Abstract
Catabolite control protein C (CcpC) belongs to the LysR-type transcriptional regulator (LTTR) family, which regulates the transcription of genes encoding the tricarboxylic acid branch enzymes of the TCA cycle by responding to a pathway-specific metabolite, citrate. The biological function of CcpC has been characterized several times, but the structural basis for the molecular function of CcpC remains elusive. Here, we report the characterization of a full-length CcpC from Bacillus amyloliquefaciens (BaCcpC-FL) and a crystal structure of the C-terminal inducer-binding domain (IBD) complexed with citrate. BaCcpC required both dyad symmetric regions I and II to recognize the citB promoter, and the presence of citrate reduced citB promoter binding. The crystal structure of CcpC-IBD shows two subdomains, IBD-I and IBD-II, and a citrate molecule buried between them. Ile100, two arginines (Arg147 and Arg260), and three serines (Ser129, Ser189, and Ser191) exhibit strong hydrogen-bond interactions with citrate molecules. A structural comparison of BaCcpC-IBD with its homologues showed that they share the same tail-to-tail dimer alignment, but the dimeric interface and the rotation between these molecules exhibit significant differences. Taken together, our results provide a framework for understanding the mechanism underlying the functional divergence of the CcpC protein.
Similar content being viewed by others
Introduction
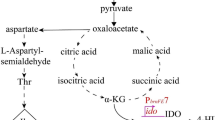
The tricarboxylic acid (TCA) cycle, also known as the Krebs cycle or the citric acid cycle (CAC), is a central metabolic pathway in the cell1. The TCA cycle provides organisms with reducing potential, energy, and three of the 13 biosynthetic intermediates2. In Bacillus, TCA activity is controlled by several important regulatory proteins, including global regulators catabolite control protein A (CcpA) and CodY and the specific regulator catabolite control protein C (CcpC), which are coordinated by fructose-1,6-bisphosphate (FBP) and glucose-6-phosphate, guanosine-5′-triphosphate (GTP) and branched-chain amino acids (BCAAs), and citrate, respectively3. CcpA and CodY are metabolite-responsive global regulators of carbon metabolism pathways4. These global regulators coordinate the expression of numerous metabolic, biosynthetic and virulence genes that respond to three metabolites5. CcpA is a member of the LacI/GalR family of transcriptional repressors, which exert both direct (through citrate synthase, citZ, and ccpC) and indirect effects on TCA branch enzyme expression6. CodY is also a repressor of the citB gene belonging to a unique family of regulators in B. subtilis and other homologues of gram-positive bacteria7. Recent studies have shown that CcpC and CcpE exclusively regulate the TCA branch enzymes of the TCA cycle (citB, aconitase; citC, isocitrate dehydrogenase; and citZ, citrate synthase) by responding to a pathway-specific metabolite for both Bacillus subtilis and Staphylococcus aureus, respectively8,9.
CcpC widely exists in prokaryotes and is classified as belonging to the LysR-type transcriptional regulator (LTTR) family10. Typical LTTR family proteins comprise approximately 330 amino acids that form structures highly similar to those of N-terminal DNA-binding domains (DBDs), which are directly involved in DNA interactions, and poorly conserved C-terminal inducer-binding domains (IBDs) and are known to adopt different oligomeric states10. The DBD is highly conserved and directly involved in DNA interactions, similar to the helix-loop-helix, zinc finger, and β–sheet-anti-parallel domains11,12,13. IBD serves as a binding site of inducer and thus plays an important role in transcriptional activation by binding to inducers14.
To reveal the molecular mechanism of LTTRs, the structure of LTTRs has long been studied. The crystal structure of the DBD in complex with recognition binding site (RBS) has been determined in BenM, which revealed that 25-bp DBD of BenM interacting with benA-RBS15. While the crystal structure of BenM-DBD complexed RBS showed the details of the interaction, the mechanism responsible for the selection of a specific sequence remains elusive15. The RBS of BenM and CbnR have high similarity16. The crystal structure of the DBD of CbnR complexed with RBS reveals the detailed mechanism of the specific interaction between CbnR and its promoter DNA and that Thr33 leads to selective interactions with DNA16. Moreover, structural information of the IBD of LTTRs obtained is equally necessary. The first crystal structure of LTTRs is IBD of CysB, and the IBD induces conformational changes, resulting in structural changes for transcriptional activation17. Therefore, the mechanism of the conformational change plays an important role in the study of LTTRs17. The conformational change of IBD upon inducer binding has been structurally analysed in AphB, BenM, CcpE, and OxyR18,19,20,21. While several models of transcriptional activation of LTTRs with significant conformational changes have been proposed, all structural changes seem to be caused by the IBD upon inducer binding18,22,23. The first full-length crystal structure of CbnR, which exists in the crystal structure as a tetramer, was determined24. Four DBDs are located at the bottom of the tetramer of CbnR and arranged in a V-shape, which provides significant insight into DNA bending24. Small angle X-ray scattering (SAXS) studies of DntR have provided clear evidence of the structural changes that the LTTR tetramer undergoes upon activation via an inducer, and have supported the ‘sliding dimer’ hypothesis concerning LTTR transcription activation mechanisms22. The crystal structure of the full-length CbnR complex with its promoter DNA showed the mechanism of the quaternary structural change caused by inducer binding14. These changes are likely to be necessary for recruiting RNA polymerase to the promoter site to initiate transcription25.
In B. subtilis, CcpC binds to two sites within the citB promoter region, a dyad symmetry element centred at position −66 that induces of the transcriptional start site and a half-dyad element located at position −27 to −3326. Two CcpC dimers interact with these sites for repression, resulting in bending of the DNA, and then blocking the access of RNA polymerase to the promoter, which results in repression of citB expression. As with other LTTR proteins, CcpC interacts with the inducer to relieve this repression; that is, citrate induces the expression of citB26. The coinducer citrate is important for the function of CcpC and appears to function as a key catabolite for coordinating the B. subtilis metabolic state by binding to and activating CcpC9. Previous studies have indicated citrate-induced derepression in the citB gene of L. monocytogenes. L. monocytogenes CcpC represses the transcription of citB and citZ27. The binding site of the citB regulatory region in L. monocytogenes CcpC is almost consistent that in B. subtilis CcpC, with a dyad symmetry element centred at positions −68 to −2827. In addition, binding to the full dyad is maintained, however, binding to the half-dyad is reduced in the presence of citrate27. Thus, the CcpC complex with citrate is a signal that morphs CcpC into a conformation that is competent for binding DNA and inducing gene transcription. Several biological functional properties of CcpC are well characterized; however, the structure-based molecular function has been elusive9.
In this study, we characterized the full-length Bacillus amyloliquefaciens CcpC and determined the crystal structure of the C-terminal IBD of B. amyloliquefaciens CcpC (BaCcpC-IBD) at 2.3 Å resolution. The citB binding properties and the oligomeric state of BaCcpC were analysed. The crystal structure of BaCcpC-IBD was compared with structures of the LTTR family members. Taken together, our findings provide insight into the citrate-responsive mechanism of CcpC.
Results and discussion
Biochemical study of BaCcpC
In B. subtilis, CcpC (BsCcpC) negatively regulates citB gene expression, which is responsible for the interconversion of citrate and isocitrate9. The BsCcpC binding region forms two dyad symmetry elements centred at positions −66 and −27 (Fig. 1A)9. This BsCcpC bind to the DNA-binding boxes “ATAA”, “TTAT”, and “TATT” in the citB promoter region9. In the B. amyloliquefaciens genome, the potential promoter region of citB (named citB-P) was found from position −73 to −20, and it shows high similarity with the same DNA-binding boxes “ATAA”, “TTAT”, and “TATT” (Fig. 1A). The DNA sequence of the BsCcpC binding box was identical to that of the BaCcpC binding box in citB promoter region I (named citB-PI, −73 to −54), but the nonbinding sequences did not match, whereas the DNA sequences of citB promoter region II (named citB-PII, −40 to −20) matched a consensus sequence (Fig. 1A). Moreover, the spacer sequence between citB protomer regions I and II was identical to those of B. subtilis and B. amyloliquefaciens, and the same finding was obtained for the DNA length.
citB promoter-binding properties of BaCcpC. (A) Comparison of the citB promoter regions from B. amyloliquefaciens (upper) and B. subtilis (bottom). DNA-binding boxes are indicated in bold and underlined. EMSA experiment of BaCcpC with (B) the citB promoter, (C) region I in the citB promoter region, (D) region II in the citB promoter region, and (E) the citB promoter with various concentrations of sodium citrate.
To verify whether CcpC regulates the predicted citB promoter region in B. amyloliquefaciens, we performed an electrophoretic mobility shift assay (EMSA) using a 56 bp citB promoter DNA fragment (from nucleotides −73 to −20 relative to the start codon of citB) as a probe (Fig. 1A). A complete shift of the free probe was observed as the concentration of BaCcpC increased (Fig. 1B). This result indicates that BaCcpC has a high binding affinity for citB promoter DNA of B. amyloliquefaciens. Next, the binding of BaCcpC to each of the citB promoter regions (I and II) for each CcpC was assessed. BaCcpC did not bind to either promoter region (Fig. 1C, D). These results indicate that both dyad symmetry promoter regions are required for BaCcpC to bind to the promoter of citB. To determine whether citrate affects the binding of CcpC to the citB promoter, an EMSA of CcpC for citB was performed with citrate. The results showed that the presence of citrate slightly suppressed the binding of CcpC to citB-P (Fig. 1E).
Oligomerization of BaCcpC
LTTRs are usually functionally active as tetramers and dependent on a coinducer28. To verify the oligomeric state of BaCcpC (MW ~ 30 kDa) in solution, we performed size exclusion chromatography coupled to multiangle light scattering (SEC-MALS). BaCcpC-FL formed oligomeric peaks that eluted in the SEC-MALS chromatogram (Fig. 2A). The calculated molecular weights of BaCcpC were ~ 60 kDa (corresponding to 2 monomers). This result indicated that BaCcpC exists as a dimer in the solution. Moreover, in equilibrium buffer with 10 mM citrate, the chromatograms remained dimeric (Fig. 2B). These results indicated that citrate cannot influence the oligomeric state of BaCcpC.
Overall structure of the IBD of BaCcpC
To better understand the molecular function of BaCcpC, we performed a crystallographic study on full-length BaCcpC; however, it was not successful. Furthermore, crystallographic studies for the DBD and IBD of BaCcpC were separately performed. Finally, we obtained crystals for the IBD of BaCcpC and determined the crystal structure of BaCcpC-IBD in complex with citrate at 2.3 Å resolution using single-wavelength anomalous diffraction (SAD) phasing. The BaCcpC-IBD crystal belonged to space group C2 and had unit-cell parameters of a = 140.96, b = 90.90, c = 105.53 Å and β = 106.18°. The Rwork and Rfree of the final model were 20.7% and 26.6%, respectively. The BaCcpC-IBD molecule is composed of two distinct regulatory domains: IBD-I (His90-Arg155 and Gly266-Gln289) and IBD-II (Asp168-Gly259) (Fig. 3A). IBD-I has three β-sheets, which are surrounded by three α-helices and 310 helices (Fig. 3B). IBD-II has four β-sheets, which are surrounded by two α-helices and two 310 helices (Fig. 3B). Both the IBD-I and IBD-II subdomains adopt the typical α/β fold, which is connected by two crossover regions that form a hinge at central regions of two antiparallel β-strands (β4 and β9) (Fig. 3A). Five BaCcpC-IBD molecules are in the asymmetric unit, and each molecule has an RMSD of 0.201–0.275 Å for the 144–180 Cα atoms, which emphasizes the similarity of their conformations. BaCcpC-IBD forms dimers with a head-to-tail arrangement in the asymmetric unit of its crystal structure, and both molecules have essentially the same overall structure (Fig. 3C). Superposition of two dimeric molecules in the asymmetric unit gives an RMSD of 0.327 Å for 321 Cα. The dimeric interface is stabilized by the main chain interactions Val122-Thr212* (2.82 Å, * denoting the partner molecule), Val122-Leu214* (2.92 Å), Leu124-Asp216* (2.80 Å), and Thr126-Asp216* (3.33 Å) between the β2 strand and β6 strand (Fig. 3D).
Overall structure of BaCcpC-IBD. (A) Monomer structure of BaCcpC-IBD. The hinge regions, consisting of β4- and β9-strands, are indicated in blue. Citrate molecules are located between the IBD-I and IBD-II subdomains. (B) Close-up view of the IBD-I and IBD-II subdomains of BaCcpC-IBD. (C) Dimer of BaCcpC-IBD. (D) Close-up view of the interactions in the dimer interface.
The citrate binding site of BaCcpC-IBD
To observe the citrate-bound state of BaCcpC-IBD, we added sodium citrate to the purification buffer during all protein purification steps. The electron density corresponding to citrate molecules is found at the positively charged interface between IBD-I and IBD-II of BaCcpC-IBD (Fig. 4A). In the electron density map, the positions of each carboxyl and hydroxyl group of citrate are clearly distinguished (Fig. 4A). Citrate is a small organic acid that includes three carboxyl groups, one hydroxyl group, and one prochiral centre. To distinguish between the terminal carboxyl groups, these were named pro-R and pro-S. The pro-R carboxyl group accepts a strong hydrogen bond from the backbone nitrogen atom of Ile100 (average distance for five molecules in the asymmetric unit: 3.12 Å). In addition, the pro-S carboxyl group also has hydrogen bonds from the side-chain NE atoms of Arg147 (2.93 Å) and Arg260 (2.80 Å). The central carboxyl group of citrate accepts hydrogen bonds from the backbone nitrogen atom of Ser129 (2.70 Å), the side-chain hydroxyl group, and Ser189 (2.59 Å) and Ser191 (2.89 Å). The hydroxyl group of citrate interacts with the side-chain hydroxyl group of Ser129 (2.76 Å). The atoms of these residues that contact the citrate molecule are ~ 3.0 Å away from the latter’s oxygen atoms, demonstrating that citrate was coordinated by extensive strong hydrogen-bonding interactions (Fig. 4A).
Citrate binding site of BaCcpC-IBD. (A) Citrate binding to the interface between IBD-I and II of BaCcpC-IBD. (Insert) Interaction between citrate and BaCcpC-IBD. Simulated annealing 2mFo-DFc omit electron (green mesh, 1σ) density map for citrate molecules. (B) Partial sequence alignment of the citrate binding sites of BaCcpC, BsCcpC, and SaCcpE. (C) Superimposition of the citrate binding sites of the IBD of BaCcpC (yellow) and SaCcpC (green).
Inducers are important for the function of LTTRs and often participate in the feedback loop of a specific metabolic/synthesis pathway29. However, citrate molecules are inducers of BaCcpC, BsCcpC, and SaCcpE8,9. Sequence alignment showed that the Arg147 and Arg260 residues of BaCcpC are highly conserved in both BsCcpC and SaCcpE, whereas Ile100, Ser129, Ser189, and Ser191 of BaCcpC are not conserved in SaCcpE and are conserved in only BsCcpC (Fig. 4B). Moreover, the citrate-binding Arg147 and Arg260 residues of BaCcpC-IBD are analogous to the citrate-binding Arg145 and Arg256 residues of CcpE, which are required for CcpE to evoke an appropriate response in the presence of citrate20,30 (Fig. 4C). Therefore, we consider these two arginine residues to also play important roles in the citrate binding and functional assembly of BaCcpC.
Comparison of BaCcpC-IBD with other IBDs from the LTTR family
To better understand the structural properties of BaCcpC-IBD, its homologues were sought using the Dali server. The IBD of CcpE from S. aureus (named SaCcpE-IBD, Z-score: 22.0, sequence identity: 28%, Protein Data Bank (PDB) code: 4QBA), C-terminal domain of a putative transcriptional regulator from Klebsiella pneumoniae (KpYneJ-CTD, 20.6, 19%, 5TPI) effector binding domain of BenM from Acinetobacter baylyi (AbBenM-EBD, 18.3, 15%, 2F6G), ligand-binding domain of CynR from Escherichia coli (EcCynR-LBD, 18.8, 16%, 3HFU) and ligand-binding domain of OccR from Agrobacterium tumefaciens (AtOccR-LBD, 19.3, 17%, 5VVH) showed structural similarity to BaCcpC. These proteins belong to the LTTR family. BaCcpC is involved in citrate metabolism, similar to SaCcpE, while AbBenM, EcCynR, AtOccR, and KpYneJ are involved in aromatic compound catabolism, cyanate detoxification, octopine catabolism, and biosynthesis of cysteine, respectively8,17,28,31,32. Although BaCcpC-IBD shared low amino acid sequence identities (less than 30%) with SaCcpE-IBD, KpYneJ-CTD, and AtOccR-LBD, it was commonly composed of two subdomains, similar to IBD-I and IBD-II of BaCcpC-IBD (Fig. 5A). The superposition of BaCcpC-IBD with SaCcpE-IBD-apo, SaCcpE-IBD-citrate, KpYneJ-CTD, AbBenM-EBD, EcCynR-LBD, and AtOccR-LBD showed structural similarity with RMSDs of 1.818 Å, 1.181 Å, 2.240 Å, 2.594 Å, 2.315 Å, and 1.941 Å, respectively.
Structural comparison of BaCcpC-IBD with the LTTR family. (A) Ribbon representations of the IBDs (or CBDs) of BaCcpC, SaCcpE, AbBenM, EcCynR, KpYneJ and AtOccR, which consist of two subdomains. (B) The tail-to-tail dimers arrangement of BaCcpC, SaCcpE, AbBenM, EcCynR, KpYneJ, and AtOccR. (C) Comparison of rotation in BaCcpC-IBD-citrate, SaCcpE-IBD-citrate, and SaCcpE-IBD-apo dimers.
SaCcpE-IBD, AbBenM-EBD, EcCynR-LBD, KpYneJ-CTD, and AtOccR-LBD also show dimers with the same tail-to-tail alignments as BaCcpC-IBD, but the dimeric interfaces and the rotations between these molecules exhibit significant differences. The dimeric interface of SaCcpE-IBD is formed by hydrogen bonds as well as some salt bridges between α1 and α5*, α1 and loop*, and loop and loop* (an asterisk indicates the partner molecule) (Fig. 5B). The dimer interface of AbBenM-EBD consists of the same interaction in α1 and α6* β2-loop*, however, EcCynR-LBD only consists of hydrogen bonds in α1-α5*, β2-β6* β2-loop*. The dimer interface of KpYneJ-CTD consists of two α-helices that interact with one β-sheet, namely, α1-α5*, β2-α5*, α5-β2* and α5-α1*, which differ from the dimer interface of AtOccR-LBD in α1-α5*, loop-loop* and β2-β7* (Fig. 5B and Supplementary Fig. 1). These findings indicated that IBD dimers are relatively stable, even after poor conservation in IBDs. Meanwhile, the rotation angles of the dimers are distinct. In addition, CcpC-IBD is functionally distinct from CcpE-IBD but also recognizes citrate molecules. The monomers of citrate-bound BaCcpC-IBD and SaCcpE-IBD dimers are rotated at angles of approximately 33° and 35°, respectively (Fig. 5C). As a result, the rotation angle of the dimer interface of citrate-bound BaCcpC-IBD is very similar to that of citrate-bound SaCcpE-IBD. On the other hand, in the citrate-free state of SaCcpE-IBD, the angle of the dimer interface is approximately 80°, indicating that there is a change in dimer formation depending on citrate binding. The conformational change between the BaCcpC-IBD domains by the citrate molecule is important for explaining the functional mechanism. To better understand the citrate-induced structural change of BaCcpC, we performed molecular dynamics (MD) simulations. However, there was no significant structural change in BaCcpC with or without citrate molecules (Supplementary Fig. 2). To confirm whether the BaCcpC-IBD domain undergoes a structural change according to the citrate molecule like the CcpE-IBD domain, the apo-state BaCcpC-IBD structure needs to be determined in further studies.
Conclusion
BaCcpC needs both dyad symmetry regions I and II to recognize the citB promoter and the presence of citrate-reduced citB binding. Citrate binds the interface between IBD-I and IBD-II of the IBD of BaCcpC. The IBD of BaCcpC shares low sequence similarity with other IBDs of the LTTR family but exhibits similarity in terms of the overall structure and dimer formation. Our results provide the framework for functional analysis of CcpC as well as the diversity and similarity of IBDs of the LTTR family.
Methods
Construction, expression, and purification
The full-length (residues 1–293; named BaCcpC-FL) and C-terminal region of the IBD (residues 88–293; named BaCcpC-IBD) of CcpC were obtained from genomic DNA of B. amyloliquefaciens by PCR. The gene was cloned into the NcoI and XhoI sites of the pPROEX-HTA vector (Invitrogen, USA), which contains a hexahistidine tag (MSYYHHHHHH), a spacer region (DYDIPTT) and a tobacco etch virus (TEV) protease cleavage site (ENLYFQ) at the N-terminus. The construct was transformed into E. coli BL21 (DE3) competent cells to obtain the target proteins. Protein expression and purification procedures were the same for BaCcpC-FL and BaCcpC-IBD. Cells were grown in 2 L of Luria–Bertani (LB) medium containing 0.5 μg ml−1 ampicillin at 310 K. When the OD600 of the culture reached 0.8, 0.5 mM isopropyl-β-d-thiogalactoside (IPTG) was added, and the culture was incubated at 303 K for 8 h. The bacterial cells were centrifuged for harvesting and resuspended in lysis buffer containing 20 mM Tris (pH 8.0), 10 mM sodium citrate, 150 mM NaCl, and 2 mM β-mercaptoethanol. Then, the cells were disrupted by sonication, and the lysate was centrifuged at 13,000 rpm for 30 min at 277 K. The lipid fractions were mixed with a nickel-nitrilotriacetic acid (Ni–NTA) affinity resin (GE Healthcare) that had been preincubated with lysis buffer and stirred for 30 min at 277 K. The resin was washed and eluted with lysis buffer containing 20 mM imidazole and 300 mM imidazole. The fractions containing BaCcpC were pooled, and β-mercaptoethanol was added to 10 mM (final concentration). To remove the hexahistidine tag, the mixture was incubated with a recombinant TEV protease at 298 K overnight. For further purification, the mixture was diluted fourfold using 20 mM Tris (pH 8.0) buffer and loaded onto a Q anion-exchange column (HiTrap-Q; GE Healthcare, USA). The fractions containing BaCcpC were purified using a HiLoad Superdex 200 gel filtration column (GE Healthcare, USA) pre-equilibrated with buffer containing 20 mM Tris (pH 8.0), 10 mM sodium citrate, 150 mM NaCl and 2 mM β-mercaptoethanol. To express selenomethionine (Se-Met)-substituted CcpC-IBD protein, the bacterial cells were cultured in 1 L of M9 medium supplemented with an amino acid mixture containing L-( +)-Se-Met at 310 K. When the OD600 was between 0.6 and 0.8, the cells were induced with 0.5 mM IPTG for 8 h. The Se-Met-substituted protein was purified under the same conditions as the native protein. To obtain crystal structures of BaCcpC-IBD bound to citrate, we added 10 mM sodium citrate throughout the whole purification process. During purification, the presence of the proteins was detected by sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS-PAGE) in a 15% gel with Coomassie blue R-250 for staining.
Electrophoretic mobility shift assay (EMSA) experiments
A chemiluminescent EMSA kit was purchased from Beyotime Biotechnology (Nan**g, China), and a biotin-labelled B. amyloliquefaciens citB promoter was synthesized by Generation (Wuhan, China). Supplementary Table S1 provides the list of oligonucleotide sequences used for EMSA analysis. EMSA experiments between BaCcpC and citB promoter DNA were performed at room temperature. For EMSA between BaCcpC and the citB protomer, various concentrations of purified BaCcpC (2.5–40 μM) protein were incubated with the the citB promoter (800 nM) for 30 min. For EMSA of BaCcpC with regions I and II of the citB promoter, purified BaCcpC (40 μM) protein was incubated with each citB promoter (800 nM). To determine the effect of citrate on binding between BaCcpC and the citB promoter, BaCcpC (40 μM) protein was incubated with citB promoter (800 nM) at various citrate concentrations (0–70 μM) for 30 min. After incubation, the reaction mixture was placed in a 6% acrylamide gel on ice using 0.5 × Tris/Borate/EDTA (TBE) buffer. The product was analysed by chemiluminescence detection with the Tanon 4600 Chemiluminescent Imaging system (Tanon, China).
Size-exclusion chromatography coupled to multiangle light
The oligomer states of BaCcpC-FL were analysed by size-exclusion chromatography coupled to multiangle light scattering. One hundred microlitres of the BaCcpC-FL protein that had been incubated or not incubated with 10 mM sodium citrate was loaded into a Superdex 200 Increase 10/300 GL column (GE Healthcare) at 295 K with a flow rate of 0.5 ml min−1 The aqueous mobile phase consisted of 20 mM Tris buffer (pH 8.0) containing 150 mM NaCl and 2 mM 2-mercaptoethanol. Data were collected and processed using ASTRA 6 (Wyatt Technology, USA).
Crystallization, data collection, structure determination, and refinement
BaCcpC-IBD was concentrated to 20 mg ml−1 using a Vivaspin centrifugal concentrator (Cut-off: 10 kDa, Millipore, USA). The initial crystallization was performed using the sitting-drop vapour-diffusion method at 295 K using a Crystal Screen HT high-throughput reagent kit (Hampton Research, USA). Crystals of BaCcpC-IBD were grown in 10% polyethylene glycol 6000, 5% 2 methyl-2,4-pentanediol (MPD), and 0.1 M HEPES (pH 7.5) using a 1:1 ratio of protein to mother liquor at 287 K. Finally, crystals of BaCcpC-IBD were obtained in sitting drops over 8% (w/v) polyethylene glycol 6000, 6% MPD, and 0.1 M HEPES (pH 7.5) using a 1:1 ratio of protein to mother liquor at 287 K. Immediately after the single crystals were taken from their drop, they were soaked for 5 s in cryoprotectant solution consisting of the mother liquor solution containing 25% (v/v) glycerol and subsequently flash-cooled in liquid nitrogen.
The dataset was collected at 100 K using an ADSC Q310 CCD detector at Beamline 7A, Pohang Accelerator Laboratory (Pohang, Republic of Korea). The peak wavelength of Se-Met in BaCcpC was determined to be 0.9826 Å by a fluorescence scan. Data were collected using the inverse beam method with an oscillation range of 1° per frame over a 360° rotation, and the exposure time was 5 s per frame. The crystal of the BaCcpC-IBD protein was diffracted to 2.3 Å resolution. The diffraction data were processed, merged, and scaled using the HKL-2000 program33. Initial phases were obtained using AUTOSOL in the software package PHENIX 1.15.2_3472 (phenix-online.org)34. Refinement was performed using the Crystallographic Object-Oriented Toolkit (COOT) and phenix.refine35,36. The data collection and refinement statistics are given in Table 1. Structural images were generated by PyMOL37.
Molecular dynamics
Simulations of the CcpC-IBD domain (apo state of monomer and apo state of dimer) were performed using the GROMACS GROMACS 2020.3 manual (www.gromacs.org)38. For the simulation, the Amber99sb-ildn force field and explicit solvent based on the TIP3P model were employed39,40. The structures were solvated with explicit water in periodic rectangular boxes under normal (150 mM) saline conditions. The LINCS algorithm was used to constrain all bond lengths41. Long-range electrostatic interactions were treated with the particle-mesh Ewald method40. For the MD calculations, the nonbonded (electrostatic and VDW) cut-off range was 10 Å and the time step was 2 fs.
Before each MD simulation, the entire system was first minimized by a 1000-step steepest descent calculation followed by a 50,000-step conjugate gradient optimization. The total computer simulation time for BaCcpC-IBD was 1 ns.
Data availability
The atomic coordinates and structure factors for BaCcpC-IBD (PDB ID 7DMW) have been deposited in the RCSB Protein Data Bank (www.rcsb.org).
References
Krebs, H. A., Kay, J. & Weitzman, P. D. J. Krebs’ Citric Acid Cycle : Half A Century and Still Turning (Biochemical Society, 1987).
Strasters, K. C. & Winkler, K. C. Carbohydrate metabolism of Staphylococcus aureus. J. Gen. Microbiol. 33, 213–229. https://doi.org/10.1099/00221287-33-2-213 (1963).
Sonenshein, A. L. Control of key metabolic intersections in Bacillus subtilis. Nat. Rev. Microbiol. 5, 917–927. https://doi.org/10.1038/nrmicro1772 (2007).
Fujita, Y. Carbon catabolite control of the metabolic network in Bacillus subtilis. Biosci. Biotechnol. Biochem. 73, 245–259. https://doi.org/10.1271/bbb.80479 (2009).
Richardson, A. R., Somerville, G. A. & Sonenshein, A. L. Regulating the intersection of metabolism and pathogenesis in gram-positive bacteria. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.MBP-0004-2014 (2015).
Kim, H. J., Roux, A. & Sonenshein, A. L. Direct and indirect roles of CcpA in regulation of Bacillus subtilis Krebs cycle genes. Mol. Microbiol. 45, 179–190. https://doi.org/10.1046/j.1365-2958.2002.03003.x (2002).
Sonenshein, A. L. CodY, a global regulator of stationary phase and virulence in Gram-positive bacteria. Curr. Opin. Microbiol. 8, 203–207. https://doi.org/10.1016/j.mib.2005.01.001 (2005).
Hartmann, T. et al. Catabolite control protein E (CcpE) is a LysR-type transcriptional regulator of tricarboxylic acid cycle activity in Staphylococcus aureus. J. Biol. Chem. 288, 36116–36128. https://doi.org/10.1074/jbc.M113.516302 (2013).
Jourlin-Castelli, C., Mani, N., Nakano, M. M. & Sonenshein, A. L. CcpC, a novel regulator of the LysR family required for glucose repression of the citB gene in Bacillus subtilis. J. Mol. Biol. 295, 865–878. https://doi.org/10.1006/jmbi.1999.3420 (2000).
Maddocks, S. E. & Oyston, P. C. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 154, 3609–3623. https://doi.org/10.1099/mic.0.2008/022772-0 (2008).
Huffman, J. L. & Brennan, R. G. Prokaryotic transcription regulators: More than just the helix-turn-helix motif. Curr. Opin. Struct. Biol. 12, 98–106 (2002).
Aravind, L., Anantharaman, V., Balaji, S., Babu, M. M. & Iyer, L. M. The many faces of the helix-turn-helix domain: Transcription regulation and beyond. FEMS Microbiol. Rev. 29, 231–262. https://doi.org/10.1016/j.femsre.2004.12.008 (2005).
Pérez-Rueda, E. & Collado-Vides, J. Common history at the origin of the position–function correlation in transcriptional regulators in archaea and bacteria. J. Mol. Evol. 53, 172–179 (2001).
Giannopoulou, E. A. et al. Crystal structure of the full-length LysR-type transcription regulator CbnR in complex with promoter DNA. FEBS J. https://doi.org/10.1111/febs.15764 (2021).
Alanazi, A. M., Neidle, E. L. & Momany, C. The DNA-binding domain of BenM reveals the structural basis for the recognition of a T-N11-A sequence motif by LysR-type transcriptional regulators. Acta Crystallogr. D Biol. Crystallogr. 69, 1995–2007. https://doi.org/10.1107/S0907444913017320 (2013).
Koentjoro, M. P., Adachi, N., Senda, M., Ogawa, N. & Senda, T. Crystal structure of the DNA-binding domain of the LysR-type transcriptional regulator CbnR in complex with a DNA fragment of the recognition-binding site in the promoter region. FEBS J. 285, 977–989. https://doi.org/10.1111/febs.14380 (2018).
Tyrrell, R. et al. The structure of the cofactor-binding fragment of the LysR family member, CysB: A familiar fold with a surprising subunit arrangement. Structure 5, 1017–1032. https://doi.org/10.1016/s0969-2126(97)00254-2 (1997).
Taylor, J. L. et al. The crystal structure of AphB, a virulence gene activator from Vibrio cholerae, reveals residues that influence its response to oxygen and pH. Mol. Microbiol. 83, 457–470. https://doi.org/10.1111/j.1365-2958.2011.07919.x (2012).
Ezezika, O. C., Haddad, S., Clark, T. J., Neidle, E. L. & Momany, C. Distinct effector-binding sites enable synergistic transcriptional activation by BenM, a LysR-type regulator. J. Mol. Biol. 367, 616–629. https://doi.org/10.1016/j.jmb.2006.09.090 (2007).
Chen, J. et al. Structural and biochemical analysis of the citrate-responsive mechanism of the regulatory domain of catabolite control protein E from Staphylococcus aureus. Biochemistry 57, 6054–6060. https://doi.org/10.1021/acs.biochem.8b00671 (2018).
Jo, I. et al. Structural details of the OxyR peroxide-sensing mechanism. Proc. Natl. Acad. Sci. USA 112, 6443–6448. https://doi.org/10.1073/pnas.1424495112 (2015).
Lerche, M. et al. The solution configurations of inactive and activated DntR have implications for the sliding dimer mechanism of LysR transcription factors. Sci. Rep. 6, 19988. https://doi.org/10.1038/srep19988 (2016).
Monferrer, D. et al. Structural studies on the full-length LysR-type regulator TsaR from Comamonas testosteroni T-2 reveal a novel open conformation of the tetrameric LTTR fold. Mol. Microbiol. 75, 1199–1214. https://doi.org/10.1111/j.1365-2958.2010.07043.x (2010).
Muraoka, S. et al. Crystal structure of a full-length LysR-type transcriptional regulator, CbnR: Unusual combination of two subunit forms and molecular bases for causing and changing DNA bend. J. Mol. Biol. 328, 555–566. https://doi.org/10.1016/s0022-2836(03)00312-7 (2003).
Lochowska, A. et al. Identification of activating region (AR) of Escherichia coli LysR-type transcription factor CysB and CysB contact site on RNA polymerase alpha subunit at the cysP promoter. Mol. Microbiol. 53, 791–806. https://doi.org/10.1111/j.1365-2958.2004.04161.x (2004).
Mittal, M. et al. Dual role of CcpC protein in regulation of aconitase gene expression in Listeria monocytogenes and Bacillus subtilis. Microbiology (Reading) 159, 68–76. https://doi.org/10.1099/mic.0.063388-0 (2013).
Kim, H. J., Mittal, M. & Sonenshein, A. L. CcpC-dependent regulation of citB and lmo0847 in Listeria monocytogenes. J. Bacteriol. 188, 179–190. https://doi.org/10.1128/JB.188.1.179-190.2006 (2006).
Maddocks, S. E. & Oyston, P. C. F. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology (Reading) 154, 3609–3623. https://doi.org/10.1099/mic.0.2008/022772-0 (2008).
Picossi, S., Belitsky, B. R. & Sonenshein, A. L. Molecular mechanism of the regulation of Bacillus subtilis gltAB expression by GltC. J. Mol. Biol. 365, 1298–1313. https://doi.org/10.1016/j.jmb.2006.10.100 (2007).
Ding, Y. et al. Metabolic sensor governing bacterial virulence in Staphylococcus aureus. Proc. Natl. Acad. Sci. U S A 111, E4981-4990. https://doi.org/10.1073/pnas.1411077111 (2014).
Bundy, B. M., Collier, L. S., Hoover, T. R. & Neidle, E. L. Synergistic transcriptional activation by one regulatory protein in response to two metabolites. Proc. Natl. Acad. Sci. U S A 99, 7693–7698. https://doi.org/10.1073/pnas.102605799 (2002).
Kim, Y. et al. Crystal structure of the ligand-binding domain of a LysR-type transcriptional regulator: Transcriptional activation via a rotary switch. Mol. Microbiol. 110, 550–561. https://doi.org/10.1111/mmi.14115 (2018).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326. https://doi.org/10.1016/S0076-6879(97)76066-X (1997).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
Emsley, P. & Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. Sect. D Biol. Crystallogr. 60, 2126–2132. https://doi.org/10.1107/S0907444904019158 (2004).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367. https://doi.org/10.1107/S0907444912001308 (2012).
DeLano, W. L. Use of PYMOL as a communications tool for molecular science. Abstr. Pap. Am. Chem. S 228, U313–U314 (2004).
Van Der Spoel, D. et al. GROMACS: Fast, flexible, and free. J. Comput. Chem. 26, 1701–1718. https://doi.org/10.1002/jcc.20291 (2005).
Duan, Y. et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012. https://doi.org/10.1002/jcc.10349 (2003).
Price, D. J. & Brooks, C. L. 3rd. A modified TIP3P water potential for simulation with Ewald summation. J. Chem. Phys. 121, 10096–10103. https://doi.org/10.1063/1.1808117 (2004).
Hess, B. P-LINCS: A parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4, 116–122. https://doi.org/10.1021/ct700200b (2008).
Acknowledgements
The authors acknowledge the staff at beamline 5C of the Pohang Light Source (PLS, Pohang, Republic of Korea) for their help with data collection. The authors acknowledge the professor Nam-Chul Ha at Seoul National University for he help with SEC-MALS experiment. This work was supported by the Natural Science Foundation of Liaoning province (Grant No. 2019-MS-065 to Y. Xu), the key lab of marine bioactive substance and modern analytical technique, SOA (Grant No. MBSMAT-2019-05 to L. **), the Program for Liaoning Excellent Talents in University (Grant No. LJQ2015030 to Y. Xu), the Dalian High level Talents Innovation support Plan Project (Grant No. 2019CT09 to C. Quan), the Fundamental Research Funds for the Central Universities (Grant No. DC201502020203 to Y. Xu, Grant No. DC201502020201 to C. Quan), and a National Research Foundation of Korea (NRF) (Grant Nos. NRF-2017M3A9F6029736 and NRF-2021R1I1A1A01050838).
Author information
Authors and Affiliations
Contributions
Y.X., K.H.N., and C.Q. conceived the study and designed experiments; W.L., and, J.C. performed the experiments for protein crystallization and solved the structure. L.J., and K.H.N. analyzed the data. Y.X. and K.H.N. wrote the manuscript; Z.Y.L., M.L., G.J., and Q.Y. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, W., Chen, J., **, L. et al. Functional and structural analysis of catabolite control protein C that responds to citrate. Sci Rep 11, 20285 (2021). https://doi.org/10.1038/s41598-021-99552-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-99552-x
- Springer Nature Limited