Abstract
Transcriptionally silent chromatin often localizes to the nuclear periphery. However, whether the nuclear envelope (NE) is a site for post-transcriptional gene repression is not well understood. Here we demonstrate that Schizosaccharomyces pombe Lem2, an NE protein, regulates nuclear-exosome-mediated RNA degradation. Lem2 deletion causes accumulation of RNA precursors and meiotic transcripts and de-localization of an engineered exosome substrate from the nuclear periphery. Lem2 does not directly bind RNA but instead interacts with the exosome-targeting MTREC complex and its human homolog PAXT to promote RNA recruitment. This pathway acts largely independently of nuclear bodies where exosome factors assemble. Nutrient availability modulates Lem2 regulation of meiotic transcripts, implying that this pathway is environmentally responsive. Our work reveals that multiple spatially distinct degradation pathways exist. Among these, Lem2 coordinates RNA surveillance of meiotic transcripts and non-coding RNAs by recruiting exosome co-factors to the nuclear periphery.
Similar content being viewed by others
Main
Eukaryotic genomes are pervasively transcribed, giving rise to sense and antisense RNAs from intra- and intergenic regions and repetitive elements. Accumulation of cryptic unstable transcripts (CUTs) may cause genome instability through RNA-DNA hybridization1. Other coding and non-coding RNAs are continuously transcribed but function during specific developmental stages; for example, fission yeast meiotic mRNAs are expressed but rapidly degraded in vegetative cells2,3,4,5. Constant surveillance is required to prevent the aberrant accumulation of these transcripts, which is mediated by both nuclear and cytosolic pathways. However, how RNA-degradation pathways are coordinated in the nucleus is not well understood. Although the role of perinuclear anchoring in transcriptional silencing has been intensely studied, the effect of nuclear organization on post-transcriptional repression remains unclear.
Eukaryotic nuclear RNA degradation is mediated by the nuclear exosome, a multiprotein complex containing two 3′→5′ exoribonucleases, Rrp6 (ribosomal RNA processing 6) and Dis3/Rrp44 (ref. 6). RNA-surveillance pathways contribute to substrate specificity and RNA processing, assisting exosome-targeting complexes through polyadenylation and RNA helicase activities that unwind RNA secondary structures6. In Schizosaccharomyces pombe, these targeting complexes include TRAMP (Trf4/5–Air1/2–Mtr4 polyadenylation) and MTREC (Mtl1–Red1 core). The TRAMP complex comprises a non-canonical poly(A) polymerase (Cid14; Trf4/Trf5 in Saccharomyces cerevisiae), a zinc-knuckle RNA-binding protein (Air1), and an RNA helicase (Mtr4). Fission yeast TRAMP degrades transcripts derived from pericentromeric repeats and plays a minor role in CUT elimination and small nucleolar RNA (snoRNA) processing7,8,9. The MTREC complex, also known as NURS (nuclear RNA silencing), comprises the Mtr4-like helicase Mtl1 and the zinc-finger domain protein Red1 (RNA elimination defective 1). MTREC mediates turnover of CUTs and meiotic and non-spliced transcripts9,10,11 and is orthologous to the human PAXT (poly(A) tail exosome targeting) complex12. MTREC assembles into an 11-subunit ‘super complex,’ in which Red1 scaffolds different submodules and recruits Rrp6 via Mtl1 (refs. 10,11,13). The submodules have different activities and include the canonical Poly(A) polymerase Pla1 and the complexes Red5–Pab2–Rmn1, Ars2–Cbc1–Cbc2, and Iss10–Mmi1 (refs. 9,14,15,16,17,18,19).
Exosome targeting and function are best understood in the context of meiotic transcript turnover in S. pombe. The YTH (YT521-B homology) protein Mmi1 recognizes hexanucleotide motifs (UNAAAC) known as DSR (determinant of selective removal) sequences2,3,20. Substrate binding requires Mmi1 dimerization and interaction with its partner Erh1 (enhancer of rudimentary homolog 1) to form the tetrameric Erh1–Mmi1 complex (EMC). EMC sequesters RNA substrates, preventing their nuclear export and translation21,22,23,24,7c). Because Mei2 dot assembly during early meiosis causes inactivation of Mmi1-dependent elimination2, we also tested whether the accumulation of exosome targets in lem2∆ cells was indirectly caused by stabilizing sme2. However, deleting sme2+ did not suppress the accumulation of meiotic transcripts or snoRNAs in lem2∆ cells (Extended Data Fig. 7d), further arguing that Lem2 plays a direct role in their degradation. Together, these results imply that RNA degradation is coordinated through distinct degradation pathways that depend on Lem2 and Iss10 and differ in their substrate specificity.
a,b, Left, scheme of the subunit that was mutated on its own or together with Lem2. Right, transcript levels of ssm4, mei4, sme2, mei3, sno20, and snR42, quantified by RT–qPCR in the indicated strains (a, erh1∆; b, iss10∆). Substrates controlled in an additive or non-additive manner are indicated above. Data normalized to act1 transcript levels are shown relative to WT (n = 4 independent biological replicates, except mei4 transcripts in combination with iss10∆, n = 2). The individual replicates are shown in a floating bar plot, and the line depicts the median. Statistical analysis was performed using one-way ANOVA, and letters denote groups with significant differences from Tukey’s post hoc tests at P < 0.05. c, K-means clustering of RNA-seq data. Genes were grouped using K-means clustering on the basis of their differential expression in the indicated mutants. Eight clusters were generated, and the clusters with most upregulated transcripts in lem2∆ (clusters 1–4) were analyzed with AnGeLi with a two-tailed Fisher’s exact test and false-discovery rate (FDR) = 0.05 (ref. 46). The red–blue color scale represents the log2(fold change expression) relative to WT. d, Left, scheme showing the experimental setup and medium shift. Right, transcript levels of sme2 and ade2 quantified by RT–qPCR in the indicated strains (gray, WT; red, lem2∆) and medium (YES or EMM-N). e, Left, scheme showing the experimental setup and media shift. Right, transcript levels of sme2 and ade2, quantified by RT–qPCR in the indicated strains (gray, WT; red, lem2∆), time points (t = 0, 3, 6 or 9 hours) and medium (YES or EMM-N). For d and e, data from independent biological replicates are normalized to euchromatin expression and shown relative to the WT strain in YES (d; n = 4) or the WT strain at t = 0 in YES (e; n = 3).
Previously, Lem2-dependent heterochromatin assembly was reported to be regulated by nutrient availability43. Heterochromatic transcripts are no longer repressed in a Lem2-dependent manner when growth conditions are restricted using EMM (Edinburgh minimal medium; Extended Data Fig. 7e). Since nitrogen starvation promotes sexual differentiation and entry into meiosis, we tested whether EMM lacking nitrogen (EMM-N) impacts Lem2 functions. Intriguingly, we found that sme2 transcript levels increased in WT cells in EMM and EMM-N, similar to what is seen in lem2∆ cells, suggesting that Lem2 becomes inactivated under these growth conditions (Fig. 6d). To further investigate whether Lem2 activity is regulated during meiotic onset, we grew mating-competent h90 cells in rich medium and analyzed sme2 transcripts upon transfer to EMM-N in a time-course experiment (Fig. 6e). sme2 transcripts steadily increased in WT cells upon nitrogen starvation, whereas this transcript was already upregulated in lem2∆ cells in rich medium and did not further increase upon starvation (Fig. 6e). Although these meiotic transcripts did not accumulate in WT cells to the same level as in lem2∆ cells, it is possible that some cells within the population may not have entered the meiotic program, likely explaining the weaker phenotype in WT cells. On the basis of these results, we propose that nutrient-dependent inactivation of Lem2 contributes to the fine-tuning of meiotic transcript accumulation during early meiosis. Beyond this role in meiosis, Lem2 is key to a spatially and functionally distinct pathway that operates in post-transcriptional regulation at the NE.
Discussion
Transcriptionally silent chromatin often localizes to the nuclear periphery, which is thought to provide a specialized compartment for gene repression57. However, whether the NE influences broader modes of gene regulation is largely unknown. Here, we demonstrate that the conserved INM protein Lem2 collaborates with the nuclear exosome to control and process ncRNAs and meiotic transcripts. Several lines of evidence argue that this Lem2-mediated regulation occurs post-transcriptionally and is distinct from its previously described function in heterochromatin silencing38,43: (1) although lem2+ deletion causes accumulation of various targets (ncRNA, LTRs, and heterochromatic transcripts), Pol II abundance is increased at heterochromatin but not at ncRNA genes; (2) repression of targets co-regulated by Lem2 and the exosome is unaffected by silencing factors (Clr4/HP1, RNAi, SHREC) that work with Lem2 at heterochromatin; (3) mutants lacking Lem2 and exosome factors (Rrp6, MTREC) display overlap** but non-additive phenotypes in the repression of several meiotic transcripts; (4) Lem2 physically interacts with the MTREC subunit Red1 via its MSC domain; (5) Lem2 does not affect chromatin binding of Mmi1 and Red1; (6) instead, Lem2 promotes Mmi1 and Red1 binding to RNA substrates; (7) Lem2 is critical for the localization and association of a DRS-containing transcript with a subnuclear pool of Mmi1 at the NE. On the basis of these findings, we propose that Lem2 recruits exosome co-factors to the nuclear periphery to coordinate post-transcriptional RNA processing and degradation, a function independent of heterochromatin silencing.
Despite major differences in transcriptional and post-transcriptional regulation, Lem2 plays a general role in increasing the local concentration of factors involved in these processes at the nuclear periphery. As previously shown, Lem2 promotes the localization of the repressor complex SHREC to heterochromatin38. Similarly, we find that Lem2 facilitates the association of MTREC with RNA substrates (Fig. 5) and interacts with Red1 through its MSC domain (Fig. 3). Lem2 therefore employs a common mechanism for interaction with its partners, which is in agreement with the MSC-dependent recruitment of members of the ESCRT pathways in NE repair and other functions reported for Lem2 homologs58,59,60,61,62. Thus, Lem2 may provide a general INM recruitment platform for interaction with various partners. How Lem2 specifically coordinates these different functions remains unknown.
Various membrane-less nuclear bodies have been assigned to specific functions, such as the Cajal body involved in snRNA and snoRNA modification and assembly63. RNA turnover has also been proposed to occur within subnuclear foci formed by exosome factors in an Iss10-dependent manner13,24,26,28. This raises the question of how these nuclear foci relate to Lem2-mediated regulation. Although an engineered exosome substrate localizes more frequently at the nuclear periphery (Fig. 4), such preferential localization was not seen for Red1, Erh1, or Mmi1 foci (Extended Data Fig. 7). Nonetheless, we found that a subpopulation of Mmi1 foci co-localizing with this substrate also localized to the NE in a Lem2-dependent manner (Fig. 4). Interestingly, Mmi1 and another nuclear RNA-processing complex, CCR4–NOT, have been shown to associate with the nuclear rim protein Amo1 (ref. 39), implying that multiple spatially distinct degradation pathways exist. Indeed, lem2∆ mutants that also lack the EMC or TRAMP complexes show synergistic upregulation of various transcripts (Fig. 6). This likely explains why the phenotype is weaker than that of red1∆ and rrp6∆ mutants (see model, Fig. 7). The impact on distinct pathways would explain why Iss10 is critical for nuclear foci formation while having only a minor role in meiotic transcript turnover in vegetative cells10,26,28,29. We speculate that separate pathways allow differential regulation and distinct substrate specificity. We further observe that Lem2 mediates some MTREC-independent function in snoRNA processing (Fig. 2). Further work should elucidate Lem2’s specific role in the broad spectrum of its substrates.
Left, Lem2 cooperates with the nuclear exosome machinery at the nuclear periphery by directly interacting with Red1 and assisting in substrate binding by Mmi1 and MTREC. Absence or inactivation of Lem2 results in impaired DSR substrate location at the periphery, reduced binding to exosome-targeting factors, and eventually an increase in untimely transcript expression due to lower RNA degradation (Lem2-dependent). Lem2 may also interact with other factors associated with Mmi1. Right, nuclear foci are formed by multiple exosome factors in an Iss10-dependent manner and likely represent additional sites of RNA degradation (Lem2-independent). Both pathways show partial redundancy for most meiotic transcripts, whereas ncRNAs are mainly degraded by the Lem2-dependent pathway.
The expression of meiotic genes is toxic during vegetative growth and therefore tightly regulated64. Under nutrient starvation, cells mate and undergo meiosis, which requires precise orchestration of the meiotic gene expression program65. Nitrogen starvation is signaled by TOR pathway inactivation, resulting in the dephosphorylation and degradation of Iss10 (ref. 29). Similarly, cells shifted from rich to minimal medium show upregulation of key meiotic regulators in a Lem2-dependent manner (Fig. 6e). Hence, we propose that Lem2 is part of a regulatory circuit that fine-tunes gene expression in response to environmental cues. How different growth conditions alter Lem2 activity remains unknown. Lem2 transcript (Extended Data Fig. 7f) and protein levels43 are unaltered under minimal growth conditions, suggesting that Lem2 may undergo post-translational modifications that impact association with the NE or downstream factors. Thus, it will be interesting to address how nutritional cues affect Lem2 function and to elucidate the underlying signaling cascade.
Our data demonstrate that RNA degradation is not a generic process, but a spatially specific mode of regulation that is critical in the biological response to environmental changes. Since both Lem2 and exosome-targeting complexes are found in higher eukaryotes, we propose that this pathway is broadly conserved. Indeed, the nuclear exosome localizes to the NE in other organisms, including Drosophila66. Moreover, Lem2 binding to MTREC is conserved for its human homolog, the PAXT complex (Fig. 3b). Further studies examining the nuclear location of the exosome in fission yeast and higher eukaryotes may shed light on the mechanisms that collaborate to regulate this complex machinery.
Methods
Yeast techniques, plasmids, and strains
A list of the strains used in this study can be found in Supplementary Tables 1 and 2. All plasmids used in this study are listed in Supplementary Table 3.
Standard medium and genome engineering methods were used. Cells were grown in rich medium (YE5S aka YES) except for the data shown in Figures 3d and 6a and Extended Data Figure 6e,f (cells were initially grown in minimal medium (EMM) then shifted for 12 hours into YES) and Figure 6c (cells were initially grown in YES then shifted into EMM-N). Strains expressing constructs derived from pREP81 vectors (shown in Extended Data Fig. 3a) were grown in EMM-leu.
Strains expressing epitope-tagged proteins were generated through homologous recombination using pFA6a or pYM-based vectors67 and expressed from the chromosomal locus using the endogenous promoter. For Lem2-GFP fusions expressing strains, the respective fragments were cloned into a pJK210 vector and integrated into lem2∆ background strains in the endogenous locus. The soluble MSC-GFP overexpression construct was expressed under control of the TEF (Ashbya gossypii) promoter.
RNA-seq library preparation and data analysis
For RNA-seq, 1 μg of RNA was used as starting material to prepare libraries, following the manufacturer’s instructions for NEBNext Ultra Directional RNA Library Prep Kit for Illumina (NEB). Two or more biological replicates were used for generating libraries in parallel. For man1∆ and clr4∆, three biological replicates were processed. Single-end, 50-bp sequencing of libraries was performed on a HiSeq1500 sequencer in the LAFUGA core facility at the Gene Center, LMU Munich. Raw reads were de-multiplexed using Je (v1.2). Adapter-trimmed reads were aligned to the S. pombe reference genome (ASM294v2.27) and a custom GTF file using STAR (v2.7.3a), then processed using RSEM (v1.3.3). Differential expression was analyzed using the DESeq2 (v1.22.2) and tximport (v1.10.1) R libraries. For PCA plotting, data were batch normalized using the sva (v3.30.1) R library. Bedgraph coverage files for plus and minus strands were generated using genomecov (bedtools v2.29.1). RNA-seq data have been deposited in Gene Expression Omnibus (GEO) database under the accession number GSE174347. The full code for all NGS-related workflow is available at: https://github.com/Tsvanemden/Martin_Caballero_et_al_2021.
RT–qPCR analyses
RT–qPCR experiments were performed as previously described68. Briefly, cells were lysed by bead beating (Precellys 24, Bertin instruments) using TRIzol reagent and zirconia/silica beads (BioSpec), followed by centrifugation at 13,000 r.p.m. for 15 minutes at 4 °C. Recovered supernatant was extracted with chloroform and reprecipitated with isopropyl alcohol. Resuspended RNA was treated with DNaseI, and 10 mg of RNA was used in standard RT reactions using oligo((dT)20-N) primers. cDNAs were quantified by qPCR using primaQUANT SYBR Master mix (Steinbrenner Laborsysteme) and a QuantStudio 3 or QuantStudio 5 Real-Time PCR system (Applied Biosystems/Thermo Fisher). Data from 2–6 independent biological replicates are shown as individual data points together with the median.
For northern blotting and RT–qPCR analysis of snoRNAs (Fig. 2), total RNAs were prepared using the hot acid phenol method and treated with DNAse (Ambion). Concentrations were measured with a Nanodrop. Two micrograms of DNAse-treated RNAs were denatured at 65 °C for 5 minutes in the presence of strand-specific primers. Reactions were carried out with 100 units Maxima H minus Reverse Transcriptase (Thermo Fisher Scientific) at 50 °C for 30 minutes. The enzyme was then denatured at 85 °C for 5 minutes, and reactions were diluted to 1:10 ratio. Experiments included negative controls without Reverse Transcriptase. Samples were analyzed by qPCR with the SYBR Green Master Mix (Roche) and a LightCycler LC480 apparatus (Roche). Quantification was performed using the ∆Ct method.
Primers used for qPCR are listed in Supplementary Table 4. Expression values for WT and mutants were calculated by normalizing to act1 and then dividing by the mean of all samples from the same experiment (group normalization), as previously described69. When analyzing single and double mutants for epistatic interactions, statistical testing was performed using R. Multiple testing was performed using ANOVA followed by a Tukey’s post hoc test at a 0.05 significance level.
Northern blotting
Three micrograms of RNA were separated on a 2% agarose gel and transferred overnight by capillarity on a nylon membrane (GE Healthcare) in SSC 10× buffer. RNAs were then UV-crosslinked to the membrane using a Stratalinker apparatus. In vitro-transcribed dig-labeled RNA probes were generated and incubated with the membrane using the Dig-Northern-Starter Kit (Roche), following the manufacturer’s instructions. Membranes were washed twice in 2× SSC 0.1% SDS and once in 1× SSC 0.1% SDS, for 10 minutes at 65 °C. Revelation was done according to the kit instructions using Chemidoc Imaging MP detection device (Biorad). Oligonucleotides used to generate DNA templates for RNA probes are listed in Supplementary Table 4.
Y2H assays
Constructs were cloned into either pGADT7 or pGBKT7 vectors (Clontech). The S. cerevisiae strain Y2H Gold (Takara) was used to co-transform the plasmids, following the manufacturer’s instructions. Spotting assays were performed 3–5 days after transformation of the plasmids. Different dropout mixes were used to assess the strength of the interaction: SDC-Leu-Trp (Formedium), SDC-Leu-Trp-His (Formedium) and SDC-Leu-Trp-His-Ade (Formedium).
Co-immunoprecipitation assays
CoIP assays were performed following a previously described protocol70 with a few modifications in the lysis buffer. The lysis buffer contained 50 mM HEPES pH 7.4, 100 mM NaCl, 10% glycerol, 1 mM EDTA pH 8, 2.5 mM MgCl2, 0.5% NP-40, 1× complete EDTA-free protease inhibitor cocktail (Roche), 2 mM PMSF (Serva), 20 mM N-ethylmaleimide (NEM, Sigma). Cell lysates were prepared by resuspending the pellets from 150 to 200 OD600 units in 800 μl lysis buffer. Cells were lysed by bead beating (Precellys 24, Bertin instruments) with zirconia/silica beads (BioSpec), and lysates were cleared by centrifugation (800g, 5 min). Clarified extracts were incubated with pre-equilibrated GFP-Trap or Myc-Trap (Chromotek) for 1.5 hours at 4 °C. Following immunoprecipitation, the bound material was incubated with RNase (Roche) or benzonase (Sigma) in the reactions, as indicated. Beads were washed four times with lysis buffer and two times with wash buffer (50 mM Tris pH 7.5, 100 mM NaCl, 1 mM EDTA pH 8, 2.5 mM MgCl2). Proteins were eluted with 30 μl HU buffer (8 M urea, 5% SDS, 200 mM Tris-HCl pH 6.8, 20 mM dithiothreitol (DTT) and bromophenol blue 1.5 mM) and analyzed by immunoblotting.
Immunoblotting
Cells corresponding to OD600 = 1 (≈2 × 107 cells) were pelleted from a suspension culture grown to mid-log phase. Total protein extracts were made using trichloroacetic acid (TCA, Sigma) precipitation68. Proteins were solubilized in HU buffer (8 M urea, 5% SDS, 200 mM Tris-HCl pH 6.8, 20 mM dithiothreitol (DTT, Sigma) and bromophenol blue 1.5 mM). Proteins were resolved on NuPAGE 4–12% gradient gels (Invitrogen) or self-made 8% gels. Proteins were transferred onto PVDF membranes (polyvinylidene fluoride membranes, GE Healthcare) and analyzed by standard immunoblotting techniques using specific antibodies. H3 antibody was used as a loading control.
Live-cell microscopy
Live-cell imaging was essentially performed as described38. In brief, cells were grown overnight on rich medium (YES) to the logarithmic phase. Prior to imaging, cells were attached with lectin (Sigma) to glass-bottom dishes with a micro well (MatTek). Cells were imaged on a Zeiss AxioObserver Z1 confocal spinning disk microscope with an EMM-CCD camera (Photometrics, Evolve 512) through a Zeiss Alpha Plan/Apo ×100/1.46 oil DIC M27 objective lens. Z-stacks were obtained at focus intervals of 0.4 μm. FiJi/ImageJ software was used to measure the distances between the foci and the periphery.
For the imaging of cells expressing CFP-Mmi1, the following setup was used: confocal microscopy was performed at the Core Facility Bioimaging of the Biomedical Center (LMU Munich) with an inverted Leica SP8 X WLL microscope, equipped with 405-nm laser, WLL2 laser (470–670 nm), and acusto-optical beam splitter. Images were acquired with a HC PL APO ×93/1.30 GLYC motCORR-STED WHITE objective, and Z-stacks were obtained at focus intervals of 0.25 μm. Images were deconvolved using the SVI Huygens suite and FiJi/ ImageJ software was used to measure the distances between the foci and the periphery.
Single molecular RNA fluorescence in situ hybridization
smFISH was performed as described in the previous report with slight modification71. Cells of WT (H1N2330), lem2Δ (H1N2324), and red1Δ (H1N2328) were pre-cultured in minimum medium (EMMG5S) for overnight at 30 °C, then transferred into rich medium (YES) and cultured for 16 hours. The cells (~1 × 108) were mixed with one-tenth of meiosis-induced PSB1940 (YY548-13C) cells (~1 × 107) as a positive control for smFISH, and then fixed with 4% formaldehyde (Polysciences) at 30 °C for 30 minutes. The cells were labeled with Quasar 570-labeled RNA probe set for sme2 (See Supplementary Table 4 in ref. 71 in detail), then mounted in ProLong Glass Antifade Mountant (Thermo Fisher Scientific).
The cells were observed using ×60 PlanApo N OSC oil-immersion objective lens (numerical aperture (NA) = 1.4, Olympus) on the DeltaVision Elite system (GE Healthcare) equipped with pco.edge 4.2 sCMOS camera (PCO). Chromatic shifts were corrected using Chromagnon software (v0.87) using a bleed-through fluorescence image as a reference72. The images were denoised by the ND-safir program73, deconvolved using the built-in SoftWoRx software (v7.0.0), and then projected by maximum-intensity projection. The brightness of the images was adjusted using the Fiji software74 for better visualization, without changing the gamma settings.
RIP assays
Cell lysates prepared from equal amounts of cells (between 100–165 OD600) were fixed with 1% formaldehyde (Sigma) for 15 minutes at RT, followed by quenching 5 minutes at RT with 125 mM glycine (Sigma). Cultures were spun down, washed once with 1×PBS and frozen in liquid nitrogen. Cells pellets were resuspended in lysis buffer (250 mM KCl, 1% Triton X-100, 0.1% SDS, 0.1% Na-deoxycholate, 50 mM HEPES pH 7.5, 2 mM EDTA, 2 mM EGTA, 5 mM MgCl2, 0.1% Nonidet P-40, 20% glycerol) and lysed with glass beads (Roth) in a bead beater (Precellys 24, Peqlab). Fragmented material was sonicated (Qsonica Q800R1) for 1 hour with cycles of 30 seconds ON/OFF at 4 °C. The lysate was cleared and used for immunoprecipitation with 15 μl GFP-trap or Myc-trap (Chromotek). Beads were washed with lysis buffer, and bound material was eluted from beads with elution buffer (50 mM Tris-HCl pH 8.0, 10 mM EDTA, 1% SDS) with 15 minutes of incubation at RT and 15 minutes at 65 °C. RIP samples along with inputs were de-crosslinked at 95 °C for 10 minutes. Samples were then incubated with 40 μg of proteinase K (Sigma) for 4 hours at 37 °C. RNA was recovered with a phenol-chloroform-isoamyl alcohol extraction (Thermo Fisher Scientific) followed by precipitation with sodium acetate, isopropanol, and glycogen (Thermo Fisher Scientific). Precipitated RNA was digested with DNase I (Thermo Fisher Scientific) for 2 hours at 37 °C. Purified RNA was used for reverse transcription following manufacturer’s instructions (Superscript III, Thermo Fisher Scientific) and used for qPCR, as described for RT–qPCR samples. The primers used are listed in Supplementary Table 4.
Pol II-S5P, H3K9me2, and Red1-Myc ChIP–qPCR assays
ChIP was performed as previously described38 with minor modifications, as follows: 100 ml of a 0.5 OD600 cell suspension was crosslinked with 1% formaldehyde (Roth) and quenched with 125 mM glycine (Sigma). Following lysis and sonication, solubilized chromatin corresponding to approximately 5–6 × 108, 4 × 108, and 9 × 108 cells was immunoprecipitated with antibodies against Pol II-S5P (25 μl supernatant, kindly provided by A. Ladurner), H3K9me2 (2 μl), and Myc-trap (10 μl, Chromotek), respectively.
RT–qPCR experiments were performed as described above, using the primers listed in Supplementary Table 4. The IP values were divided by the input and corrected for variation by normalizing to the mean of three euchromatin loci (act1+, tef3+, ade2+). The data were shown as relative to the untagged strain.
Antibodies
Rat monoclonal anti-HA (3F10, 1:1,000) and mouse monoclonal anti-GFP (B-2, 1:1,000) antibodies were purchased from Roche and Santa Cruz Biotechnology, respectively. Mouse monoclonal anti-H3 (1B1-B2, 1:5,000) was obtained from Active Motif. Rabbit polyclonal anti-Myc (ab9106, 1:2,000, Abcam) or mouse monoclonal anti-Pol II-S5P (3E8, generated by the lab of D. Eick) antibodies were kindly provided by M. Spletter and by A. Ladurner, respectively. Mouse monoclonal H3K9me2 ChIP grade (ab1220) was purchased from Abcam. Secondary antibodies fused to HRP were used for detection (goat anti-mouse HRP 1:3,000, BioRad; goat anti-rat HRP 1:3,000, Merck Millipore; goat anti-rabbit HRP 1:3,000, BioRad)
Statistics and reproducibility
Representative results of at least two independent experiments were presented in all of the figure panels for all blots. No statistical methods were performed to predetermine the sample size, and the number of biological replicates was based on similar studies for each experiment. Analyses of the variance were performed, and pairwise differences were evaluated with Tukey’s post hoc test using R statistical language (R Development Core Team, 2008); different groups are marked with letters at the 0.05 significance level. For all graphs, data the individual replicates are shown in a floating bar plot, and the line depicts the median. P values were generated using two-tailed Student’s t-tests or chi-square (χ2) analyses; n.s., P ≥ 0.05, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All sequencing data that support the findings of this study have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (GEO) and are accessible through the GEO Series accession number GSE174347. Source data are provided with this paper.
Code availability
Full code for all NGS-related workflow is available at: https://github.com/Tsvanemden/Martin_Caballero_et_al_2021.
References
Wahba, L., Gore, S. K. & Koshland, D. The homologous recombination machinery modulates the formation of RNA–DNA hybrids and associated chromosome instability. eLife 2, e00505 (2013).
Harigaya, Y. et al. Selective elimination of messenger RNA prevents an incidence of untimely meiosis. Nature 442, 45–50 (2006).
Hiriart, E. et al. Mmi1 RNA surveillance machinery directs RNAi complex RITS to specific meiotic genes in fission yeast. EMBO J. 31, 2296–2308 (2012).
Tashiro, S., Asano, T., Kanoh, J. & Ishikawa, F. Transcription-induced chromatin association of RNA surveillance factors mediates facultative heterochromatin formation in fission yeast. Genes Cells 18, 327–339 (2013).
Zofall, M. et al. RNA elimination machinery targeting meiotic mRNAs promotes facultative heterochromatin formation. Science 335, 96–100 (2012).
Schmid, M. & Jensen, T. H. Nuclear quality control of RNA polymerase II transcripts. Wiley Interdiscip. Rev. RNA 1, 474–485 (2010).
Bühler, M., Spies, N., Bartel, D. P. & Moazed, D. TRAMP-mediated RNA surveillance prevents spurious entry of RNAs into the Schizosaccharomyces pombe siRNA pathway. Nat. Struct. Mol. Biol. 15, 1015–1023 (2008).
Zhang, K. et al. Clr4/Suv39 and RNA quality control factors cooperate to trigger RNAi and suppress antisense RNA. Science 331, 1624–1627 (2011).
Zhou, Y. et al. The fission yeast MTREC complex targets CUTs and unspliced pre-mRNAs to the nuclear exosome. Nat. Commun. 6, 7050 (2015).
Egan, E. D., Braun, C. R., Gygi, S. P. & Moazed, D. Post-transcriptional regulation of meiotic genes by a nuclear RNA silencing complex. RNA 20, 867–881 (2014).
Lee, N. N. et al. Mtr4-like protein coordinates nuclear RNA processing for heterochromatin assembly and for telomere maintenance. Cell 155, 1061–1074 (2013).
Ogami, K., Chen, Y. & Manley, J. RNA surveillance by the nuclear RNA exosome: mechanisms and significance. Noncoding RNA 4, 8–21 (2018).
Sugiyama, T. & Sugioka-Sugiyama, R. Red1 promotes the elimination of meiosis-specific mRNAs in vegetatively growing fission yeast. EMBO J. 30, 1027–1039 (2011).
Dobrev, N. et al. The zinc-finger protein Red1 orchestrates MTREC submodules and binds the Mtl1 helicase arch domain. Nat. Commun. 12, 3456 (2021).
Lemay, J.-F. et al. The nuclear poly(A)-binding protein interacts with the exosome to promote synthesis of noncoding small nucleolar RNAs. Mol. Cell 37, 34–45 (2010).
Sugiyama, T., Wanatabe, N., Kitahata, E., Tani, T. & Sugioka-Sugiyama, R. Red5 and three nuclear pore components are essential for efficient suppression of specific mRNAs during vegetative growth of fission yeast. Nucleic Acids Res. 41, 6674–6686 (2013).
Thillainadesan, G. et al. Conserved protein Pir2ARS2 mediates gene repression through cryptic introns in lncRNAs. Nat. Commun. 11, 1–11 (2020).
Yamanaka, S., Yamashita, A., Harigaya, Y., Iwata, R. & Yamamoto, M. Importance of polyadenylation in the selective elimination of meiotic mRNAs in growing S. pombe cells. EMBO J. 29, 2173–2181 (2010).
St-André, O. et al. Negative regulation of meiotic gene expression by the nuclear poly(a)-binding protein in fission yeast. J. Biol. Chem. 285, 27859–27868 (2010).
Yamashita, A. et al. Hexanucleotide motifs mediate recruitment of the RNA elimination machinery to silent meiotic genes. Open Biol. 2, 120014 (2012).
Andric, V. et al. A scaffold lncRNA shapes the mitosis to meiosis switch. Nat. Commun. 12, 770 (2021).
Hazra, D., Andrić V, Palancade, B., Rougemaille, M. & Graille, M. Formation of S. pombe Erh1 homodimer mediates gametogenic gene silencing and meiosis progression. Sci. Rep. 10, 1034 (2020).
Shichino, Y., Otsubo, Y., Kimori, Y., Yamamoto, M. & Yamashita, A. YTH-RNA-binding protein prevents deleterious expression of meiotic proteins by tethering their mRNAs to nuclear foci. eLife 7, 999 (2018).
Sugiyama, T. et al. Enhancer of rudimentary cooperates with conserved RNA-processing factors to promote meiotic mRNA decay and facultative heterochromatin assembly. Mol. Cell 61, 747–759 (2016).
**e, G. et al. A conserved dimer interface connects ERH and YTH family proteins to promote gene silencing. Nat. Commun. 10, 1–15 (2019).
Yamashita, A., Takayama, T., Iwata, R. & Yamamoto, M. A novel factor Iss10 regulates Mmi1-mediated selective elimination of meiotic transcripts. Nucleic Acids Res. 41, 9680–9687 (2013).
Yamanaka, S. et al. RNAi triggered by specialized machinery silences developmental genes and retrotransposons. Nature 493, 557–560 (2013).
Shichino, Y., Otsubo, Y., Yamamoto, M. & Yamashita, A. Meiotic gene silencing complex MTREC/NURS recruits the nuclear exosome to YTH-RNA-binding protein Mmi1. PLoS Genet. 16, e1008598–19 (2020).
Wei, Y. et al. TOR targets an RNA processing network to regulate facultative heterochromatin, developmental gene expression and cell proliferation. Nat. Cell Biol. 23, 243–256 (2021).
Shichino, Y., Yamashita, A. & Yamamoto, M. Meiotic long non-coding meiRNA accumulates as a dot at its genetic locus facilitated by Mmi1 and plays as a decoy to lure Mmi1. Open Biol. 4, 140022 (2014).
Ding, D.-Q. et al. Meiosis-specific noncoding RNA mediates robust pairing of homologous chromosomes in meiosis. Science 336, 732–736 (2012).
Shimada, T., Yamashita, A. & Yamamoto, M. The fission yeast meiotic regulator Mei2p forms a dot structure in the horse-tail nucleus in association with the sme2 locus on chromosome II. Mol. Biol. Cell 14, 2461–2469 (2003).
Harr, J. C., Gonzalez-Sandoval, A. & Gasser, S. M. Histones and histone modifications in perinuclear chromatin anchoring: from yeast to man. EMBO Rep. 17, 139–155 (2016).
Nakayama, J., Rice, J. C., Strahl, B. D., Allis, C. D. & Grewal, S. I. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292, 110–113 (2001).
Bannister, A. J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001).
Sadaie, M. et al. Balance between distinct HP1 family proteins controls heterochromatin assembly in fission yeast. Mol. Cell. Biol. 28, 6973–6988 (2008).
Sugiyama, T. et al. SHREC, an effector complex for heterochromatic transcriptional silencing. Cell 128, 491–504 (2007).
Barrales, R. R., Forn, M., Georgescu, P. R., Sarkadi, Z. & Braun, S. Control of heterochromatin localization and silencing by the nuclear membrane protein Lem2. Genes Dev. 30, 133–148 (2016).
Holla, S. et al. Positioning heterochromatin at the nuclear periphery suppresses histone turnover to promote epigenetic inheritance. Cell 180, 150–164 (2019)
Iglesias, N. et al. Native chromatin proteomics reveals a role for specific nucleoporins in heterochromatin organization and maintenance. Mol. Cell 77, 51–66 (2019).
Banday, S., Farooq, Z., Rashid, R., Abdullah, E. & Altaf, M. Role of inner nuclear membrane protein complex Lem2–Nur1 in heterochromatic gene silencing. J. Biol. Chem. 291, 20021–20029 (2016).
Hirano, Y. et al. Lem2 is retained at the nuclear envelope through its interaction with Bqt4 in fission yeast. Genes Cells 14, 943–14 (2018).
Tange, Y. et al. Inner nuclear membrane protein Lem2 augments heterochromatin formation in response to nutritional conditions. Genes Cells 21, 812–832 (2016).
Lock, A. et al. PomBase 2018: user-driven reimplementation of the fission yeast database provides rapid and intuitive access to diverse, interconnected information. Nucleic Acids Res. 201, 403–407 (2018).
Watanabe, Y. & Yamamoto, M. S. pombe mei2+ encodes an RNA-binding protein essential for premeiotic DNA synthesis and meiosis I, which cooperates with a novel RNA species meiRNA. Cell 78, 487–498 (1994).
Bitton, D. A., Schubert, F. & Dey, S. AnGeLi: A tool for the analysis of gene lists from fission yeast. Front. Genet. 6, 330 (2015).
Steglich, B., Filion, G. J., Steensel, Bvan & Ekwall, K. The inner nuclear membrane proteins Man1 and Ima1 link to two different types of chromatin at the nuclear periphery in S. pombe. Nucleus 3, 77–87 (2012).
Cam, H. P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37, 809–819 (2005).
Gallagher, P. S. et al. Iron homeostasis regulates facultative heterochromatin assembly in adaptive genome control. Nat. Struct. Mol. Biol. 25, 372–383 (2018).
Yamamoto, M. The selective elimination of messenger RNA underlies the mitosis–meiosis switch in fission yeast. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 86, 788–797 (2010).
Chen, H.-M., Futcher, B. & Leatherwood, J. The fission yeast RNA binding protein Mmi1 regulates meiotic genes by controlling intron specific splicing and polyadenylation coupled RNA turnover. PLoS ONE 6, e26804 (2011).
Hu, C. et al. Structural insights into chromosome attachment to the nuclear envelope by an inner nuclear membrane protein Bqt4 in fission yeast. Nucleic Acids Res. 47, 1573–1584 (2019).
Hediger, F., Taddei, A., Neumann, F. R. & Gasser, S. M. Methods for visualizing chromatin dynamics in living yeast. Meth. Enzymol. 375, 345–365 (2004).
Caputo, S. et al. The carboxyl-terminal nucleoplasmic region of MAN1 exhibits a DNA binding winged helix domain. J. Biol. Chem. 281, 18208–18215 (2006).
Brachner, A., Reipert, S., Foisner, R. & Gotzmann, J. LEM2 is a novel MAN1-related inner nuclear membrane protein associated with A-type lamins. J. Cell Sci. 118, 5797–5810 (2005).
Yamashita, A., Shichino, Y. & Yamamoto, M. The long non-coding RNA world in yeasts. Biochim. Biophys. Acta 1859, 147–154 (2016).
Akhtar, A. & Gasser, S. M. The nuclear envelope and transcriptional control. Nat. Rev. Genet. 8, 507–517 (2007).
Appen, A. et al. LEM2 phase separation promotes ESCRT-mediated nuclear envelope reformation. Nature 582, 1–25 (2020).
Capella, M., Caballero, L. M., Pfander, B., Braun, S. & Jentsch, S. ESCRT recruitment by the S. cerevisiae inner nuclear membrane protein Heh1 is regulated by Hub1-mediated alternative splicing. J. Cell Sci. 133, jcs250688 (2020).
Gu, M. et al. LEM2 recruits CHMP7 for ESCRT-mediated nuclear envelope closure in fission yeast and human cells. Proc. Natl Acad. Sci. USA 114, E2166–E2175 (2017).
Pieper, G. H., Sprenger, S., Teis, D. & Oliferenko, S. ESCRT-III/Vps4 controls heterochromatin-nuclear envelope attachments. Dev. Cell 53, 27–41.e6 (2020).
Thaller, D. J. et al. An ESCRT–LEM protein surveillance system is poised to directly monitor the nuclear envelope and nuclear transport system. eLife 8, e45284 (2019).
Mao, Y. S., Zhang, B. & Spector, D. L. Biogenesis and function of nuclear bodies. Trends Genet. 27, 295–306 (2011).
Harigaya, Y. & Yamamoto, M. Molecular mechanisms underlying the mitosis-meiosis decision. Chromosome Res. 15, 523–537 (2007).
Mata, J., Lyne, R., Burns, G. & Bähler, J. The transcriptional program of meiosis and sporulation in fission yeast. Nat. Genet. 32, 143–147 (2002).
Graham, A. C., Kiss, D. L. & Andrulis, E. D. Differential distribution of exosome subunits at the nuclear lamina and in cytoplasmic foci. Mol. Biol. Cell 17, 1399–1409 (2006).
Janke, C. et al. A versatile toolbox for PCR‐based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Braun, S. et al. The Cul4–Ddb1–Cdt2 ubiquitin ligase inhibits invasion of a boundary-associated antisilencing factor into heterochromatin. Cell 144, 41–54 (2011).
Georgescu, P. R., Capella, M., Burkart, S. F. & Braun, S. The euchromatic histone mark H3K36me3 preserves heterochromatin through sequestration of an acetyltransferase complex in fission yeast. Micro. Cell 7, 80–92 (2020).
Capella, M. et al. Nucleolar release of rDNA repeats for repair involves SUMO-mediated untethering by the Cdc48/p97 segregase. Nat. Commun. 12, 1–12 (2021).
Ding, D.-Q. et al. Chromosome-associated RNA protein complexes promote pairing of homologous chromosomes during meiosis in Schizosaccharomyces pombe. Nat. Commun. 10, 1–12 (2019).
Matsuda, A., Schermelleh, L., Hirano, Y., Haraguchi, T. & Hiraoka, Y. Accurate and fiducial-marker-free correction for three-dimensional chromatic shift in biological fluorescence microscopy. Sci. Rep. 8, 7583 (2018).
Boulanger, J. et al. Patch-based nonlocal functional for denoising fluorescence microscopy image sequences. IEEE Trans. Med. Imaging 29, 442–454 (2010).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank members of the Braun lab and A. Ladurner, P. Korber, and M. Halic for fruitful discussions during the study. We thank S. Lall (Life Science Editors) for editorial assistance. We further thank A. Ladurner (LMU Munich, Germany), M. Smolle (LMU Munich, Germany), and M. Halic (St. Jude Children’s Research Hospital, Memphis, TN, USA) for strains, plasmids and reagents; A. Yamashita; C. Brönner, M. Halic, and M. Murawska for protocols; T. Straub and T. Schauer from the BMC Bioinformatics core facility for training and counseling; G. Timinszky, J. Preisser, A. Thomae, and the BMC Bioimaging core facility for training and counseling in confocal microscopy and data deconvolution; D. Q. Ding and K. Okamasa for helpful discussion for smFISH; S. Stöcker for assistance in library preparation for RNA-seq analysis; G. Schermann for support in antisense transcript analysis. Schemes in figures were created with BioRender.com. S. B. was supported by the German Research Foundation (DFG) through the collaborative research center CRC 1064 (project ID 213249687-SFB 1064) and the research grant BR 3511/4-1. I. S. was supported by the Leibniz program of the DFG (SI586/6-1). Additional support was provided by the Australian Government through the Australian Research Council’s Discovery Projects funding scheme awarded to T. F. (project DP190100423); by the Centre National pour la Recherche Scientifique and the Agence Nationale de la Recherche to M. R. (ANR-16-CE12-0031); and by the JSPS KAKENHI grants to Y. Hirano (JP19K06489, JP20H05891), Y. K. (JP19K23725), and Y. Hiraoka (JP18H05533, JP19K22389).
Funding
Open access funding provided by Justus-Liebig-Universität Gießen.
Author information
Authors and Affiliations
Contributions
L. M. C., R. R. B., and S. B. conceived the study. L. M. C., R. R. B., and T. v. E. performed RNA-seq experiments, and T. v. E. generated the RNA-seq analysis pipeline. N. D., M. C., and L. M. C. performed Y2H assays. L. M. C. and V. N. S. S. performed western blots. M. C. and L. M. C. conducted coIP experiments. L. M. C., V. N. S. S., and S. F.-B. performed RT–qPCR experiments. A. N. and M. R. performed snoRNA northern blots and RT–qPCR experiments. Y. Hirano conducted smFISH experiments. All other experiments were performed by L. M. C. All authors analyzed the data. S.B., Y. Hiraoka, I.S., T.F. and M.R. supervised experiments. L. M. C. and S. B. conceived and wrote the manuscript. Y. Hirano, Y. K., and Y. Hiraoka made original observations confirming independently several key findings. All authors contributed to editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Francois Bachand and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Anke Sparmann, and Carolina Perdigoto, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Lem2 represses non-coding RNAs and meiotic genes.
a, b, Volcano plots depicting statistical significance (Y axis) against fold change (X axis) from the RNA-seq data of man1∆ vs WT (a) and clr4∆ vs WT (b). Genes significantly up- (red) or downregulated (blue) are highlighted (log2 fold change > 1 or < -1 with P adj. value < 0.01 by the Wald test, as implemented within the DESeq2 framework). c, Pie charts showing of ncRNA, LTR, protein coding, other transcripts (pseudogene, rRNA, snoRNA, snRNA, tRNA). Left: genome-wide distribution of transcript features in a WT genome. Right: transcript feature distribution of the significantly upregulated transcripts in the indicated mutants (log2 fold change > 1 with P adj. value < 0.01 by the Wald test). d, Transcript levels of mei4 and mei3 analyzed by RT-qPCR qPCR (n = 4 independent biological replicates; except lem2∆: n = 3). e, Transcript levels of ssm4, SPAC212.08c, cen dg, and ade2 analyzed by RT-qPCR (n = 4 independent biological replicates; except clr2∆, ago1∆: n = 3). For (d, e), data are normalized to act1 transcript levels or the average of selected euchromatic genes (act1+, tef3+, ade2+), respectively, and shown relative to WT. The individual replicates are shown in a floating bar plot and the line depicts the median. HC, heterochromatin; EC, euchromatin.
Extended Data Fig. 2 Lem2 cooperates with different exosome factors.
a, PCA plot from the RNA-seq libraries generated of the indicated strains. Each dot represents a biological replicate. b, c, Scatterplots of genome-wide log2 fold change expression from transcripts in red1∆ vs lem2∆ (b) or air1∆ vs lem2∆, erh1∆ vs lem2∆ and ccr4∆ vs lem2∆ (c), each of them relative to WT. The linear regression line is depicted together with the Pearson correlation coefficient value (R). d, Genome-wide plot showing the log2 coverage of all annotated antisense mRNAs (Y axis, normalized antisense reads). All units are normalized to 1200 bp (X axis, gene location). The colors depict different strains (see legend on the left). e, Table with selected results from gene list enrichment analysis from the clusters (#) 1-5 from Fig. 2b. The Bähler Lab AnGeLi tool with a two-tailed Fisher’s exact test and a false discovery rate of 0.05 was used for this analysis (Ref. 46). Freq., frequency; corr., corrected; gene exp., gene expression; GO biol., Gene Ontology biological process; mei., meiotic; mid., middle, sporul. mod., sporulation module. f, Table showing the linear expression values of multiple exosome subunits, as shown in Fig. 2a in lem2∆ cells. Data were retrieved from RNA-seq analyses and are shown as log2 fold change of lem2∆ over WT.
Extended Data Fig. 3 Lem2 collaborates with distinct RNA degradation pathways.
a, Expression levels of Red1-dependent (left, red) and Red1-independent (right, blue) islands in lem2∆ cells. Gene names are indicated below the graph. Data were retrieved from RNA-seq analyses and are shown as log2 fold change of lem2∆ over WT. b, ChIP-qPCR analysis of Red1-Myc enrichment at mei4+ and tef3+ in the indicated strains (n = 7 independent biological replicates, except sample Red1-myc iss10∆, n = 3). c, ChIP-qPCR analysis of GFP-Mmi1 enrichment at mei4+ and tef3+ in the indicated strains (n = 4 independent biological replicates). d, ChIP-qPCR analysis of H3K9me2 enrichment of mei4+ (heterochromatin island), tf2-5+ (HOOD) in the indicated strains (n = 2 independent biological replicates). For (b-d), data are divided by the input and normalized to euchromatin levels (act1+, tef3+, ade2+). e, sno20 and snR42 transcript levels quantified by RT-qPCR in the indicated strains. Data from independent biological replicates (sno20, n = 6; snR42, n = 5) are normalized to act1 transcript levels and shown relative to WT on a log2 scale. Statistical analysis was performed using one-way ANOVA, and letters denote significant differences between groups with Tukey’s post hoc tests at P < 0.05. For (b-e), the individual replicates are shown in a floating bar plot and the line depicts the median.
Extended Data Fig. 4 Lem2 interacts with the nuclear exosome through the MSC domain and functions in a location-dependent manner.
a, Co-immunoprecipitation of Red1-6xHA with Lem2-GFP WT (Lem2-GFP FL) or truncation mutants lacking the N terminal (Lem2-GFP ∆N) or MSC domain (Lem2-GFP ∆MSC). Histone H3 served as loading control. Proteins were expressed from a pREP81 based overexpression vector transformed in lem2∆ strains. b, Y2H analysis of Red1, Iss10, Rrp6, Pab2 and Mtl1 with MSC-Lem2 or Mtl1, grown for 3 days on medium with different auxotrophies (SDC-Leu-Trp, SDC-Leu-Trp-His-Ade). Fusions with Gal4-activating domain (pGADT7-AD) or Gal-4-DNA-binding domain (pGBKT7-BD) are shown. c, Schematic representation of Lem2 truncation constructs. Protein domains and positions (amino acids) are indicated. All constructs were C-terminally GFP-tagged and inserted in the endogenous locus in lem2∆ background cells. d, Representative images from live-cell microscopy of the constructs in Extended Data Fig. 4c. Left panels show the GFP excited channel and right panels show the corresponding bright field images. A projection of several Z-stacks with maximum intensity is shown. For some cells, Lem2-GFP is out of plane and not visible. Scale bar = 10 μm. e, Immunoblot of Lem2-GFP constructs in Extended Data Fig. 4c. In here, two different promoters were used to overexpress the soluble MSC fragment (left = pGPD, right = pTEF). H3 served as loading control. Asterisk (*) indicates unspecific band. f, sme2, ssm4 and ade2 transcript levels quantified by RT-qPCR in Lem2 truncation mutants. Data from n = 4 independent biological replicates were normalized to act1 transcript levels and shown relative to WT. The individual replicates are shown in a floating bar plot and the line depicts the median. For (a, d, and e), blots shown are representative examples of experiments reproduced at least three times.
Extended Data Fig. 5 Exosome substrates localize to the nuclear periphery.
a, Top: smFISH analysis for sme2 RNA using sequence-specific probes labeled with Quasar®570. The sme2 locus was visualized using a lacO/LacI-GFP system. Cut11-CFP was used to mark the NE. Scale bar, 2 μm. Bottom: smFISH color merge panels showing multiple cells. Arrows indicate meiotic cells, where the sme2 RNA signal collapses to a single dot. Scale bar, 10 μm. For top and bottom: color merge images show the smFISH signal (magenta), the lacO/LacI-GFP (yellow) and Cut11-CFP (blue). b, Transcript levels of luciferase and tef3 quantified by RT-qPCR on a strain encoding 0×DSR copies (0×DSR), 14×DSR copies (14×DSR), 14×DSR and the NE marker in a WT (14×DSR Cut11-mCherry), iss10∆ background (14×DSR Cut11-mCherry iss10∆) or red1∆ background (14×DSR Cut11-mCherry red1∆). Data from n = 4 independent biological replicates are normalized to act1 transcript levels and shown relative to the 14×DSR strain. The individual replicates are shown in a floating bar plot and the line depicts the median. c, Top: Schematic representation of the live-cell imaging acquisition method. Z-stacks were acquired from cell nuclei detecting DSR dots and the NE (Cut11-mCherry). To assess localization, DSR dots were assigned to one out of three concentrical zones with equal areas within the nucleus, as measured by the distance to the NE. Bottom: Live-cell microscopy representative images of the DSR containing strain in a WT or lem2∆ background. Cut11-mCherry marks the NE. A single z-stack is shown. The DSR location was quantified in WT and lem2∆ backgrounds relative to the periphery expressed in percentage of dots. Statistical analysis was performed using χ2 test. ****, P < 0.0001. Scale bar = 1 μm. d, Left: Live-cell microscopy representative image of the sme2+ locus (sme2::ura4::lacOp, his7+::LacI-GFP) in a WT background. Cut11-mCherry was used as a marker for the NE. A single z-stack is shown. Scale bar = 1 μm. Right: quantification of sme2+ locus location in WT and background relative to the periphery expressed in percentage of dots. For (c-d), n denotes the number of cells counted in two independent experiments.
Extended Data Fig. 6 Lem2 regulates transcript binding by the exosome machinery.
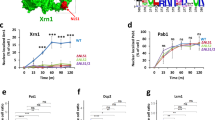
a, Transcript levels of sme2 quantified by RT-qPCR in the input from RIP samples in the indicated strains (Lem2-GFP, n = 3; GFP-Mmi1, n = 4, Red1-Myc, n = 3 independent biological replicates). Data are normalized to act1 transcript levels and shown relative to the untagged strain. b, Transcript binding to GFP-Mmi1 analyzed by RIP-qPCR analysis in WT, red1∆ and lem2∆ cells. Data from n = 4 (except GFP-Mmi1 in WT background, n = 3) biological replicates are divided by the input and shown relative to the median of the untagged strain. c, Transcript levels of sme2 and ssm4 quantified by RT-qPCR in the input of the indicated RIP strains (n = 4; except GFP-Mmi1 in WT background, n = 3). Data were normalized to act1 transcript levels and shown relative to the untagged strain. The individual replicates are shown in a floating bar plot and the line depicts the median.
Extended Data Fig. 7 RNA regulation by Lem2 at the nuclear periphery occurs independently of exosome factors associated with nuclear foci.
a, Left: schematic representation of the S. pombe nucleus divided into three equal areas designated I-III. Right: Representative live-cell microscopy images of GFP-Mmi1, Erh1-GFP and Red1-GFP in a WT or lem2∆ background. Cut11-mCherry was used as a marker for the NE. A single z-stack is shown. The quantification of protein localization is shown in WT and lem2∆ backgrounds relative to the periphery expressed in percentage of dots. n = number of cells counted in two independent experiments. n.s.= not significant from χ2 test analysis. Scale bar = 1 μm. b, Co-immunoprecipitation of Red1-6xHA with GFP-Mmi1 in WT or lem2∆ background. H3 served as loading control. Proteins were expressed from their endogenous loci. Two independent experiments were performed with similar results. c, Transcript levels of mei4, sme2, mei3 and sno20 quantified by RT-qPCR in the indicated strains (n = 3). d, Transcript levels of mei4, ssm4, mei3 and sno20 quantified by RT-qPCR in the indicated strains (n = 3). e, Transcript levels of mat3 and cen dg quantified by RT-qPCR in the indicated strains (n = 3-4). f, Transcript levels of lem2 and ade2 quantified by RT-qPCR in the indicated strains (n = 3). For (c-f), data from independent biological replicates were normalized to act1 transcript levels and shown relative to the WT strain (c, d), WT in YES (e) or the WT in EMM (f). Individual replicates are shown in a floating bar plot and the line depicts the median. For (c, d), statistical analysis was performed using one-way ANOVA, and letters denote significant differences between groups with Tukey’s post hoc tests at P < 0.05. For (e, f), blue shadowing indicates minimal media (EMM).
Supplementary information
Supplementary Information
Supplementary Tables 1–4
Supplementary Data 1
GO terms of upregulated transcripts in lem2∆ analyzed by AnGeLi
Supplementary Data 2
GO terms in top clusters containing upregulated transcripts in lem2∆, rrp6∆ and red1 mutants analyzed by AnGeLi
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 2
Unprocessed gels
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 3
Unprocessed Western Blots
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 3
Unprocessed Western Blots
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 7
Unprocessed Western Blots
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Martín Caballero, L., Capella, M., Barrales, R.R. et al. The inner nuclear membrane protein Lem2 coordinates RNA degradation at the nuclear periphery. Nat Struct Mol Biol 29, 910–921 (2022). https://doi.org/10.1038/s41594-022-00831-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00831-6
- Springer Nature America, Inc.
This article is cited by
-
Nuclear mRNA decay: regulatory networks that control gene expression
Nature Reviews Genetics (2024)