Abstract
Tissue specificity is a fundamental property of an organ that affects numerous biological processes, including aging and longevity, and is regulated by the circadian clock. However, the distinction between circadian-affected tissue specificity and other tissue specificities remains poorly understood. Here, using multi-omics data on circadian rhythms in mice, we discovered that approximately 35% of tissue-specific genes are directly affected by circadian regulation. These circadian-affected tissue-specific genes have higher expression levels and are associated with metabolism in hepatocytes. They also exhibit specific features in long-reads sequencing data. Notably, these genes are associated with aging and longevity at both the gene level and at the network module level. The expression of these genes oscillates in response to caloric restricted feeding regimens, which have been demonstrated to promote longevity. In addition, aging and longevity genes are disrupted in various circadian disorders. Our study indicates that the modulation of circadian-affected tissue specificity is essential for understanding the circadian mechanisms that regulate aging and longevity at the genomic level.
Similar content being viewed by others
Introdution
Tissue specificity refers to the unique genes expressed in different tissues. Many heritable diseases are tissue-specific, including aging-associated diseases1. The process of aging has been conceptualized as the greatest risk factor for diseases2, and this process is heterogeneous across different tissues3. Tissue-specific transcriptomic signatures have been reported in aging organs from human to Caenorhabditis elegans4,5,6,7. Recent studies have indicated that tissue specificity is lost during aging, leading to the phenomenon of coordinated global aging behavior across tissues7,8,9.
Circadian regulation has been shown to influence aging and longevity10,11,12. The circadian regulatory process involves several proteins that comprise a negative autoregulatory feedback loop13,14. In mice, the PER and CRY proteins are active in the late evening or at night, while the BMAL1 and CLOCK protein are active during the daytime15. The CLOCK–BMAL1 complex promotes the transcription of Per and Cry through binding to E-boxes. In turn, PER and CRY dimerize and interact with BMAL1-CLOCK complex to repress their own transcription at night14. The deficiency of CLOCK or BMAL1 protein can decrease the lifespan of mice16,17, and aging can impact the amplitude and phase of circadian rhythms18,19,20.
Circadian disturbances such as jet leg, have a significant effect on older mice compared to adult mice21 and can lead to increased mortality22. Shift work increases blood pressure and the risk of cardiovascular disease23. Many genes related to longevity are regulated by circadian genes24,25. Circadian rhythmicity is known to weaken with age, characterized by a decrease in the circadian amplitude, an advance or delay of the circadian phase, and a disturbances in cell and tissue synchronization26,27,28. Studies have shown that controlled feeding regimens can reset circadian rhythms and improve health29,30 and phase-aligned feeding activities can effectively delay aging and promote longevity31.
Tissue specificity can also be regulated by the circadian clock as well32,33,34,35,36 Each of the core circadian transcriptional factors binds to thousands of target genes across the genome14,37,38, coordinating the temporal functional organization of different organs39,40. However, The target genes of these circadian regulatory proteins vary in different organs in mice, as well as in other species like baboon35,36,41,42,43, owing to factors such as tissue-specific DNA binding events mediated by the transcription factor BMAL140,44 and tissue-specific chromatin activities45,46. Meanwhile, the binding targets of other core clock transcription factors, such as CLOCK, PERs, CRYs, also differ tremendously14. Although the suprachiasmatic nucleus (SCN) is understood as the principal circadian pacemaker, the circadian regulation system is present in almost all tissues and cells47,48, suggesting it is involved in the regulation of tissue-specificity as a common molecular mechanism.
Although the relationship between tissue specificity and circadian regulation is well-established, the molecular mechanisms that underlie this connection remain elusive. In the present study, we defined tissue specificity as the differential transcriptomic signatures identified in a given tissue compared to other tissues. To quantify the relationship between circadian clock and tissue specificity, we conducted a large-scale sequencing of multiple peripheral circadian tissues (forebrain, cerebellum, liver, kidney) in both normal and sleep disrupted conditions. We firstly accessed whether tissue specificity and circadian oscillation are linked in both normal and sleep deprivation conditions. We then investigated the temporal organization of tissue-specific genes in liver and kidney across 24 h and identified distinct classes of tissue-specific genes that are affected to varying degrees by circadian rhythms. Based on these classes, we found that tissue-specific genes affected by circadian clock are involved in metabolism-related functions, enriched in specific cell types, and highly enriched certain co-expression modules. Finally, we explored the property of circadian-affected tissue-specific genes, and revealed that they were linked to aging and longevity.
Results
The circadian rhythmicity of gene expression is highly correlated with tissue-specificity
To investigate the relationship between circadian rhythm and tissue specificity at the transcriptomic level, we explored the mouse circadian atlas35. We employed three data analysis methods — the tau index (Tau), specificity measure (SPM) and expression enrichment (EE) — to calculate the tissue specificity of genes expressed in 12 organs. We paid particular attention to how the expression of specific genes varied across different tissues over the 24 h circadian cycle. For each of the 12 organs examined, gene’s tissue specificity score oscillates across different time points (Fig. 1a). A significant positive correlation was detected between the number of tissue-specific genes and circadian genes (R = 0.65, R = 0.84 and 0.73, Fig. 1b, Pearson correlation). Liver and kidney, which were shown to have a higher number of cycling genes compared to other 10 organs, also had the most abundant tissue-specific genes (235 and 465 genes, respectively, compared to 7–119 genes in other organs).
a The heatmaps display the tissue-specificity (measured by SPM) of circadian genes, and the pie charts show the proportion of them in tissue-specific genes (top 10% SPM score). R function pheatmap with the parameter scale = row was used to display the SPM changes of circadian genes. b The number of tissue-specific genes was positively correlated with the number of circadian genes in the 12 examined organs, as seen with tissue-specificity indicators including Tau, EE, and SPM. c Enrichment analysis of circadian Tau genes and rhythmically expressed genes. JTK_CYCLE was used to analyze Tau values of genes at different time points, and genes with Benjamini-Hochberg adjusted P < 0.05 were considered circadian Tau genes. Tissue circadian genes from the mouse circadian transcriptome atlas were also analyzed by JTK_CYCLE. d Enrichment analysis of tissue-specific genes (SPM) and tissue-specific genes (Tau). Tissue-specific genes were identified using the SPM value and Tau value calculated from the average gene expression in the mouse circadian transcriptome atlas and a cutoff value > 0.8. Tissue specific genes (Tau) is classified according to the tissue with the highest gene expression. e Enrichment analysis of biological processes of tissue-specific genes across circadian time in the liver. Both odds ratio and significance level (p) are indicated. f The heatmaps display the expression of enriched biological pathways across circadian time in the liver. g Scatter plots showing the relationship between the amplitudes of circadian genes and tissue specificity under normal conditions. In all four organs, tissue-specificity indicators (such as SPM) from the mouse circadian transcriptome atlas are significantly positively correlated with the amplitude of gene expression of the transcriptome (AMP), even in sleep deprivation conditions. Each dot represents a gene, and the Pearson correlation coefficient and its significance level (P) are displayed in each plot. h Scatter plots showing the relationship between the amplitudes of circadian genes and tissue specificity under sleep deprivation conditions.
Our analysis revealed that the tissue specificity of the identified genes exhibited temporal oscillations at different time points. We utilized JTK_CYCLE to calculate the circadian of gene expression in mouse 12 tissues (Jonckheere-Terpstra-Kendall test, q-value < 0.05). Notably, we found that the genes whose Tau values showed temporal oscillations were significantly enriched with the rhythmic genes of the tissue, with odds ratios greater than 3 and P values less than 2.2 x 10−6 for all tissues analyzed (Fig. 1c). Consistent results were obtained using another two-alternative gene-tissue specificity calculation methods, SPM and EE. In fact, the tissue specificities defined by the three methods are highly similar to each other (R > 0.8, Fig. 1d and Fig. S1). At the pathway level, these tissue specific genes are enriched in metabolic functional pathways (Fig. 1e), which also exhibited rhythmic oscillation signals throughout the day (Fig. 1f). So circadian regulation and tissue specificity are highly correlated throughout the body. Similarly, we analyzed the functional differences between tissues based on circadian genes. Specifically, we examined the top 10 biological processes of Gene Ontology (GO) annotation associated with circadian genes from the mouse circadian atlas. Our analysis revealed that circadian genes exhibit functional diversity, with significant associations to metabolism and biological rhythms (Fig. S2). Furthermore, we employed the Weighted Gene Co-expression Network Analysis (WGCNA) to explore overall gene expression patterns. This network was partitioned into several modules (Fig. S3a), and we identified those modules enriched with circadian genes. Notably, the liver and kidney exhibited the highest enrichment in circadian gene modules (Fig. S3b, c).
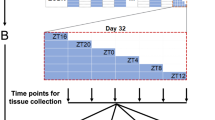
Circadian disturbances such as acute sleep deprivation have been shown to induce tissue-specific transcription and DNA methylation of human core clock genes49. To explore how circadian disturbance affects the relationship between circadian rhythm and tissue specificity, we conducted RNA-sequencing on 96 samples from both normal and sleep-deprived (circadian disrupted) mouse organs, including the liver, kidney, forebrain, and cerebellum. These organs were selected due to their previous demonstration of containing robust circadian expressed transcripts, and the forebrain and cerebellum are recognized as top influenced tissues by the circadian systems within the brain35. For sleep-deprived samples, mice were subjected to a 10 h sleep deprivation, as severe sleep deprivation can affect the transcriptome of mouse tissues (Maret et al. 2007). We collected two replicates every 4 hours from male mice, under constant darkness. We chose to sequence the samples at high depth, with an average of 54 million reads sequenced per sample. The reads were aligned to the mouse genome GRCm38 using HISAT2, and the transcription information for each annotated genomic region was estimated, facilitating downstream analysis50. This allowed us to access the expression signature for both protein-coding and noncoding transcripts. We detected a total of 18,531 transcripts with abundant transcriptional signatures, expressed in all samples, which were used for downstream analysis. To validate the accuracy of the sleep deprivation experiment, we analyzed rhythmic genes in each tissue. The findings revealed varied degrees of alteration in the expression of rhythmic transcriptomes and key circadian genes (Fig. S4).
Next, we analyzed the relationship between rhythmic expression signals in the four organs and the tissue-specific signals of the genes as measured by the specificity measure (SPM) from the mouse circadian transcriptome atlas. Although sleep deprivation altered the pattern of gene oscillations, we observed that the correlation between circadian amplitude (AMP) and tissue specificity of rhythmically expressed genes remained significant in both control and sleep-deprived conditions (Fig. 1g, h), particularly in the liver. We note that similar results were obtained when expressed genes were defined using different cutoffs (see methods). Importantly, when comparing the tissue specificity of rhythmically expressed genes with that of non-rhythmically expressed genes in calorie-restricted feeding states, we observed a significant difference. In both adult and old mice, the tissue specificity of rhythmically expressed genes was markedly greater than that of non-rhythmically expressed genes (Fig. S5). This finding suggests that the rhythmic expression of genes may play a crucial role in coordinating tissue-specific responses to calorie-restricted feeding.
Both differentially expressed protein-coding and noncoding genes are temporally organized
To quantify the relationship between circadian regulation and tissue specificity, we focused on the circadian transcriptome of the liver, utilizing kidney as a reference organ, as these two peripheral organs contain the most robust circadian transcription patterns35. We profiled their 24 h circadian transcriptomes with four replicates collected every four hours under constant darkness, to confidently access their differentially expressed transcripts for each time point. More replicates allow us to identify the cycling genes more robustly and enable a more sophisticated computational analysis. For the protein-coding genes expressed in both organs, 3310 transcripts in the liver and 1125 transcripts in the kidney were finally identified as rhythmically expressed protein-coding genes (Jonckheere-Terpstra-Kendall test51, BH.Q adjusted P < 0.05).
We then identified the differentially expressed genes (DEGs) between the two organs to define tissue specificity, using both DESeq252 and edgeR53 algorithms, with an adjusted p < 0.05 and greater than 2-fold change used as cutoffs. In total, 9,898 DEGs were identified between liver and kidney for the 6 time points. Strikingly, only 44% (4,454 genes) of these DEGs were defined as DEGs across all 6 time points, suggesting that most DEGs are time-dependent. To further assess the prevalence of common DEGs, we conducted a permutation test across all the samples. On average, 47% of genes were found to be differentially expressed at all time points under randomization experiments. Therefore, the observed value of 44% is slightly lower than expected by chance. These differences of gene expression profiles between liver and kidney reflect the temporal organization of the transcriptome.
To assess the effect of circadian regulation on these DEGs, we further classified them into genes that are potentially affected by the circadian regulation and those that are not. The first category, circadian-affected DEGs, includes genes that are cyclic expressed in either liver or kidney with a 24-hour period, as well as genes that could be bound by the core circadian transcriptional factors (BMAL1, CLOCK, PERs, CRYs)14. The remaining DEGs can be divided into two categories: 1) constitutive DEGs, which are differentially expressed across all time points but are not affected by circadian regulation, and 2) temporal DEGs, which are differentially expressed in some time points but are not circadian-affected DEGs. We discovered that 35% (3,418) of DEGs are circadian-affected DEGs, while only 28% (2,775) of them are constitutive DEGs, demonstrating a larger contribution of circadian regulation to DEGs than expected (Fig. 2a and Supplementary Data 1).
a The pie chart displays the proportion and number of three classes of differentially expressed genes (DEGs). Among all DEGs, 35% are circadian-affected, 28% are constitutive, and 35% are temporal DEGs. b Expression abundance comparison among the three classes of DEGs. P-values were determined by Wilcoxon rank sum test. Error bars are the 95% confidence interval, the bottom and top of the box are the 25th and 75th percentiles in boxplot. c Differential transcription factors (TFs) potentially binding to distinct DEGs. Top TFs are indicated in each panel. The Venn diagram shows the number of TFs identified in overlap. d Functional annotation of the three DEG classes. Circadian-affected DEGs are enriched in metabolic functions, while other DEGs are associated with distinct biological pathways. Both enrichment odds ratio and significant level (P) are displayed. e Enrichment analysis of metabolic diseases for circadian-affected, constitutive, and temporal DEGs. + indicates significantly overrepresented, and - indicates significantly underrepresented. Different colors indicate odds ratios.
Noncoding RNAs are transcribed widely throughout the genome, and exhibit oscillation signatures and high tissue-specificity. Long noncoding RNAs (lncRNAs) are a particularly noteworthy class, but their regulatory role in circadian rhythms remains incompletely understood54. We examined if and to what extend differentially expressed noncoding genes are temporally organized. We identified 161 rhythmic lncRNA transcripts among the 2273 expressed lncRNAs in liver and kidney. The proportion of rhythmic lncRNAs (7.1%) is lower than that of protein-coding genes (24.6%), consistent with previous observations35. We focused on two classes of non-coding RNAs interested: bi-promoter transcripts and antisense transcripts (Pelechano and Steinmetz, 2013). We estimated their differential expression signatures and identified 523 differentially expressed antisense transcripts and 181 differentially expressed bidirectional transcripts. In both categories, the proportion of constitutive differentially expressed noncoding transcripts is less than one-third of all differentially expressed noncoding transcripts (Fig. S6a, S6b). Thus, the majority of tissue specificity defined by noncoding RNAs also follow a time dependent manner.
The phase of rhythmical lncRNAs is close to that of their neighboring protein-coding genes, with the majority having phase differences of less than four hours (R = 0.42, P = 0.00048, Fig. S6c). This suggests that the oscillation of lncRNAs is coordinated with that of neighboring protein-coding genes, possibly because of chromatin remodeling. This is also evidenced by the observation that the location of cycling non-coding transcripts is co-clustered with cycling coding transcripts on the chromosome (Fig. S6d).
Circadian-affected DEGs exhibit distinct functions compared to other DEGs
We discovered several key differences between circadian-affected DEGs and other DEGs. Circadian-affected DEGs typically have elevated transcription levels than other two groups of DEGs (median TPM expression level = 9.39 vs. 0.77, P < 2.2 x 10−16, Fig. 2b), but no differences in expression level were detected between constitutive DEGs and temporal DEGs (median TPM expression level = 0.77 vs. 0.98, P = 0.18, Fig. 2b). Moreover, differences in the expression levels of the three types of DEGs were observed to persist in the calorie-restricted mice, as illustrated in Figure S7a. Similar to the protein-coding genes, circadian affected DE noncoding genes also have higher expression level than other DE noncoding transcripts (median TPM expression level = 0.85 vs. 0.42, P = 1.7 x 10−08, Figure S6e). We also found that these three groups of genes are regulated by different transcription factors (Fig. 2c), with only 20 common TFs shared among the top 100 TFs (Supplementary Data 2). This implies a significant difference in the transcriptional regulatory structure of the three groups of genes. Circadian-affected DEGs were predicted to be regulated by a various circadian core clocks, including RXRA, NR1D1, BMAL1, and transcription factors such as CEBPA and CEBPB55, which are involved in liver metabolic processes. CEBPB is also the top regulator of temporal DEGs, which are also controlled by some general transcription factors involved in cell growth and replication, such as FOSL2 and RELA. In contrast, IRF1 and IRF8 with immune function were involved in the regulation of constitutive DEGs56,57,58. Finally, in combination with recently released long-read sequencing data for the liver59, our analysis identified 59 Circadian-affected DEGs that are potentially fusion genes in cancer tissue, a significantly higher number than expected (R = 1.48, p = 0.0093, Fisher’s exact test). These fusion events tend to occur more frequently in the 5’ region than in the 3’ region, as shown in supplemental Fig. S8.
Interestingly, the three types of DEGs function in biological processes. GO functional enrichment results indicated that circadian-affected DEGs are extensively involved in liver-specific metabolic processes such as fatty acid and alcohol metabolism. Similar results were obtained by the KEGG analysis (Fig. S9). Temporal DEGs, on the other hand, play a significant role in immune cell proliferation and intercellular communication, while constitutive DEGs are linked to kidney functions such as urogenital system development. Furthermore, genes involved in cell movement, including leukocyte migration and actin filament-based movement, are also well-represented among constitutive DEGs and Temporal DEGs (Fig. 2d, Supplementary Data 3). The immune-related gene expression signatures observed in the temporal set were predominantly associated with immune cells, rather than the kidney or liver cells themselves. GO functional enrichment results of circadian genes in caloric restriction were found to be similar to those of circadian-affected DEGs, rather than other DEGs, as depicted in Figure S7b. Next, we investigated whether these three types of DEGs display different patterns by examining datasets for metabolism-related diseases and cancer60. The circadian-affected DEGs were highly enriched in metabolic disorders such as hypercholesterolemia and type 2 diabetes, consistent with their biological functions. Conversely, constitutive DEGs exhibited only weak enrichment in these diseases, while temporal DEGs did not show any clear trend (Fig. 2e, Supplementary Data 4). In summary, our findings demonstrate that the three DEGs have distinct transcriptional regulatory mechanisms, biological functions, and disease tendencies.
Aging and longevity genes are enriched among the circadian-affected DEGs
Aging and longevity processes have been shown to be closely linked with circadian regulation3,10,12. Therefore, we quantified the relationship between circadian-affected DEGs and aging & longevity genes at the genome-scale. Initially, we focused on the genes that exhibit aging-related expression changes in different cell types, which were identified using the Tabula Muris Senis single-cell dataset61. We used the Fisher’s exact test to analyze the association between aging and longevity genes with circadian-affected DEGs. The genes expressed in the sample were used as the total gene set, and the aging and longevity genes were required to be expressed in the sample. The circadian-affected DEGs are enriched in cell-type-specific aging changes when compared to the constitutive DEG and the temporal DEGs (Fig. 3a). This result was validated in two additional independent aging and longevity gene collections - LongvityMap database62, and GeneAge database (Fig. 3a, Supplementary Data 5).
a Enrichment analysis of three types of DEGs with aging and longevity genes. The “enriched” and “depleted” relationships are shown in red and blue, respectively. The odds ratio is indicated by the size of the circle. Fisher’s exact test was used for enrichment analysis, and the total gene set was the genes expressed in all samples. b Enrichment analysis of three types of DEGs with genes that oscillate in response to calorie and time restriction. Eight different intervention protocols were included: 06mo_CR.night.2 h, 06mo_CR.night.12 h, 06mo_CR.day.2 h, 06mo_CR.day.12 h, 19mo_CR.night.2 h, 19mo_CR.night.12 h, 19mo_CR.day.2 h, and 19mo_CR.day.12 h. Fisher’s exact test was used for enrichment analysis, and the total gene set was the genes expressed in all samples. c Functional enrichment analysis of aging-related diseases for circadian-affected DEGs, constitutive DEGs, and temporal DEGs. The enriched and depleted relationships are shown in red and blue, respectively. The odds ratio is illustrated by different colors. Fisher’s exact test was used for enrichment analysis, and the total gene set was the genes expressed in all samples. d Comparison of correlation coefficients between fatty acid metabolic genes and other genes with age. Error bars are the 95% confidence interval in boxplot. e–h Scatter plots showing the correlation between the expression levels of four circadian-affected DEGs (Acox3, Avpr1a, Crot, and Cyp4a32) and age. The Pearson correlation coefficient was used to calculate the correlation coefficient.
As calorie restriction with aligned circadian phase can significantly promote longevity in mice31, we further focused on the genes that show circadian signals under both caloric and time restriction condition, where the oscillation is susceptible to these restrictions. We found that circadian-affected DEGs are significantly enriched in caloric & time restriction related circadian genes, whereas other DEGs are depleted in those conditions (Fig. 3b, Supplementary Data 6). Similar results were observed when we explored the reorganized liver circadian transcriptome of both young and old mice subjected to caloric restriction condition (Figure S10, P < 2.2 x 10−16 in all comparisons)63. Then we conducted a comparison of the fold change of genes among the three types of DEGs during caloric restriction (CR) and ad libitum feeding (AL). The findings revealed that only circadian-affected DEGs exhibited a significantly lower fold change compared to other genes (P = 0.0053), as illustrated in Figure S11a. These results suggest that tissues affected by circadian-driven specificity are involved in the aging process and are influenced by caloric restriction. Furthermore, we selected aging marker genes previously reported to be associated with oxidative stress effects and aging, and intersected them with liver tissue-specific genes to identify liver tissue-specific aging genes. Under caloric restriction, the expression of most of these genes exhibited a significant decrease in both mice, as depicted in Figure S11b. These results strongly indicate that the set of circadian affected DEGs is enriched for genes with functions potentially beneficial for lifespan extension25,31.
We found that Circadian-affected DEGs were enriched in various age-related diseases, as annotated by human genetic disease database9 (Fig. 3c). One of the top aging diseases that we identified was Nonalcoholic fatty liver disease, a common disease in elderly people64. Other diseases involving basal metabolic dysfunction, such as Osteonecrosis and cardiovascular complication of diabetes, were also significantly enriched in these DEGs. Interestingly, temporal DEGs exhibited the opposite trend, with diseases such as Arteriosclerosis being heavily depleted. By comparing the Spearman correlation coefficients between their expression level and age, it was found the correlation coefficients of fatty acid metabolic genes are significantly higher than those of other genes (Absolute values of Spearman: 0.21 vs. 0.14, P < 2.2 x 10−16, Fig. 3d). The expression of many genes in this function is anticorrelated with age, including Acox3, Avpr1a, Crot and Cyp4a32 (Fig. 3e-h). The decreased expression of the four genes during aging may be related to weakened fatty liver metabolism and impaired liver function in elderly individuals.
Circadian-affected DEGs are highly expressed in hepatocytes
Tissue-specificity may arise from tissue-specific cell types, given that most genes specifically expressed in a tissue are enriched in certain cell types65. At the single-cell level, both hepatocyte and other cells in the liver are under the control of the circadian clock66. To investigate the cell-type origin of the circadian tissue specificity in the liver, we compared the cell-type marker genes in each cell population66. In total, 2,587 cell-type marker genes were identified in 10 cell types, ranging from 150 to 600 genes per cell type (Fig. 4a). In hepatocyte cells, ~40% of cell markers were circadian-affected DEGs, while in other cell types (e.g., Chol and EC), only ~20% cell markers were circadian-affected DEGs (Fig. 4a). Cell marker genes tend to be circadian-affected DEGs in 4 out of 10 liver cell types (P < 2.2 x 10−16, Fig. 4b). Through analysis in the Cistrome DB database combined with large-scale CHIP-Seq data, we discovered that hepatocyte cells markers potentially bind with circadian core transcription factors such as CLOCK, BMAL1, CRY1, CRY2, while other cell types markers are more divergent (Figure S12). In addition, the circadian genes under both caloric and time restriction condition are mostly enriched in the four major hepatocyte cells (Odds ratio > 3 for all of them, P < 2.2 x 10−16), indicating that hepatocytes actively participate in the circadian oscillation of gene expression under metabolic restructuring55.
a The histogram shows the number of marker genes for different cell types, and the broken line represents the proportion of circadian-affected DEGs among the marker genes. b Enrichment of circadian-affected DEGs in marker genes for different cell types. c UMAP plot visualizing liver cell clusters based on single-cell transcriptomes. d UMAP plot visualizing the enrichment of the three classes of DEGs. The different colors indicate the cell types enriched by different classes of DEGs, while gray indicates no enrichment of any DEG. e Functional network annotation of the three classes of DEGs based on enriched cell type marker genes. The dots of different colors represent different classes of DEGs.
The analysis of DEGs and cell marker genes revealed that the circadian-affected DEGs are enriched in hepatocyte cells (Hep Zone1, P < 2.2 x 10−16; Hep Zone2, P < 2.2 x 10−16; Hep Zone3, P < 2.2 x 10−16; Hep Zone4, P < 2.2 x 10−16), while temporal DEGs are enriched in immune cells (P < 0.05), and constitutive DEGs are enriched in other cell types (P < 0.01) (Fig. 4c, d). This suggests that immune cells and epithelial cells in the liver may contribute less to the overall tissue specificity. Similarly, the functional annotation of cell-type marker genes indicated that hepatocytes cell markers were involved in metabolic functions, such as fatty acids and amino acids metabolic process, and formed a single cluster, while genes for cell interactions and immune function were shared among cell types to varying degrees (Fig. 4e). These results reveal that the three types of DEGs are derived from different cell types dynamically, suggesting that different cell types contribute to different aspects of the temporal distribution of tissue-specific gene expression.
Cycling co-expression modules are associated with aging
To understand the modular properties of circadian-affected DEGs, we next conducted the weighted gene co-expression network analysis (WGCNA) on liver circadian transcriptomes across all 24 samples. We identified a total of 13 co-expression network modules, with module size ranging from 249 to 5466 genes (Fig. 5a, b, Supplementary Data 7). We then investigated the rhythmicity of eigengene expression in each module (Fig. 5c, d). Our findings were consistent with the previous report67, which demonstrated that a significant number of network modules display a rhythmic expression pattern. Specifically, eight out of the 13 modules are cycling modules (black, greenyellow, brown, green, pink, purple, magenta, and red, P < 0.001, BHQ method; Fig. 5b), indicating that the gene regulatory network changes dynamically throughout the day.
a The co-expression network of liver samples visualized using a cluster diagram, with different types of DEGs assigned to the identified modules. b Identification of cycling modules with enrichment of circadian-affected DEGs, liver-specific aging genes, and rhythmic genes under caloric and time restriction conditions. The odds ratio is shown on the X-axis, and co-expression modules are shown on the Y-axis. Again, “enriched” is indicated by red dots, while “depleted” is indicated by blue dots. c Heatmap of the eight circadian modules ordered by circadian phase, showing module eigengene values. d Heatmap of the eight circadian modules ordered by circadian phase, showing averaged expression levels. e Functional annotation of the cycling modules, showing their involvement in processes such as response to protein folding, cell junction assembly, rRNA processing, rRNA metabolic processes, and small GTPase-mediated signal transduction.
We then assessed the enrichment of three groups of DEGs in those modules. We found that all eight circadian-expressed modules were enriched with circadian-affected DGEs (Fig. 5b, c). Moreover, the distribution of temporal DEGs and constitutive DEGs in the module was opposite to circadian-affected DEGs, with these genes being significantly depleted in cycling module. Additionally, we found that among these cycling modules, the purple, brown and red modules were also enriched with liver-specific aging-related genes. Remarkably, 7 of all 8 cycling modules were highly associated with circadian genes under both caloric and time restriction condition (Fig. 5b), indicating the robustness of oscillations of network modules under restricted dietary conditions. This suggests the robustness of oscillations of network modules under different dietary conditions and that aging-related genes also show higher levels of circadian rhythm at the module level in the gene regulatory network.
Functional annotation of these circadian modules uncovered that they were mainly involved in response to protein folding (P < 1.99 x 10−12), cell junction assembly (P < 6.50 x 10−07), rRNA processing (P < 2.2 x 10−10), rRNA metabolic process (P < 1.37 x 10−10) and small GTPase mediated signal transduction (P < 7.08 x 10−08) (Fig. 5e), which have been reported to be associated with aging and longevity in previous studies68. In summary, the results above indicate that the differential genes in different tissues are aggregated in the gene expression network in the form of modules, which show cyclic expression together and are involved in the liver aging process.
Aging genes are more disrupted in circadian misalignment environments compared with other genes
Circadian disruption is a deleterious condition that can lead to various diseases such as diabetes and cancer. We investigated whether the regulation of aging related genes is more susceptible to disturbance than other genes in the cycling transcriptome in circadian misalignment environments. To address this impact, we collected data on different circadian disruption conditions, such as sleep deprivation and an inverted diet. For both datasets, we utilized JTK_CYCLE to calculated the oscillation amplitude in the experimental and control groups and used the amplitude change between these groups to assess effect size for the examined gene sets. Briefly, we examined six sets of aging-related genes—obtained from the LongvityMap &GeneAge database—and the aging genes obtained from the Tabula Muris Senis single-cell dataset.
As displayed in Fig. 6a, the cycling amplitude of aging-related genes exhibits a significantly larger change than other genes under 10 h acute sleep deprivation condition in the liver (total amplitude difference: 1.14 vs 0.30, P < 4.5 x 10−05, Fig. 6a). Similar results were observed when we performed downsampling to equalize the number of genes between groups (Fig. S13) and a similar trend was also observed in the kidney (total amplitude difference: 2.41 vs 0.71, P < 0.05, Fig. 6b). Moreover, we compared essential genes and found that they are less affected by sleep deprivation, exhibiting more robust expression (Fig. S14). These results suggest that the circadian expression of aging-related genes in peripheral organs is susceptible to sleep behavior interference, meaning that sleep deprivation can exert pressure on the rhythmic expression of the circadian-affected DEGs in the aging process.
a A violin plot is used to show the change in circadian amplitude (AMP) of aging and longevity genes in the liver in comparison to other genes during 10 h sleep deprivation. All 6 aging and longevity gene lists are included in the plot. b The change in circadian amplitude (AMP) of aging and longevity genes in the kidney is displayed in a violin plot, compared to other genes during 10 h sleep deprivation. All 6 aging and longevity gene lists are included in the plot. c A violin plot to compare the change in circadian amplitude (AMP) of aging and longevity genes with that of other genes during a 2 h restricted inverted feeding condition. d A violin plot is used to compare the change in circadian amplitude (AMP) of aging and longevity genes with that of other genes during a 12 h restricted inverted feeding condition. e A boxplot to compare the changes in circadian AMP for circadian-affected DEGs, constitutive DEGs, temporal DEGs, and other genes during sleep deprivation. f A boxplot to compare the changes in circadian AMP for circadian-affected DEGs, constitutive DEGs, temporal DEGs, and other genes during inverted food intake condition. Both conditions exhibit similar trends. Statistical significance is denoted by ***P < 0.001; **P < 0.01; *P < 0.05; and n.s. for non-significant differences. Error bars are the 95% confidence interval, the bottom and top of the box are the 25th and 75th percentiles in boxplot.
In the context of a restricted diet that is known to improve the circadian rhythm of mouse behavior69,70,71, our findings reveal a similar pattern: the circadian amplitude of aging gene expression responds more dramatically to this intervention (total amplitude difference: 0.55 vs 0.19, P < 0.05, Fig. 6c, d). This suggests that the change may not be limited to only sleep or diet, but rather a broader molecular mechanism that underlies both and plays a role in the amplitude sensitivity of aging genes. In addition, circadian-affected DEGs had larger amplitude changes in the two organs than the other DEGs and than the non-DEGs (DEGs amplitude difference: 0.81 vs 0.32 vs 0.25, P < 2.2 x 10−16, Fig. 6e). This is consistent with the results of aging genes, supporting the close relationship between aging genes and DEG. We also noted similar observations in the inverted feeding condition (2 h, or 12 h inverse in adult mice, Fig. 6f)31 in the diet disruption condition, which was also validated by analyzing the recently released inverse-feeding peripheral tissues transcriptome datasetSleep deprivation experiment The sleep deprivation experiment followed previously reported protocols75. We utilized male C57BL/6 mice, aged between 6 and 8 weeks, which were kept in an environment with a 12 h light/dark cycle. Water and food were provided ad libitum. Mice were randomly assigned to either the experimental or control group. The experimental group was subjected to a sleep deprivation instrument, which involved forced locomotion through a slowly rotating drum (40 cm in diameter, 0.4 m per min) while having access to food and water. All mice in experimental group were placed in the sleep deprivation device for 10 h (6:00 AM–4:00 PM) during their normal resting phase while the control animals were undisturbed, because acute sleep deprivation, lasting between 6 and 10 h, can leads to noticeable alterations in mice’s circadian rhythms and significantly affects the rhythmic expression of numerous genes75,76,77. Then we collected two replicates of 4 peripheral circadian tissues (forebrain, cerebellum, liver, kidney) were obtained at next six circadian time (CT) points with an interval of 4 h (CT4, CT8, CT12, CT16, CT20, and CT24). We extracted total RNA from tissue samples using Trizol (Invitrogen). After separating the RNA, DNA, and protein layers with chloroform, we precipitated the RNA using a standard isopropanol/ethanol procedure. The resulting RNA pellet was washed, and then resuspended in RNase-free water. We assessed the quantity and quality of the total RNA using the Qubit RNA HS Assay (Life Technologies) and RNA 6000 Nano Assay on a Bioanalyzer 2100 (Agilent). For RNA-seq library construction, we followed the Illumina TruSeq Total RNA stranded protocol. We used 1 μg of total RNA and depleted rRNA using the ribo-zero rRNA gold removal kit. We then chemically fragmented the rRNA-depleted RNA for three minutes and prepared a paired-end 150-bp strand-specific RNA-seq library using the NEB Next Ultra II Directional RNA Library Prep Kit (NEB 7760), following the manual. We performed deep sequencing using the Illumina HiSeq X platform. All our new samples have passed RNA sequencing quality control (RNAseq QC). We deposited all raw sequencing data in the Gene Expression Omnibus for free access accession. We used Trim Galore to assess quality and remove adapters from the FastQ files (parameters: --phred33 --length 50 --stringency 3 --paired) (https://github.com/FelixKrueger/TrimGalore, v0.6.4). We aligned the reads to the mouse reference genome (GRCm38, downloaded from GENCODE) using HISAT2 (version 2.1.0)50 with parameters -t -p 8 --dta --rna-strandness RF. We extracted high-quality alignment results from BAM files using Samtools (version 1.9)78 with parameters view -b -q 60. After alignment, we calculated read counts using featureCounts (version v1.6.4)79 with parameters -T 16 -p -s 2. We then created a final list of genes that were expressed (TPM > 0) in all samples. We used raw counts to calculate differentially expressed genes. Differential expression analysis was performed using both DESeq2 (version 1.26.0)52 and EdgeR (version 3.28.1)53 between liver and kidney at each time point. We considered a gene to be differentially expressed if the adjusted p-value was less than 0.05 in both DESeq2 and EdgeR, which was selected as the significance cutoff. We used the Jonckheere-Terpstra-Kendall algorithm from the MetaCycle R package (version 1.20)80 to identify oscillating genes from the cycling transcriptome. The algorithm was run with TPM as input, and we considered only genes with a Benjamini-Hochberg adjusted p-value of less than 0.05 as confidently circadian expressed genes. To identify circadian genes under caloric restriction conditions, we used the method adopted in the corresponding articles of each dataset to identify circadian rhythm genes. We employed the JTK algorithm from the MetaCycle R package (version 1.20)80 with a p-value cutoff of 0.01, using data from Sato et al.63. We selected genes that showed rhythmic expression only under caloric restriction conditions as our gene list. To identify circadian genes that oscillate exclusively under both caloric and time-restricted conditions, we used data from Acosta-Rodríguez et al.31, and selected cycling genes that were significant in all three different algorithms (JTK_CYCLE, ARSER, and RAIN) with a p-value cutoff of 0.05 and FDR < 0.05. We selected our final gene sets by filtering genes that were rhythmically expressed exclusively under both caloric and time-restricted conditions, but not under ad libitum or CR-spread conditions. To evaluate tissue-specificity of the transcriptome, we calculated Tau for each gene, using the following formula81: Where, xi is the expression level of a given gene in tissue i and n is the number of tissues in a given time point. We used another tissue-specificity indicator, specifically the Specificity Measure (SPM), to evaluate tissue-specificity of the transcriptome. SPM was calculated for each gene using the following formula82: Tissue-specific gene was defined as genes with SPM in top 10% of all genes. We also used an additional tissue-specificity indicator, expression enrichment (EE)83, to validate our main discoveries. The EE score was calculated as following: Si summary of the expression of all genes in tissue i For the weighted gene co-expression network analysis (WGCNA)84, we used the rlog transformation in the DESeq2 package to standardize the original RNA-Seq count data, which has a similar variance stabilizing effect to the varianceStabilizingTransformation. We selected genes with a Median Absolute Deviation (MAD) > 0.1. We then constructed co-expression networks using the Bicor method to calculate expression similarities. The softpower parameter was chosen as 14, resulting in 13 co-expression modules containing 17,070 expressed genes. To determine whether a module was rhythmic, we analyzed both the module eigengene and the mean expression of the module using JTK_CYCLE. Modules with a Benjamini-Hochberg adjusted P-value < 0.05 were considered circadian modules. We used Landscape In Silico Deletion Analysis (LISA v2.2.5)85 to predict potential transcription factors. For each analysis, we input the top 500 genes with the highest fold-change among each type of differentially expressed genes (DEGs). We performed functional annotation of genes using ClusterProfiler (version 4.0)86. Each term was ranked according to its false discovery rate (FDR), and the significance threshold was set to < 0.05. Since many enriched terms had intersections, we clustered similar terms based on their semantic similarity using a method described by Wang et al.87. Metabolic-related diseases were identified based on a previous study60. Cancer-associated genes were obtained from experimental data reported in the cancer gene census or from significantly enriched driver genes included in the Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/). Disease genes associated with metabolism were extracted from the Human Gene Mutation Database (HGMD)88 by selecting gene sets with at least one or more disease-associated mutations. We obtained aging-related diseases from a previous report9. Related disease gene sets were obtained from the MalaCards Human Disease Database89,90,91. To identify the corresponding mouse orthologs, we used the Ensembl Biomart92 to convert the human genes. Marker genes for peak-based clustering were identified using Seurat’s FindAllMarkers() function93 on the gene activity matrix with the following parameters: min.pct = 0.25 and logfc.threshold = 0.25. We calculated the correlations of time-course data between circadian lncRNAs and neighboring protein coding genes by selecting only those lncRNA clusters and their nearby protein-coding genes that were circadian expressed. For each lncRNA cluster, we calculated Pearson’s correlation coefficients between the phases of circadian lncRNA transcripts and circadian protein-coding genes located within a 50 kb region. The median of these correlation coefficients was used as the final correlation coefficient between this lncRNA and adjacent protein-coding genes in this region. All the analyses were performed in R (v4.0.2). Statistical significance was determined at Benjamini-Hochberg adjusted P-value < 0.05. Gene set enrichment analysis was performed using a two-sided Fisher’s exact test. Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.RNA-sequencing experiment
RNA-sequencing data processing
Differential gene expression analysis
Identification of circadian expressed genes
Identification of genes that oscillate exclusively under both caloric and time restriction condition
Estimation of tissue-specificity and tissue-specific gene
Gene co-expression network analysis
Transcriptional factor (TF) prediction
Biological function enrichment analysis
Gene sets for metabolism-related diseases
Gene sets for aging-related diseases
Single cell marker gene identification
Calculating the phase correlations between circadian lncRNAs and protein-coding genes
Statistics and reproducibility
Reporting summary
Data availability
All raw and processed sequencing data generated in this study have been submitted to the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) under accession number GSE240693, which will be freely accessible upon the publication of the manuscript.
Code availability
We used R for most of the analyses. All original code is available upon request.
References
Hekselman, I. & Yeger-Lotem, E. Mechanisms of tissue and cell-type specificity in heritable traits and diseases. Nat. Rev. Genet. 21, 137–150 (2020).
Harman, D. The aging process: major risk factor for disease and death. Proc. Natl Acad. Sci. USA 88, 5360–5363 (1991).
Yamazaki, S. et al. Effects of aging on central and peripheral mammalian clocks. Proc. Natl Acad. Sci. USA 99, 10801–10806 (2002).
Wang, X. et al. Ageing induces tissue-specific transcriptomic changes in Caenorhabditis elegans. EMBO J. 41, e109633 (2022).
Chatsirisupachai, K., Palmer, D., Ferreira, S. & de Magalhaes, J. P. A human tissue-specific transcriptomic analysis reveals a complex relationship between aging, cancer, and cellular senescence. Aging Cell 18, e13041 (2019).
Srivastava, A. et al. Tissue-specific Gene Expression Changes Are Associated with Aging in Mice. Genomics Proteom. Bioinforma. 18, 430–442 (2020).
Yamamoto, R. et al. Tissue-specific impacts of aging and genetics on gene expression patterns in humans. Nat. Commun. 13, 5803 (2022).
Izgi, H. et al. Inter-tissue convergence of gene expression during ageing suggests age-related loss of tissue and cellular identity. Elife 11. https://doi.org/10.7554/eLife.68048 (2022).
Zhang, M. J., Pisco, A. O., Darmanis, S. & Zou, J. Mouse aging cell atlas analysis reveals global and cell type-specific aging signatures. Elife 10. https://doi.org/10.7554/eLife.62293 (2021).
Acosta-Rodriguez, V. A., Rijo-Ferreira, F., Green, C. B. & Takahashi, J. S. Importance of circadian timing for aging and longevity. Nat. Commun. 12, 2862 (2021).
Kondratova, A. A. & Kondratov, R. V. The circadian clock and pathology of the ageing brain. Nat. Rev. Neurosci. 13, 325–335 (2012).
Chaudhari, A., Gupta, R., Makwana, K. & Kondratov, R. Circadian clocks, diets and aging. Nutr. Healthy Aging 4, 101–112 (2017).
Gekakis, N. et al. Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569 (1998).
Koike, N. et al. Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 338, 349–354 (2012).
Lee, C., Etchegaray, J. P., Cagampang, F. R., Loudon, A. S. & Reppert, S. M. Posttranslational mechanisms regulate the mammalian circadian clock. Cell 107, 855–867 (2001).
Kondratov, R. V., Kondratova, A. A., Gorbacheva, V. Y., Vykhovanets, O. V. & Antoch, M. P. Early aging and age-related pathologies in mice deficient in BMAL1, the core componentof the circadian clock. Genes Dev. 20, 1868–1873 (2006).
Dubrovsky, Y. V., Samsa, W. E. & Kondratov, R. V. Deficiency of circadian protein CLOCK reduces lifespan and increases age-related cataract development in mice. Aging 2, 936–944 (2010).
Valentinuzzi, V. S., Scarbrough, K., Takahashi, J. S. & Turek, F. W. Effects of aging on the circadian rhythm of wheel-running activity in C57BL/6 mice. Am. J. Physiol. 273, R1957–R1964 (1997).
Weinert, D. Age-dependent changes of the circadian system. Chronobiol. Int. 17, 261–283 (2000).
Wolff, C. A. et al. Defining the age-dependent and tissue-specific circadian transcriptome in male mice. Cell Rep. 42, 111982 (2023).
Sellix, M. T. et al. Aging differentially affects the re-entrainment response of central and peripheral circadian oscillators. J. Neurosci. 32, 16193–16202 (2012).
Davidson, A. J. et al. Chronic jet-lag increases mortality in aged mice. Curr. Biol. 16, R914–R916 (2006).
Morris, C. J., Purvis, T. E., Hu, K. & Scheer, F. A. Circadian misalignment increases cardiovascular disease risk factors in humans. Proc. Natl Acad. Sci. USA 113, E1402–E1411 (2016).
Lu, J. Y. et al. Comparative transcriptomics reveals circadian and pluripotency networks as two pillars of longevity regulation. Cell Metab. 34, 836–856 e835 (2022).
Lopez-Otin, C., Galluzzi, L., Freije, J. M. P., Madeo, F. & Kroemer, G. Metabolic Control of Longevity. Cell 166, 802–821 (2016).
Hu, K., Van Someren, E. J., Shea, S. A. & Scheer, F. A. Reduction of scale invariance of activity fluctuations with aging and Alzheimer’s disease: Involvement of the circadian pacemaker. Proc. Natl Acad. Sci. USA 106, 2490–2494 (2009).
Witting, W., Kwa, I. H., Eikelenboom, P., Mirmiran, M. & Swaab, D. F. Alterations in the circadian rest-activity rhythm in aging and Alzheimer’s disease. Biol. Psychiatry 27, 563–572 (1990).
Zhang, W. et al. Emerging Insight Into the Role of Circadian Clock Gene BMAL1 in Cellular Senescence. Front Endocrinol. (Lausanne) 13, 915139 (2022).
Froy, O. Circadian aspects of energy metabolism and aging. Ageing Res Rev. 12, 931–940 (2013).
Chang, H. C. & Guarente, L. SIRT1 mediates central circadian control in the SCN by a mechanism that decays with aging. Cell 153, 1448–1460 (2013).
Acosta-Rodríguez, V. et al. Circadian alignment of early onset caloric restriction promotes longevity in male C57BL/6J mice. Science 376, 1192–1202 (2022).
Yeung, J. & Naef, F. Rhythms of the Genome: Circadian Dynamics from Chromatin Topology, Tissue-Specific Gene Expression, to Behavior. Trends Genet 34, 915–926 (2018).
Menet, J. S. & Hardin, P. E. Circadian clocks: the tissue is the issue. Curr. Biol. 24, R25–R27 (2014).
Talamanca, L., Gobet, C. & Naef, F. Sex-dimorphic and age-dependent organization of 24-hour gene expression rhythms in humans. Science 379, 478–483 (2023).
Zhang, R., Lahens, N. F., Ballance, H. I., Hughes, M. E. & Hogenesch, J. B. A circadian gene expression atlas in mammals: implications for biology and medicine. Proc. Natl Acad. Sci. USA 111, 16219–16224 (2014).
Qu, M., Qu, H., Jia, Z. & Kay, S. A. HNF4A defines tissue-specific circadian rhythms by beaconing BMAL1::CLOCK chromatin binding and sha** the rhythmic chromatin landscape. Nat. Commun. 12, 6350 (2021).
Rey, G. et al. Genome-wide and phase-specific DNA-binding rhythms of BMAL1 control circadian output functions in mouse liver. PLoS Biol. 9, e1000595 (2011).
Yoshitane, H. et al. CLOCK-controlled polyphonic regulation of circadian rhythms through canonical and noncanonical E-boxes. Mol. Cell Biol. 34, 1776–1787 (2014).
Lowrey, P. L. & Takahashi, J. S. Mammalian circadian biology: elucidating genome-wide levels of temporal organization. Annu. Rev. Genomics Hum. Genet. 5, 407–441 (2004).
Beytebiere, J. R. et al. Tissue-specific BMAL1 cistromes reveal that rhythmic transcription is associated with rhythmic enhancer-enhancer interactions. Genes Dev. 33, 294–309 (2019).
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002).
Storch, K. F. et al. Extensive and divergent circadian gene expression in liver and heart. Nature 417, 78–83 (2002).
Mure, L. S. et al. Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 359. https://doi.org/10.1126/science.aao0318 (2018).
Qu, M., Duffy, T., Hirota, T. & Kay, S. A. Nuclear receptor HNF4A transrepresses CLOCK:BMAL1 and modulates tissue-specific circadian networks. Proc. Natl Acad. Sci. USA 115, E12305–E12312 (2018).
Yeung, J. et al. Transcription factor activity rhythms and tissue-specific chromatin interactions explain circadian gene expression across organs. Genome Res 28, 182–191 (2018).
Menet, J. S., Pescatore, S. & Rosbash, M. CLOCK:BMAL1 is a pioneer-like transcription factor. Genes Dev. 28, 8–13 (2014).
Welsh, D. K., Takahashi, J. S. & Kay, S. A. Suprachiasmatic nucleus: cell autonomy and network properties. Annu Rev. Physiol. 72, 551–577 (2010).
Hastings, M. H., Reddy, A. B. & Maywood, E. S. A clockwork web: circadian timing in brain and periphery, in health and disease. Nat. Rev. Neurosci. 4, 649–661 (2003).
Cedernaes, J. et al. Acute sleep loss results in tissue-specific alterations in genome-wide DNA methylation state and metabolic fuel utilization in humans. Sci. Adv. 4, eaar8590 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genoty** with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Hughes, M. E., Hogenesch, J. B. & Kornacker, K. JTK_CYCLE: an efficient nonparametric algorithm for detecting rhythmic components in genome-scale data sets. J. Biol. rhythms 25, 372–380 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Mosig, R. A. & Kojima, S. Timing without coding: How do long non-coding RNAs regulate circadian rhythms? Semin. cell Developmental Biol. 126, 79–86 (2022).
Greco, C. M. et al. Integration of feeding behavior by the liver circadian clock reveals network dependency of metabolic rhythms. Sci. Adv. 7, eabi7828 (2021).
Renoux, F. et al. The AP1 Transcription Factor Fosl2 Promotes Systemic Autoimmunity and Inflammation by Repressing Treg Development. Cell Rep. 31, 107826 (2020).
Ronin, E. et al. The NF-κB RelA Transcription Factor Is Critical for Regulatory T Cell Activation and Stability. Front. Immunol. 10, 2487 (2019).
Tamura, T., Yanai, H., Savitsky, D. & Taniguchi, T. The IRF family transcription factors in immunity and oncogenesis. Annu. Rev. Immunol. 26, 535–584 (2008).
Kiyose, H. et al. Comprehensive analysis of full-length transcripts reveals novel splicing abnormalities and oncogenic transcripts in liver cancer. PLoS Genet. 18, e1010342 (2022).
Zhou, Y. et al. A network medicine approach to investigation and population-based validation of disease manifestations and drug repurposing for COVID-19. PLoS Biol. 18, e3000970 (2020).
Schaum, N. et al. Ageing hallmarks exhibit organ-specific temporal signatures. Nature 583, 596–602 (2020).
Budovsky, A. et al. LongevityMap: a database of human genetic variants associated with longevity. Trends Genet 29, 559–560 (2013).
Sato, S. et al. Circadian Reprogramming in the Liver Identifies Metabolic Pathways of Aging. Cell 170, 664–677.e611 (2017).
Bertolotti, M. et al. Nonalcoholic fatty liver disease and aging: epidemiology to management. World J. Gastroenterol. 20, 14185–14204 (2014).
Karlsson, M. et al. A single-cell type transcriptomics map of human tissues. Sci. Adv. 7. https://doi.org/10.1126/sciadv.abh2169 (2021).
Guan, D. et al. The hepatocyte clock and feeding control chronophysiology of multiple liver cell types. Science 369, 1388–1394 (2020).
Li, J., Nie, P., Turck, C. W. & Wang, G. Z. Gene networks under circadian control exhibit diurnal organization in primate organs. Commun. Biol. 5, 764 (2022).
Paxman, J. et al. Age-dependent aggregation of ribosomal RNA-binding proteins links deterioration in chromatin stability with challenges to proteostasis. Elife 11. https://doi.org/10.7554/eLife.75978 (2022).
Desmet, L., Thijs, T., Mas, R., Verbeke, K. & Depoortere, I. Time-Restricted Feeding in Mice Prevents the Disruption of the Peripheral Circadian Clocks and Its Metabolic Impact during Chronic Jetlag. Nutrients 13. https://doi.org/10.3390/nu13113846 (2021).
Charlot, A., Hutt, F., Sabatier, E., and Zoll, J. Beneficial Effects of Early Time-Restricted Feeding on Metabolic Diseases: Importance of Aligning Food Habits with the Circadian Clock. Nutrients 13. https://doi.org/10.3390/nu13051405 (2021).
Hou, T. et al. Time-restricted feeding protects the blood pressure circadian rhythm in diabetic mice. Proc Natl Acad Sci USA. 118. https://doi.org/10.1073/pnas.2015873118 (2021).
**n, H. et al. A multi-tissue multi-omics analysis reveals distinct kineztics in entrainment of diurnal transcriptomes by inverted feeding. iScience 24, 102335 (2021).
Sinturel, F. et al. Diurnal Oscillations in Liver Mass and Cell Size Accompany Ribosome Assembly Cycles. Cell 169, 651–663 e614 (2017).
Hoekstra, M. M., Jan, M., Katsioudi, G., Emmenegger, Y., Franken, P. The sleep-wake distribution contributes to the peripheral rhythms in PERIOD-2. Elife 10. https://doi.org/10.7554/eLife.69773 (2021).
Lu, Y., Liu, B., Ma, J., Yang, S. & Huang, J. Disruption of Circadian Transcriptome in Lung by Acute Sleep Deprivation. Front. Genet. 12, 664334 (2021).
Challet, E., Turek, F. W., Laute, M. & Van Reeth, O. Sleep deprivation decreases phase-shift responses of circadian rhythms to light in the mouse: role of serotonergic and metabolic signals. Brain Res. 909, 81–91 (2001).
Barclay, J. L. et al. Circadian desynchrony promotes metabolic disruption in a mouse model of shiftwork. PloS one 7, e37150 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Wu, G., Anafi, R. C., Hughes, M. E., Kornacker, K. & Hogenesch, J. B. MetaCycle: an integrated R package to evaluate periodicity in large scale data. Bioinformatics 32, 3351–3353 (2016).
Yanai, I. et al. Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification. Bioinformatics 21, 650–659 (2005).
**ao, S. J., Zhang, C., Zou, Q. & Ji, Z. L. TiSGeD: a database for tissue-specific genes. Bioinformatics 26, 1273–1275 (2010).
Yu, X., Lin, J., Zack, D. J. & Qian, J. Computational analysis of tissue-specific combinatorial gene regulation: predicting interaction between transcription factors in human tissues. Nucleic Acids Res 34, 4925–4936 (2006).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 9, 559 (2008).
Qin, Q. et al. Lisa: inferring transcriptional regulators through integrative modeling of public chromatin accessibility and ChIP-seq data. Genome Biol. 21, 32 (2020).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov. (Camb. (Mass)) 2, 100141 (2021).
Wang, J. Z., Du, Z., Payattakool, R., Yu, P. S. & Chen, C. F. A new method to measure the semantic similarity of GO terms. Bioinformatics 23, 1274–1281 (2007).
Stenson, P. D. et al. Human Gene Mutation Database (HGMD): 2003 update. Hum. Mutat. 21, 577–581 (2003).
Rappaport, N. et al. MalaCards: an integrated compendium for diseases and their annotation. Database (Oxf.) 2013, bat018 (2013).
Rappaport, N. et al. MalaCards: A Comprehensive Automatically-Mined Database of Human Diseases. Curr Protoc Bioinformatics 47, 1 24 21-19. https://doi.org/10.1002/0471250953.bi0124s47 (2014).
Rappaport, N. et al. MalaCards: an amalgamated human disease compendium with diverse clinical and genetic annotation and structured search. Nucleic Acids Res 45, D877–D887 (2017).
Cunningham, F. et al. Ensembl 2022. Nucleic Acids Res 50, D988–D995 (2022).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Acknowledgements
This work is supported by grants from the National Natural Science Foundation of China (Grant Nos. 81827901 and 32170567) to G-Z, W. Y, X, is supported by the Lingang Laboratory & National Key Laboratory of Human Factors Engineering Joint Grant (LG-TKN-202203-01).
Author information
Authors and Affiliations
Contributions
Conceptualization, G-Z.W.; Formal Analysis, R.C., J.M. and Z.Z.; Experiment, B.L., Z.H., G.H., J.H. and Y.X.; Writing – Review & Editing, R.C., Z.Z. and G-Z.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Joao Valente.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, R., Zhang, Z., Ma, J. et al. Circadian-driven tissue specificity is constrained under caloric restricted feeding conditions. Commun Biol 7, 752 (2024). https://doi.org/10.1038/s42003-024-06421-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06421-0
- Springer Nature Limited