Abstract
Lung cancer is one of the most dangerous malignant tumors affecting human health. Lung adenocarcinoma (LUAD) is the most common subtype of lung cancer. Both glycolytic and cholesterogenic pathways play critical roles in metabolic adaptation to cancer. A dataset of 585 LUAD samples was downloaded from The Cancer Genome Atlas database. We obtained co-expressed glycolysis and cholesterogenesis genes by selecting and clustering genes from Molecular Signatures Database v7.5. We compared the prognosis of different subtypes and identified differentially expressed genes between subtypes. Predictive outcome events were modeled using machine learning, and the top 9 most important prognostic genes were selected by Shapley additive explanation analysis. A risk score model was built based on multivariate Cox analysis. LUAD patients were categorized into four metabolic subgroups: cholesterogenic, glycolytic, quiescent, and mixed. The worst prognosis was the mixed subtype. The prognostic model had great predictive performance in the test set. Patients with LUAD were effectively typed by glycolytic and cholesterogenic genes and were identified as having the worst prognosis in the glycolytic and cholesterogenic enriched gene groups. The prognostic model can provide an essential basis for clinicians to predict clinical outcomes for patients. The model was robust on the training and test datasets and had a great predictive performance.
Similar content being viewed by others
Introduction
Although lung cancer is the second most common cancer globally, it is the leading cause of cancer deaths, accounting for an annual estimated total of two million new cases and 1.76 million deaths1,2. Lung cancer can be broadly grouped into small-cell lung cancer (SCLC, 15%) and non-small-cell lung cancer (NSCLC, 85%), and lung adenocarcinoma (LUAD) is the most common subtype of NSCLC2,3,4. The treatment of NSCLC has changed dramatically over the past decade, primarily due to advances in biomarkers that allow for targeted and immune-based therapies for specific patients with significant success5. However, the vast majority of advanced NSCLC become resistant to current treatments and eventually progress6. Therefore, searching for new predictors to predict and improve the prognosis of LUAD is imminent.
Reprogramming of cell metabolism is an essential feature of malignancy, as shown by abnormal uptake of glucose and amino acids and dysregulation of glycolysis7,8. Glycolysis is a specific metabolic pattern of tumor cells, which meets the requirements of tumor cells for ATP, etc9. Mitochondrial pyruvate carrier (MPC) consists of MPC1, and MPC2 is responsible for the import of pyruvate from the cytoplasmic matrix into the mitochondrial matrix, which can affect glycolysis, and damaged MPC function may induce tumors with solid capabilities for proliferation, migration, and invasion10,11. Pyruvate is converted to acetyl coenzyme A, which is further changed to citric acid, a precursor substance required for lipogenesis, including the synthesis of cholesterol12. There is growing evidence for a close relationship between cholesterol metabolism and some types of cancer, such as allosteric interactions in the microenvironment of tumors, cancer cell spreading and metastasis forming, and lipid metabolism in tumor-initiating cells (TICs)13,14.
In recent decades, some studies have investigated potential prognostic signatures of LUAD using only bulk RNA- seq data, which principally provides data on the average of the total number of cells in the sample15,16. Single-cell sequencing is a powerful instrument for dissecting the cellular and molecular landscape with single-cell resolution, revolutionizing our comprehension of the biological features and dynamics within cancer pathologies17. Single-cell RNA-seq technology can comprehensively characterize the heterogeneity of the tumor microenvironment and help dissect the complex cell type compositions and expressive heterogeneity in TME, and the Tumor Immune Single Cell Hub (TISCH) can assist us with a simple analysis18.
The analysis of large volumes of complex biomedical data through computer algorithms, driven by the ongoing development of computer hardware and enormous amounts of data, offers substantial advantages for advancing biology and accurately estimating patient conditions19,20. Machine learning (ML) is a scientific discipline focusing on how computers learn from data and build predictive models. It is becoming an embedded part of modern research in biology, but its “black box” nature is an additional challenge21,22. Many algorithms are widely used to analyze complex biomedical data, such as extreme gradient boosting (XGBoost)23, Random Forest Classifier (RFC)24, Logistic Regression (LR)25, Support Vector Machine (SVM)26, and K-Nearest Neighbors (KNN)27. Interpretation is an integral branch of method development, with Shapley additive explanation (SHAP) being an integral approach28. The SHAP, which explains the model outcome by computing the contribution of each input feature for all samples, was applied to study the effects of different variables7C).
Analysis of the correlation between prognostic genes and TME
Given the role of TME in tumor development and its impact on prognosis, we used a NSCLC_GSE117570 dataset from the TISCH database to analyze the expression of some prognostic genes in TME-associated cells. We then examined the dataset, which is categorized into ten cell types. Figure 8A shows the number of cells of each cell type and presents the distribution of each type of TME-associated cells. In this dataset, malignant cells were the most abundant (n = 2721). We found that COL4A3, FOLR1, KRT6A, and SLC22A3 had higher expression in malignant cells compared to other types of TME-associated cells (Fig. 8B). These results support the association of prognostic genes with lung cancer.
Discussion
Lung cancer is one of the most deadly malignancies in humans, and most patients with advanced lung cancer experience recurrence and treatment resistance. The abnormal metabolism of cancer cells characterized by high glycolysis that occurs even in the presence of high amounts of oxygen, a metabolic reprogramming called the Warburg effect or aerobic glycolysis, has been recognized as a new hallmark of cancer34. Inhibition of glycolysis is considered a therapeutic option for aggressive cancers, including lung cancer, and related genes can be used as potential targets for metabolic therapy against cancer cells, such as ARID1A and circ-ENO135,36,37. Altered metabolism is not limited to cellular energy pathways but also includes alterations in lipid biosynthesis and other pathways (e.g., polyamine processing) in lung cancer and can affect its surrounding microenvironment38. It has been shown that lung cancer tissues demonstrate elevated cholesterol levels because the proliferation of cancer cells depends heavily on its availability. Strategies to reduce cholesterol synthesis or inhibit cholesterol uptake have been proposed as potential antineoplastic therapies39,40. Therefore, it is essential to clarify the metabolic pathways of lung cancer for its prevention and treatment.
In this study, based on 93 glycolysis and cholesterol synthesis genes, in order to find the most representative genes, consistent clustering was used to minimize the gene numbers, yielding 7 cholesterogenesis and glycolysis co-expressed genes, respectively. Based on these genes, the samples were classified into four subtypes: glycolytic, cholesterogenic, quiescent, and mixed. Although cholesterol plays a crucial role in tumors, survival analysis showed that cholesterol subtypes have a better prognosis than other subtypes, and randomized controlled trials could not support a survival benefit through lipid lowering in lung cancer patients, the reasons for which deserve further investigation41.
Pyruvate is central to carbohydrate, fat, and amino acid metabolism. Pyruvate is appealing as a therapeutic target against cancer because it promotes respiratory reserve capacity and mitochondrial oxygen consumption, which may contribute to the aggressive disease phenotype42. Mitochondrial pyruvate carrier(MPC) is one of the critical enzymes responsible for pyruvate transport and oxidation11. Low or absent MPC1 and MPC2 levels lead to metabolic disorders and alterations in tumor metabolism, and their restored expression inhibits tumor growth, invasiveness, metastasis, and stemness43. By analyzing the expression of MPC1/2, the results showed that there were significantly different expressions of MPC1/2 among different subtypes of metabolism, suggesting that the MPC complex affects the metabolic pathway and thus participates in the malignant progression of lung cancer by regulating the amount of pyruvate entering the mitochondria.
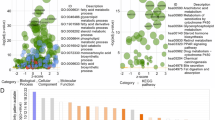
We identified DEGs between the best and worst prognosis subtypes and performed a functional enrichment analysis. The results showed significant enrichment of DEGs between the mixed and cholesterogenic subtypes in terms of p53 signaling pathways, microRNAs in cancer, and cell cycle. Then we decided to build the model using ML, with DEGs as features and ending events as labels. It performs best in the test set based on XGBoost’s powerful ability to handle complex classification problems. SHAP was then used to select the most important nine features. Then the most important nine features which SHAP selected were used to construct a prognostic model using a multivariate cox regression model. And this makes it possible to combine the excellent classification power of ML with the interpretability of the prognostic model.
Nine prognostic genes were included four non-coding RNA genes (RP11.87E22.2, RP11.789C1.1, AL589743.1, and MIR31HG) and five coding protein genes (COL4A3, SLC22A3, FOLR1, ESPL1, and KRT6A). After reviewing the kinds of literature, we found that the role of many prognostic genes has been studied concerning lung cancer and has been revealed to impact tumorigenesis and progression. Specifically, ESPL1 expression was positively correlated with SHAP values, and high expression of ESPL1 has been previously shown to be associated with poor prognosis in lung cancer by Zhao et al.44. Similarly, KRT6A, MIR31HG, and FOLR1 have been found to enhance lung cancer proliferation and may be potential therapeutic targets45,46,47. Analyzing the association between these genes and TME, we discovered that some prognostic genes were highly expressed in malignant cells using a single-cell sequencing database, which contributed to our construction of a better prognostic model. Subsequently, all samples were classified into high and low risk groups, and the clinical characteristics of the different risk groups were analyzed. Consistent results with the training set were observed in this test set. The model was robust on the training and test datasets and had a great predictive performance.
There are also some limitations to this study. Firstly, the performance of our model has not been tested externally, and there are doubts about its availability for large-scale use. Secondly, the biological associations between the selected prognostic genes remain to be investigated, and their biological explanations with prognostic profiles are to be explored. Future experimental verification is needed. Finally, using the median risk score as a cutoff value to classify high and low risk needs to be optimized.
Conclusions
Patients with LUAD were effectively typed by glycolytic and cholesterogenic genes and were identified as having the worst prognosis in the glycolytic and cholesterogenic enriched gene groups. Prognostic genes selected by the XGboost algorithm and SHAP analysis can be used to analyze patient prognosis. The prognostic models can provide an important basis for clinicians to predict clinical outcomes for patients. Of course, our model also has some challenges. In clinical practice, a large proportion of patients with LUDA do not undergo genetic testing. We hope to design a more popular model in the future.
Data availability
A publicly available dataset was analysed in this study. The data sets analyzed during the current study are available in UCSC Xena browser (https://xenabrowser.net/ (accessed on 1 July 2022)) and the Molecular Signatures Database v7.5 (http://www.gsea-msigdb.org/gsea/msigdb/ (accessed on 1 July 2022)).
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA. Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Thai, A. A., Solomon, B. J., Sequist, L. V., Gainor, J. F. & Heist, R. S. Lung Cancer. Lancet Lond. Engl. 398, 535–554. https://doi.org/10.1016/S0140-6736(21)00312-3 (2021).
Gridelli, C. et al. Non-small-cell lung cancer. Nat. Rev. Dis. Primer 1, 1–16. https://doi.org/10.1038/nrdp.2015.9 (2015).
MP, C.; B, E.; HR, S.; H, S.; J, F.; M, H.; P, B. Cancer Incidence in Five Continents Volume IX; ISBN 978-92-832-2160-9.
Herbst, R. S., Morgensztern, D. & Boshoff, C. The biology and management of non-small cell lung cancer. Nature 553, 446–454. https://doi.org/10.1038/nature25183 (2018).
Wang, M., Herbst, R. S. & Boshoff, C. Toward personalized treatment approaches for non-small-cell lung cancer. Nat. Med. 27, 1345–1356. https://doi.org/10.1038/s41591-021-01450-2 (2021).
Pavlova, N. N. & Thompson, C. B. The emerging hallmarks of cancer metabolism. Cell Metab. 23, 27–47. https://doi.org/10.1016/j.cmet.2015.12.006 (2016).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: The next generation. Cell 144, 646–674. https://doi.org/10.1016/j.cell.2011.02.013 (2011).
Warburg, O. On the origin of cancer cells. Science 123, 309–314. https://doi.org/10.1126/science.123.3191.309 (1956).
Ruiz-Iglesias, A. & Mañes, S. The importance of mitochondrial pyruvate carrier in cancer cell metabolism and tumorigenesis. Cancers 13, 1488. https://doi.org/10.3390/cancers13071488 (2021).
Herzig, S. et al. Identification and functional expression of the mitochondrial pyruvate carrier. Science 337, 93–96. https://doi.org/10.1126/science.1218530 (2012).
Baggetto, L. G. Deviant energetic metabolism of glycolytic cancer cells. Biochimie 74, 959–974. https://doi.org/10.1016/0300-9084(92)90016-8 (1992).
Snaebjornsson, M. T., Janaki-Raman, S. & Schulze, A. Greasing the wheels of the cancer machine: The role of lipid metabolism in cancer. Cell Metab. 31, 62–76. https://doi.org/10.1016/j.cmet.2019.11.010 (2020).
Luo, J., Yang, H. & Song, B.-L. Mechanisms and regulation of cholesterol homeostasis. Nat. Rev. Mol. Cell Biol. 21, 225–245. https://doi.org/10.1038/s41580-019-0190-7 (2020).
Yang, X. et al. Identification and validation of an immune cell infiltrating score predicting survival in patients with lung adenocarcinoma. J. Transl. Med. 17, 217. https://doi.org/10.1186/s12967-019-1964-6 (2019).
Chen, Z. et al. Identification of differentially expressed genes in lung adenocarcinoma cells using single-cell RNA sequencing not detected using traditional RNA sequencing and microarray. Lab. Investig. J. Tech. Methods Pathol. 100, 1318–1329. https://doi.org/10.1038/s41374-020-0428-1 (2020).
Lei, Y. et al. Applications of single-cell sequencing in cancer research: Progress and perspectives. J. Hematol. Oncol. 14, 91. https://doi.org/10.1186/s13045-021-01105-2 (2021).
Sun, D. et al. TISCH: A comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res. 49, D1420–D1430. https://doi.org/10.1093/nar/gkaa1020 (2020).
Min, S., Lee, B. & Yoon, S. Deep learning in bioinformatics. Brief. Bioinform. 18, 851–869. https://doi.org/10.1093/bib/bbw068 (2017).
Ngiam, K. Y. & Khor, I. W. Big data and machine learning algorithms for health-care delivery. Lancet Oncol. 20, e262–e273. https://doi.org/10.1016/S1470-2045(19)30149-4 (2019).
Camacho, D. M., Collins, K. M., Powers, R. K., Costello, J. C. & Collins, J. J. Next-generation machine learning for biological networks. Cell 173, 1581–1592. https://doi.org/10.1016/j.cell.2018.05.015 (2018).
Deo, R. C. Machine learning in medicine. Circulation 132, 1920–1930. https://doi.org/10.1161/CIRCULATIONAHA.115.001593 (2015).
Hou, N. et al. Predicting 30-days mortality for MIMIC-III patients with sepsis-3: A machine learning approach using XGboost. J. Transl. Med. 18(1), 462. https://doi.org/10.1186/s12967-020-02620-5 (2020).
Al-Barakati, H., Newman, R. H., Kc, D. B. & Poole, L. B. Bioinformatic analyses of peroxiredoxins and RF-Prx: A random forest-based predictor and classifier for Prxs. Methods Mol. Biol. 2499, 155–176. https://doi.org/10.1007/978-1-0716-2317-6_8 (2022).
Książek, W., Gandor, M. & Pławiak, P. Comparison of various approaches to combine logistic regression with genetic algorithms in survival prediction of hepatocellular carcinoma. Comput. Biol. Med. 134, 104431. https://doi.org/10.1016/j.compbiomed.2021.104431 (2021).
Huang, S. et al. Applications of support vector machine (SVM) learning in cancer genomics. Cancer Genom. Proteom. 15(1), 41–51. https://doi.org/10.21873/cgp.20063 (2018).
Tjärnberg, A. et al. Optimal tuning of weighted kNN- and diffusion-based methods for denoising single cell genomics data. PLoS Comput. Biol. 17(1), e1008569. https://doi.org/10.1371/journal.pcbi.1008569 (2021).
Zou, J. et al. A primer on deep learning in genomics. Nat. Genet. 51, 12–18. https://doi.org/10.1038/s41588-018-0295-5 (2019).
Lundberg, S. & Lee, S.-I. A unified approach to interpreting model predictions. Ar**v170507874 Cs Stat 2017.
Lundberg, S. M. et al. Explainable machine-learning predictions for the prevention of hypoxaemia during surgery. Nat. Biomed. Eng. 2, 749–760. https://doi.org/10.1038/s41551-018-0304-0 (2018).
Karasinska, J. M. et al. Altered gene expression along the glycolysis-cholesterol synthesis axis is associated with outcome in pancreatic cancer. Clin. Cancer Res. 26, 135–146. https://doi.org/10.1158/1078-0432.CCR-19-1543 (2020).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. ClusterProfiler: An R package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 16, 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Schell, J. C. et al. A role for the mitochondrial pyruvate carrier as a repressor of the warburg effect and colon cancer cell growth. Mol. Cell 56, 400–413. https://doi.org/10.1016/j.molcel.2014.09.026 (2014).
Hua, Q. et al. LINC01123, a c-Myc-activated long non-coding RNA, promotes proliferation and aerobic glycolysis of non-small cell lung cancer through MiR-199a-5p/c-Myc axis. J. Hematol. Oncol. 12, 91. https://doi.org/10.1186/s13045-019-0773-y (2019).
Smolle, E. et al. Distribution and prognostic significance of gluconeogenesis and glycolysis in lung cancer. Mol. Oncol. 14, 2853–2867. https://doi.org/10.1002/1878-0261.12780 (2020).
Liu, X. et al. Chromatin remodeling induced by ARID1A loss in lung cancer promotes glycolysis and confers JQ1 vulnerability. Cancer Res. 82, 791–804. https://doi.org/10.1158/0008-5472.CAN-21-0763 (2022).
Zhou, J. et al. CircRNA-ENO1 promoted glycolysis and tumor progression in lung adenocarcinoma through upregulating its host gene ENO1. Cell Death Dis. 10, 885. https://doi.org/10.1038/s41419-019-2127-7 (2019).
Fahrmann, J. F., Vykoukal, J. V. & Ostrin, E. J. Amino acid oncometabolism and immunomodulation of the tumor microenvironment in lung cancer. Front. Oncol. 10, 276. https://doi.org/10.3389/fonc.2020.00276 (2020).
Eggers, L. F. et al. Lipidomes of lung cancer and tumour-free lung tissues reveal distinct molecular signatures for cancer differentiation, age, inflammation, and pulmonary emphysema. Sci. Rep. 7, 11087. https://doi.org/10.1038/s41598-017-11339-1 (2017).
Hoppstädter, J. et al. Dysregulation of cholesterol homeostasis in human lung cancer tissue and tumour-associated macrophages. EBioMedicine 72, 103578. https://doi.org/10.1016/j.ebiom.2021.103578 (2021).
**a, D.-K., Hu, Z.-G., Tian, Y.-F. & Zeng, F.-J. Statin use and prognosis of lung cancer: A systematic review and meta-analysis of observational studies and randomized controlled trials. Drug Des. Devel. Ther. 13, 405–422. https://doi.org/10.2147/DDDT.S187690 (2019).
Martinez-Outschoorn, U. E., Peiris-Pagés, M., Pestell, R. G., Sotgia, F. & Lisanti, M. P. Cancer metabolism: A therapeutic perspective. Nat. Rev. Clin. Oncol. 14, 11–31. https://doi.org/10.1038/nrclinonc.2016.60 (2017).
Cui, J. et al. A novel KDM5A/MPC-1 signaling pathway promotes pancreatic cancer progression via redirecting mitochondrial pyruvate metabolism. Oncogene 39, 1140–1151. https://doi.org/10.1038/s41388-019-1051-8 (2020).
Liu, X., Zeng, W., Zheng, D., Tang, M. & Zhou, W. Let-7c-5p restrains cell growth and induces apoptosis of lung adenocarcinoma cells via targeting ESPL1. Mol. Biotechnol. https://doi.org/10.1007/s12033-022-00511-2 (2022).
Che, D. et al. KRT6A promotes lung cancer cell growth and invasion through MYC-regulated pentose phosphate pathway. Front. Cell Dev. Biol. 9, 694071. https://doi.org/10.3389/fcell.2021.694071 (2021).
Qin, J. et al. LncRNA MIR31HG overexpression serves as poor prognostic biomarker and promotes cells proliferation in lung adenocarcinoma. Biomed. Pharmacother. Biomedecine Pharmacother. 99, 363–368. https://doi.org/10.1016/j.biopha.2018.01.037 (2018).
Han, X. et al. Identification of proteins related with pemetrexed resistance by ITRAQ and PRM-based comparative proteomic analysis and exploration of IGF2BP2 and FOLR1 functions in non-small cell lung cancer cells. J. Proteom. 237, 104122. https://doi.org/10.1016/j.jprot.2021.104122 (2021).
Acknowledgements
The work was supported by the Major Research Project of Science Technology, Department of Zhejiang Province (grant number: 2021C03124).
Author information
Authors and Affiliations
Contributions
Conceptualization, J,J. and B.Q.; formal analysis, J.J. and B.Q.; investigation, B.Q. and Y.G.; data curation, Y.G.; writing—original draft preparation, J.J.; writing—review and editing, B.Q. and Y.G.; visualization, Z.H.; supervision, J.J. and Z.H. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, J., Qian, B., Guo, Y. et al. Identification of subgroups and development of prognostic risk models along the glycolysis–cholesterol synthesis axis in lung adenocarcinoma. Sci Rep 14, 14704 (2024). https://doi.org/10.1038/s41598-024-64602-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-64602-7
- Springer Nature Limited