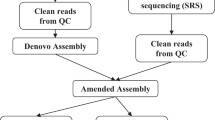
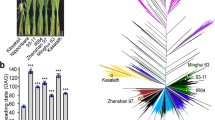
Abstract
Background
Tongil (IR667-98-1-2) rice, developed in 1972, is a high-yield rice variety derived from a three-way cross between indica and japonica varieties. Tongil contributed to the self-sufficiency of staple food production in Korea during a period known as the `Korean Green Revolution'. We analyzed the nucleotide-level genome structure of Tongil rice and compared it to those of the parental varieties.
Results
A total of 17.3 billion Illumina Hiseq reads, 47× genome coverage, were generated for Tongil rice. Three parental accessions of Tongil rice, two indica types and one japonica type, were also sequenced at approximately 30x genome coverage. A total of 2,149,991 SNPs were detected between Tongil and Nipponbare varieties. The average SNP frequency of Tongil was 5.77 per kb. Genome composition was determined based on SNP data by comparing Tongil with three parental genome sequences using the sliding window approach. Analyses revealed that 91.8% of the Tongil genome originated from the indica parents and 7.9% from the japonica parent. Copy numbers of SSR motifs, ORF gene distribution throughout the whole genome, gene ontology (GO) annotation, and some yield-related QTLs or gene locations were also comparatively analyzed between Tongil and parental varieties using sequence-based tools. Each genetic factor was transferred from the parents into Tongil rice in amounts that were in proportion to the whole genome composition.
Conclusions
Tongil was derived from a three-way cross among two indica and one japonica varieties. Defining the genome structure of Tongil rice demonstrates that the Tongil genome is derived primarily from the indica genome with a small proportion of japonica genome introgression. Comparative gene distribution, SSR, GO, and yield-related gene analysis support the finding that the Tongil genome is primarily made up of the indica genome.
Similar content being viewed by others
Background
Rice (Oryza sativa L.) is a staple food for more than half of the world's population, providing about 19 percent of the world's and 29 percent of Asia's caloric supply (IRRI [2009]). Although demands on the nutritional and industrial functionality of rice are increasing, especially to improve human health and quality of life, improving the yield potential of rice is still a major challenge for rice breeders, who must address the rapid growth of the world population along with dramatic reductions in the amount of cultivated land (Khush [1999]), as well as environmental challenges (Nelson, International Food Policy Research Institute [2009]). Asian varieties of cultivated rice include two major subspecies, O. sativa indica and O. s. japonica, which are differentiated based on morphological and physiological characteristics and geographical distribution (Morishima and Oka [1981]; Sano and Morishima [1992]). O. s. indica cultivars have higher genetic diversity (Lu et al. [2002]), a broader cultivation range, and stronger resistance to prominent diseases and insect pests compared to O. s. japonica cultivars (Chung and Heu [1991]). Inter-subspecific hybridization between indica and japonica rice cultivars may enrich allelic variation and facilitate hybrid vigor by creating new genetic recombinations (Cheng et al. [2007]). In spite of these advantages, the introduction of desirable indica traits into the japonica variety has not been successful due to reproductive barriers and the incorporation of undesirable characteristics, such as low eating quality for people who prefer the taste of japonica rice (Chung and Heu [1991]).
Tongil rice (IR667-98-1-2) is the first semi-dwarf variety obtained by a three-way cross of indica/japonica varieties as part of a collaborative research project between the International Rice Research Institute (IRRI) and the government of South Korea (Figure 1). The development of Tongil rice resulted in a significant yield increase from 4 to 5 t ha-1, corresponding to a 30% yield increase relative to the leading japonica varieties grown in Korea (Chung and Heu [1980]). After the introduction of Tongil rice in 1972, Korean rice production significantly increased and the South Korean government announced the achievement of agricultural self-sufficiency (the so-called `Green Revolution') in 1977. However, the genome characterization and structure of Tongil rice have never been analyzed.
Morphological comparison of Tongil and parental lines. From left to right: Tongil, Yukara, IR8, and TN1. (A) The plant architecture of Tongil, its japonica parent (Yukara), and its indica parents (IR8 and TN1). (B) The panicle phenotype of Tongil and its parents. (C) The grain shapes and brown rice shapes of Tongil and its parents. Scale bars are included in each panel.
Rice is a useful model crop for studying genome structure due to its relatively small genome. Furthermore, its genetic and physical data have been extensively analyzed by the International Rice Genome Sequencing Project (IRGSP) (International Rice Genome Sequencing P [2005]). The recent improvement of next-generation sequencing (NGS) technology has enabled high-throughput genoty** and elucidation of genome structures of various rice cultivars (Huang et al. [2009]; Huang et al. [2012]). Most sequence-based rice genome analyses are based on DNA polymorphisms, single nucleotide polymorphisms (SNPs) and insertion-deletions (InDels). SNP detection is the first step for comparing DNA variation and is an effective tool to elucidate genome structure and composition (Feltus et al. [2004]; McNally et al. [2009]; Shen et al. [2010]; Chen et al. [2014]).
In this study, we sequenced the whole genomes of Tongil rice (Oryza sativa L.) and its parental varieties to analyze the genome structure of Tongil in detail and to identify regions of the indica and japonica parental genomes that introgressed in the Tongil genome. In addition, we analyzed previously reported yield-related genes (Gn1a, Ghd7, sd1, GS3 and qSW5), SSRs, GO annotation, and other genetic characteristics of the Tongil genome.
Results
Genome structure of Tongil
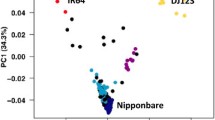
The whole genomes of Tongil and its three parental varieties, Yukara, IR8, and TN1 (Taichung Native 1), were sequenced on the Illumina-GAII platform. A large number of short reads were mapped onto the reference Nipponbare genome and then assembled into a consensus sequence. A total of 199,543,820 reads of the Tongil genome, corresponding to 17,339,883,560 bp (17.3 Gb), were generated, representing a 47-fold sequence depth and covering 88.8% of the Nipponbare pseudomolecules (Table 1 and Additional file 1: Table S1). We detected a total of 2,149,991 SNPs between Tongil and Nipponbare sequences (Additional file 2: Table S2). The two indica parents of Tongil, IR8 and TN1, had 6.22 and 6.04 SNPs per kb, respectively, whereas the japonica parent of Tongil, Yukara, had only 0.49 SNP per kb (Additional file 2: Table S2). Using the SNP data sets from Tongil and its parents, we defined the genomic origins of regions of the Tongil genome by SNP calling (Additional file 3: Figure S1; Additional file 4: Table S3; see also the SNP calling section in the Materials and Methods), and then performed a SEG-Map analysis (Zhao et al. [2010b]) of Tongil (Figure 2). The whole genome of Tongil consisted of an average contribution of 91.8% from indica, 7.9% from japonica, and 0.3% unknown (i.e., not defined as indica or japonica regions) (Figure 2 and Table 2). The contribution of indica to the Tongil genome varied across chromosomes, from 74% (Chr. 2) to 100% (Chr. 12). A relatively high proportion of the japonica genome was found on chromosomes 1, 2, and 3, whereas the japonica sequences were barely detectable on chromosomes 8 and 12. In addition, there were no differences in gene density between the indica- and japonica-derived genome regions of Tongil (Figure 2 and Table 2).
Gene distribution and gene ontology analysis of Tongil
We analyzed the gene content of Tongil to understand the relationship between the composition of the genome and genes (open reading frames: ORFs), and also to elucidate the distribution of indica- and japonica- originated genes (alleles) within the Tongil genome. The gene distribution ratio according to indica or japonica genome composition was similar to the genome distribution ratio (Table 2 and Additional file 5: Table S4). The origins of genes from the indica and japonica parents were 88.3% and 11.4%, respectively, suggesting that the average gene composition was similar to the genome composition ratio of Tongil, although the distribution of parental origin varied across chromosomes. We performed gene ontology (GO) analysis of the Tongil genome according to three categories to identify biological patterns using a list of genes derived from indica, japonica, and unknown genomes: cellular components, molecular functions, and biological processes (Additional file 6: Figure S2; Additional file 7: Figure S3; Additional file 8: Figure S4). The results of GO analysis revealed that the average contribution of the indica or japonica genome to each GO category was almost identical to the gene and genome distribution ratios. O. s. indica and O. s. japonica contributed 86.8% and 12.7% of the cellular components, 87.4% and 12.2% of the molecular functions, and 87.3% and 12.2% of the biological processes, respectively, to the Tongil genome. However, in the `molecular functions' category, all 17 genes related to channel regulator activity were derived from indica regions, whereas all adhesion-related genes in the biological processes category were derived solely from japonica regions.
Simple sequence repeats (SSRs) in the Tongil genome
A total of 177 distinctive motif families were annotated on the Tongil genome (Additional file 9: Figure S5; Additional file 10: Figure S6). Di-nucleotide repeats were predominant among the classified repeats, and AT/TA repeats were the most abundant motifs in both indica- (29.09%) and japonica- derived (21.8%) regions within the Tongil genome. The next most abundant motif relative to AT/TA was CT/GA, and CGC was the most abundant motif among tri-nucleotide repeats. The di-, tri-, and tetra-nucleotide repeat patterns were different from that of the reference Nipponbare genome (McCouch et al. [2002]; Zhou et al. [2005]), and also differed from that of wheat (Weng et al. [2005]). A total of 90.1% of SSR motifs in the Tongil genome were from indica, 9.6% were from japonica, and 0.3% were from an unknown genome (Additional file 10: Figure S6).
Distribution of yield-related genes in the Tongil genome
One of the most important aims of this study was to explore which regions of the indica and japonica parental genomes have introgressed into the Tongil variety to provide its high-yield potential. Tongil is morphologically characterized by short plant height, lodging resistance, open plant architecture, medium-long erect leaves, thick leaf sheaths and culms, relatively long panicles, and easily shattered grain (Chung and Heu [1980]) (Figure 1). Although these phenotypic characteristics affect Tongil's high-yield potential, to date we have no molecular genetic evidence regarding the nature of these traits, with the exception of semi-dwarf gene 1 (sd1) (Chung and Heu [1980]). Therefore, we analyzed several well-characterized genes associated with high yield potential in the Tongil genome: sd1 (Nagano et al. [2005]; Sasaki et al. [2002]; Monna et al. [2002]), Ghd7 (Liu et al. [2009]). In each window, the proportion of SNPs originating from each parent was examined for genotype calling. Huang et al. determined optimum window size by calculating the probability of finding a specific number of japonica SNPs in a window based on SNP error rates. Recent improvements in sequencing technology, however, resulted in fewer errors in SNP identification. Thus, the method suggested by Huang et al. ([2009]) was not directly applicable in this study. Even with a window size of 2, for example, calling accuracy could reach 99.99%. Instead of calculating this probability, the optimum window size was determined iteratively by comparing the portion of japonica SNPs (O) and the portion of the genome originating from japonica (P). Tongil was resequenced to obtain SNPs originating from its parents and to calculate the percentage of japonica SNPs in each chromosome. SEG-Map software (Huang et al. [2009]) was also used for genotype calling on each chromosome. Because the optimum window size was unknown, a range of window sizes from 1 to 199 was used. Then, the Nash-Sutcliffe efficiency (E) between O and P was calculated as follows:
Here, an individual chromosome is denoted by i. The average percentage of japonica SNPs on each chromosome is denoted by Om. The optimal window size was defined as that with a maximum value of E; values of E ranged from -29 to 0.963. This maximum value of E occurred with a window size of 9. The percentage of indica SNPs was at its second highest (0.966) with a window size of 9. At a window size of 10, the E value dropped rapidly for japonica SNPs (0.037) and indica SNPs (-0.018). Thus, a window size of 9 was selected as the optimum for data analysis (Additional file 7: Figure S3).
Parental genome composition of Tongil
We compared DNA variation between the parental and Tongil genomes. Genomic regions originating from the japonica (Yukara) and indica (TN1 or IR8) parents were identified by comparing the Tongil genome sequence to parental sequences. Estimated indica and japonica regions in the Tongil genome sequence were calculated based on the methods of Zhao et al. (Zhao et al. [2010a]).
Gene ontology and classification
Annotated Nipponbare reference genes were classified based on parental origin in the Tongil genome and assigned to the three main GO-term categories (cellular component, molecular function, and biological process) using BLAST2GO software (www.blast2go.com) (Conesa et al. [2005]).
Simple sequence repeats (SSRs)
SSR loci were searched using SSR search software (Initiative [2000]) and classified with respect to their parental origin.
Authors' contributions
BK and HK conceived of the study and participated in its design. IC and BC performed bioinformatic analysis and data processing. BK and JL collected samples and phenotype data. DK, BK, GL, and JS analyzed the data and helped to draft the manuscript. TY, KK, DK, and JC helped to revise the manuscript. All authors read and approved the final manuscript.
Additional files
Abbreviations
- SNP:
-
Single nucleotide polymorphism
- NGS:
-
Next generation sequencing
- IRGSP:
-
International rice genome sequencing project
- SSR:
-
Simple sequence repeat
- GO:
-
Gene ontology
- SEG-map:
-
Sequencing enabled genoty** for map** recombination populations
References
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, Angeles ER, Qian Q, Kitano H, Matsuoka M: Cytokinin oxidase regulates rice grain production. Science 2005, 309(5735):741–745. doi: 10.1126/science.1113373 doi: 10.1126/science.1113373 10.1126/science.1113373
Chen H, ** array for rice biology and molecular breeding. Mol Plant 2014, 7(3):541–553. doi:10.1093/mp/sst135 doi:10.1093/mp/sst135 10.1093/mp/sst135
Cheng SH, Zhuang JY, Fan YY, Du JH, Cao LY: Progress in research and development on hybrid rice: a super-domesticate in China. Ann Bot 2007, 100(5):959–966. doi:10.1093/aob/mcm121 doi:10.1093/aob/mcm121 10.1093/aob/mcm121
Chung GS, Heu MH: Status of japonica-indica hybridization in Korea. In Selected papers from the 1979 International Rice Research. International Rice Research Institute, Manila; 1980:135–152.
Chung GS, Heu MH: Improvememt of Tongil-Type Rice Cultivars from Indica Japonica Hybridization in Korea. In Biotechnology in Agriculture and Forestry 14. Springer-Verlag Berlin Heidelberg, Germany; 1991:105–112.
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M: Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21(18):3674–3676. doi:10.1093/bioinformatics/bti610 doi:10.1093/bioinformatics/bti610 10.1093/bioinformatics/bti610
Fan C, **ng Y, Mao H, Lu T, Han B, Xu C, Li X, Zhang Q: GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet 2006, 112(6):1164–1171. doi:10.1007/s00122–006–0218–1 doi:10.1007/s00122-006-0218-1 10.1007/s00122-006-0218-1
Feltus FA, Wan J, Schulze SR, Estill JC, Jiang N, Paterson AH: An SNP resource for rice genetics and breeding based on subspecies indica and japonica genome alignments. Genome Res 2004, 14(9):1812–1819. doi:10.1101/gr.2479404 doi:10.1101/gr.2479404 10.1101/gr.2479404
Harushima Y, Kurata N, Yano M, Nagamura Y, Sasaki T, Minobe Y, Nakagahra M: Detection of segregation distortions in an indica-japonica rice cross using a high-resolution molecular map. Theor Appl Genet 1996, 92(2):145–150. doi:10.1007/BF00223368 doi:10.1007/BF00223368 10.1007/BF00223368
Huang X, Feng Q, Qian Q, Zhao Q, Wang L, Wang A, Guan J, Fan D, Weng Q, Huang T, Dong G, Sang T, Han B: High-throughput genoty** by whole-genome resequencing. Genome Res 2009, 19(6):1068–1076. doi:10.1101/gr.089516.108 doi:10.1101/gr.089516.108 10.1101/gr.089516.108
Huang X, Kurata N, Wei X, Wang ZX, Wang A, Zhao Q, Zhao Y, Liu K, Lu H, Li W, Guo Y, Lu Y, Zhou C, Fan D, Weng Q, Zhu C, Huang T, Zhang L, Wang Y, Feng L, Furuumi H, Kubo T, Miyabayashi T, Yuan X, Xu Q, Dong G, Zhan Q, Li C, Fujiyama A, Toyoda A, Lu T, Feng Q, Qian Q, Li J, Han B: A map of rice genome variation reveals the origin of cultivated rice. Nature 2012, 490(7421):497–501. doi:10.1038/nature11532 doi:10.1038/nature11532 10.1038/nature11532
Initiative TAG: Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 2000, 408(6814):796–815. 10.1038/35048692
IRRI (2009) IRRI World Rice Statistic. , [http://ricestat.irri.org:8080/wrs2/entrypoint.htm] IRRI (2009) IRRI World Rice Statistic.
IRRI (2009) IRRI World Rice Statistic. , [http://ricestat.irri.org:8080/wrs2/entrypoint.htm] IRRI (2009) IRRI World Rice Statistic.
Khush GS: Green revolution: preparing for the 21st century. Genome 1999, 42(4):646–655. 10.1139/g99-044
Li Q, Li L, Yang X, Warburton ML, Bai G, Dai J, Li J, Yan J: Relationship, evolutionary fate and function of two maize co-orthologs of rice GW2 associated with kernel size and weight. BMC Plant Biol 2010, 10: 143. doi:10.1186/1471–2229–10–143 doi:10.1186/1471-2229-10-143 10.1186/1471-2229-10-143
Lin SY, Ikehashi H, Yanagihara S, Kawashima A: Segregation distortion via male gametes in hybrids between Indica and Japonica or wide-compatibility varieties of rice (Oryza sativa L). Theoret Appl Genetics 1992, 84(7-8):812–818. doi:10.1007/BF00227389 doi:10.1007/BF00227389
Liu T, Liu H, Zhang H, **ng Y (2013) Validation and characterization of Ghd7.1, a major QTL with pleiotropic effects on spikelets per panicle, plant height, and heading date in rice (Oryza sativa L.). J Integr Plant Biol, doi:10.1111/jipb.12070.
Lu BR, Zheng KL, Qian HR, Zhuang JY: Genetic differentiation of wild relatives of rice as assessed by RFLP analysis. Theor Appl Genet 2002, 106(1):101–106. doi:10.1007/s00122–002–1013–2 doi:10.1007/s00122-002-1013-2
McCouch SR, Teytelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, ** of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 2002, 9(6):199–207. 10.1093/dnares/9.6.199
McNally KL, Childs KL, Bohnert R, Davidson RM, Zhao K, Ulat VJ, Zeller G, Clark RM, Hoen DR, Bureau TE, Stokowski R, Ballinger DG, Frazer KA, Cox DR, Padhukasahasram B, Bustamante CD, Weigel D, Mackill DJ, Bruskiewich RM, Ratsch G, Buell CR, Leung H, Leach JE: Genomewide SNP variation reveals relationships among landraces and modern varieties of rice. Proc Natl Acad Sci U S A 2009, 106(30):12273–12278. doi:10.1073/pnas.0900992106 doi:10.1073/pnas.0900992106 10.1073/pnas.0900992106
Miura K, Ikeda M, Matsubara A, Song XJ, Ito M, Asano K, Matsuoka M, Kitano H, Ashikari M: OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat Genet 2010, 42(6):545–549. doi:10.1038/ng.592 doi:10.1038/ng.592 10.1038/ng.592
Monna L, Kitazawa N, Yoshino R, Suzuki J, Masuda H, Maehara Y, Tanji M, Sato M, Nasu S, Minobe Y: Positional cloning of rice semidwarfing gene, sd-1: rice "green revolution gene" encodes a mutant enzyme involved in gibberellin synthesis. DNA Res 2002, 9(1):11–17. 10.1093/dnares/9.1.11
Morishima H, Oka HI: Phylogenetic Differentiation of Cultivated Rice.22. Numerical Evaluation of the Indica-Japonica Differentiation. Jpn J Breed 1981, 31(4):402–413. 10.1270/jsbbs1951.31.402
Nagano H, Onishi K, Ogasawara M, Horiuchi Y, Sano Y: Genealogy of the "Green Revolution" gene in rice. Genes Genet Syst 2005, 80(5):351–356. 10.1266/ggs.80.351
Climate change: impact on agriculture and costs of adaptation. Food policy report, International Food Policy Research Institute, Washington, D.C; 2009.
Sano R, Morishima H: Indica-Japonica Differentiation of Rice Cultivars Viewed from Variations in Key Characters and Isozymes, with Special Reference to Landraces from the Himalayan Hilly Areas. Theor Appl Genet 1992, 84(3-4):266–274.
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, Ishiyama K, Saito T, Kobayashi M, Khush GS, Kitano H, Matsuoka M: Green revolution: a mutant gibberellin-synthesis gene in rice. Nature 2002, 416(6882):701–702. doi:10.1038/416701a doi:10.1038/416701a 10.1038/416701a
Shen Y, Wan Z, Coarfa C, Drabek R, Chen L, Ostrowski EA, Liu Y, Weinstock GM, Wheeler DA, Gibbs RA, Yu F: A SNP discovery method to assess variant allele probability from next-generation resequencing data. Genome Res 2010, 20(2):273–280. doi:10.1101/gr.096388.109 doi:10.1101/gr.096388.109 10.1101/gr.096388.109
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M: Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet 2008, 40(8):1023–1028. doi:10.1038/ng.169 doi:10.1038/ng.169 10.1038/ng.169
Song XJ, Huang W, Shi M, Zhu MZ, Lin HX: A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet 2007, 39(5):623–630. doi: 10.1038/ng2014 doi: 10.1038/ng2014 10.1038/ng2014
Takano-Kai N, Jiang H, Kubo T, Sweeney M, Matsumoto T, Kanamori H, Padhukasahasram B, Bustamante C, Yoshimura A, Doi K, McCouch S: Evolutionary history of GS3, a gene conferring grain length in rice. Genetics 2009, 182(4):1323–1334. doi:10.1534/genetics.109.103002 doi:10.1534/genetics.109.103002 10.1534/genetics.109.103002
Weng DX, Xu SC, Lin RM, Wan AM, Li JP, Wu LR: Microsatellite marker linked with stripe rust resistant gene Yr9 in wheat. Yi Chuan Xue Bao 2005, 32(9):937–941.
Xue W, **ng Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X, Zhang Q: Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 2008, 40(6):761–767. doi:10.1038/ng.143 doi:10.1038/ng.143 10.1038/ng.143
Yan S, Zou G, Li S, Wang H, Liu H, Zhai G, Guo P, Song H, Yan C, Tao Y: Seed size is determined by the combinations of the genes controlling different seed characteristics in rice. Theor Appl Genet 2011, 123(7):1173–1181. doi:10.1007/s00122–011–1657-x doi:10.1007/s00122-011-1657-x 10.1007/s00122-011-1657-x
Zhao K, Wright M, Kimball J, Eizenga G, McClung A, Kovach M, Tyagi W, Ali ML, Tung CW, Reynolds A, Bustamante CD, McCouch SR: Genomic diversity and introgression in O. sativa reveal the impact of domestication and breeding on the rice genome. PLoS One 2010, 5(5):e10780. 10.1371/journal.pone.0010780
Zhao QA, Huang XH, Lin ZX, Han B: SEG-Map: A Novel Software for Genotype Calling and Genetic Map Construction from Next-generation Sequencing. Rice 2010b, 3(2-3):98–102. doi:10.1007/s12284–010–9051-x doi:10.1007/s12284-010-9051-x 10.1007/s12284-010-9051-x
Zhou LJ, Ao GH, Wu XJ, Li SG: SSR markers linked with early stability in rice. Yi Chuan Xue Bao 2005, 32(8):837–845.
Acknowledgments
This work was supported by a grant from the Next-Generation BioGreen 21 Program (Plant Molecular Breeding Center No. PJ008125), Rural Development Administration, Republic of Korea.
Accession codes
Raw sequence data obtained in our study have been submitted to the NCBI Short Read Archive (Takano-Kai et al.) with the following accession numbers: Tongil [SRA: SRR923809, SRA: SRR923810], IR8 [SRA: SRR921498], TN1 [SRA: SRR921505], and Yukara [SRA: SRR925387].
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors have no potential competing interests to declare.
Electronic supplementary material
12284_2014_22_MOESM3_ESM.docx
Additional file 3: Figure S1.: Determination of window size followed by E-value calculation. The x-axis is the window size and the y-axis is the calculated E-value. (DOCX 24 KB)
12284_2014_22_MOESM6_ESM.docx
Additional file 6: Figure S2.: GO analysis according to the cellular components category of Tongil genes corresponding to indica/japonica sequences. (DOCX 22 KB)
12284_2014_22_MOESM7_ESM.docx
Additional file 7: Figure S3.: GO analysis according to the molecular functions category of Tongil genes corresponding to indica/japonica sequences. (DOCX 22 KB)
12284_2014_22_MOESM8_ESM.docx
Additional file 8: Figure S4.: GO analysis according to the biological processes category of Tongil genes corresponding to indica/japonica sequences. (DOCX 22 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, B., Kim, DG., Lee, G. et al. Defining the genome structure of `Tongil' rice, an important cultivar in the Korean "Green Revolution". Rice 7, 22 (2014). https://doi.org/10.1186/s12284-014-0022-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12284-014-0022-5