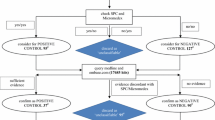
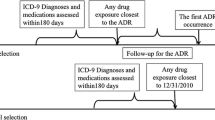
The processing pipeline for the construction of CRESCENDDI included the following steps: (1) web data extraction of information related to DDIs and single-drug ADRs from 4 different online resources for DDIs, single-drug ADRs, and drug indications; (2) map** (normalization) of drug names appearing in the extracted data; (3) extraction of the intersection of the DDI online resources; (4) manual annotation and map** of English language text descriptions for DDIs, single-drug ADRs, and drug indications to MedDRA concepts; (5) generation of positive controls for DDIs using the normalized intersection of the DDI online resources; (6) generation of negative controls using drugs and AEs from the positive control set combined with PubMed search; and (7) aggregation of information on single-drug ADRs and drug indications for the drugs appearing in the DDI reference set (i.e., positive and negative control sets) and generation of negative controls for single drugs with a process similar to the one followed in the previous step for DDI negative controls.
Web data extraction
DDI data were derived from the following online resources: the British National Formulary (BNF) website25, the French National Drug Safety Institute (ANSM) Portable Document Format (PDF) file (Thesaurus)26 and the Micromedex platform27. For BNF and Micromedex, web data extraction tools in Python 3.628 enabled the extraction of the relevant fields into a Comma Separated Values (CSV) file (June-August 2018). For Thesaurus, the R package IMThesaurusANSM29 was used and the resulting R dataframe from the 2019 update was converted to a CSV file.
The tables contained the following fields:
-
interacting drug name 1 (D1) (e.g., Metoprolol Tartrate);
-
interacting drug name 2 (D2) (e.g., Lidocaine);
-
text description for the DDI (e.g., Lidocaine is predicted to increase the risk of cardiovascular adverse effects when given with metoprolol. Manufacturer advises use with caution or avoid);
-
severity label (e.g., Severe);
-
evidence label (if available) (e.g., Study).
For single-drug ADRs, the following sources were considered: the BNF website25 and SIDER dataset30. For drug indications, SIDER was used. BNF ADR data were extracted in a similar way as previously with DDI data (automated web data extraction) into a CSV file, while SIDER data for ADRs and drug indications were already available in CSV files.
A table containing the following fields was constructed:
-
drug name (e.g., Metoprolol Tartrate);
-
event text description (e.g., Bradycardia);
-
event type (e.g., ADR);
-
source (e.g., SIDER).
Drug name map**
To facilitate usability and ensure compatibility, a standardization process was followed such that we could provide a resource with normalized concepts to standard terminologies for drugs and medical events. Specifically, the Observational Health Data Sciences and Informatics (OHDSI) Vocabulary version 5 was selected for map** the drug names occurring in each of the DDI online resources into RxNorm and RxNorm Extension standard codes (at the Ingredient level) using OHDSI Usagi31.
We removed combination drugs (as DDIs of their constituent drug ingredients were separately mentioned), vaccines, vitamins, herbal medicines, food, beverages, supplements, tobacco, and lab tests. Also, generic drug classes (e.g., combined hormonal contraceptives, hormonal replacement therapy) appearing in the BNF were not mapped to their individual drug ingredients, as there was no table on the BNF website specifying the drugs belonging in each drug class. We mapped the remaining unique drug names occurring in the DDI resources to OHDSI standard vocabulary concept identifiers. For example, Metoprolol Tartrate was mapped to the RxNorm Ingredient concept metoprolol. For Thesaurus, a native French speaker (pharmacist) confirmed the drug map**s of French drug names to English language OHDSI concepts.
A similar process was followed for drugs that appear in the single-drug data (ADRs and indications).
Intersection of DDI online resources
By matching drug names to their mapped drug ingredients in the extracted DDI data tables, we obtained the set of common drug pairs across the tables and generated a new table that contains only the DDIs and associated information which could be found in each of the DDI online resources under consideration. Cases where the interacting drug map** of D1 and D2 were swapped in the original data tables (i.e., (D1,D2) and (D2,D1)) were considered equivalent.
The final table contained the following fields:
-
drug_1 concept name (e.g., metoprolol);
-
drug_2 concept name (e.g., lidocaine);
-
bnf description (e.g., Lidocaine is predicted to increase the risk of cardiovascular adverse effects when given with metoprolol. Manufacturer advises use with caution or avoid);
-
micromedex description (e.g., lidocaine toxicity (anxiety, myocardial depression, cardiac arrest));
-
bnf severity (e.g., Severe);
-
ansm severity (e.g., Précautions d’emploi (Precautions for use));
-
micromedex severity (e.g., major);
-
bnf evidence (e.g., Study);
-
micromedex evidence (e.g., probable).
Adverse event and indication map**s
For DDI-related text descriptions in English from BNF and Micromedex that could be found in the DDI intersection table, a drug name blinding process was performed by replacing the interacting drug names with a common token in all cases (i.e., ‘X’). In this way, the number of unique descriptions was reduced, thus facilitating the map** process that followed. For example, the descriptions:
Both dexibuprofen and ibuprofen can increase the risk of nephrotoxicity.and
Both polymyxins and streptomycin can increase the risk of nephrotoxicity.
were both mapped to the following blinded description:
Both X and X can increase the risk of nephrotoxicity.
The set of blinded text descriptions for BNF and Micromedex was extracted from the table and a semi-automated map** process using OHDSI Usagi mapped them to MedDRA PT concepts. We explicitly focused on text descriptions that included clinical manifestations of DDIs, e.g., X may increase the risk of hypoglycaemia when taken with X. Text descriptions containing a potential mechanism of the interaction were left unmapped. In some cases, a single text description was linked to multiple concepts. For example:
Interaction Effect: An increased risk of cardiotoxicity (QT prolongation, torsades de pointes, cardiac arrest).
includes 3 different MedDRA PTs.
Also, serotonin syndrome was not mapped to its corresponding MedDRA PT and was not further considered as an AE for inclusion in the reference set.
Text descriptions from the BNF regarding single-drug ADRs were mapped to MedDRA PTs (where possible), but only for the drug ingredients that could be found in the DDI pair intersection table. SIDER ADR and indication data for the same list of drugs were also mapped to OHDSI concepts; however, for this resource, MedDRA PT codes were already available.
Positive controls
The set of positive controls was derived from the DDI intersection table, using map**s of text descriptions to AEs that were generated in the previous step. It contained 10,286 drug-drug-event (DDE) triplets, 454 unique individual drug ingredients, and 179 unique AEs (as OHDSI concepts) in total.
Negative controls
The set of negative controls was generated by randomly pairing two drug ingredients from the 454 unique drug ingredients that can be found in the positive controls and, in case the random drug pair did not appear in any of the DDI online resources, then it was randomly paired with an AE from the 179 unique AEs present in the positive control set. The choice of generating negative controls with common drug ingredients and AEs as the ones appearing in positive controls aimed to ensure the generation of a balanced reference set that does not contain added biases by design. For each of the created DDE triplets, a customized query (“(DRUG_1_CONCEPT_NAME) AND (DRUG_2_CONCEPT_NAME) AND ((EVENT_CONCEPT_NAME) OR (interaction))”, e.g., “(Oxazepam) AND (Naproxen) AND ((Hyperkalaemia) OR (interaction))”) was submitted to PubMed in an automated fashion and, if the search returned no results, the triplet was added to the negative control set. This process aimed to provide more confidence, to the best of our ability, about the absence of literature evidence of a potential DDI for the triplet under consideration, rather than definitive evidence to support the lack of a potential association.
The process was repeated until the number of negative controls with non-zero counts in the US Food and Drug Administration (FDA) Adverse Event Reporting System (FAERS) database (see Usage Notes) was similar in size (N = 4,544) compared to the equivalent subset of positive controls. The negative control set included 161 unique AEs and 435 unique drug ingredients.
Single-drug ADRs, indications, and negative controls
By replacing text descriptions from BNF (N = 1,538) and MedDRA PT codes to their corresponding mapped OHDSI concepts, a table with ADR and indication information related to the drug ingredients of the DDI reference set was generated. The table included: 438 unique drug ingredients, which could be found in at least one of the resources under consideration (i.e., BNF and SIDER), 3,492 AEs and 1,557 indication terms (as OHDSI concepts).
BNF and SIDER jointly contained 69,721 single-drug ADRs, with 12,318 common instances; this set could be utilized as a source for single-drug positive controls. This set covered 381 unique drugs and 835 unique AE concepts. Random pairing of those drugs and AE concepts followed by submission of a customized query (“(DRUG_CONCEPT_NAME) AND (EVENT_CONCEPT_NAME) AND ((adverse event) OR (adverse drug reaction))’”, e.g., “(oxazepam) AND (myoclonus) AND ((adverse event) OR (adverse drug reaction))”) to PubMed (to ensure absence of literature evidence of a potential ADR for the various drug-event associations) enabled the generation of a negative control set for single drugs (N = 12,141).