Abstract
Waddlia chondrophila is a possible cause of fetal death in humans. This Chlamydia-related bacterium is an emergent pathogen that causes human miscarriages and ruminant abortions, which results in financial losses. Despite the years of efforts, the underlying mechanism behind the pathogenesis of W. chondrophila is little known which hindered the development of novel treatment options. In the framework of current study, computational approaches were used to identify novel inhibitors (phytocompounds) and drug targets against W. chondrophila. At first, RNA polymerase sigma factor SigA and 3-deoxy-d-manno-octulosonic acid transferase were identified through subtractive proteomics pipeline. Afterwards, extensive docking and simulation analyses were conducted to optimize potentially novel phytocompounds by assessing their binding affinity to target proteins. A 100ns molecular dynamics simulation well complimented the compound's binding affinity and indicated strong stability of predicted compounds at the docked site. The calculation of binding free energies with MMGBSA corroborated the significant binding affinity between phytocompounds and target protein binding sites. The proposed phytocompounds may be a viable treatment option for patients infected with W. chondrophila; however, further research is required to ensure their safety.
Similar content being viewed by others
Introduction
Outbreaks of Re-Emerging Infectious Diseases (REID) and Emerging Infectious Diseases (EID) have increased since the turn of the millennium which are communicable threats to public health1. Another Chlamydia-like abortigenic agent, Waddlia chondrophila, has also been linked to poor pregnancy outcomes in humans2,3. W. chondrophila is regarded as a potential cause of embryonic death in humans because it causes poor reproduction in females who have had infrequent recurrent abortions4. Additionally, lung samples from people who had pneumonia also revealed the involvement of W. chondrophila. Despite years of research, nothing is known about the biology or pathogenicity of W. chondrophila.
Recent studies provide evidence that W. chondrophila is responsive to azithromycin, doxycycline, and fluoroquinolones in cell cultures and resistant to lactams, but there is currently no approved therapy against W. chondrophila5. Regrettably, bacteria often develop resistance to antibiotics, making them difficult or even impossible to treat6. To tackle this issue, computational approaches are widely used to understand infectious diseases by establishing the latest research trends that seize advantages from the advancements made in both structural and molecular biology. Recently, subtractive proteomics has proven to be an efficient approach for the identification of species-specific vaccine candidates as well as prospective therapeutic targets against a variety of harmful bacteria7,8. Subtractive proteomics is thought to be the panacea for identifying novel therapeutic targets in pathogens9. The confirmation of target proteins acquired via subtractive proteomics underscores the pivotal role these identified targets play in the disease pathology. Furthermore, molecular dynamics (MD) simulation surpasses docking by integrating a spectrum of physiological parameters, crucial for accurately predicting the authentic mode of molecular interactions. This advanced computational approach provides a nuanced understanding of the dynamic behavior and structural changes within biological systems, offering invaluable insights for drug design and therapeutic interventions.
Understanding the mechanisms of drug resistance in Waddlia chondrophila is essential for devising effective treatment strategies. Current studies reveal that resistance mechanisms include genetic mutations altering antibiotic targets, efflux pump activity expelling antibiotics, and phenotypic changes like entering a dormant state. Furthermore, bioinformatics analysis of protein expression differences between drug-sensitive and drug-resistant strains could unveil potential therapeutic targets. Targeting proteins upregulated in resistant strains or unique to sensitive strains may offer avenues for novel treatments, enhancing antibiotic efficacy and combating resistance. Overall, elucidating resistance mechanisms and identifying potential targets through proteomic analysis are pivotal in develo** strategies to tackle W. chondrophila infections.
Phytochemicals are compounds that are primarily produced by plants and have biological function. Plants are a good source of many active compounds in the pharmaceutical sector. They have pharmacological properties that can be used to treat bacterial and fungal infections, as well as chronic-degenerative disorders like cancer and diabetes. Many studies in recent years have shown that phytochemicals exert antibacterial activity via various mechanisms of action, such as suppression of virulence factors and bacterial membrane damage, including inhibition of toxin and enzyme activity, and bacterial biofilm formation10.
To effectively manage the pathogenesis of infectious diseases in the wake of all these changes, computational approaches are being developed. The study uses a multi-target strategy to conduct virtual screening utilizing molecular docking and dynamic simulation to find potential phytochemicals that can reduce the pathogenicity of diseases. Regarding this, a subtractive proteomics process was utilized to identify the target proteins, which were then further explored with a library of phytochemicals using molecular docking and dynamics simulations. Furthermore, we anticipate that this work may open the way for the development of promising and effective drugs candidates against W. chronophilia.
Methodology
Sequence retrieval, filtering and identification of paralogous sequences
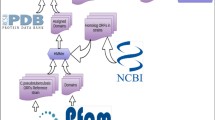
The complete proteome of the Waddlia chondrophila was collected from UniProt in FASTA format11. UniProt is a central hub of protein sequence and functional information. It offers a user-friendly interface for interconnecting and storing valuable protein-related information from large and disparate sources12. One copy of each protein is sufficient to carry out an optimal function since redundant proteins are not necessary. The entire proteome was then subjected to CD-Hit suit for removal of non-redundant sequences13. The cutoff criteria for screening of non-redundant sequences was set to 80%. Only those proteins which met the precise criteria of screening were used for further analysis.
Non-homologous proteins identification
Non-homology with host proteins is required for the anticipated non-redundant proteins with unique metabolic pathways. It is important to remember that predicted proteins may trigger immunological reactions if they are comparable to the host.14. To discover the non-homologous protein sequences of W. chondrophila, essential proteins were submitted to BlastP against the human proteome with standard parameter settings15. BLASTp was performed against non-redundant proteins, and proteins with query cover > 30% and identity > 70% were considered as non-homologous proteins.
Protein subcellular localization prediction
Identifying the protein subcellular localization is crucial to grasp the structure and function of the cell as a whole as well as the function of specific proteins. For a large variety of proteins, bioinformatics predictors of localization can swiftly offer this information16. Regarding this, CELLO17 and pSORTb server18 were used for the identification of cytoplasmic proteins from the pool of predicted non-homologous proteins. The proper functioning of certain proteins is determined on their subcellular localization. Both PSORTb and CELLO server used support vector machine algorithm to expand the prediction of bacterial protein subcellular localization.
Comparative analysis of metabolic pathways
Screened cytoplasmic proteins were processed to compare metabolic pathways. This research is carried out in order to identify therapeutic targets based on pathway enzymes that are both common and essential to bacteria19. The metabolic pathways of Waddlia chondrophila were discovered by comparing Waddlia chondrophila with Homo sapiens pathways in the Kyoto Encyclopedia of Genes and Genomes (KEGG) database20. Those metabolic pathways that are unique to Waddlia chondrophila and not found in humans were chosen. The purpose of this study is to identify potential therapeutic targets based on the similarities and differences between the metabolic pathways that bacteria and humans share. Therefore, only those proteins with unique metabolic pathways were taken into consideration for the rest of the investigation.
Drugability analysis
The druggability of predicted cytoplasmic proteins was also investigated. The nature of the target, or the target's "druggability," essentially limits the effectiveness of many drug design endeavours. Here, we define "druggability" as the capacity of a target to be altered by strong, little molecules that act like drugs (which are often suitable for oral delivery). Early target assessment can therefore be a potent portfolio management tool by allocating resources to "druggable" targets that are more likely to produce clinical candidates21. BLAST heuristic search with an e-value of 10−5 is utilized by the user-friendly cheminformatics program known as DrugBank. This search is used to merge qualitative drug data with in knowledge of therapeutic strategies22.
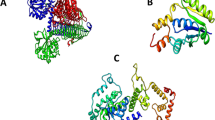
Prediction and evaluation of 3D-structure of targeted proteins
After successful completion of the protein sequence analysis, and evaluation, the targeted proteins were sent to the structure prediction tool for further consideration. AlphaFold was applied to forecast the 3D structure of targeted proteins23. AlphaFold is a neural network-based approach for precise protein structure prediction. The quality of enhanced pharmaceutical targets was evaluated using four distinct technologies (ERRAT, ProCheck, Verify 3D, and ProsA-web)24,25,26,27.
Library preparation of phytochemicals
To investigate the potential repressive effect on the targeted proteins, 1,000 identified phytochemicals were gathered using an in silico technique from numerous databases, including PubChem, MPD3, and Zinc28,29,30. The plant-based compounds were chosen for their medicinal potential in accordance with the results of the literature review31. Phytochemicals belonging to classes sterols and alkaloids are considered as medicinally key active compounds of plants. The phytochemicals alkaloids and sterols were the most frequently selected. Chem Draw was used to create a visual representation and predict the stereochemistry of compounds32. To perform, virtual screening using molecular docking, current study relied on Molecular operating environment (MOE) software (written in a scientific vector language) for energy minimization by picking the MMFF94x force-field to create a fully prepared library of compounds33. At 310 K and pH 7, the target protein structures were refined to add partial charges using the Protonate3D tool. The target proteins' active sites were found by the Site Finder tool in the MOE software. In order to make use of the compound in the MOE ligand database were optimized with the energy of Protonate3D was reduced prior to accessing the database34.
Molecular docking analysis
Molecular docking techniques are commonly used in current drug design to analyze ligand conformations within macromolecular target binding sites. This method calculates the free energy of binding between the ligand and receptor by evaluating critical components of the intermolecular recognition process35. The MOE (Molecular operating environment) tool was utilized to screen a library of 1000 phytochemicals against the targeted proteins using a molecular docking approach. In order to acquire accurate findings, all docking experiments were carried out with the default parameters36. To rescore simulated poses, MOE's London dG scoring system was employed. Protonation and energy minimization was also done. After docking, the best and most favorable phytochemicals were selected on the basis of binding affinity37. LigX is a tool for visualizing the best-docked complexes and examining the 2D structure of ligand-receptor interactions38.
Druglikeness and ADMET profiling
Drug-likeness evaluation is a vital stage in the drug development process that should not be disregarded39. Chemical and physical parameters such as molecular weight, the logP coefficient (miLogP), hydrogen bond acceptors, hydrogen bond donors were all considered40. In order to determine the druggability of the top-docked ligands, we used the Molinspiration online tool (retrieved on May 8, 2021))41. It is generally recognized that ADMET profiling of candidate compounds are among the most important major shortcomings of drug development; therefore, it is imperative that ADMET studies be performed as early in the drug development process as possible42. Evaluation models to find the adverse drug-drug interaction have been established as an alternative tool to aid medicinal chemists in the creation and optimization of lead compounds. The ADMET lab 2.0 server was used to evaluate the ADMET properties73. Hence, the novel drug targets discovered in this work should be very helpful in the drug therapeutic industry for finding inhibitors and devising new drug formulations to combat W. chondrophila infections, albeit more experimental research are required to confirm these drug targets.
Conclusion
In conclusion, the urgent global challenge of antimicrobial resistance has spurred intensive research efforts to identify new therapeutic avenues. Through the exploration of metabolic pathways specific to pathogens and leveraging successful targeting strategies in other bacterial species, this study has unveiled two promising targets within W. chondrophila. This discovery marks a notable advancement in our understanding and potential treatment of infections caused by this pathogen. Moving forward, it is imperative for future research endeavors to delve deeper into these identified targets, assessing their impact on the longevity and virulence of W. chondrophila. By doing so, we can pave the way for the development of effective antibacterial agents, offering hope in the ongoing battle against antimicrobial.
References
Woolhouse, M. E. Population biology of emerging and re-emerging pathogens. Trends Microbiol. 10, s3–s7 (2002).
Ligon, B.L. Infectious diseases that pose specific challenges after natural disasters: A review. In Proceedings of the Seminars in pediatric infectious diseases, pp. 36–45 (2006).
Millar, B. C. & Moore, J. E. Emerging pathogens in infectious diseases: Definitions, causes and trends. Rev. Med. Microbiol. 17, 101–106 (2006).
Lamoth, F., Pillonel, T. & Greub, G. Waddlia: An emerging pathogen and a model organism to study the biology of chlamydiae. Microbes Infect. 17, 732–737 (2015).
Henning, K. et al. Neospora caninum and Waddlia chondrophila strain 2032/99 in a septic stillborn calf. Vet. Microbiol. 85, 285–292 (2002).
Kebbi-Beghdadi, C., Cisse, O. & Greub, G. Permissivity of Vero cells, human pneumocytes and human endometrial cells to Waddlia chondrophila. Microbes Infect. 13, 566–574 (2011).
Vasilevsky, S. et al. Waddlia chondrophila induces systemic infection, organ pathology, and elicits Th1-associated humoral immunity in a murine model of genital infection. Front. Cell. Infect. Microbiol. 5, 76 (2015).
Hu, S. et al. Salivary proteomics for oral cancer biomarker discovery. Clin. Cancer Res. 14, 6246–6252 (2008).
Goy, G., Croxatto, A., Posfay-Barbe, K., Gervaix, A. & Greub, G. Development of a real-time PCR for the specific detection of Waddlia chondrophila in clinical samples. Eur. J. Clin. Microbiol. Infect. Dis. 28, 1483–1486 (2009).
Mendoza, N. & Silva, E. M. E. Introduction to phytochemicals: secondary metabolites from plants with active principles for pharmacological importance. Phytochem. Source Antioxid. Role Dis. Prev. 25, 1 (2018).
Consortium U. UniProt: A hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Consortium U. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Nunez-Castilla, J., Stebliankin, V., Baral, P., Balbin, C. A., Sobhan, M., Cickovski, T., Mondal, A. M., Narasimhan, G., Chapagain, P., & Mathee, K. Potential autoimmunity resulting from molecular mimicry between SARS-CoV-2 Spike and human proteins. BioRxiv 2021.2008. 2010.455737 (2022).
Mount, D.W. Using the basic local alignment search tool (BLAST). Cold Spring Harbor Protocols 2007, pdb. top17 (2007).
Yu, C. S., Chen, Y. C., Lu, C. H. & Hwang, J. K. Prediction of protein subcellular localization. Proteins Struct. Funct. Bioinf. 64, 643–651 (2006).
Yu, C.-S. et al. CELLO2GO: A web server for protein subCELlular LOcalization prediction with functional gene ontology annotation. PloS One 9, e99368 (2014).
Yu, N. Y. et al. PSORTb 3.0: Improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26, 1608–1615 (2010).
Anishetty, S., Pulimi, M. & Pennathur, G. Potential drug targets in Mycobacterium tuberculosis through metabolic pathway analysis. Comput. Biol. Chem. 29, 368–378 (2005).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Wishart, D. S. et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Borrel, A. & Fourches, D. RealityConvert: A tool for preparing 3D models of biochemical structures for augmented and virtual reality. Bioinformatics 33, 3816–3818 (2017).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 1–8 (2008).
Dym, O., Eisenberg, D., & Yeates, T. ERRAT (2012).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: A program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Eisenberg, D., Lüthy, R., & Bowie, J. U. VERIFY3D: assessment of protein models with three-dimensional profiles. In Methods in enzymology, Volume 277, pp. 396–404 (Elsevier, 1997).
Wiederstein, M. & Sippl, M. J. ProSA-web: Interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 35, W407–W410 (2007).
Kim, S. et al. PubChem substance and compound databases. Nucleic Acids Res. 44, D1202–D1213 (2016).
Mumtaz, A. et al. MPD3: A useful medicinal plants database for drug designing. Nat. Prod. Res. 31, 1228–1236 (2017).
Irwin, J. J. & Shoichet, B. K. ZINC—a free database of commercially available compounds for virtual screening. J. Chem. Inf. Model. 45, 177–182 (2005).
Mani, J. S. et al. Antioxidative and therapeutic potential of selected Australian plants: A review. J. Ethnopharmacol. 268, 113580 (2021).
Mills, N. ChemDraw Ultra 10.0 CambridgeSoft, 100 Cambridge Park Drive, Cambridge, MA 02140. www.cambridgesoft.com. Commercial Price: 1910fordownload, 2150 for CD-ROM; Academic Price: 710fordownload, 800 for CD-ROM (2006).
Vilar, S., Cozza, G. & Moro, S. Medicinal chemistry and the molecular operating environment (MOE): Application of QSAR and molecular docking to drug discovery. Curr. Top. Med. Chem. 8, 1555–1572 (2008).
Labute, P. Protonate3D: Assignment of ionization states and hydrogen coordinates to macromolecular structures. Proteins Struct. Funct. Bioinf. 75, 187–205 (2009).
Morris, G. M. & Lim-Wilby, M. Molecular docking 365–382 (Springer, 2008).
Bursulaya, B. D., Totrov, M., Abagyan, R. & Brooks, C. L. Comparative study of several algorithms for flexible ligand docking. J. Comput. Aid. Mol. Des. 17, 755–763 (2003).
Yusuf, D., Davis, A. M., Kleywegt, G. J. & Schmitt, S. An alternative method for the evaluation of docking performance: RSR vs RMSD. J. Chem. Inf. Model. 48, 1411–1422 (2008).
Khalifa, I., Zhu, W., Nafie, M. S., Dutta, K., Li, C. Anti-COVID-19 effects of ten structurally different hydrolysable tannins through binding with the catalytic-closed sites of COVID-19 main protease: An in-silico approach. 1, 1 (2020).
Ursu, O., Rayan, A., Goldblum, A. & Oprea, T. I. Understanding drug-likeness. Wiley Interdiscip. Rev. Comput. Mol. Sci. 1, 760–781 (2011).
Lipinski, C. A. Lead-and drug-like compounds: The rule-of-five revolution. Drug Discov. Today Technol. 1, 337–341 (2004).
Jarrahpour, A. et al. Petra, Osiris and Molinspiration (POM) together as a successful support in drug design: Antibacterial activity and biopharmaceutical characterization of some azo Schiff bases. Med. Chem. Res. 21, 1984–1990 (2012).
Li, A. P. Screening for human ADME/Tox drug properties in drug discovery. Drug Discov. Today 6, 357–366 (2001).
**ong, G. et al. ADMETlab 2.0: An integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucleic Acids Res. 49, W5–W14 (2021).
Saager, B. & Fischer, J. Predictive power of effective intermolecular pair potentials: MD simulation results for methane up to 1000 MPa. Fluid Phase Equ. 57, 35–46 (1990).
Desmond, C. Project management tools-software tools. IEEE Eng. Manag. Rev. 45, 24–25 (2017).
Shivakumar, D., Harder, E., Damm, W., Friesner, R. A., & Sherman, W. J. Improving the prediction of absolute solvation free energies using the next generation OPLS force field. 8, 2553–2558 (2012).
Price, D. J., & Brooks, C. L. A modified TIP3P water potential for simulation with Ewald summation. 121, 10096–10103 (2004).
Rehman, A. et al. In silico core proteomics and molecular docking approaches for the identification of novel inhibitors against streptococcus pyogenes. Int. J. Environ. Res. Public Health 18, 11355 (2021).
Rehman, A. et al. Integrated core proteomics, subtractive proteomics, and immunoinformatics investigation to unveil a potential multi-epitope vaccine against schistosomiasis. Vaccines 9, 658 (2021).
Rehman, A., Ashfaq, U. A., Shahid, F., Noor, F. & Aslam, S. The Screening of phytochemicals against NS5 Polymerase to treat Zika Virus infection: Integrated computational based approach. Combin. Chem. High Throughput Screen. 1, 1 (2021).
Bhavsar, R. B., Makley, L. N. & Tsonis, P. A. The other lives of ribosomal proteins. Human Genom. 4, 1–18 (2010).
Xu, L. et al. Biological effect of ribosomal protein L32 on human breast cancer cell behavior. Mol. Med. Rep. 22, 2478–2486 (2020).
Baud, D., Regan, L. & Greub, G. Emerging role of Chlamydia and Chlamydia-like organisms in adverse pregnancy outcomes. Curr. Opin. Infect. Dis. 21, 70–76 (2008).
Baud, D. & Greub, G. Intracellular bacteria and adverse pregnancy outcomes. Clin. Microbiol. Infect. 17, 1312–1322 (2011).
Baud, D. et al. Waddlia chondrophila: From bovine abortion to human miscarriage. Clin. Infect. Dis. 52, 1469–1471 (2011).
Kebbi-Beghdadi, C. et al. Identification of immunogenic proteins of Waddlia chondrophila. PloS One 7, e28605 (2012).
Verweij, S. et al. Waddlia chondrophila and Chlamydia trachomatis antibodies in screening infertile women for tubal pathology. Microb. Infect. 17, 745–748 (2015).
Lienard, J. et al. Undressing of Waddlia chondrophila to enrich its outer membrane proteins to develop a new species-specific ELISA. New Microbes New Infect. 2, 13–24 (2014).
Baud, D., Zufferey, J., Hohlfeld, P. & Greub, G. Performance of an automated multiplex immunofluorescence assay for detection of Chlamydia trachomatis immunoglobulin G. Diagn. Microbiol. Infect. Dis. 78, 217–219 (2014).
De Barsy, M. & Greub, G. Waddlia chondrophila: from biology to pathogenicity. Microb. Infect. 15, 1033–1041 (2013).
Dilbeck-Robertson, P., McAllister, M. M., Bradway, D. & Evermann, J. F. Results of a new serologic test suggest an association of Waddlia chondrophila with bovine abortion. J. Vet. Diagn. Investig. 15, 568–569 (2003).
Borel, N. et al. Parachlamydia spp. and related Chlamydia-like organisms and bovine abortion. Emerg. Infect. Dis. 2007, 13 (1904).
Blumer, S. et al. Waddlia, Parachlamydia and Chlamydiaceae in bovine abortion. Vet. Microbiol. 152, 385–393 (2011).
Murugan, N. A., Kumar, S., Jeyakanthan, J. & Srivastava, V. Searching for target-specific and multi-targeting organics for Covid-19 in the Drugbank database with a double scoring approach. Sci. Rep. 10, 1–16 (2020).
Ogata, H., Goto, S., Fujibuchi, W. & Kanehisa, M. Computation with the KEGG pathway database. Biosystems 47, 119–128 (1998).
Kesharwani, R. K. et al. Role of ADMET tools in current scenario: application and limitations 71–87 (Springer, 2020).
Daina, A., Michielin, O. & Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 7, 1–13 (2017).
Mahmood, A. et al. Structural and dynamics insights into the GBA variants associated with Parkinson’s disease. J. Biomol. Struct. Dyn. 1, 1–13 (2023).
Alotaibi, B. S. et al. New drug target identification in Vibrio vulnificus by subtractive genome analysis and their inhibitors through molecular docking and molecular dynamics simulations. Heliyon 1, e17650 (2023).
Ajmal, A., Ali, Y., Khan, A., Wadood, A. & Rehman, A. U. Identification of novel peptide inhibitors for the KRas-G12C variant to prevent oncogenic signaling. J. Biomol. Struct. Dyn. 1, 1–10 (2022).
Shami, A. et al. In silico subtractive proteomics and molecular docking approaches for the identification of novel inhibitors against Streptococcus pneumoniae strain D39. Life 13, 1128 (2023).
Kumar, A., Thotakura, P. L., Tiwary, B. K. & Krishna, R. Target identification in Fusobacterium nucleatum by subtractive genomics approach and enrichment analysis of host-pathogen protein-protein interactions. BMC Microbiol. 16, 1–12 (2016).
Shahid, F., Alghamdi, Y. S., Mashraqi, M., Khurshid, M. & Ashfaq, U. A. Proteome based map** and molecular docking revealed DnaA as a potential drug target against Shigella sonnei. Saudi J. Biol. Sci. 29, 1147–1159 (2022).
Acknowledgements
HMA lab is supported by the Researchers Project Number (RSPD2024R1083) King Saud University, Riyadh, Saudi Arabia. The BBDP has received financial support from the National Institute on Aging (P30 AG019610 and P30AG072980, Arizona Alzheimer’s Disease Center).
Funding
The BBDP has received financial support from the National Institute on Aging (P30 AG019610 and P30AG072980, Arizona Alzheimer’s Disease Center).
Author information
Authors and Affiliations
Contributions
S.A: conceptualization, validation, H.M.A: formal analysis, writing—original draft preparation. M.M and GS reviewed the manuscript and help in analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aslam, S., Aljawdah, H.M., Murshed, M. et al. Pharmacophore modelling based virtual screening and molecular dynamics identified the novel inhibitors and drug targets against Waddlia chondrophila. Sci Rep 14, 13472 (2024). https://doi.org/10.1038/s41598-024-63555-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-63555-1
- Springer Nature Limited