Abstract
Emerging machine-learned models have enabled efficient and accurate prediction of compound formation energy, with the most prevalent models relying on graph structures for representing crystalline materials. Here, we introduce an alternative approach based on sparse voxel images of crystals. By develo** a sophisticated network architecture, we showcase the ability to learn the underlying features of structural and chemical arrangements in inorganic compounds from visual image representations, subsequently correlating these features with the compounds’ formation energy. Our model achieves accurate formation energy prediction by utilizing skip connections in a deep convolutional network and incorporating augmentation of rotated crystal samples during training, performing on par with state-of-the-art methods. By adopting visual images as an alternative representation for crystal compounds and harnessing the capabilities of deep convolutional networks, this study extends the frontier of machine learning for accelerated materials discovery and optimization. In a comprehensive evaluation, we analyse the predicted convex hulls for 3115 binary systems and introduce error metrics beyond formation energy error. This evaluation offers valuable insights into the impact of formation energy error on the performance of the predicted convex hulls.
Similar content being viewed by others
Introduction
Machine learning has emerged as an effective approach for develo** predictive models for high-throughput screening of materials1,2,3,4,5,6,7,8. For example, machine-learned models for formation energy prediction can construct a convex hull for a rapid assessment of the thermodynamic stability of compounds at a fraction of the computation cost and time needed for density functional theory (DFT)-calculated convex hulls with reasonable accuracy9. In materials research, a machine learning model can be characterized by two aspects; the representation of the material as a readable entity (or input) to the learning algorithm and the learning algorithm itself. Several machine learning approaches have investigated a variety of representations as simple as a pool of physicochemical attributes (e.g., atomic number, cohesive energy, band gap, and heat of melting), and composition vectors10,11,12,13,14,15,16,17 up to more advanced graph representations of composition and structure of crystal compounds19,20,21,22,23,24,25. The use of image representations for machine learning, however, has been less explored in the materials research community. Image representation can be especially useful because of the significant advancements that have been made in pattern recognition (or representation learning) of visual images in the field of computer vision (a field of computer science that deals with processing and understanding visual data like images or videos). These advancements are largely because of the evolution towards more sophisticated architectures of convolutional neural networks (e.g., Residual Neural Network (ResNet)26, EfficientNet27, U-Net28) which has enabled adopting increasingly deeper networks. Inspired by this untapped opportunity for materials representation learning, we develop a sparse voxel image representation of crystalline materials that is input into a very deep convolutional neural network (CNN) with a sophisticated architecture inspired by ResNet.
We use the formation energy prediction of crystalline compounds as a platform for demonstrating the performance of our deep-learning model on voxel images of crystals. Formation energy is an ideal platform because large databases of DFT-calculated formation energies are available (e.g., Materials Project29 and AFLOW (Automatic Flow)30), which provide the large amount of data needed for training our deep CNN. Additionally, there are several available machine learning approaches for formation energy prediction with which we compare the performance of our model. We show that our model’s formation energy predictive performance is comparable to the state-of-the-art machine learning models’ prediction. We present a thorough comparison of 3115 binary convex hulls constructed from our model’s formation energy against DFT-calculated binary convex hulls in the Materials Project database. By introducing multiple error metrics for assessing binary convex hulls, we showcase how the error in the formation energy prediction is projected into the performance of a predicted convex hull.
Among machine learning methods for formation energy prediction of crystal compounds, graph neural networks have shown promising performance because the graph data structure can efficiently capture the physical, compositional, and structural information of crystal compounds19,20,21,22,23,24,25. In their pioneering work, ** skip connections). These skip connections enable the transfer of lower-level information from earlier layers to deeper layers, providing better conditioning for the optimization problem and facilitating easier learning26.
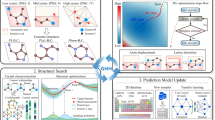
In our approach, we adopt the architecture of residual blocks to construct a 15-layer CNN with 7 skip connections. The overall architecture, as depicted in Fig. 1, consists of a deep CNN followed by a fully connected neural network for the prediction of formation energy using sparse voxel images of crystals. The deep CNN part of the architecture is employed for feature learning of voxel crystal images. These learned features are then flattened and passed as input to the fully connected neural network, which performs the final prediction of the formation energy. In our network design, we deliberately delay the introduction of pooling layers in our CNN. The first pooling layer is introduced only after the fifth convolutional kernel, with subsequent pooling layers added after the eleventh and fifteenth kernels, respectively. A detailed description of our CNN architecture can be found in Methods. In the context of materials representation learning, the use of skip connections in our CNN allows for the bypassing of local atomic features discovered in the shallower layers, while progressively learning more global features of crystal compounds across the layers of the deep network. This hierarchical learning approach facilitates the extraction of relevant abstractions, enabling the model to capture both local and global features within the crystal structures.
Our CNN, inspired by the ResNet architecture described in ref. 26, incorporates slight modifications to better suit our specific task. In contrast to the original design, we choose not to adopt the batch normalization technique in our residual blocks. This decision is based on the observation that batch normalization hampers the training of our CNN, likely due to the intrinsic differences between sparse crystal images and natural images (such as those in ImageNet41). Consequently, the batch normalization process may not yield the intended benefits for our crystal image representation. Furthermore, we adjust the way in which we handle the number of channels within our network. Instead of doubling the number of channels after each convolution layer, as outlined in the original ResNet design, we increase the number of channels, after each pooling, by concatenating the side skip connections with the output of the convolution layer. This alternative approach allows for a more effective utilization of information from both the skip connections and the convolutional layers, promoting better feature representation within our network. By tailoring the ResNet-inspired architecture to the characteristics of our crystal images, we optimize the training process and enhance the performance of our CNN for the specific task of crystal compound formation energy prediction.
Data Sets
We obtained a data set of 139,367 crystal structures along with their corresponding DFT-calculated formation energies (the target variables) from Materials Project (v2021.05.13)29. From this, 15,354 structures are excluded because they either require a high resolution or a large image (more details in Methods). To train our model, we split the data into train (60%), validation (20%), and test (20%) sets. During the data pre-processing stage, we removed 9175 crystal structures from the train set that either contain two atoms occupying the same voxel or have a unit cell that does not fit in the 17-Å cubic box, as described in detail in Methods. During training, we employ data augmentation by randomly rotating each crystal image before feeding it into the model at each epoch (see Supplementary Fig. S1). This technique helps alleviate overfitting (see Supplementary Fig. S4) and enhances the predictive performance of our model. Data augmentation is particularly beneficial as it effectively increases the size of the train data and implicitly enforces the rotation-invariance of crystal compounds with respect to their formation energy, as explained further below. To monitor the training process and prevent overfitting, we use predictions on the validation data. Once the model is trained, we evaluate its overall performance using the test data, as outlined below. In the Discussion section, we delve into the significance of data augmentation and skip connections in our CNN architecture, highlighting their role in improving the model’s performance.
Formation Energy Prediction Assessment
In this section, we examine the performance of our model’s prediction. As detailed in Methods, we employ an ensemble averaging technique for predicting the formation energy. Figure 2a shows the parity plot of the formation energy prediction of our model against the DFT-calculated formation energies on both the train and test sets. The results indicate an MAE of 0.042 eV per atom and 0.046 eV per atom on the train and test sets, respectively. Over 89% of the samples in the test set exhibit absolute errors below 0.1 eV per atom, and only about 2% of the samples have absolute errors exceeding 0.2 eV per atom (see Supplementary Fig. S2b). The formation energy prediction error (i.e., predicted formation energy - DFT formation energy) shows a slightly positive skew normal distribution with a median and mean value of 0.003 eV per atom and -0.003 eV per atom on the test set (see Supplementary Fig. S2b). As shown in Fig. 2b, c, our model tends to exhibit higher errors for crystal compounds with more positive and larger formation energies. This trend has also been observed in other studies16,42. To exemplify this trend, we analyze four equally populated subsets of our test set sorted by the formation energy with respective formation energy ranges of (−4.47, −2.39), (−2.39, −1.47), (−1.47, −0.46), and (−0.46, 5.33) eV per atom with calculated MAEs of 0.037, 0.039, 0.046, and 0.064 eV per atom, respectively. The relatively diminished prediction performance observed for larger, positive-value ranges of formation energy can be attributed to an inherent bias in the existing dataset. The data available in the Materials Project predominantly comprises chemically stable structures characterized by negative formation energies. In contrast, the occurrence of chemically unstable crystal structures with positive formation energies remains a minority within this dataset. Notably, less than 10% of all samples possess positive formation energy (see Supplementary Fig. S2a). Pandey et al.43 have elucidated how this disparity in data distribution impacts the model’s predictive capabilities.
a The parity plot for samples in the train and test sets. The MAE of formation energy prediction for the test and train data is reported in the legend. b, c Distribution of the prediction error of test data over different ranges of formation energy. b Box and whisker representation of prediction error (i.e., predicted Ef - DFT Ef) for different intervals of DFT formation energy. The left side, middle line, and right side of each box show respectively the first quartile, median, and third quartile of the error. The whisker line shows the minimum and maximum of the error. c The scatter plot of samples in the test set showing the DFT formation energy versus prediction error.
We conducted a comparative analysis of our model’s predictive performance with state-of-the-art machine learning models, including ElemNet16 and Roost (Representation Learning from Stoichiometry)17 as the best models based on compositional features, and ALIGNN23 and CGCNN54 employ skip connections. Traditionally, skip connections are recognized for their role in alleviating optimization challenges by producing smoother loss functions, facilitating easier training52. However, our work sheds light on an additional aspect of skip connections beyond their optimization benefits. We demonstrate that skip connections serve as a mechanism to capture the essential physicochemical information at different levels. By allowing the outputs of different layers (both shallow and deep) to bypass through identity map**, skip connections enable the network to leverage local atomic fingerprints from shallower layers while simultaneously learning abstract, generalized features from deeper layers. In this way, skip connections facilitate the integration of both local and global information, leading to improved performance in formation energy prediction.
Methods
Data collection and voxel image preparation
We gather crystal structure information in Crystallographic Information File (CIF) format and the corresponding DFT-calculated formation energies from the Materials Project database (v2021.05.13)29. To extract the structural information, we utilize the Atomic Simulation Environment (ASE) package55. Our in-house Python code is then employed to generate sparse voxel images of the crystals. In the voxelization process, we repeat the crystal unit cell (cubic or non-cubic) in space to fill a cubic box with an edge size of 17 Å. We eliminate a crystal structure if its unit cell does not fit in the cubic box. The box is then voxelized using a 32 × 32 × 32 grid, resulting in images with dimensions of 32 × 32 × 32 voxels. To ensure that each voxel contains at most one atom, we set the minimum interatomic distance to be greater than the diagonal of a voxel, dv, calculated as \({d}_{v}=(17/32)\times \sqrt{3}=0.92\) Å. Consequently, crystal structures with minimum interatomic distances smaller than 0.92 Å are filtered out. The 3D sparse voxel images of crystals are color-coded using three channels, similar to an RGB image. These channels represent the normalized atomic number, group number, and period number. For lanthanides and actinides, we assign a group number of 3.5. During training, to introduce variability and enhance generalization, we apply a random rotation to each crystal image at each epoch. Rather than applying a direct rotation to the unit cell and subsequently executing the computationally intensive task of filling the 17 Å box - a method which becomes intractably repetitive - we initially construct a larger ‘encompassing’ box with an edge equal to the diagonal of the 17-Å cubic box. During the data pre-processing stage, we fill the larger box by replicating the crystal unit cell in all directions only once. Consequently, whenever an instance of an crystal structure input is requested, either for training or prediction, we perform a random rigid-body rotation to the larger box, while the 17-Å box remains unchanged and consistently populated after each rotation. Thereafter, we perform the voxelization of the 17-Å box to generate the final sparse voxel images. Supplementary Fig. S1 visually details the rotation methodology.
Convolutional neural network
We develop a 15-convolutional-layer network consisting of 7 residual blocks and 3 average pooling layers, followed by a fully connected neural network (see Supplementary Fig. S11). Each residual block consists of two convolutional layers, each followed by a rectified linear unit (ReLU) activation layer and a skip connection that connects the beginning of the block to its end. In each convolutional layer, we use a kernel of size 3 and padding of type SAME with stride 1 to ensure that the filter is applied to all the voxels of the input. To merge a skip connection (i.e., side stream) with the mainstream coming from the convolutional layer, we either use addition or concatenation. We use the concatenation of outputs only before a pooling layer in order to double the number of channels while reducing the image size during pooling. The addition of outputs is used elsewhere as the method of merging in the residual blocks.
The deep convolutional network consists of three distinct segments, each containing a different image size and ending with a pooling layer. The first segment consists of a single convolutional layer, followed by an activation layer. In this layer, we increase the number of channels from 3 to 32. This single layer is followed by two residual blocks, each consisting of two convolutional and activation layers, outputting 32 channels. We utilize concatenation to combine the outputs of the mainstream and skip connection, rendering the number of channels of the output of this segment equal to 64. This segment ends with an average pooling layer, reducing the image size by half (16 × 16 × 16). The second segment consists of three residual blocks, followed by an average pooling. The images passing through this segment have 64 channels, and at the end of the segment, their size is reduced by half (8 × 8 × 8) and their channels are doubled (128). The last segment consists of two residual blocks and an average pooling layer, but in this case, the last block uses addition instead of concatenation, kee** the channels as 128 and reducing the size to 4 × 4 × 4. A detailed schematic of the network is shown in Supplementary Fig. S11.
The last pooling layer is flattened to a vector of size (4 × 4 × 4 × 128 = 8192) and is connected to a fully connected network with a node architecture of 16-16-1 with linear activation functions. The Keras package56 is used to build and train this network. The 3D images of the train set are randomly rotated in 3D space and input to the network for 500 epochs in batches of size 32. The mean squared error (MSE) is used as the loss function. To train the network, we use the Adam optimizer with a learning rate of 0.001, the exponential decay rates of 0.9 and 0.999 for the first and second moment estimates, respectively, and a machine precision threshold (or ϵ) of 1e-07.
Rotational ensemble averaging
Once the model is trained, we employ an ensemble averaging method for predictiong the formation energy. Once a crystal sample is input into the trained model, a ensemble of 50 randomly rotated instances of the sample is generated and the formation energy prediction is averaged over the ensemble. The ensemble averaging methods improves the prediction accuracy and robustness of our model, as detailed in the Results section Fig. 3, and Supplementary Figs. S6,S7.
Error metrics
The evaluation of the formation energy prediction and the constructed convex hull is performed using the following error metrics:
Formation Energy Mean Absolute Error (MAE): The MAE is calculated using the formula:
where yi represents the true formation energy of sample i (DFT-calculated formation energy obtained from the Materials Project database), \({\hat{y}}_{i}\) corresponds to the model’s prediction of the formation energy for sample i, and the sum runs over total of n samples. When computing the MAE for a binary convex hull prediction, only crystal compounds (or samples) from that specific binary system are included.
Depth error for Convex Hull: The depth error for the convex hull measures the difference in the confined area between the predicted and true convex hulls, and is defined as:
where Apredicted and Atrue represent the areas enclosed by the predicted and true (or DFT-calculated) convex hulls, respectively.
Accuracy of Convex Hull Prediction: The accuracy of the convex hull prediction is calculated as the percentage of correctly predicted crystal samples on the hull with respect to the crystal samples on the DFT-calculated hull. In other words, the hull accuracy measures the percentage of predictions on the hull that matches the DFT-calculated samples on the hull. Accordingly, if our model mistakenly predicts a crytal sample to be on the hull while the DFT-calculated sample is above the hull, the hull accuracy measure will not be affected (e.g., see Fig. 4a).
Data availability
The data developed or used by this study is available on our GitHub repository at: https://github.com/kadkhodaei-research-group/XIE-SPP/tree/main/training/formation-energy/data_sets.
Code availability
The codes developed or utilized in this study are openly accessible to support transparency and facilitate further research. They can be found in our GitHub repository at: https://github.com/kadkhodaei-research-group/XIE-SPP.
References
Butler, K. T., Davies, D. W., Cartwright, H., Isayev, O. & Walsh, A. Machine learning for molecular and materials science. Nature 559, 547–555 (2018).
Pilania, G. Machine learning in materials science: from explainable predictions to autonomous design. Comput. Mater. Sci. 193, 110360 (2021).
Rodrigues, J. F., Florea, L., de Oliveira, M. C. F., Diamond, D. & Oliveira, O. N. Big data and machine learning for materials science. Discov. Mater. 1, 12 (2021).
Schmidt, J., Marques, M. R. G., Botti, S. & Marques, M. A. L. Recent advances and applications of machine learning in solid-state materials science. NPJ Comput. Mater. 5, 83 (2019).
Stanev, V., Choudhary, K., Kusne, A. G., Paglione, J. & Takeuchi, I. Artificial intelligence for search and discovery of quantum materials. Commun. Mater. 2, 105 (2021).
Wei, J. et al. Machine learning in materials science. InfoMat 1, 338–358 (2019).
Morgan, D. & Jacobs, R. Opportunities and challenges for machine learning in materials science. Ann. Rev. Mater. Res. 50, 71–103 (2020).
Gu, G. H., Noh, J., Kim, I. & Jung, Y. Machine learning for renewable energy materials. J. Mater. Chem. A 7, 17 096–17 117 (2019).
Bartel, C. J. et al. A critical examination of compound stability predictions from machine-learned formation energies. NPJ Comput. Mater. 6, 97 (2020).
Faber, F., Lindmaa, A., von Lilienfeld, O. A. & Armiento, R. Crystal structure representations for machine learning models of formation energies. Int. J. Quantum Chem. 115, 1094–1101 (2015).
Ward, L. et al. Including crystal structure attributes in machine learning models of formation energies via voronoi tessellations. Phys. Rev. B 96, 024104 (2017).
Meredig, B. et al. Combinatorial screening for new materials in unconstrained composition space with machine learning. Phys. Rev. B 89, 094104 (2014).
Ward, L., Agrawal, A., Choudhary, A. & Wolverton, C. A general-purpose machine learning framework for predicting properties of inorganic materials. NPJ Comput. Mater. 2, 16028 (2016).
Dunn, A., Wang, Q., Ganose, A., Dopp, D. & Jain, A. Benchmarking materials property prediction methods: the matbench test set and automatminer reference algorithm. NPJ Comput. Mater. 6, 138 (2020).
Peterson, G. G. C. & Brgoch, J. Materials discovery through machine learning formation energy. J. Phys. Energy 3(mar), 022002 (2021).
Jha, D. et al. Elemnet: Deep learning the chemistry of materials from only elemental composition. Sci. Rep. 8, 17593 (2018).
Goodall, R. E. A. & Lee, A. A. Predicting materials properties without crystal structure: deep representation learning from stoichiometry. Nat. Commun. 11, 6280 (2020).
**e, T. & Grossman, J. C. Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties. Phys. Rev. Lett. 120, 145301 (2018).
Park, C. W. & Wolverton, C. Develo** an improved crystal graph convolutional neural network framework for accelerated materials discovery. Phys. Rev. Mater. 4, 063801 (2020).
Zhan, H., Zhu, X., Qiao, Z. & Hu, J. Graph neural tree: A novel and interpretable deep learning-based framework for accurate molecular property predictions. Analytica Chimica Acta, p. 340558, (2022). [Online]. Available: https://www.sciencedirect.com/science/article/pii/S0003267022011291.
Kaundinya, P. R., Choudhary, K. & Kalidindi, S. R. Prediction of the electron density of states for crystalline compounds with atomistic line graph neural networks (alignn). JOM 74, 1395–1405 (2022).
Meyer, P. P., Bonatti, C., Tancogne-Dejean, T. & Mohr, D. Graph-based metamaterials: deep learning of structure-property relations. Mater. Design 223, 111175 (2022).
Choudhary, K. & DeCost, B. Atomistic line graph neural network for improved materials property predictions. NPJ Comput. Mater. 7, 185 (2021).
Schütt, K. T., Sauceda, H. E., Kindermans, P.-J., Tkatchenko, A. & Müller, K.-R. Schnet - a deep learning architecture for molecules and materials. J. Chemi. Phys. 148, 241722 (2018).
Chen, C., Ye, W., Zuo, Y., Zheng, C. & Ong, S. P. Graph networks as a universal machine learning framework for molecules and crystals. Chem. Mater. 31, 3564–3572 (2019).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. CoRR, vol. abs/1512.03385, (2015). [Online]. Available: http://arxiv.org/abs/1512.03385.
Tan, M. & Le, Q. V. Efficientnet: Rethinking model scaling for convolutional neural networks. CoRR, vol. abs/1905.11946, 2019. [Online]. Available: http://arxiv.org/abs/1905.11946.
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015, N. Navab, J. Hornegger, W. M. Wells, and A. F. Frangi, Eds.Cham: Springer International Publishing, 2015, pp. 234–241. [Online]. Available: https://doi.org/10.1007/978-3-319-24574-4_28.
Jain, A. et al. Commentary: The materials project: A materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Curtarolo, S. et al. Aflow: An automatic framework for high-throughput materials discovery. Comput. Mater. Sci. 58, 218–226 (2012).
Chen, Z., Li, X. & Bruna, J. Supervised community detection with line graph neural networks. (2017). [Online]. Available: https://arxiv.org/abs/1705.08415.
Choudhary, K. The atomistic line graph neural network. https://github.com/usnistgov/alignn.git, (2021).
Long, T. et al. Constrained crystals deep convolutional generative adversarial network for the inverse design of crystal structures. NPJ Comput. Mater. 7, 66 (2021).
Hoffmann, J. et al. Data-driven approach to encoding and decoding 3-d crystal structures. (2019). [Online]. Available: https://arxiv.org/abs/1909.00949.
Kaundinya, P. R., Choudhary, K. & Kalidindi, S. R. Machine learning approaches for feature engineering of the crystal structure: application to the prediction of the formation energy of cubic compounds. Phys. Rev. Mater. 5, 063802 (2021).
Davariashtiyani, A., Kadkhodaie, Z. & Kadkhodaei, S. Predicting synthesizability of crystalline materials via deep learning. Commun. Mater. 2, 115 (2021).
Kajita, S., Ohba, N., **nouchi, R. & Asahi, R. A Universal 3D voxel descriptor for solid-state material informatics with deep convolutional neural networks. Sci. Rep. 7, 16991 (2017). [Online]. Available: https://doi.org/10.1038/s41598-017-17299-w.
Noh, J. et al. Inverse design of solid-state materials via a continuous representation. Matter 1, 1370–1384 (2019).
Kim, S., Noh, J., Gu, G. H., Aspuru-Guzik, A. & Jung, Y. Generative adversarial networks for crystal structure prediction. ACS Central Sci. 6, 1412–1420 (2020). pMID: 32875082.
Sanchez-Lengeling, B. & Aspuru-Guzik, A. Inverse molecular design using machine learning: generative models for matter engineering. Science 361, 360–365 (2018).
Deng, J. et al. Imagenet: A large-scale hierarchical image database. in 2009 IEEE conference on computer vision and pattern recognition. IEEE, (2009), pp. 248–255. [Online]. Available: https://doi.org/10.1109/CVPR.2009.5206848.
Jiang, Y. et al. Topological representations of crystalline compounds for the machine-learning prediction of materials properties. NPJ Comput. Mater. 7, 28 (2021).
Pandey, S., Qu, J., Stevanović, V., St John, P. & Gorai, P. Predicting energy and stability of known and hypothetical crystals using graph neural network. Patterns 2, 100361 (2021).
Cohen, T. S. & Welling, M. Group equivariant convolutional networks. CoRR, vol. abs/1602.07576, (2016). [Online]. Available: http://arxiv.org/abs/1602.07576.
Thomas, N. et al. Tensor field networks: Rotation- and translation-equivariant neural networks for 3d point clouds. CoRR, vol. abs/1802.08219, (2018). [Online]. Available: http://arxiv.org/abs/1802.08219.
Geiger, M. & Smidt, T. e3nn: Euclidean neural networks. (2022). [Online]. Available: https://arxiv.org/abs/2207.09453.
Smidt, T. E., Geiger, M. & Miller, B. K. Finding symmetry breaking order parameters with euclidean neural networks Phys. Rev. Res. 3 (2021). [Online]. Available: https://doi.org/10.1103/physrevresearch.3.l012002.
Chen, Z. et al. Machine learning on neutron and x-ray scattering and spectroscopies. Chem. Phys. Rev. 2, 031301 (2021).
Cheng, Y. et al. Direct prediction of inelastic neutron scattering spectra from the crystal structure*. Mach. Learning Sci. Technol. 4, 015010 (2023).
Okabe, R. et al. Virtual node graph neural network for full phonon prediction (2023). [Online]. Available: https://arxiv.org/abs/2301.02197.
Batzner, S. et al. E(3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials. Nat. Commun. 13, (2022). [Online]. Available: https://doi.org/10.1038/s41467-022-29939-5.
Li, H., Xu, Z., Taylor, G. & Goldstein, T. Visualizing the loss landscape of neural nets. CoRR, vol. abs/1712.09913, (2017). [Online]. Available: http://arxiv.org/abs/1712.09913.
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. vol. abs/1512.03385, (2015). [Online]. Available: http://arxiv.org/abs/1512.03385.
He, K., Zhang, X., Ren, S. & Sun, J. Identity map**s in deep residual networks. CoRR, vol. abs/1603.05027, (2016). [Online]. Available: http://arxiv.org/abs/1603.05027.
Larsen, A. H. et al. The atomic simulation environment–a python library for working with atoms. Journal of Physics: Condensed Matter 29, 273002 (2017).
Chollet, F. et al. (2015) Keras. [Online]. Available: https://github.com/fchollet/keras.
Acknowledgements
This research is based upon work supported by the National Science Foundation (NSF) under Award Numbers DMR-2119308. We used resources at the Electronic Visualization Laboratory (EVL) at UIC available through the NSF Award CNS-1828265. Additionally, we would like to thank Zahra Kadkhodaie for helpful suggestions regarding the design of the convolutional network.
Author information
Authors and Affiliations
Contributions
A.D. and S.K. conceptulazied the presented idea, analyzed the data and wrote the manuscript. A. D. designed the networks and the computational framework. S.K. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Materials thanks Nathan Szymanski and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Reinhard Maurer and Aldo Isidori.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Davariashtiyani, A., Kadkhodaei, S. Formation energy prediction of crystalline compounds using deep convolutional network learning on voxel image representation. Commun Mater 4, 105 (2023). https://doi.org/10.1038/s43246-023-00433-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43246-023-00433-9
- Springer Nature Limited