdiff with an FDR < 0.05. The line thickness represents the strength of the data to support the interaction, including text mining, experimental, database, co-expression, neighborhood, gene fusion, and co-occurrence sources. The minimum required interaction score was set to 0.4. Shell parameters were set to “None”. Disconnected nodes are not shown. Cluster colors are based on the Markov Cluster Algorithm with the inflation parameter set to 2
RT-qPCR validation
Eight genes were selected for subsequent data validation by RT-qPCR based on their predicted functions and assumption of relevance to vocal fold or laryngeal physiology; they consist of ENSOCUG00000003548, annotated as an epithelial chloride channel protein which will be referred to as “ECCP”, cadherin related family member 4 (CDHR4), corneodesmosin (CDSN), macrophage cationic peptide 1 (MCP1), matrix metallopeptidase 12 (MMP12), suprabasin (SPBN), zinc activated cationic channel (ZACN), and mucin 21 (MUC21), although the absolute value of log2 FC for MUC21 by RNA-Seq was only 0.79.
Of the eight genes tested, significant differences in relative expression were validated for ECCP (p = 0.028), MCP1 (p = 0.030), and MMP12 (p = 0.045) and were marginally non-significant for SPBN (p = 0.067) and ZACN (p = 0.066). The most prominent fold changes between the low and moderate humidity groups was observed for MMP12 (FC = 6.8), MCP1 (FC = 5.2), and ZACN (FC = 2.76). ECCP exhibited the largest downregulation (FC = 3.74). The remaining genes exhibited non-significant changes despite differential expression by RNA-Seq analysis (Fig. 5). Comparison of data from RNA-Seq and RT-qPCR are provided in Table 2.
RT-qPCR validation. Relative quantification for each gene was determined by the ΔΔCt method. All reactions were run in triplicate. The level of expression of each tested gene was standardized to the housekee** gene HPRT1, and ΔΔCt was calculated using the average of the ΔCts from the control group for the respective gene. ECCP, MCP1 and MMP12 were significantly different (p < 0.05) and SPBN and ZACN marginally non-significant (p = 0.06). Differences between groups as determined by the Welch t-test. Results represent 5–7 samples/group for each gene after the removal of outlier values as determined by the iterative application of a two-tailed Grubb’s test. Error bars represent the SEM for relative quantification within the respective humidity group
In silico analysis of ENSOCUG00000003548 gene (ECCP)
ENSOCUG00000003548 maps to NCBI gene accession number 100352679, annotated as epithelial chloride channel protein. This gene lies downstream of LOC100338755 (calcium-activated chloride channel regulator 4-like), calcium-activated chloride channels 4, 2, and 1 (CLCA4, CLCA2, CACL1).
Discussion
The transcriptional changes observed in this study indicate that just 8 h exposure to a low humidity environment can adversely affect vocal fold biology. To the best of our knowledge, this is the first study to demonstrate the effects of surface dehydration on vocal fold tissue in vivo. Important to our methodology, evaluation of the change in PCV following experimental challenge ruled out systemic dehydration as an unintended confounding factor in our analysis. There is considerable evidence that systemic dehydration negatively impacts phonation [20,21,22,23]. Surface dehydration represents a loss of water from the mucosal surface of the larynx, and while some level of local tissue water loss may be experienced through compensatory rehydration of the epithelial surface, we would not expect systemic dehydration to result. We hypothesize that the homeostatic responses to surface and systemic dehydration are governed by different cellular mechanisms, thus we used % PCV change to control for unintended systemic consequences of low humidity exposure with the concomitant withholding of food and water.
We developed a method to efficiently challenge rabbits to low humidity. We achieved average low relative humidity of approximately 20%, representing physiologically-realistic and substandard occupational conditions per Occupational Safety and Health Administration (OSHA) recommendations [51]. Moderate humidity control exposures were conducted in the same chamber with all compartments open to room air of variable temperature within housing guidelines for rabbits. Low humidity challenge and moderate humidity exposure could not be conducted at the same time because preliminary tests demonstrated that a fully closed air circuit that is needed to lower humidity in the chamber measurably increased the interior temperature of the compartments. By separating them, we successfully maintained appropriate ambient temperatures for the low humidity exposures [52] and maintained a 2-fold increase in moderate humidity exposures.
It is noteworthy that exposure to low relative humidity below the Occupational Health and Safety Administration (OHSA) recommended limit of 20% induced transcriptional changes within functional gene categories including inflammation, ion transport, and keratinocyte development. The most robust functional enrichments identified by STRING were stress, defense, and inflammatory responses. Additionally, outside of the STRING analysis, various genes for immunoglobulin chains were identified, three of which were downregulated and one that was upregulated. Interestingly, this cluster presents two opposing interpretations of innate immune dampening and possible macrophage activation.
While none of these genes or corresponding proteins are described within the larynx, the downregulated cluster can be interpreted as a dampening of acute inflammatory response. ORM1 and SAA1 are both acute phase proteins. ORM1 is an acute phase protein that has been shown to polarize M2 macrophage differentiation [53] and to enhance epithelial integrity in a culture model of the blood-brain barrier [54]. While ORM1 exhibits anti-inflammatory activity and its downregulation may allow for the development of a more robust inflammatory process, it may also be interpreted as indicative of surface dehydration not contributing to an activating inflammatory event. SAA1 is also an acute phase protein and is associated with a variety of pathological conditions, but it has also been shown to positively influence keratinocyte activity [55]. The S100 proteins are diverse with involvement in several cellular processes, but both S100A9 [56] and S100A12 [57] have been described as damage associated molecular patterns in the literature. Taken together, these results suggest that either surface dehydration is not inducing inflammatory pathways or that there is active repression of pro-inflammatory mediators. The latter is substantiated by the increase of IL1RN which encodes the IL-1 receptor antagonist (IL1RA). IL1RN was upregulated in the posterior cricoarytenoid muscle 1 week following transection of the recurrent laryngeal nerve in a rat model [58], and IL1RA was significantly increased following 8 h of industrial exposure to respirable and inhalable dust in humans [59]. Together this substantiates a role for the increased IL1RN we observed and of a possible active innate immunity repression in response to the low humidity challenge.
Conversely, the upregulation of MMP12 and MCP1 genes may suggest the activation of inflammatory macrophages. MMP12 was the most significantly upregulated gene in this study by RNA-Seq and RT-qPCR. MMP12 exhibits proteolytic activity on multiple ECM components including elastin, fibronectin, entactin, and type IV collagen [60], all of which are expressed within the vocal folds. Although called “macrophage elastase”, it is also expressed in human vocal fold fibroblasts [61] and bronchial epithelial cells in vitro [62], and in both superficial and deep epidermal layers of the skin in response to ultraviolet radiation [63]. MMP12 has a potential role in the development of dysphonia following low humidity exposure since type IV collagen and elastin play an important role in the viscoelasticity and phonatory function of the vocal folds [64, 65]. MMP12 may contribute directly to inflammation though epidermal growth factor receptor (EGFR) dependent induction of IL-8 from the respiratory epithelium [66]. Interestingly, MMP12 has been shown to positively influence wound healing following epithelial injury to the cornea [67], so it is unclear if the upregulated response to low humidity would be deleterious or influence a reparative response in the vocal folds. MCP1 is an α-defensin expressed in the lungs of fetal and adult rabbits [68]; it is secreted from neutrophils and rabbit lung macrophages and exhibits broad antimicrobial activity In our study, the expression of MCP1 was novelly detected in the rabbit larynx, and its upregulation in repsose to low humidity warrants further investiation including targeted anaylsis of differential expression between inflammatory cells and the larygeal tissue.
It is not surprising to find evidence of a pro-inflammatory response with surface dehydration as other environmental stressors such as simulated acidic reflux [69], hypertonic challenge [38], and phonotrauma [47, 70] can perturb the epithelial tight junctions of the vocal folds—indicative of the activation of proinflammatory pathways. As we did not investigate for cell-specific gene expression in this study, we are limited to conclude if the upregulation of these genes reflects activation of macrophages or activity of the epithelium or lamina propria fibroblasts, and further study is warranted. An intriguing hypothesis for a case of macrophage activation would be altered response to local microbiome or pathogens resulting from changes to the laryngeal microenvironment following dehydration.
The perturbation of ion transport or other lubrication mechanisms is anticipated as a response to the altered hydration state of the laryngeal surface [71]. Although no gene or protein interaction enrichment cluster was identified within the 103 DEGs analyzed, presumably due to the diversity of substrate and transporter type, a considerable set of ion and solute transporter related genes were identified by RNA-Seq, including ECCP, SLC5A1, SLC13A5, SLC23A1, SLC27A2, and ZACN. All SLC family members were downregulated. This set represents predominantly ion transport, with SLC13A5 and SLC27A2 being involved in glucose transport and fatty acid ligation. In vitro studies of human nasal epithelial cells [36] and human bronchial cell culture [72] demonstrated that apical osmotic pressure can result in altered epithelial electrolyte transport; however, studies with canine tracheal and bronchial cell culture [73] and an in vivo canine model [74] concluded that not all epithelial fluid flux is coupled to electrolyte transport. This evidence suggests that the epithelium may respond to either aberrant electrolyte concentrations or non-ionic osmotic pressure. It is not surprising to find evidence of altered chloride secretion specifically, as balanced sodium and chloride ion secretion is attributed to volume regulation of the airway surface fluid, but the contribution of transport of other ionic and non-ionic species is not well described for airway surface fluid regulation. Our results suggest the pertinence of future targeted study of noncanonical secretion products in the respiratory tract.
Although the ECCP is annotated as an epithelial chloride channel protein, the translation product for ECCP is neither well characterized nor has a direct ortholog in humans. It may belong to the calcium-activated chloride channel proteins (CLCA) family as identified by conserved functional domains, although it exhibits limited homology to the rabbit CLCA proteins. The genes for CLCA1, CLCA2, and CLCA4 lie within the same genetic neighborhood as ECCP but were identified by RNA-Seq with FDR > 0.99, indicating they are not differentially expressed in our model of surface dehydration (Additional file 1: Table S1). This suggests a distinct role for ECCP and its downregulation that warrant further investigation as an ion channel protein newly described in the context laryngeal surface dehydration. In contrast to ECCP, ZACN was upregulated in low humidity compared to moderate humidity but failed to reach statistical significance by RT-qPCR. ZACN is a cation channel expressed in the human trachea and other tissues and demonstrates permeability to potassium ions but not to chloride ions [75]; there is no discussion of its expression in the vocal folds in the literature, and it is unclear if it may also be sodium ion permeable. Taken together with the SLC family members identified, these results support a potential role for solute flux as a homeostatic response to surface dehydration. Interestingly, however, the downregulation of chloride transportation would be a counterintuitive response to surface dehydration at the apical membrane as chloride is generally directed out of the cell and aberrant chloride transport can be detrimental in the airways as seen in cystic fibrosis. There is a distinction between the respiratory epithelium of the airways and the nonkeratinized stratified squamous epithelium of the vocal folds, so care must be taken with direct translations of actions between the two.
The mucins are equally important to maintain satisfactory hydration of the laryngeal surface as ion and fluid flux. MUC12, MUC21, and TFF1 were identified as downregulated by RNA-Seq. Both mucins are members of the cell-surface associated mucin family, and as such, should originate directly from the epithelial cells. The first exon of MUC12 exhibited increased expression in laryngeal epithelium from laryngeal reflux patients compared to reflux negative patients [76]. Exogenous surface expression of MUC21 in in vitro cell culture reduced intercellular adhesion and adhesion to extracellular components [77]. It is interesting then to observe all three to be downregulated. However, in addition to roles as epithelial protectants, mucins and related proteins also serve roles in cell signaling with physiological consequences. This is recently shown for MUC21 overexpression as influencing the development of lung adenocarcinoma [78] and TFF1 influencing epithelial-mesenchymal transition. Together, this may be a contributing factor to the STRING cluster of keratinocyte differentiation factors discussed below, but further study is warranted to determine which cell types are expressing these genes and which cell signaling may be impacted.
Although there was no gross inflammation observed, some level of epithelial cellular response to surface dehydration is expected. The vocal folds are covered by a non-keratinized stratified squamous epithelium for which some aspects of development are well understood, such as embryological developmental factors and differentially expressed structural components [79, 80], but a comprehensive molecular description is not available as for other epithelia like the epidermis. It is interesting that several keratinocyte developmental and epithelial structural factors were identified with RNA-Seq, enriched in the low humidity group by GSEA, and as a protein interaction cluster in the STRING analysis: CDSN, CNFN, CRNN, KRT80, KRTDAP, and TGM3. Also identified by RNA-Seq were SPBN, another keratinocyte factor, and CDHR4, a cell interaction mediator. All of these were downregulated. SPBN, CDHR4, and CDSN were selected for RT-qPCR validation. All three gene products may be involved in maintaining the integrity of the stratified squamous epithelium, though none have been described specifically within the vocal folds until this study. SPBN is expressed in the suprabasal layers of tongue, stomach, and epidermis [81]. It is required for keratinocyte differentiation in an in vitro skin model [82] and skin development in murine embryos [83]. The specific activity of CDHR4 is not described in the literature, but family member CDHR2 is expressed in gastrointestinal epithelial cells and is associated with microvillus development [84], while family member CHDR3 is expressed in ciliated respiratory epithelial cells and is associated with ciliary development and intercellular interactions [85]. CDSN is expressed in the stratum granulosum of human skin and appears to participate in cellular cohesion at this level, with its loss associated with desquamation [86, 87]. That the entire cluster was downregulated substantiates surface dehydration as capable to influence vocal fold epithelial maintenance. Further study is required to elucidate the specific roles of these proteins within the vocal folds, as this epithelium is distinct from the epidermis.
Limitations
A limitation of designing an environmental chamber as described here was that it precluded the provision of relative humidity lower than 15%. While environmental rooms and chambers are commercially available, they are cost prohibitive and their small size precludes the use of certain animal models, such as rabbits. Another limitation of the study is that we only observed a single time point after low humidity exposure. It has been shown that local response to vocal fold injury is transient and time-dependent [43, 49, 88]. Further studies specifically observing for inflammatory response at multiple times points within a single challenge or within repeated or chronic challenges would be helpful in further characterization of vocal fold biology. Finally, the dissected vocal fold tissue included striated muscle and small amounts of respiratory epithelium immediately above and below the region of the vocal folds. Therefore, genes associated with muscle or respiratory epithelium were not selected for the discussion.
Conclusions
In this study, we investigated whether surface dehydration induced by low humidity would affect vocal fold biology. We successfully developed an efficient and cost-effective environmental chamber to induce surface dehydration. Humidity was maintained at occupationally relevant levels to observe for physiological impact. Low humidity challenge did not induce systemic dehydration as evaluated with PCV. High-throughput RNA-Seq provided evidence of a biologic response to low humidity challenge, including changes within stimulus-response, lubricative mechanisms, and potential alterations in epithelial maintenance. This study successfully serves as a framework for targeted investigations of molecular response to surface dehydration in the larynx.
Methods
Animals
All experiments were conducted in accordance with the guidelines and after approval of the Purdue Animal Care and Use Committee (Protocol # 1606001428). Animals were obtained from Envigo Global (Indianapolis, IN) and acclimatized for at least 1 week. Male New Zealand White rabbits, six to 8 months of age, and approximately 3 Kg were used for all experiments. Each experiment was run with two rabbits at a time randomly assigned to either the low (n = 8) or moderate (n = 6) humidity group. Samples sizes were selected based on recommendation from the Purdue Bioinformatics Core to ensure ideal minimum samples for statistical validity of RNA-Seq (n = 6 from each group). Changes to PCV were examined for all rabbits. RT-qPCR validation was conducted with 13 rabbits (low n = 7, moderate n = 6); one rabbit was excluded due to poor quality of RNA obtained after repeat extraction. Food and water were withheld during experiments under both humidity conditions. To encourage consistent, baseline hydration, all animals were pre-hydrated with 0.1 M sucrose in water ad libitum for the 2 days preceding the experiment. Animals were euthanized at the end of the experimental exposure with a single 1.0 mL IV dose of Beuthanasia-D Special (Schering Plough Animal Health Corp., Union, NJ).
Humidity challenge protocol
Eight hour low humidity and moderate humidity exposure were conducted in a specially fabricated environmental chamber. The chamber interior was segmented into three similar compartments, each with dimensions approximately 61 cm × 61 cm × 46 cm (Fig. 1a, b). Two compartments were sealed to limit the influx of room air and were intended for low humidity exposure, whereas the third compartment was left open to room air and was intended for a moderate humidity control. Gated duct caps were included within the wall of the low humidity compartment to allow for titration of room air as necessary.
Low humidity was achieved with a 70-pint commercial dehumidifier (Hisense DH70K1G: Qingdao, China) set to High Continuous attached to the chamber via 4-in. ducting. Moderate humidity exposure was achieved by opening the chamber airspace to room air without conditioning from the dehumidifier. Internal relative humidity and temperature were tracked using a HOBO Data Logger with a 12-bit Temperature/Relative Humidity Smart Sensor (U14–002, S-THB-M002: ONSET, Bourne, MA) at one-minute intervals.
Blood collection and analysis
Blood was collected in heparinized tubes at the beginning of the 8-h experiment and immediately prior to euthanasia via venipuncture of the lateral ear vein to minimize trauma and distress of collection. Packed cell volume (PCV) was measured manually by visual assessment using a microhematocrit reader card following centrifugation.
Sample collection and RNA extraction
The larynx and proximal trachea were excised from each animal immediately following euthanasia. The larynx was bisected posteriorly along the sagittal midline and pinned onto wax to expose the laryngeal lumen. Full-thickness soft tissue was microdissected bilaterally at the level of the glottis under magnification with microdissection scissors. Sections approximately 2-3 mm in any dimension collectively representing the vocal fold and surrounding tissue were immediately stored in RNAlater® Stabilization Solution (Invitrogen, Waltham, MA), stored at 4 °C overnight, and at − 80 °C until processing. Total RNA was extracted with the RNeasy Fibrous Tissue Mini Kit following the manufacturer protocol (QIAGEN®, Hilden, Germany).
RNA sequencing (RNA-Seq)
RNA quality was assessed by RNA Eukaryotic Pico Chip (Agilent Technologies Inc., Santa Clara, CA) and used to construct poly-A derived cDNA libraries with the Universal Plus mRNA-Seq kit (NuGEN Technologies, Inc., Redwood City, CA). High throughput sequencing was completed with an Illumina® NovaSeq™ 6000 Sequencing System (Illumina Inc., San Diego, CA) by 100 million reads, paired, of 150 bases per sample. Differential gene expression analysis was conducted by the Purdue Bioinformatics Core using Cuffdiff with default parameters [89]. Data were submitted to the NCBI GEO database under accession number GSE148588.
Quality control and read map**
Sequence quality was assessed using FastQC (v0.11.7) (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) for all samples, and quality trimming was done using FASTX-Toolkit (v 0.0.14) (http://hannonlab.cshl.edu/fastx_toolkit/) to remove bases with Phred33 score of less than 30, while retaining the resulting reads of at least 50 bases in length. The quality trimmed reads were mapped against the reference genome of Oryctolagus cuniculus using STAR (v 2.5.4b) [90].
Differential gene expression analysis and annotation
Differential gene expression analysis between low and moderate-humidity groups was carried out using ‘R’ (v 3.5.1; http://www.r-project.org/). STAR map** (bam) files were used for analysis by the Cuffdiff from Cufflinks (v 2.2.1) [89] suite of programs that perform differential expression analysis based on FPKM values. Syntax is provided in Additional file 4. Cuffdiff uses bam files to calculate Fragments per Kilobase of exon per Million fragments mapped (FPKM) values, from which differential gene expression between the pairwise comparisons can be ascertained. FPKM obtained were used for principal component analysis comparing individual rabbits; low expression genes were not removed. The gene annotations were retrieved from BioMart databases using biomartr package in ‘R’.
Functional enrichment analysis and predicted protein interactions
Differential gene expression data were filtered for a false discovery rate (FDR) of less than or equal to 0.05. Log2 FC positive values imply upregulation, while negative values imply downregulation of genes in the vocal folds exposed to low humidity versus moderate humidity challenge. The set of differentially expressed genes provided by Cufflinks meeting the FDR criterion (n = 103) was used as input by Ensembl gene ID for DAVID (v6.8) [91] to obtain functional annotation analysis with 86 being found within the database. The same set of genes was used as input for STRING (Search Tool for the Retrieval of Interacting Genes/Proteins) v.11.0 (https://string-db.org) for prediction of protein interaction analysis, providing a full, supplemented network of 100 nodes. In parallel, genes were ranked in descending order based on -log10 P-value multiplied by the sign of log2 transformed FC as input for GSEAPreranked (versions 7.2.0) that was used to perform gene set enrichment analysis using GO gene sets.
GO and KEGG enrichments were obtained from DAVID with default settings. STRING analysis parameters were set with line thickness representing the strength of the data to support interaction including text mining, experimental, database, co-expression, neighborhood, gene fusion, and co-occurrence sources, the minimum required interaction score set to 0.4, shell parameters set to “None”, and disconnected nodes to be hidden. Clusters were generated based on the Markov Cluster Algorithm with the inflation parameter set to 2. The parameters used and full datasets from GSEA are provided in Additional file 4.
Quantitative reverse transcription PCR (RT-qPCR)
Total RNA was used to generate cDNA with SuperScript™ IV VILO™ Master Mix (Invitrogen) using 374 ng of RNA as the template. RT-qPCR was performed in triplicate using SYBR Green 2x PCR Master Mix (Applied Biosystems, Waltham, MA) with 0.1 M of each primer and 2.5 μL of template cDNA in a 25 μL reaction volume using a QuantStudio 3 System (Applied Biosystems) thermocycler. Data was collected over 40 cycles by QuantStudio Design & Analysis Software v1.5.1. Primers used in this study are listed in Additional file 3: Table S3. Relative expression quantification of each gene was calculated using the 2(−ΔΔCt) method [92].
Statistical analysis
Statistical analysis was completed and visualized using RStudio™ Version 1.2.1335 (RStudio Inc., Boston, MA) with libraries Tidyverse [93] and outliers [94]. Changes in PCV were evaluated with Mann-Whitney nonparametric test. Relative gene expressions from RT-qPCR were tested with Welch two-sample t-test following removal of outlier values as determined by Grubb’s test. A p-value < 0.05 was considered statistically significant for all analyses.
Availability of data and materials
The RNA Sequencing dataset supporting the conclusions of this article are available in the NCBI GEO database under accession number GSE148588 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE148588). Datasets including a full list of genes identified and Cufflinks-calculated differential expression, the functional enrichment analyses of differentially expressed genes, a list of primers used in the RT-qPCR validation, the full GSEA data set, and the syntax used for Cufflinks analysis are provided in Additional files 1, 2, 3 and 4, respectively.
Abbreviations
- CDHR4:
-
Cadherin related family member 4
- CDSN:
-
Corneodesmosin
- ECCP:
-
Epithelial chloride channel protein
- DAVID:
-
Database for Annotation Visualization and Integrated Discovery v6.8
- DEGs:
-
Differentially expressed genes
- FC:
-
Linear fold change
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- MCP1:
-
Macrophage cationic peptide 1
- MMP12:
-
Matrixmetalloprotease 12
- MUC21:
-
Mucin-21
- PCV:
-
Packed cell volume
- RT-qPCR:
-
Reverse transcriptase quantitative polymerase chain reaction
- SPBN:
-
Suprabasin
- STRING:
-
Search Tool for Recurring Instances of Neighboring Genes
- ZACN:
-
Zinc activated chloride channel
References
Fourie K, Richardson M, van Der Linde J, Abdoola S, Mosca R. The reported incidence and nature of voice disorders in the private healthcare context of Gauteng. J Voice. 2017;31(6):774.e23.
Lai Y-T, Wang Y-H, Yen Y-C, Yu T-Y, Chao P-Z, Lee F-P, et al. The epidemiology of benign voice disorders in Taiwan: a Nationwide population-based study. Ann Otol Rhinol Laryngol. 2019;128(5):406–12.
Mohammadzadeh A, Sandoughdar N. Prevalence of voice disorders in Iranian primary school students. J Voice. 2017;31(2):263.e13.
National Institute on Deafness and Other Communication Disorders ib. Taking care of your voice. Updating April 2014. Bethesda: U.S. Department of Health and Human Services, National Institutes of Health, National Institute on Deafness and Other Communication Disorders; 2014.
Naunheim MR, Goldberg L, Dai JB, Rubinstein BJ, Courey MS. Measuring the impact of dysphonia on quality of life using health state preferences. The laryngoscope; 2019.
Pernambuco L, Espelt A, Góis ACB, de Lima KC. Voice disorders in older adults living in nursing homes: prevalence and associated factors. J Voice. 2017;31(4):510.e15.
Roy N, Merrill RM, Thibeault S, Parsa RA, Gray SD, Smith EM. Prevalence of voice disorders in teachers and the general population. J Speech Language Hearing Res. 2004;47(2):281.
Bhattacharyya N. The prevalence of voice problems among adults in the United States. Laryngoscope. 2014;124(10):2359–62.
Bainbridge KE, Roy N, Losonczy KG, Hoffman HJ, Cohen SM. Voice disorders and associated risk markers among young adults in the United StatesClinical report. Laryngoscope. 2017;127(9):2093.
Byeon H. Occupational risks for voice disorders: evidence from a Korea national cross-sectional survey. Logopedics Phoniatrics Vocol. 2017;42(1):39.
Da Rocha LM, de Lima BS, Do Amaral PL, Behlau M, de Mattos Souza LD. Risk factors for the incidence of perceived voice disorders in elementary and middle school teachers. J Voice. 2017;31(2):258.e7.
Mori MC, Francis DO, Song PC. Identifying occupations at risk for laryngeal disorders requiring specialty voice care. Otolaryngol Head Neck Surg. 2017;157(4):670.
Trinite B. Epidemiology of voice disorders in Latvian school teachers. J Voice. 2017;31(4):508.e1–9.
Cohen SM, Kim J, Roy N, Asche C, Courey M. Direct health care costs of laryngeal diseases and disorders. Laryngoscope. 2012;122(7):1582–8.
de Souza CM, Granjeiro RC, de Castro MP, RdC I, GMGF O. Outcomes of teachers away from work for voice disorders, state secretariat for education, Federal District, 2009–2010/ Desfecho dos professores afastados da Secretaria de Estado de Educacao do Distrito Federal por disturbios vocais entre 2009–2010.(ARTIGO ORIGINAL). Revista Brasileira de Med do Trabalho. 2017;15(4):324.
Treatment of Voice Disorders, 2nd Edition. Beaverton: Ringgold Inc; 2017.
Alves M, Krüger E, Pillay B, van Lierde K, van Der Linde J. The effect of hydration on voice quality in adults: a systematic review. J Voice. 2019;33(1):125.e13–28.
Behlau M, Oliveira G. Vocal hygiene for the voice professional. Curr Opin Otolaryngol Head Neck Surg. 2009;17(3):149–54.
Benninger MS, Murry T, Johns MM. The performer’s voice. 2nd ed. San Diego: Plural Publishing; 2016.
Brozmanova A, Jochem J, Javorka K, Zila I, Zwirska-Korczala K. Diuretic-induced dehydration/hypovolemia inhibits thermal panting in rabbits. Respir Physiol Neurobiol. 2006;150(1):99–102.
Fisher KV, Ligon J, Sobecks JL, Roxe DM. Phonatory effects of body fluid removal. Journal of speech, language, and hearing research. JSLHR. 2001;44(2):354.
Jiang J, Verdolini K, Jennie N, Aquino B, Hanson D. Effects of dehydration on phonation in excised canine larynges. Ann Otol Rhinol Laryngol. 2000;109(6):568–75.
Verdolini K, Min Y, Titze IR, Lemke J, Brown K, van Mersbergen M, et al. Biological mechanisms underlying voice changes due to dehydration. (statistical data included). J Speech Language Hearing Res. 2002;45(2):268.
Fujiki R, Chapleau A, Sundarrajan A, McKenna V, Sivasankar MP. The interaction of surface hydration and vocal loading on voice measures. J Voice. 2017;31(2):211–7.
Li L, Zhang Y, Maytag AL, Jiang JJ. Quantitative study for the surface dehydration of vocal folds based on high-speed imaging. J Voice. 2015;29(4):403–9.
Mahalingam S, Boominathan P. Effects of steam inhalation on voice quality-related acoustic measures. Laryngoscope. 2016;126(10):2305–9.
Patel RR, Walker R, Sivasankar PM. Spatiotemporal quantification of vocal fold vibration after exposure to superficial laryngeal dehydration: a preliminary study. J Voice. 2016;30(4):427–33.
Ayache S, Ouaknine M, Dejonkere PH, Prindere P, Giovanni A. Experimental study of the effects of surface mucus viscosity on the glottic cycle. J Voice. 2004;18(1):107–15.
RJB lH, Wieneke GH, Lebacq J, Dejonckere PH. Laryngeal mucosa elasticity and viscosity in high and low relative air humidity. Eur Arch Otorhinolaryngol. 2001;258(3):125–9.
Witt RE, Taylor LN, Regner MF, Jiang JJ. Effects of surface dehydration on mucosal wave amplitude and frequency in excised canine larynges. Otolaryngol Head Neck Surg. 2011;144(1):108–13.
Prazma J, Coleman CC, Shockley WW, Boucher RC. Tracheal vascular response to hypertonic and hypotonic solutions. J Appl Physiol (Bethesda, Md : 1985). 1994;76(6):2275.
Warren N, Crampin E, Tawhai M. The role of airway epithelium in replenishment of evaporated airway surface liquid from the human conducting airways. Ann Biomed Eng. 2010;38(12):3535–49.
Wells UM, Hanafi Z, Widdicombe JG. Osmolality alters tracheal blood flow and tracer uptake in anesthetized sheep. J Appl Physiol (Bethesda, Md : 1985). 1994;77(5):2400.
Tabka Z, Jebria AB, Guénard H. Effect of breathing dry warm air on respiratory water loss at rest and during exercise. Respir Physiol. 1987;67(2):115–25.
Anderson SD, Holzer K. Exercise-induced asthma: is it the right diagnosis in elite athletes? 2000. p. 419–28.
Willumsen NJ, Davis CW, Boucher RC. Selective response of human airway epithelia to luminal but not serosal solution hypertonicity. Possible role for proximal airway epithelia as an osmolality transducer. J Clin Invest. 1994;94(2):779.
Levendoski EE, Sivasankar MP. Vocal fold ion transport and Mucin expression following Acrolein exposure. J Membr Biol. 2014;247:5.
Sivasankar M, Erickson E, Rosenblatt M, Branski RC. Hypertonic challenge to porcine vocal folds: effects on epithelial barrier function. Otolaryngol Head Neck Surg. 2010;142(1):79–84.
Sivasankar M, Fisher K. Vocal fold epithelial response to luminal osmotic perturbation. J Speech Language Hearing Res. 2007;50(4):886–98.
Ajlan AM, Al-Khatib T, Al-Sheikah M, Jastaniah S, Salih A, Althubaiti A, et al. Helical computed tomography scanning of the larynx and upper trachea in rabbits. Acta Vet Scand. 2015;57:1.
Loewen MS, Walner DL. Dimensions of rabbit subglottis and trachea. Lab Anim. 2001;35(3):253.
Sato K, Chitose SI, Umeno H. Dimensions and morphological characteristics of human newborn glottis. Laryngoscope. 2015;125(5):E186–E9.
Branski RC, Rosen CA, Verdolini K, Hebda PA. Biochemical markers associated with acute vocal fold wound healing: a rabbit model. J Voice. 2005;19(2):283.
Hansen JK, Thibeault SL, Walsh JF, Shu XZ, Prestwich GD. In vivo engineering of the vocal fold extracellular matrix with injectable hyaluronic acid hydrogels: early effects on tissue repair and biomechanics in a rabbit model. Ann Otol Rhinol Laryngol. 2005;114(9):662–70.
** H-J, Lee S-H, Lee S-U, Lee H-S, ** S-M, Kim D-H, et al. Morphological and histological changes of rabbit vocal fold after steroid injection. Otolaryngol Head Neck Surg. 2013;149(2):277–83.
Kojima T, Garrett C, Novaleski C, Rousseau B. Quantification of acute vocal fold epithelial surface damage with increasing time and magnitude doses of vibration exposure. PLoS One. 2014;9(3):e91615.
Kojima T, Valenzuela CV, Novaleski CK, Van Deusen M, Mitchell JR, Garrett CG, et al. Effects of phonation time and magnitude dose on vocal fold epithelial genes, barrier integrity, and function. Laryngoscope. 2014;124(12):2770–8.
Kojima T, Mitchell JR, Garrett CG, Rousseau B. Recovery of vibratory function after vocal fold microflap in a rabbit model. Laryngoscope. 2014;124(2):481–6.
Mitchell JR, Kojima T, Wu H, Garrett CG, Rousseau B. Biochemical basis of vocal fold mobilization after microflap surgery in a rabbit model. Laryngoscope. 2014;124(2):487–93.
Okhovat S, Milner TD, Clement WA, Wynne DM, Kunanandam T. Validation of animal models for simulation training in pediatric Laryngotracheal reconstruction. Ann Otol Rhinol Laryngol. 2020;129(1):46–54.
OSHA. Reiteration of existing OSHA policy on indoor air quality: office temperature/humidity and environmental tobacco smoke; 2003.
NRC, Council NR. Guide for the care and use of laboratory animals. 8th ed. Washington, D.C: National Academy Press; 2011.
Nakamura K, Ito I, Kobayashi M, Herndon DN, Suzuki F. Orosomucoid 1 drives opportunistic infections through the polarization of monocytes to the M2b phenotype. Cytokine. 2015;73(1):8–15.
Zhang S, Mark KS. α1-acid glycoprotein induced effects in rat brain microvessel endothelial cells. Microvasc Res. 2012;84(2):161–8.
Yu N, Zhang S, Lu J, Li Y, Yi X, Tang L, et al. Serum amyloid a, an acute phase protein, stimulates proliferative and proinflammatory responses of keratinocytes. Cell Prolif. 2017;50(3):1.
Tsai S, Segovia J, Chang T, Morris IR, Berton M, Tessier PA, et al. DAMP molecule S100A9 acts as a molecular pattern to enhance inflammation during influenza a virus infection: role of DDX21-TRIF-TLR4-MyD88 pathway. PLoS Pathog. 2014;10:1.
Kallinich T, Wittkowski H, Keitzer R, Roth J, Foell D. Neutrophil-derived S100A12 as novel biomarker of inflammation in familial Mediterranean fever. Ann Rheum Dis. 2010;69(4):677.
Bijangi-Vishehsaraei K, Blum K, Zhang H, Safa AR, Halum SL. Microarray analysis gene expression profiles in laryngeal muscle after recurrent laryngeal nerve injury. Ann Otol Rhinol Laryngol. 2016;125(3):247–56.
Hedbrant A, Andersson L, Eklund D, Westberg H, Särndahl E, Persson A, et al. Quartz dust exposure affects NLRP3 Inflammasome activation and plasma levels of IL-18 and IL-1Ra in iron foundry workers. Mediat Inflamm. 2020;2020:1.
Gronski TJ, Martin RL, Kobayashi DK, Walsh BC, Holman MC, Huber M, et al. Hydrolysis of a broad spectrum of extracellular matrix proteins by human macrophage elastase. J Biol Chem. 1997;272(18):12189.
Adams GM, Xu CC, Chan RW. Senescence of human vocal fold fibroblasts in primary culture.(report). J Biomed Sci Eng (JBiSE). 2010;3(2):148.
Lavigne MC, Thakker P, Gunn J, Wong A, Miyashiro JS, Wasserman AM, et al. Human bronchial epithelial cells express and secrete MMP-12. Biochem Biophys Res Commun. 2004;324(2):534–46.
Tewari A, Grys K, Kollet J, Sarkany R, Young AR. Upregulation of MMP12 and its activity by UVA1 in human skin: potential implications for Photoaging. J Investig Dermatol. 2014;134(10):2598–609.
Chan R, Fu M, Young L, Tirunagari N. Relative contributions of collagen and elastin to elasticity of the vocal fold under tension. J Biomed Eng Soc. 2007;35(8):1471–83.
Rohlfs A-K, Goodyer E, Clauditz T, Hess M, Kob M, Koops S, et al. The anisotropic nature of the human vocal fold: an ex vivo study. Eur Arch Otorhinolaryngol. 2013;270(6):1885.
Le Quément C, Guénon I, Gillon J-Y, Lagente V, Boichot E. MMP-12 induces IL-8/CXCL8 secretion through EGFR and ERK1/2 activation in epithelial cells. Am J Phys Lung Cell Mol Phys. 2008;294(6):L1076.
Wolf M, Maltseva I, Clay SM, Pan P, Gajjala A, Chan MF. Effects of MMP12 on cell motility and inflammation during corneal epithelial repair. Exp Eye Res. 2017;160:11–20.
Zhu QZ, Hu J, Mulay S, Esch F, Shimasaki S, Solomon S. Isolation and structure of corticostatin peptides from rabbit fetal and adult lung. Proc Natl Acad Sci U S A. 1988;85(2):592.
Erickson E, Sivasankar M. Simulated reflux decreases vocal fold epithelial barrier resistance. Laryngoscope. 2010;120(8):1569–75.
Rousseau B, Suehiro A, Echemendia N, Sivasankar M. Raised intensity phonation compromises vocal fold epithelial barrier integrity. Laryngoscope. 2011;121(2):346.
Freed AN, Davis MS. Hyperventilation with dry air increases airway surface fluid osmolality in canine peripheral airways. Am J Respir Crit Care Med. 1999;159(4 Pt 1):1101.
Matsui H, Davis CW, Tarran R, Boucher RC. Osmotic water permeabilities of cultured, well-differentiated normal and cystic fibrosis airway epithelia. J Clin Invest. 2000;105(10):1419–27.
Grubb B, Schiretz F, Boucher RC. Volume transport across tracheal and bronchial airway epithelia in a tubular culture system. Am J Physiol-Cell Physiol. 1997;273(1):C21–C9.
Chen BT, Yeates DB. Differentiation of ion-associated and osmotically driven water transport in canine airways. Am J Respir Crit Care Med. 2000;162(5):1715.
Davies P, Wang W, Hales T, Kirkness E. A novel class of ligand-gated Ion Channel is activated by Zn super (2+). J Biol Chem. 2003;278(2):712–7.
Samuels TL, Handler E, Syring ML, Pajewski NM, Blumin JH, Kerschner JE, et al. Mucin gene expression in human laryngeal epithelia: effect of Laryngopharyngeal reflux. Ann Otol Rhinol Laryngol. 2008;117(9):688–95.
Yi Y, Kamata-Sakurai M, Denda-Nagai K, Itoh T, Okada K, Ishii-Schrade K, et al. Mucin 21/epiglycanin modulates cell adhesion. J Biol Chem. 2010;285(28):21233.
Yoshimoto T, Matsubara D, Soda M, Ueno T, Amano Y, Kihara A, et al. Mucin 21 is a key molecule involved in the incohesive growth pattern in lung adenocarcinoma. Cancer Sci. 2019;110(9):3006–11.
Lungova V, Verheyden JM, Herriges J, Sun X, Thibeault SL. Ontogeny of the mouse vocal fold epithelium. Dev Biol. 2015;399(2):263.
Lungova V, Chen X, Wang Z, Kendziorski C, Thibeault SL. Human induced pluripotent stem cell-derived vocal fold mucosa mimics development and responses to smoke exposure. Nat Commun. 2019;10(1):4161.
Park GT, Lim SE, Jang S-I, Morasso MI. Suprabasin, a novel epidermal differentiation marker and potential cornified envelope precursor. J Biol Chem. 2002;277(47):45195.
Aoshima M, Phadungsaksawasdi P, Nakazawa S, Iwasaki M, Sakabe J-I, Umayahara T, et al. Decreased expression of suprabasin induces aberrant differentiation and apoptosis of epidermal keratinocytes: possible role for atopic dermatitis. J Dermatol Sci. 2019;95(3):107.
Nakazawa S, Aoshima M, Funakoshi A, Shimauchi T, Asakawa S, Hirasawa N, et al. 650 Suprabasin-deficient mice show limited but discernible defective barrier in both skin and upper digestive tract. J Invest Dermatol. 2018;138(5):S111.
Pinette JA, Mao S, Millis BA, Krystofiak ES, Faust JJ, Tyska MJ. Brush border protocadherin CDHR2 promotes the elongation and maximized packing of microvilli in vivo. Mol Biol Cell. 2019;30(1):108.
Lutter R, Ravanetti L. Cadherin-related family member 3 (CDHR3) drives differentiation of ciliated bronchial epithelial cells and facilitates rhinovirus C infection, although with a little help. J Allergy Clin Immunol. 2019;144(4):926–7.
Lundström A, Serre G, Haftek M, Egelrud T. Evidence for a role of corneodesmosin, a protein which may serve to modify desmosomes during cornification, in stratum corneum cell cohesion and desquamation. Arch Dermatol Res. 1994;286(7):369.
Oji V, Eckl K-M, Aufenvenne K, Naetebus M, Tarinski T, Ackermann K, et al. Loss of Corneodesmosin leads to severe skin barrier defect, pruritus, and Atopy: unraveling the peeling skin disease. Am J Hum Genet. 2010;87(2):274–81.
Rousseau B, Ge PJ, Ohno T, French LC, Thibeault SL. Extracellular matrix gene expression after vocal fold injury in a rabbit model. Ann Otol Rhinol Laryngol. 2008;117(8):598–603.
Cole T, Brian AW, Geo P, Ali M, Gordon K, Marijke JVB, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28(5):511.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29(1):15–21.
Huang DW, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37(1):1.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25(4):402–8.
Hadley Wickham MA, Bryan J, Chang W, D’Agostino McGowan L, François R, Grolemund G, et al. Welcome to the Tidyverse. J Open Source Softw. 2019;4:43.
Komsta L. Outliers v0.14 R documentation; 2011. Available from: https://www.rdocumentation.org/packages/outliers.
Acknowledgements
The authors acknowledge Jessica Engen and Chenwei Duan for invaluable support to animal husbandry and sample collection, Dr. Mara Varvil for assistance with physical assessment of animals and technical support, Norvin Bruns from the Purdue Biomedical Engineering Machine Shop for assistance in the construction of the environmental chamber, Alison Sorg formally from the Purdue Genomics Core, and Kenneth Price for the schematic illustration of the environmental chamber.
Funding
Funding was provided from R01DC0115545 (National Institutes of Health/National Institute on Deafness and other Communication Disorders). The NIH/NIDCD did not participate in any component of the study design, execution, or analysis.
Author information
Authors and Affiliations
Contributions
TWB was responsible for the conception and design of the study, data acquisition, data analysis and interpretation, and manuscript writing. APS supervised the study, provided resources, reviewed and edited the manuscript. NCN was involved in the design of the study, data analysis, and reviewing and editing the manuscript. SX and JT were responsible for data analysis and interpretation and writing, reviewing, and editing the manuscript. MPS was responsible for conception and supervision of the study, funding, reviewing and editing the manuscript. AC was responsible for conception and supervision of the study, project administration, and reviewing and editing the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All experiments were conducted in accordance with the guidelines of the Purdue Animal Care and Use Committee (Protocol # 1606001428).
Consent for publication
Not applicable.
Competing interests
The authors identity no competing or conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S1.
Complete list of genes identified by RNA-Seq analysis and differentially expressed genes (DEG) identified in the larynges of low humidity exposure group by Cufflinks. Excel table of all DEGs obtained by Cufflinks analysis, including Ensembl gene IDs, statistics, and Biomart annotations. Color code Ensembl_ID: green (FDR < 0.05), yellow (0.05 < FDR ≤ 0.1), red (0.1 < FDR ≤ 0.2).
Additional file 2: Table S2.
Functional enrichment analyses of differentially expressed genes identified in the larynges of low humidity exposure group. Excel table containing the functional annotation enrichments divided into GO categories and KEGG obtained with DAVID (first tab), the functional annotation enrichments obtained by STRING (second tab), and the enrichement data for moderate and low humidity groups by GSEA.
Additional file 3: Table S3.
List of primers used in the RT-qPCR validation.
Additional file 4 Cuffdiff Cmd.
Syntax used for Cufflinks analysis.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Bailey, T.W., dos Santos, A.P., do Nascimento, N.C. et al. RNA sequencing identifies transcriptional changes in the rabbit larynx in response to low humidity challenge. BMC Genomics 21, 888 (2020). https://doi.org/10.1186/s12864-020-07301-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-020-07301-7