Abstract
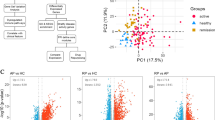
Patients with systemic lupus erythematosus (SLE) display a complex blood transcriptome whose cellular origin is poorly resolved. Using single-cell RNA sequencing, we profiled ~276,000 peripheral blood mononuclear cells from 33 children with SLE with different degrees of disease activity and 11 matched controls. Increased expression of interferon-stimulated genes (ISGs) distinguished cells from children with SLE from healthy control cells. The high ISG expression signature (ISGhi) derived from a small number of transcriptionally defined subpopulations within major cell types, including monocytes, CD4+ and CD8+ T cells, natural killer cells, conventional and plasmacytoid dendritic cells, B cells and especially plasma cells. Expansion of unique subpopulations enriched in ISGs and/or in monogenic lupus-associated genes classified patients with the highest disease activity. Profiling of ~82,000 single peripheral blood mononuclear cells from adults with SLE confirmed the expansion of similar subpopulations in patients with the highest disease activity. This study lays the groundwork for resolving the origin of the SLE transcriptional signatures and the disease heterogeneity towards precision medicine applications.








Similar content being viewed by others
Data availability
scRNA-seq data from child and adult cohorts reported in this study have been deposited in the dbGaP database under accession code phs002048.v1.p1. The processed data can be viewed using an interactive R Shiny App: https://scrnaseq-sle.jax.org/.
Code availability
Jupyter notebooks used for the analysis, including processing, clustering, batch effect correction and subclustering, are available on GitHub: https://github.com/dnehar/SingleCells_SLE_paper.
References
Arbuckle, M. R. et al. Development of autoantibodies before the clinical onset of systemic lupus erythematosus. N. Engl. J. Med. 349, 1526–1533 (2003).
Elkon, K. B. & Wiedeman, A. Type I IFN system in the development and manifestations of SLE. Curr. Opin. Rheumatol. 24, 499–505 (2012).
Tsokos, G. C. Systemic lupus erythematosus. N. Engl. J. Med. 365, 2110–2121 (2011).
Navarra, S. V. et al. Efficacy and safety of belimumab in patients with active systemic lupus erythematosus: a randomised, placebo-controlled, phase 3 trial. Lancet 377, 721–731 (2011).
Baechler, E. C. et al. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc. Natl Acad. Sci. USA 100, 2610–2615 (2003).
Bennett, L. et al. Interferon and granulopoiesis signatures in systemic lupus erythematosus blood. J. Exp. Med. 197, 711–723 (2003).
Chiche, L. et al. Modular transcriptional repertoire analyses of adults with systemic lupus erythematosus reveal distinct type I and type II interferon signatures: modular interferon signatures and systemic lupus erythematosus. Arthritis Rheumatol. 66, 1583–1595 (2014).
Banchereau, R. et al. Personalized immunomonitoring uncovers molecular networks that stratify lupus patients. Cell 165, 551–565 (2016).
Chaussabel, D. et al. A modular analysis framework for blood genomics studies: application to systemic lupus erythematosus. Immunity 29, 150–164 (2008).
Kirou, K. A. et al. Activation of the interferon-α pathway identifies a subgroup of systemic lupus erythematosus patients with distinct serologic features and active disease. Arthritis Rheumatol. 52, 1491–1503 (2005).
Langefeld, C. D. et al. Transancestral map** and genetic load in systemic lupus erythematosus. Nat. Commun. 8, 16021 (2017).
Rice, G. I. et al. Assessment of type I interferon signaling in pediatric inflammatory disease. J. Clin. Immunol. 37, 123–132 (2017).
Omarjee, O. et al. Monogenic lupus: dissecting heterogeneity. Autoimmun. Rev. 18, 102361 (2019).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Schlitzer, A. et al. Identification of cDC1- and cDC2-committed DC progenitors reveals early lineage priming at the common DC progenitor stage in the bone marrow. Nat. Immunol. 16, 718–728 (2015).
Shalek, A. K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Gaublomme, J. T. et al. Single-cell genomics unveils critical regulators of Th17 cell pathogenicity. Cell 163, 1400–1412 (2015).
Li, H. et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 49, 708–718 (2017).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Polański, K. et al. BBKNN: fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Blanco, P., Palucka, A. K., Gill, M., Pascual, V. & Banchereau, J. Induction of dendritic cell differentiation by IFN-α in systemic lupus erythematosus. Science 294, 1540–1543 (2001).
Rodriguez-Pla, A. et al. IFN priming is necessary but not sufficient to turn on a migratory dendritic cell program in lupus monocytes. J. Immunol. 192, 5586–5598 (2014).
Guarda, G. et al. Type I interferon inhibits interleukin-1 production and inflammasome activation. Immunity 34, 213–223 (2011).
Gill, M. A. et al. Blood dendritic cells and DC-poietins in systemic lupus erythematosus. Hum. Immunol. 63, 1172–1180 (2002).
Guiducci, C. et al. TLR recognition of self nucleic acids hampers glucocorticoid activity in lupus. Nature 465, 937–941 (2010).
Sisirak, V. et al. Genetic evidence for the role of plasmacytoid dendritic cells in systemic lupus erythematosus. J. Exp. Med. 211, 1969–1976 (2014).
Farkas, L., Beiske, K., Lund-Johansen, F., Brandtzaeg, P. & Jahnsen, F. L. Plasmacytoid dendritic cells (natural interferon- α/β-producing cells) accumulate in cutaneous lupus erythematosus lesions. Am. J. Pathol. 159, 237–243 (2001).
Tucci, M. et al. Glomerular accumulation of plasmacytoid dendritic cells in active lupus nephritis: role of interleukin-18. Arthritis Rheumatol. 58, 251–262 (2008).
Villani, A.-C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Arce, E. et al. Increased frequency of pre-germinal center B cells and plasma cell precursors in the blood of children with systemic lupus erythematosus. J. Immunol. 167, 2361–2369 (2001).
Jacobi, A. M. et al. Activated memory B cell subsets correlate with disease activity in systemic lupus erythematosus: delineation by expression of CD27, IgD, and CD95. Arthritis Rheumatol. 58, 1762–1773 (2008).
Tipton, C. M. et al. Diversity, cellular origin and autoreactivity of antibody-secreting cell population expansions in acute systemic lupus erythematosus. Nat. Immunol. 16, 755–765 (2015).
Jenks, S. A. et al. Distinct effector B cells induced by unregulated Toll-like receptor 7 contribute to pathogenic responses in systemic lupus erythematosus. Immunity 49, 725–739 (2018).
Wang, S. et al. IL-21 drives expansion and plasma cell differentiation of autoreactive CD11chiT-bet+ B cells in SLE. Nat. Commun. 9, 1758 (2018).
Sanz, I. et al. Challenges and opportunities for consistent classification of human B cell and plasma cell populations. Front. Immunol. 10, 2458 (2019).
Rivero, S. J., Díaz-Jouanen, E. & Alarcón-Segovia, D. Lymphopenia in systemic lupus erythematosus. Clinical, diagnostic, and prognostic significance. Arthritis Rheumatol. 21, 295–305 (1978).
Becker, A. M. et al. SLE peripheral blood B cell, T cell and myeloid cell transcriptomes display unique profiles and each subset contributes to the interferon signature. PLoS ONE 8, e67003 (2013).
Jiang, S. H. et al. Functional rare and low frequency variants in BLK and BANK1 contribute to human lupus. Nat. Commun. 10, 2201 (2019).
Karnell, J. L. et al. Mycophenolic acid differentially impacts B cell function depending on the stage of differentiation. J. Immunol. 187, 3603–3612 (2011).
See, P. et al. Map** the human DC lineage through the integration of high-dimensional techniques. Science 356, eaag3009 (2017).
Gough, D. J., Messina, N. L., Clarke, C. J. P., Johnstone, R. W. & Levy, D. E. Constitutive type I interferon modulates homeostatic balance through tonic signaling. Immunity 36, 166–174 (2012).
Matsui, T. et al. CD2 distinguishes two subsets of human plasmacytoid dendritic cells with distinct phenotype and functions. J. Immunol. 182, 6815–6823 (2009).
Allantaz, F. et al. Blood leukocyte microarrays to diagnose systemic onset juvenile idiopathic arthritis and follow the response to IL-1 blockade. J. Exp. Med. 204, 2131–2144 (2007).
Banchereau, R. et al. Host immune transcriptional profiles reflect the variability in clinical disease manifestations in patients with Staphylococcus aureus infections. PLoS ONE 7, e34390 (2012).
Hong, S. et al. Longitudinal profiling of human blood transcriptome in healthy and lupus pregnancy. J. Exp. Med. 216, 1154–1169 (2019).
Chen, J. et al. PBMC fixation and processing for chromium single-cell RNA sequencing. J. Transl. Med. 16, 198 (2018).
Scharer, C. D. et al. Epigenetic programming underpins B cell dysfunction in human SLE. Nat. Immunol. 20, 1071–1082 (2019).
Arazi, A. et al. The immune cell landscape in kidneys of patients with lupus nephritis. Nat. Immunol. 20, 902–914 (2019).
Gatto, M., Zen, M., Iaccarino, L. & Doria, A. New therapeutic strategies in systemic lupus erythematosus management. Nat. Rev. Rheumatol. 15, 30–48 (2019).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Stoeckius, M. et al. Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Jang, K.-J. et al. Mitochondrial function provides instructive signals for activation-induced B-cell fates. Nat. Commun. 6, 6750 (2015).
Weisberg, S. P. et al. Tissue-resident memory T cells mediate immune homeostasis in the human pancreas through the PD-1/PD-L1 pathway. Cell Rep. 29, 3916–3932 (2019).
Acknowledgements
We are grateful to the patients with SLE, their families, the healthy individuals who participated in our study and the members of the Pediatric Rheumatology Clinics at Texas Scottish Rite Hospital for Children and the Children’s Medical Center in Dallas, TX and M. Sansone, T. Jackson and F. Jawed for help with recruiting adults with SLE. We gratefully acknowledge the contribution of Flow Cytometry, Single Cell Biology and Genome Technologies Scientific Services at The Jackson Laboratory, which are partially supported by NCI P30CA034196, and thank them for expert assistance with the work described in this publication. In addition, we thank JAX Research IT (particularly Sandeep Namburi) for the support with maintaining the R Shiny application. This work was supported by grants NIAMS CORT P50AR070594 Center for Lupus Research (to V.P. and J.F.B.), NIAID NIH U19 AI082715 (to V.P.) and U01 AI131386 (to O.R. and J.F.B.) and start-up funds from the Jackson Laboratory and the Drukier Institute for Children’s Health at Weill Cornell Medicine.
Author information
Authors and Affiliations
Contributions
D.N.-B. processed the data, performed the analyses and wrote the manuscript. S.H. analyzed the data, performed the SC and patient stratification analysis and wrote the manuscript. R.M. handled the samples (with R.J.R., C.-H.H. and R.P.H.) and performed flow experiments in the cSLE and aSLE cohorts. G.C. helped D.N.-B. to analyze the data. M.B. performed some of the cSLE single-cell experiments and preprocessing. P.S. helped interpret the data. W.-F.F. preprocessed some of the data. T.W. selected the patients in the cSLE group. J.B. coordinated the study. L.W. collected the cSLE clinical data. O.R. and A.M. provided and selected the cHD controls. J.-A.T.-G. and N.K. collected the adult lupus samples, under the supervision of S.L. M.A.C. coordinated data acquisition and submission. A.L.L. helped edit the manuscript. J.F.B. and V.P. conceived, designed and supervised the study with O.R., P.R. and K.P. and wrote the manuscript with D.N.-B. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
V.P. has acted as a consultant for Sanofi and Astra Zeneca and is the recipient of a research award from Sanofi. J.F.B. is a member of the BOD and SAB of Neovacs.
Additional information
Peer review information Peer reviewer reports are available. L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 childhood SLE (cSLE) dataset overview and batch effect correction.
a, Number of cells per individual (n = 44). cSLE (n = 33, in purple) and cHD (n = 11; in green). b, Number of genes per individual. cSLE (n = 33, in purple) and cHD (n = 11; in green). c, Number of cells before and after filtration (which includes multiplet removal and other filtration steps that are described in Methods), across the 44 individuals (33 cSLE and 11 cHD). d, Bar plot highlighting the cell abundances across clusters (n = 27) for cSLE and cHD (left panel) and 10X run batches (right panel) before batch effect correction. e,f, Bar plot highlighting the cell abundances across clusters (n = 20) for cSLE and cHD (left panel) and 10X run batches (right panel) after BBKNN (e), or Harmony (f), batch effect correction.
Extended Data Fig. 2 Global information after BBKNN batch correction and ISG scores throughout clusters/subclusters.
a, Bar plot displaying the cell composition of the 20 clusters. b, Bar plot highlighting the individual (n = 44) cell abundances across clusters (n = 20) after BBKNN batch effect correction. c,d, Heatmap representing Pearson correlation between individuals (n = 44; c) or SLEDAI categories (d) based on overall scRNA seq dataset. The hierarchical clustering was based on the first 50 PCs. Red and blue colors, indicate positive and negative correlation, respectively. e,f, ISG scores across the clusters (n = 20; e) or SCs (n = 37; f). Based on the average expression IFN modules M1.2, M3.4, M5.12, ISG scores have been calculated for each cell, across the clusters (e), or SCs (f). Erythrocyte cluster was a negative control.
Extended Data Fig. 3 Individuals’ UMAP for Monocytes and B cells subclusters.
a, Individual UMAP plots from 11 cHD (left), 33 cSLE (right) based on monocyte SCs (Mono-SCs, n = 8). Each color represents a distinct SC. b, Flow cytometry detection of ISG15 in PBMCs gated on CD14 + monocytes from 17 cSLE patients with different SLEDAI scores, as well as 14 cHD. c, Percentage of ISG15 + CD14 + monocytes in cHD and cSLE as categorized based on SLEDAI (cHD cells ‘none’ in gray, SLEDAI < = 4, in yellow; n = 9, and SLEDAI > 4, in red; n = 8). T-test was used for statistical analysis. P-values are shown for the respective comparisons. d, Individual UMAP plots from 11 cHD (left), 33 cSLE (right) based on B cells SCs (B-SCs, n = 7). Each color represents a distinct SC.
Extended Data Fig. 4 Individuals’ UMAP for T and NK cells subclusters.
a, Individual UMAP plots from 11 cHD (left), 33 cSLE (right) based on T cells SCs (T-SCs, n = 6). Each color represents a distinct SC. b, Percentage of GzB + , or Perforin+ CD8 + T cells within cHD and cSLE as categorized based on SLEDAI categories (cHD cells ‘none’ in gray, SLEDAI < = 4 in yellow; n = 9 and SLEDAI > 4, in red; n = 8). (c). Flow cytometry detection of GzB and perforin proteins in PBMCs, gated on CD8 + T cells, from cSLE (n = 17) with different SLEDAI scores and cHD (n = 14). T-test was used for statistical analysis. P-values are shown for the respective comparisons. d, Individual UMAP plots from 11 cHD (left), 33 cSLE (right) based on NK SCs (NK-SCs, n = 4). Each color represents a distinct SC. e, Flow staining of ISG15 on PBMCs gated on CD57 + NK cells from six cSLE patients (in purple) with different SLEDAI scores and nine matched cHD (in green). MFI values are represented. f, Boxplots representing the ESR (left panel), C4 (middle panel) and C3 (right panel) levels across the six cSLE subcluster groups (SCGs) depicted in Fig. 8b. g, Dotplot representing the correlation between hemoglobin (HGB) levels (g/dL) and proportion of cells from the Erythrocyte cluster across the cSLE samples.
Extended Data Fig. 5 Overview of cSLE and aSLE combined (caSLE) dataset.
a, Number of detected genes across cells in cSLE (n = 33), aSLE (n = 8), cHD (n = 11) and aHD (n = 6). Vertical lines represent the mean. b,c, Bar plot highlighting the cell abundances across cluster (n = 26) for cSLE, aSLE, cHD and aHD groups (left panel) and 10X run batches (right panel) before (b) and after (c) BBKNN batch effect correction. Each color represents groups (left) and batch (right). d, UMAP plot representing the 21 clusters across 340,629 PBMCs from cSLE (n = 33), aSLE (n = 8), cHD (n = 11) and aHD (n = 6). Each color represents a distinct cluster. e, Cluster annotation. Dot plot representing expression values of selected genes (x-axis) across each cluster (y-axis). Dot size represents the percentage of cells expressing the marker of interest. Color intensity indicates the mean expression within expressing cells.
Extended Data Fig. 6 caSLE monocytes analysis.
a, UMAP plots representing caSLE monocyte subclusters (Mono-caSCs, n = 6), groups (cSLE, cHD, aSLE or aHD) and SLEDAI categories. b, Bar plot highlighting the cell abundances across Mono-caSCs (n = 6) for cSLE, cHD, aSLE or aHD groups (left panel) and SLEDAI categories (right panel). c, Heatmap representing scaled expression values of the top 10 genes defining each of the Mono-SCs (n = 6). d, Individual UMAP plots from cSLE (n = 33), aSLE (n = 8), 11 cHD (n = 11), or aHD (n = 6), based on Mono-caSCs (n = 6). Each color represents a distinct caSC.
Extended Data Fig. 7 caSLE pDC and cDC cells analysis.
a, UMAP plots representing caSLE pDC subclusters (pDC−caSCs, n = 5), groups (cSLE, cHD, aSLE or aHD), SLEDAI categories and selected genes. b, Bar plot highlighting the cell abundances across pDC-caSCs (n = 5) for cSLE, cHD, aSLE or aHD groups (top panel) and SLEDAI categories (bottom panel). c, Heatmap representing scaled expression values of the top 10 genes defining each of the pDC-SCs (n = 5). d, UMAP plots representing cDC-caSCs (n = 4), groups (cSLE, cHD, aSLE or aHD), SLEDAI categories and selected genes. e, Bar plot highlighting the cell abundances across cDC-caSCs (n = 4) for cSLE, cHD, aSLE or aHD groups (top panel) and SLEDAI categories (bottom panel). f, Heatmap representing scaled expression values of the top 10 genes defining each of the cDC-caSCs (n = 4).
Extended Data Fig. 8 caSLE B cells and plasma cells analysis.
a, UMAP plots representing caSLE B cell subclusters (B-caSCs, n = 8), groups (cSLE, cHD, aSLE or aHD) and SLEDAI categories. b, Bar plot highlighting the cell abundances across B-caSCs (n = 8) of cSLE, cHD, aSLE or aHD groups (left panel) and SLEDAI categories (right panel). c, Heatmap representing scaled expression values of the top 10 genes defining each of the B-caSCs (n = 8). d, Individual UMAP plots from cSLE (n = 33), aSLE (n = 8), 11 cHD (n = 11), or aHD (n = 6), based on B-caSCs (n = 8). Each color represents a distinct caSC. e, UMAP plots representing PC SCs (n = 2), Groups (cSLE, cHD, aSLE or aHD), SLEDAI categories and selected genes. f, Bar plot highlighting the cell abundances across PC SCs (n = 2) for cSLE, cHD, aSLE or aHD groups (left panel) and SLEDAI categories (right panel). g, Heatmap representing scaled expression values of the top 10 genes defining each of the PC-caSC (n = 2).
Extended Data Fig. 9 caSLE T cells analysis.
a, UMAP plots representing caSLE T subclusters (T-caSCs, n = 8), groups (cSLE, cHD, aSLE or aHD), and SLEDAI categories. b, Bar plot highlighting the cell abundances across T-caSCs (n = 8) for cSLE, cHD, aSLE or aHD (Groups; upper panel) and SLEDAI categories (lower panel). c, Individual UMAP plots from cSLE (n = 33), aSLE (n = 8), 11 cHD (n = 11), or aHD (n = 6), based on SCs (n = 8). Each color represents a distinct caSCs. d, Heatmap representing scaled expression values of the top 10 genes defining each of the T-caSC (n = 8).
Extended Data Fig. 10 caSLE NK cells analysis, correlation of cSC and caSC, and boxplots of caSC abundance throughout caSCGs.
a, UMAP plots representing caSLE NK subclusters (NK-caSCs, n = 4), groups (cSLE, cHD, aSLE or aHD) and SLEDAI categories. b, Bar plot highlighting the cell abundances across NK-caSCs (n = 4) for cSLE, cHD, aSLE or aHD (Groups; left panel) and SLEDAI categories (right panel). c, Individual UMAP plots from cSLE (n = 33), aSLE (n = 8), 11 cHD (n = 11), or aHD (n = 6), based on NK-caSCs (n = 4). Each color represents a distinct caSC. d, Heatmap representing scaled expression values of the top 10 genes defining each of the NK-caSC (n = 4). e, Correlation plot of cluster memberships for each single cell in cSLE vs. caSLE datasets. SC membership information from the caSLE combined dataset (excluding aSLE samples) was correlated with cSLE dataset. f, Boxplots representing the proportion of ISGhi Monocytes (Mono-caSC0), ISGhi NK cells (NK-caSC3), ISG + AXL + cDCs(cDC-caSC2), ISGhi pDCs (pDC-caSC3), ISGhi T cells (T-caSC6), CD4 + Memory T cells (T-caSC3), ISGhi PCs (PC-caSC0), ISGhi B cells (B-caSC3), CD8 + Memory T cells (T-caSC5), IL1B + ISGhi Monocytes (Mono-caSC4), DN2 B cells (B-caSC5), NK-SC0 (NK-caSC1) SCs across the four caSCGs. Patients with Mono-caSC0 > 0.08, NK-caSC3 > 0.01, cDC-caSC2 > 0.02, pDC-caSC3 > 0.0005, T-caSC6 > 0.05, PC-caSC0 > 0.002, B-caSC3 > 0.01, T-caSC5 > 0.05, Mono-caSC4 > 0.025, B-caSC5 > 0.01, NK-caSC1 > 0.022 are labelled with their sample names. *, P < 0.05; **, P < 0.01; ***, P < 0.001: ****, P < 0.0001.
Supplementary information
Rights and permissions
About this article
Cite this article
Nehar-Belaid, D., Hong, S., Marches, R. et al. Map** systemic lupus erythematosus heterogeneity at the single-cell level. Nat Immunol 21, 1094–1106 (2020). https://doi.org/10.1038/s41590-020-0743-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0743-0
- Springer Nature America, Inc.
This article is cited by
-
T cell expressions of aberrant gene signatures and Co-inhibitory receptors (Co-IRs) as predictors of renal damage and lupus disease activity
Journal of Biomedical Science (2024)
-
Advancing precision rheumatology through tissue and blood profiling
Nature Reviews Rheumatology (2024)
-
Vascular damage in systemic lupus erythematosus
Nature Reviews Nephrology (2024)
-
Active juvenile systemic lupus erythematosus is associated with distinct NK cell transcriptional and phenotypic alterations
Scientific Reports (2024)
-
Single-cell analysis in rheumatic and allergic diseases: insights for clinical practice
Nature Reviews Immunology (2024)