Abstract
Single-cell multi-omics are powerful means to study cell-to-cell heterogeneity. Here, we present a single-tube, bisulfite-free method for the simultaneous, genome-wide analysis of DNA methylation and genetic variants in single cells: epigenomics and genomics of single cells analyzed by restriction (epi-gSCAR). By applying this method, we obtained DNA methylation measurements of up to 506,063 CpGs and up to 1,244,188 single-nucleotide variants from single acute myeloid leukemia-derived cells. We demonstrate that epi-gSCAR generates accurate and reproducible measurements of DNA methylation and allows to differentiate between cell lines based on the DNA methylation and genetic profiles.
Similar content being viewed by others
Introduction
Single-cell multi-omics are revolutionizing our understanding of cell-to-cell variability1,2,3,4. These techniques offer to link genomic, epigenomic, and transcriptomic information from the same cell, and therefore allow to study cell-to-cell variability at unprecedented resolution. However, the simultaneous analysis of genetic variants and DNA methylation in single cells remains challenging. The analysis of single-cell DNA methylation typically relies on methylation-sensitive restriction enzymes (MSRE) or bisulfite conversion. The latter is considered the gold standard for genome-wide methylation analysis, but its application to single cells is hampered by DNA degradation5, resulting in high dropout levels4. Moreover, the bisulfite-induced C > T substitutions impact the ability to concurrently detect gene variants6. MSRE-based single-cell approaches typically rely on PCR-based readout and are thus limited in the number of evaluable loci per cell2.
Here, we report an MSRE-based method with genome-wide readout, which facilitates simultaneous analysis of DNA methylation and genetic variants of the same cell at base-pair resolution (epigenomics and genomics of single cells analyzed by restriction; epi-gSCAR). We devised epi-gSCAR as a multistep single-tube workflow which minimizes DNA loss and reduces the risk of contamination. epi-gSCAR features accurate and reproducible characterization of DNA methylation, while preserving the vast majority of genetic information with moderate incidence of dropouts. Thus, epi-gSCAR allows to identify cell-to-cell differences in the DNA methylation profile, and to assign these differences to a given genotype. The latter is of particular importance for the analyses of leukemias and other cancer specimens, since the malignant cells usually differ from healthy cells and among themselves by the genetic aberrations acquired (i.e., clonal heterogeneity).
Results
epi-gSCAR workflow
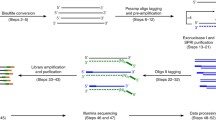
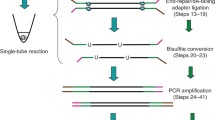
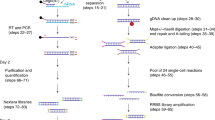
Analogous to previously published methods2,7, epi-gSCAR employs digestion using the MSRE HhaI, which results in cleavage of unmethylated recognition sites, while methylated sites stay intact (Fig. 1a). Terminal deoxynucleotidyl transferase (TdT) efficiently adds a 3′ poly(d)A tail to the generated DNA ends, which carry the genome-wide information of unmethylated recognition sites. The resulting tagged restriction enzyme scars serve as priming sites for GAT-oligo(dT)12-adapters8 containing a constant nucleotide 5′ sequence10,11; and (iii) the human genome contains 1.69 million HhaI recognition sites, providing superior genome-wide and feature-specific coverage when compared to the Infinium HumanMethylation450 BeadChip (450 K) or MethylationEPIC Kit array (Supplementary Fig. 1). In particular, CpG islands (CGIs) and transcription start sites (TSSs) are strongly enriched for HhaI sites (Fig. 1b). CGI shores, shelves and Fantom5 enhancers show HhaI coverage that is comparable to the aforementioned conventional cell-bulk assays (Supplementary Fig. 1).
Application of epi-gSCAR to measure site-specific CpG methylation
First, we applied 27 single cells of the human acute myeloid leukemia (AML) cell line Kasumi-1 to two variants of the epi-gSCAR workflow (Fig. 1a). For all single cells subjected to the epi-gSCAR assay, we could verify successful amplification of library DNA by agarose gel electrophoresis (Supplementary Fig. 2a and Supplementary Data 1). Product quality and fragment size distribution were additionally assessed on a Bioanalyzer (Agilent) High-Sensitivity DNA chip for selected reactions (Supplementary Fig. 2b).
In this first set of cells, we tested whether methylation of individual CpG sites can be assessed by targeted amplification of the loci of interest from the epi-gSCAR library. For this, we utilized step-out PCR, which facilitates isolation of amplicon ends regardless whether the target fragment contains intact HhaI sites or scar-tagged DNA ends12. This enables a convenient and cost-effective targeted readout of single-cell DNA methylation by conventional sequencing. We determined the DNA methylation status of two individual CGIs located within the promoters of the long and short isoforms of DLX4 in single cell K_05 (Fig. 1c). CGI1 was determined to be largely unmethylated, while CGI2 showed strong methylation (Fig. 1d). The results compared well with the DNA methylation levels of six other single cells analyzed by NGS (discussed below), and with Kasumi-1 bulk data derived from 450 K arrays or previously published whole-genome bisulfite sequencing (WGBS)13 (Fig. 1d, e).
Application of epi-gSCAR to measure genome-wide CpG methylation
To examine the potential of epi-gSCAR to measure genome-wide DNA methylation, we applied NGS to 7 of the 27 libraries (K_01–K_07). These libraries were sequenced at low depth (10.15–20.78 million mapped reads per cell; 0.41×–0.82× mean depth), achieving up to 18.8% genome coverage at ≥1× depth (Supplementary Data 2). NGS data were then analyzed using a custom bioinformatic pipeline (Supplementary Fig. 3).
To assess the overall quality of the data obtained by epi-gSCAR, we first analyzed the genome-wide methylation profiles of CGIs, gene bodies and five histone marks (Supplementary Fig. 4). The derived profiles were in line with those described in the literature14,15. We also compared the single-cell methylation datasets with cell-bulk methylome data obtained from 450 K arrays and from WGBS of Kasumi-1 and observed that the single-cell datasets well resembled the profiles of the cell-bulk controls (Supplementary Fig. 4). HhaI digestion efficiency as assessed by the analysis of non-methylated spike-in DNA was ≥98.3% (Supplementary Fig. 5b–d). Conversely, we confirmed complete digestion blockage of methylated spike-in DNA (Supplementary Fig. 5a).
Assessment of epigenetic heterogeneity using epi-gSCAR
We next assessed the ability of epi-gSCAR to differentiate between different cell lines based on the DNA methylation profiles. In addition to Kasumi-1, we selected OCI-AML3 as a second cell line which harbors a common, AML typical gene mutation in the DNMT3a gene and features a pronounced hypomethylation phenotype13. In order to directly compare both cell lines, we applied epi-gSCAR to 80 single cells of each cell line, which resulted in successful amplification of library DNA for all reactions as assessed by agarose gel electrophoresis. Based on the visual verification of single-cell deposition (Supplementary Data 3), we selected 20 cells (K_08–K_27 and O_01–O_20) for NGS analysis using our low-coverage approach (13.67–30.85 million mapped reads per cell; 0.62×–1.37× mean depth; Supplementary Data 2). The generated NGS datasets were used to assess and directly compare DNA methylation and genetic variant features of the cell lines.
For all analyzed cells, we obtained data on 214,634–506,063 CpG dinucleotides (mean: 373,058), corresponding to 0.78–1.85% (mean: 1.36%) of all informative CpG dinucleotides and 13.3–31.6% (mean: 23.2%) of HhaI sites (Supplementary Data 2). For both Kasumi-1 and OCI-AML3, covered CpG dinucleotides provided information on various genomic features, including CGI promoters, non-CGI promoters, orphan CGIs, gene bodies, and intergenic regions, which closely resembled the theoretical distribution of HhaI sites (Supplementary Fig. 6).
Next, we analyzed the sequencing coverage bias and found that the bias was slightly higher than that observed for published MALBAC datasets, while the coverage was more uniform than that of published multiple displacement amplification datasets (Supplementary Fig. 7).
For the second batch of single cells (K_08–K_27 and O_01–O_20), HhaI digestion efficiency was assessed by the analysis of non-methylated random spike-in control DNA (Supplementary Data 2). Reads containing spike-in DNA and covering the unmethylated HhaI control site could be detected in 22 of 40 single-cell libraries. All 22 libraries only contained spike-in amplicons with tailed HhaI scars, which confirmed complete digestion (Supplementary Data 2). Complete digestion for all processed single-cell libraries can be assumed, since incomplete digestion would inevitably result in stochastic concordance decrease (see below).
We next generated and compared single-cell DNA methylation profiles for both cell lines across different histone marks, CGIs, and gene bodies, and could identify clearly distinct methylation profiles for both OCI-AML3 and Kasumi-1, respectively (Fig. 2a, b).
a Averaged epi-gSCAR methylation profiles (K_08–K_27 and O_01–O_20; light blue lines) and corresponding pseudo-bulk (light blue lines) and cellbulk profiles (WGBS, red lines; 450 K array, green lines) across five histone marks. b Averaged epi-gSCAR methylation profiles (K_08–K_27 and O_01–O_20; light blue lines) for CGIs, all genes (all GENCODE basic genes) and genes grouped into three groups (0–20%, >20–60%, and >60–100%) based on their RNA expression level in cell bulk as FPKM (fragments per million mapped reads per kilobase exon), and corresponding pseudo-bulk (light blue lines) and cell-bulk profiles (WGBS, red lines; 450 K array, green lines). For plotting of pseudo-bulk datasets, we used HhaI sites covered in at least 5 of 20 single cells in order to reduce coverage bias. Shown is the mean methylation across 150 bp windows for each feature set and 3 kb upstream and downstream.
Activating histone marks (H3K9ac, H3K4me3, H3K4me2, and H3K27ac) were associated with low levels of methylation in both cell lines, which is in accordance with previous reports15. As expected, the lowest methylation levels were present at H3K4me3 and H3K9ac peaks, which are enriched at active promoters and associated with increased activation of promoter or enhancer regions. For both cell lines, the highest methylation values were measured for Polycomb repression-associated H3K27me3 peaks (Fig. 2a)14.
Analysis of CGIs revealed the expected depletion of methylation in Kasumi-1 and OCI-AML3 single cells. All analyzed single-cell methylomes exhibited low methylation levels around TSSs and high methylation levels within gene bodies (Fig. 2b).
We then analyzed gene body methylation in correlation with genome-wide gene expression levels by grou** genes based on their genome-wide RNA expression levels in cell bulk into three groups (0–20%, >20–60%, and >60%; Fig. 2b). For both, Kasumi-1 and OCI-AML3 single cells, methylation profiles were in line with the described relationship between gene expression and DNA methylation in gene promoters and gene bodies, i.e., that the depletion of DNA methylation around TSSs and enrichment of methylation in gene bodies correlates with higher expression rates14. Indeed, the most highly expressed genes consistently showed the lowest levels of methylation around the TSS and the strongest enrichment of methylation toward the 3′-end of the gene body, although the latter was less prominent when comparing the gene groups of >20–60% and >60% expression in OCI-AML3.
Interestingly, overall DNA methylation levels of profiles for histone marks, CGIs, and gene bodies were apparently lower for OCI-AML3 in comparison to Kasumi-1 single cells.
To compare our single-cell data with cell-bulk methylome data obtained from 450 K arrays and WGBS of Kasumi-1 and OCI-AML3 cells, we generated synthetic pseudo-bulk samples from the single-cell datasets (Supplementary Fig. 9). Although the number of covered CpGs did not reach saturation, 79.38% (n = 1,277,093) and 74.56% (n = 1,198,840) of all informative HhaI sites were covered, using 20 single cells of Kasumi-1 and OCI-AML3, respectively (K_08–K_27 and OC_01–OC_20; Supplementary Fig. 9).
Next, we analyzed how well the pseudo-bulk methylomes (K_08–K_27 and O_01–O_20) resembled the profiles measured in cell-bulk samples. We observed that the pseudo-bulk profiles largely resembled those derived from bulk WGBS and 450 K array data across all genetic features analyzed (Fig. 2a, b). As stated above, discrepancies in the methylation level between the pseudo-bulk and the cell-bulk controls (WGBS and 450 K array) are likely explained by HhaI-based coverage and local sequence context bias (Fig. 1b and Supplementary Fig. 1).
Analysis of single-cell mean methylation values across the entire genome confirmed that OCI-AML3 is strongly hypomethylated when compared with Kasumi-1 (58.8% vs. 79.8%), which is in line with the cell-bulk WGBS and 450 K array data and a previous study13 (Fig. 3a). To study variation among the single cells, we determined the pairwise CpG concordance across the single-cell libraries, separately for each cell line. Among Kasumi-1 single cells, the concordance was 80.3–93.9% (mean: 87.7%; Fig. 3b), and among OCI-AML3 single cells 77.6–85.2% (mean: 81.0%; Fig. 3c).
a Global single-cell methylation levels of Kasumi-1 (red dots) and OCI-AML3 (blue dots), and corresponding mean methylation levels (black lines) in comparison to mean methylation levels of WGBS (violet line) and 450 K array (green line) from cell-bulk samples. b Pairwise CpG concordance for all analyzed Kasumi-1 single cells. c Pairwise CpG concordance for all analyzed OCI-AML3 single cells. CpG concordance was calculated for all overlap** CpGs between each single-cell pair of each cell line as the fraction of CpGs with the identical methylation state (0, 0.5, or 1). Calculations are based on 66,588–297,161 CpGs for Kasumi-1 single-cell pairs and 148,932–257,837 CpGs for OCI-AML3 single-cell pairs. d Correlation between the mean pseudo-bulk methylation and the cell-bulk 450 K array, and WGBS datasets for Kasumi-1 and OCI-AML3. Comparisons consider genome-wide methylation of individual CpGs covered in ≥15 single cells (Kasumi-1 450 K: n = 6,607; Kasumi-1 WGBS: n = 20,000 of 72,142 CpGs covered by epi-gSCAR in ≥15 single cells; OCI-AML3 450 K: n = 11,511; OCI-AML3 WGBS: n = 20,000 of 129,153 CpGs covered by epi-gSCAR in ≥15 single cells). e Circos plot representation of genome-wide methylation profiles of randomly selected single cells, the pseudo-bulk datasets, and WGBS controls. The heatmaps show average methylation levels for 200 kb windows. Heatmap colors indicate methylation levels from low (blue) to high (red). Tracks from inside to outside represent single cells O_01, O_05, O_11, K_11, K_16, and K_17, OCI-AML3 pseudo-bulk (O_01–O_20), OCI-AML3 cell-bulk WGBS, Kasumi-1 pseudo-bulk (K_08–K_27), and Kasumi-1 cell-bulk WGBS. f Hierarchical clustering analysis based on Pearson correlation coefficients for single cells K_01–K_07 (yellow), K_08–K_27 (red), and O_01–O_20 (blue) across 200 kb windows. g Multidimensional scaling analysis using UMAP, in which each dot represents a single cell (K_01–K_07, yellow; K_08–K_27, red; and O_01–O_20, blue). Cells are clustered based on the methylation levels across 200 kb windows covered in all single cells (top; n = 10,555) or based on genetic variants called at positions covered in all single cells (bottom; n = 7,027). For genetic variant clustering, SNV data was converted into a categorical numeric matrix as an input to compute UMAP with the R package ggplot.
To assess global similarities at the CpG level, we compared the synthetic pseudo-bulk methylomes with the 450 K array, and WGBS bulk methylation datasets of Kasumi-1 and OCI-AML3. By calculating Pearson correlation coefficients (R), we found that the synthetic bulk methylomes highly correlated with the profiles derived from both cell-bulk assays (Kasumi-1, 450 K R = 0.95, WGBS R = 0.89; OCI-AML3, 450 K R = 0.93, WGBS R = 0.81; Fig. 3d). Circos plot representation of genome-wide methylation profiles confirmed a high concordance of the pseudo-bulk datasets and the respective WGBS cell-bulk datasets, and demonstrated remarkably distinct methylation landscapes for Kasumi-1 and OCI-AML3 at the pseudo-bulk and single-cell level (Fig. 3e).
To demonstrate that epi-gSCAR can assign a single cell to its cell line of origin based on the methylation patterns, we first assessed global similarities by calculating pairwise Pearson correlation coefficients across all single-cell datasets. Hierarchical clustering demonstrated that single cells of the respective cell line clustered together (Fig. 3f). We further confirmed this by dimension reduction using UMAP16 to project the single cells in a two-dimensional space. This again revealed clearly defined Kasumi-1 and OCI-AML3 clusters, and demonstrated that epi-gSCAR can distinguish cells based on their DNA methylation signatures (Fig. 3g).
Evaluation of genetic heterogeneity using epi-gSCAR
In order to show that epi-gSCAR can be applied to identify single cells not only based on their individual methylation signature, but also based on genetic variant information we searched for single-nucleotide variants (SNVs) in cells K_08–K_27 and O_01–O_20 using monovar17. On average, over 800,000 SNVs (range 498,097–1,244,188) were detected per single cell with ≥10× coverage (Supplementary Data 2). Based on the genome-wide SNV data, we performed UMAP clustering and identified two distinct single-cell clusters corresponding to the two analyzed cell lines (Fig. 3g).
Allelic dropout (ADO) rates were estimated to be as low as 20.7% and comparable to ADO rates achieved by commercially available MALBAC Kits18,Preparation of unmethylated and methylated spike-in DNA amplicons For K_01–K_07, two spike-in amplicons containing each a single HhaI recognition site were PCR-amplified from lambda DNA and then purified, using the Monarch PCR & DNA Cleanup Kit (NEB), according to the manufacturer’s recommendations (see Supplementary Data 2 for oligonucleotide sequences). For use as a methylated control, 1 µg of the spike-in meth lambda PCR product was methylated in 50 µl containing 2 µl M.SssI CpG Methyltransferase (4 units/µl; NEB) in 1× NEBuffer 4, supplemented with 0.3 µl 200× SAM. The reaction was incubated at 37 °C for 16 h, and another 0.3 µl of 200× SAM and 2 µl M.SssI CpG Methyltransferase were added, and the incubation was continued for further 6 h. The amplicon was purified and complete protection from HhaI cleavage was controlled by standard restriction digest protocol with HhaI followed by agarose gel electrophoresis. For K_08–K_27 and O_01–O_20 a random 300 bp spike-in oligonucleotide (eurofins genomics) containing a single HhaI recognition site was amplified and then purified, using the Monarch PCR and DNA Cleanup Kit (NEB), according to the manufacturer’s recommendations. Methylated and unmethylated exogenous spike-in DNA was used to control for digestion efficiency. Each amplicon contained one HhaI site. For K_01–K_07 we spiked in 30 ag (assay variant A, single cell K_01–K_04) or 3 ag (assay variant B, single cell K_05–K_07) of unmethylated and methylated lambda spike-in amplicon. This corresponds to ~110 or 11 and 60 or 6 molecules of unmethylated and methylated amplicons, respectively. For K_08–K_27 and O_01–O_20, we spiked in 10 ag of randomly unmethylated control DNA corresponding to ~30 molecules (Supplementary Data 2). For calculation of the digestion efficiency, the all-read files were filtered for reads matching corresponding amplicons using BBDuk, and aligned to the enterobacteria phage lambda reference genome or the random control oligonucleotide sequence with BWA-MEM. Then, local coverage of scar or intact reads at the control HhaI site was calculated. Quality of raw sequence data was controlled with FastQC and sequencing adapters were trimmed with Trim Galore in automatic detection mode, using standard settings. Preprocessed reads were aligned to the human genome GRCh37 (hg19) using Bowtie2 and peaks were identified with MACS2 callpeak. Peaks were called relative to the respective input controls when available. Publicly available MALBAC and MDA datasets were downloaded from the European Nucleotide Archive (SRS2062840; MALBAC, Yikon, Single Cell YK5) and the Sequence Read Archive (SRR617646; MALBAC, sw480 single cellKasumi-1 and OCI-AML3 bulk RNA-seq analysis Quality of raw sequence data was tested with FastQC and sequencing adapters were trimmed with Cutadapt, using standard settings. Preprocessed reads were aligned to the human genome GRCh37 (hg19) using RNA STAR, and fragments per million mapped reads per kilobase exon were determined using FPKM count of the RSeQC package. Expressed genes were grouped based on their FPKM distribution into three groups (0–20%, 20–60%, and >60% expression). Illumina 450k Infinium methylation array data was normalized with the beta mixture quantile algorithm and further analyzed using routines from the RnBeads software package30,31. For OCI-AML3, publicly available 450 K array data from the Gene Expression Omnibus (GEO) database were used (GSM1670296). WGBS of Kasumi-1 and OCI-AML3 was performed, as described elsewhere13, the data were kindly provided by David H. Spencer (Washington University School of Medicine, St. Louis, MO, USA). For single cells K_01–K_07 mapped all-read epi-gSCAR bam files were used as inputs for variant detection with FreeBayes in simple diploid calling mode. Only mapped reads with map** quality >30 and base quality score >20 were utilized for variant identification. Minimum coverage was set to ≥10× for single-cell libraries. Calculation of ADO estimates was based on Human SNP Array 6.0 (Affymetrix) data of Kasumi-1 (ref. 21). SNPs with variant allele frequencies ≥0.45 and ≤0.55 were defined as heterozygous. ADO estimates were calculated as the fraction of SNPs called as heterozygous in SNP array data of Kasumi-1 cell-bulk DNA though called homozygous in single cells for loci with coverage ≥10×. ADO rate estimates were additionally calculated for loci with coverage ≥3× and ≥6× in single cells. For genoty** of single cells K_08–K_27 and O_01–O_20, we processed the all-read epi-gSCAR bam files with monovar using standard settings with consensus-filtering step enabled, which removes variants with support from only one cell. In order to calculate ADO rate estimates, we filtered Kasumi-1 and OCI-AML3 SNP 6.0 datasets for heterozygous SNPs with high confidence values (0.999). For OCI-AML3, publicly available Human SNP Array 6.0 array data was downloaded from the GEO database under accession GSM888549. Genotype calling of SNP 6.0 data was performed using the R package CRLMM. ADO estimates were calculated as the fraction of SNPs called as heterozygous in SNP 6.0 datasets, while called homozygous in single cells for loci with ≥20× coverage. For the generation of Fig. 3f, Pearson correlation coefficients (R) were calculated using the tool plotCorrelation.py of the deepTools 3.3 package32. For the calculation of Pearson correlation coefficients (R) for Fig. 3d, we used the function stat_cor in package ggpubr R package (v 0.4.0/R version 3.5.2). We applied epi-gSCAR to a total of 214 single cells in four independent experiments and obtained reproducible results among the epi-gSCAR libraries subjected to sequencing (27 single Kasumi-1 single cells and 20 OCI-AML3 single cells). Further information on research design is available in the Nature Research Reporting Summary linked to this article.Evaluation of digestion efficiency of spike-in amplicons
ChIP-seq analysis
Analysis of MALBAC and MDA datasets
DNA methylation analysis using the Illumina 450 K BeadChip array and WGBS
SNP array analysis, variant calling, and ADO rate estimation
Statistics and reproducibility
Reporting summary
Data availability
All sequencing data (FASTQ) and processed single-cell DNA methylation data (BEDGRAPH) have been deposited in the Gene Expression Omnibus (GEO) database under accession GSE131723. Publicly available ChIP-seq and RNA-seq datasets used in this study were obtained from the GEO data portal with the following accessions: GSE29225, GSE62847, GSE83660, GSM1844449, GSM1844483, GSM3024903, GSM3032904, GSM3024909, and GSM3032912. MALBAC and MDA datasets were obtained from the European Nucleotide Archive (SRS2062840) and the Sequence Read Archive (SRR617646 and SRR5219394). Publicly available Human SNP Array 6.0 array data was downloaded from the GEO (GSM888549). Kasumi-1 Human SNP Array 6.0 array data is available at EMBL-EBI ArrayExpress (E-MTAB-4950). List of figures that have associated raw data: Figs. 1d, e, 2a, b, and 3a–g, and Supplementary Figs. 4, 6–11.
Code availability
The bioinformatic pipeline used for readout of methylation information from epi-gSCAR libraries by NGS is illustrated in Supplementary Fig. 3. The code is available from https://github.com/wehrleju/epi-gSCAR.
References
Dey, S. S., Kester, L., Spanjaard, B., Bienko, M. & Van Oudenaarden, A. Integrated genome and transcriptome sequencing of the same cell. Nat. Biotechnol. 33, 285–289 (2015).
Cheow, L. F. et al. Single-cell multimodal profiling reveals cellular epigenetic heterogeneity. Nat. Methods 13, 833–836 (2016).
Macaulay, I. C. et al. G&T-seq: parallel sequencing of single-cell genomes and transcriptomes. Nat. Methods 12, 519–522 (2015).
Hu, Y. et al. Simultaneous profiling of transcriptome and DNA methylome from a single cell. Genome Biol. 17, 88 (2016).
Yong, W.-S., Hsu, F.-M. & Chen, P.-Y. Profiling genome-wide DNA methylation. Epigenetics Chromatin 9, 26 (2016).
Liu, Y., Siegmund, K. D., Laird, P. W. & Berman, B. P. Bis-SNP: combined DNA methylation and SNP calling for bisulfite-seq data. Genome Biol. 13, R61 (2012).
Cheow, L. F., Quake, S. R., Burkholder, W. F. & Messerschmidt, D. M. Multiplexed locus-specific analysis of DNA methylation in single cells. Nat. Protoc. 10, 619–631 (2015).
Chapman, A. R. et al. Single cell transcriptome amplification with MALBAC. PLoS ONE 10, e0120889 (2015).
Zong, C., Lu, S., Chapman, A. R. & **e, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Nygren, A. O. H. et al. Methylation-specific MLPA (MS-MLPA): simultaneous detection of CpG methylation and copy number changes of up to 40 sequences. Nucleic Acids Res. 33, 1–9 (2005).
Korch, C. & Hagblom, P. In-vivo-modified gonococcal plasmid pJD1: a model system for analysis of restriction enzyme sensitivity to DNA modifications. Eur. J. Biochem. 161, 519–524 (1986).
Matz, M. et al. Amplification of cDNA ends based on template-switching effect and step-out PCR. Nucleic Acids Res. 27, 1558–1560 (1999).
Chen, D. et al. DNMT3AR882-associated hypomethylation patterns are maintained in primary AML xenografts, but not in the DNMT3AR882COCI-AML3 leukemia cell line. Blood Cancer J. 8, 0–3 (2018).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–329 (2015).
Sharifi-Zarchi, A. et al. DNA methylation regulates discrimination of enhancers from promoters through a H3K4me1-H3K4me3 seesaw mechanism. BMC Genomics 18, 1–21 (2017).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: Uniform Manifold Approximation and Projection. J. Open Source Softw. 3, 861 (2018).
Zafar, H., Wang, Y., Nakhleh, L., Navin, N. & Chen, K. Monovar: single-nucleotide variant detection in single cells. Nat. Methods 13, 505–507 (2016).
Picher, Á. J. et al. TruePrime is a novel method for whole-genome amplification from single cells based on TthPrimPol. Nat. Commun. 7, 13296 (2016).
Huang, L., Ma, F., Chapman, A., Lu, S. & **e, X. S. Single-cell whole-genome amplification and sequencing: methodology and applications. Annu. Rev. Genomics Hum. Genet. 16, 79–102 (2015).
Beghini, A., Magnani, I., Ripamonti, C. B. & Larizza, L. Amplification of a novel c-Kit activating mutation Asn(822)-Lys in the Kasumi-1 cell line: a t(8;21)-Kit mutant model for acute myeloid leukemia. Hematol. J. 3, 157–163 (2002).
Riba, J. et al. Molecular genetic characterization of individual cancer cells isolated via single-cell printing. PLoS ONE 11, 1–15 (2016).
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805–D811 (2015).
Smallwood, S. A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Hui, T. et al. High-resolution single-cell DNA methylation measurements reveal epigenetically distinct hematopoietic stem cell subpopulations. Stem Cell Rep. 11, 578–592 (2018).
Viswanathan, R., Cheruba, E. & Cheow, L. F. DNA Analysis by Restriction Enzyme (DARE) enables concurrent genomic and epigenomic characterization of single cells. Nucleic Acids Res. 47, e122 (2019).
Han, L. et al. Bisulfite-independent analysis of CpG island methylation enables genome-scale stratification of single cells. Nucleic Acids Res. 45, 1–13 (2017).
Han, K. Y. et al. SIDR: simultaneous isolation and parallel sequencing of genomic DNA and total RNA from single cells. Genome Res. 28, 75–87 (2018).
Stosch, J. M. et al. Gene mutations and clonal architecture in myelodysplastic syndromes and changes upon progression to acute myeloid leukaemia and under treatment. Br. J. Haematol. 182, 830–842 (2018).
Börgstrom, E., Paterlini, M., Mold, J. E., Frisen, J. & Lundeberg, J. Comparison of whole genome amplification techniques for human single cell exome sequencing. PLoS ONE 12, 1–15 (2017).
Assenov, Y. et al. Comprehensive analysis of DNA methylation data with RnBeads. Nat. Methods 11, 1138–1140 (2014).
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2013).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Acknowledgements
The authors thank David H. Spencer (Washington University School of Medicine, St. Louis, MO) for providing the whole-genome bisulfite sequencing data on Kasumi-1 and OCI-AML3. Moreover, the authors acknowledge the provision of bioinformatic tools and support by the Freiburg Galaxy team (Björn Grüning and Rolf Backofen; Dept. of Computer Science, University of Freiburg). The authors also thank Gabriele Greve (Dept. of Medicine I, Medical Center — University of Freiburg) for the scientific discussions on DNA methylation patterns in cancer. This research was supported by the Translational Research Training in Hematology of the European Hematology Association and American Society of Hematology (H.B.); the German Research Foundation (Deutsche Forschungsgemeinschaft, DFG; SPP 1463 LU 429/8-2 [M.L.]; CRC992 MEDEP C04 [M.L.]; FOR 2674 BE 6461/1-1 A05 [H.B.], LU 429/16-1 A05 [M.L.], LI 2492/3-1 A06 [D.B.L.], and A09 [M.L., C.P.]), the Böhringer Ingelheim Foundation (Exploration Grant [H.B.]), the German Cancer Aid (Deutsche Krebshilfe; 111210 [H.B.]; 110461 [R.C.]; and 70112574 [D.B.L.]), the Berta Ottenstein Fellowship Program of the University of Freiburg (J.W.), the Fördergesellschaft Forschung Tumorbiologie Liquid-Biopsy Initiative (J.W.), the Federal Ministry of Education and Research (Bundesministerium für Bildung und Forschung, BMBF) Eurostars Project E!10257 (Julian Riba), and the German José Carreras Leukemia-Foundation (Deutsche José Carreras Leukämie-Stiftung; R 14/25 [M.L.], 17 R/2019 [H.B.]).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
C.N., J.W., D.B.L., J.D., M.L., and H.B. undertook conception and design of the study, C.N., Julian Riba, N.R., Janika Rhein, S.B., and J.M.S. performed or assisted with molecular and cellular experiments, C.N. and J.W. developed computational pipelines and analyzed the data, P.L. assisted with the computational analyses, R.C., J.D., C.P., P.L., D.B.L., M.L., and H.B. provided conceptual advice, M.L. and H.B. secured funding and supervised the work. C.N., J.W., and H.B. wrote the manuscript. All authors accepted the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
Julian Riba is an employee of Cytena GmbH, which produces the single-cell printer used in the study. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Niemöller, C., Wehrle, J., Riba, J. et al. Bisulfite-free epigenomics and genomics of single cells through methylation-sensitive restriction. Commun Biol 4, 153 (2021). https://doi.org/10.1038/s42003-021-01661-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-01661-w
- Springer Nature Limited