Abstract
Toxic small alarmone synthetase (toxSAS) enzymes constitute a family of bacterial effectors present in toxin–antitoxin and secretion systems. toxSASs act through either translation inhibition mediated by pyrophosphorylation of transfer RNA (tRNA) CCA ends or synthesis of the toxic alarmone adenosine pentaphosphate ((pp)pApp) and adenosine triphosphate (ATP) depletion, exemplified by FaRel2 and FaRel, respectively. However, structural bases of toxSAS neutralization are missing. Here we show that the pseudo-Zn2+ finger domain (pZFD) of the ATfaRel2 antitoxin precludes access of ATP to the pyrophosphate donor site of the FaRel2 toxin, without affecting recruitment of the tRNA pyrophosphate acceptor. By contrast, (pp)pApp-producing toxSASs are inhibited by Tis1 antitoxin domains though occlusion of the pyrophosphate acceptor-binding site. Consequently, the auxiliary pZFD of AT2faRel is dispensable for FaRel neutralization. Collectively, our study establishes the general principles of toxSAS inhibition by structured antitoxin domains, with the control strategy directly coupled to toxSAS substrate specificity.

Similar content being viewed by others
Main
Small alarmone synthetases (SASs) are a diverse group of monofunctional RelA/SpoT Homolog (RSH) enzymes that catalyze the transfer of the adenosine triphosphate (ATP)-derived pyrophosphate moiety on the 3′ ribose position of the acceptor substrate1. The acceptor substrate is commonly a purine nucleotide, either guanosine triphosphate, diphosphate or monophosphate (GTP, GDP or GMP, yielding guanosine pentaphosphate ((pp)pGpp) alarmone nucleotides)2,3,4 or their adenosine equivalents (ATP, ADP or AMP, yielding (pp)pApp)5,6. SASs also catalyze pyrophosphate transfer to the adenine moiety of the 3′ CCA end of transfer RNAs (tRNAs), yielding pyrophosphorylated tRNA (tRNA-PP)7. As such, the substrate specificity of SASs is broadly determined by their biological function; guanosine-specific SASs are bacterial stress response factors that modulate the intracellular levels of the (pp)pGpp alarmone3,8,9, while adenine-specific toxic SASs (toxSASs) are potent inhibitors of bacterial growth5,7,10,11. The accumulation of toxSAS-produced (pp)pApp abrogates ATP production5,6, while pyrophosphorylation of tRNA CCA abrogates protein synthesis by rendering the modified tRNA aminoacylation incompetent7.
Due to their extreme toxicity, the enzymatic activity of toxSASs is tightly controlled. Acting in trans, dedicated immunity proteins and antitoxins sequester toxSASs into inactive complexes5,6,7,11. Acting in cis, the pseudo-Zn2+ finger domain (pZFD) autoinhibitory antitoxin element of the monomeric fused single-polypeptide toxin–antitoxin system (TA) CapRelSJ46 inhibits its toxic enzymatic domain (toxSYNTH)10. The pZFDCapRel was predicted to block access of ATP, the pyrophosphate donor, through a conserved YXXY sequence motif that switches to a 310-helical conformation when anchored to the toxSYNTH of CapRelSJ46 for neutralization10. Furthermore, pZFDCapRel also acts a phage sensor domain that mediates the activation of the toxin via the recognition of the phage SECΦ27 major capsid protein Gp5710. The proposed disengagement of pZFDCapRel from the toxSYNTH active site enables CapRelSJ46-mediated tRNA pyrophosphorylation, ultimately resulting in translational shutoff, which quarantines the infected cell10.
A different regulatory strategy is used to control Pseudomonas aeruginosa PAO1 Tas1 (type VI secretion effector (p)ppApp synthetase 1), which contains a divergent toxSAS-related domain6. Unlike the tRNA-targeting FaRel2 and CapRelSJ46, Tas1 produces the toxic alarmone (pp)pApp, a potent inhibitor of purine biosynthesis, which depletes the cellular ATP pool6. Tas1 is neutralized by the immunity factor Tis1 (type VI secretion immunity to (p)ppApp synthetase 1). The formation of the Tas1:Tis1 heterodimer complex is proposed to block access of the pyrophosphate acceptor, not donor, ATP nucleotide to the toxSYNTH domain of Tas1 (ref. 6). The toxic TA effector Cellulomonas marina FaRel is a well-characterized (p)ppApp synthetase5. The faRel gene is encoded in an operon with a structure that is highly uncommon for typically bicistronic TAs; it is flanked by two antitoxin genes: aTfaRel, encoding a small alarmone hydrolase (SAH) enzyme that can promiscuously neutralize the toxic products of toxSASs, and aT2faRel, encoding a type II TA-like antitoxin specific to FaRel5.
The Coprobacillus sp. D7 faRel2–aTfaRel2 operon encodes a type II TA system, with the FaRel2 toxSAS toxin being specifically neutralized by the ATfaRel2 antitoxin5 (Supplementary Fig. 1a–c). FaRel2 is a protein synthesis inhibitor that pyrophosphorylates uncharged tRNAs, similarly to CapRelSJ46 (ref. 7). As the ATfaRel2 antitoxin is a distant homolog of pZFDCapRel, it is also likely to prevent the binding of ATP to the pyrophosphate donor site of the toxSYNTH domain10. However, direct structural evidence for the mode of neutralization of either fused (CapRelSJ46) or bipartite (FaRel2:ATfaRel2) translation-targeting toxSAS TAs is lacking5,7.
Here we uncover how tRNA-modifying toxSAS are neutralized. We determine the crystal structure of the individual FaRel2 toxin and ATfaRel2 antitoxin, as well as the heterotetrameric FaRel22:ATfaRel22 complex. Our structural, microbiological and biochemical evidence suggests that bipartite toxSAS TA operons use stable multimerization as an alternative to the colocalization of toxin and antitoxin domains of monomeric fused TAs, such as CapRelSJ46, to ensure efficient toxin inhibition. We directly demonstrate that ATfaRel2 neutralizes FaRel2 by blocking access of ATP to the pyrophosphate donor site of toxSYNTH. Lastly, we propose a unifying conceptual framework that connects the mechanisms of allosteric control and substrate specificity in toxSASs.
Results
ATfaRel2 is a compact, well-structured antitoxin
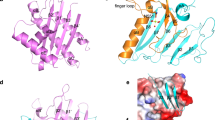
We determined the crystal structure of ATfaRel2 (residues 1–73) to 1.2-Å resolution (Fig. 1a,b and Supplementary Table 1). The antitoxin protein is monomeric in the crystal (Supplementary Fig. 2a), in good agreement with molecular weight estimates by size-exclusion chromatography (SEC) (Supplementary Fig. 2b and Supplementary Table 2). At the core of its compact, well-folded structure is an antiparallel β-sheet (β-strands β1–β3), with β2 and β3 connected by the central α-helix α1. The C-terminal extension provides an additional β-strand to the β-sheet, β4, which folds parallel to β2, as well as a second α-helix, α2, which forms a part of the protein’s hydrophobic core (Fig. 1b). Despite the lack of sequence similarity, ATfaRel2 is structurally similar to pZFDCapRel, with the two proteins superimposing with a root-mean-square deviation (r.m.s.d.) of 2.8 Å, and they both contain conserved tyrosine residues at the C terminus (Fig. 1c).
a, Cartoon representation of Coprobacillus sp. D7 ATfaRel2 structure. The pZFD core is colored light blue, the C-terminal extension elements are colored salmon and the 51YXXY54 motif is highlighted in dark blue. b, Topology representation of ATfaRel2, colored as in a. c, Superposition of the experimental ATfaRel2 structure (colored as in a) onto the AlphaFold2-predicted structure of the CapRelSJ46 antitoxin domain in the neutralizing state. pZFDCapRel is colored light green and the N-terminal extension that is part of Anchor1 is colored ocher, with the conformational switch region that contains the YXXY neutralization motif colored dark green and labeled in the figure. d, Topology representation of the CapRelSJ46 antitoxin in the neutralizing state, highlighting the circular permutation of ATfaRel2. e, Toxicity neutralization assays probing the 51YXXY54 motif of ATfaRel2 and its scaffolding structure from β1. Serial dilutions of E. coli strains expressing WT FaRel2 alone or coexpressed with ATfaRel2 (WT or V43A, I45A, A50M, Y51A and Y54A variants) were plated on solid LB medium and scored after 16 h at 37 °C. f,g, Binding of WT (f) and Y54A-substituted (g) ATfaRel2 to Y128F-substituted catalytically compromised FaRel2, monitored by ITC.
Interestingly, the predicted topologies of pZFDCapRel in the neutralizing state and ATfaRel2 are similar except for the order of structural elements at the termini. The C-terminal β/α extension of ATfaRel2 is structurally similar to the N-terminal β/α region of pZFDCapRel that connects the antitoxin with toxSYNTHCapRel via Anchor1 (Fig. 1c and Supplementary Fig. 2c–e). This is suggestive of circular permutation (compare Fig. 1b to Fig. 1d). However, as their similarities extend only to the topology level, these two small β/α elements may have evolved independently.
Changes in topology in the antitoxin elements likely reflect the in cis versus in trans neutralization of different toxSASs. Presumably, ATfaRel2 fully dissociates from FaRel2 when the toxin becomes active. Conversely, in the fused CapRels, dissociation of pZFDCapRel is not possible and activation is believed to be mediated by a conformational change10 (Supplementary Fig. 2d,e). Despite these topological variations, both domains retain strong structural similarities, including the YXXY sequence motif linked to toxSYNTH inhibition in CapRelSJ46. This suggests that ATfaRel2 and pZFDCapRel neutralize toxSYNTH domains by a common mechanism.
The YXXY motif is crucial for toxSYNTH inhibition
In CapRelSJ46, the conserved YXXY motif is located in the switch region of the pZFD and is predicted to lock the enzyme in a catalytically inactive state with both Y residues blocking the donor pyrophosphate ATP-binding site of toxSYNTH (Supplementary Fig. 2c–e). Substitutions directly or indirectly targeting this motif render CapRelSJ46 constitutively active10. In the crystal structure of the catalytically active state of CapRelSJ46, the YXXY motif assumes an extended conformation10. Conversely, in the AlphaFold2-generated neutralized state model, the YXXY motif is predicted to fold into a 310-helix that locks into the donor site (Supplementary Fig. 2d,e).
The structure of the free ATfaRel2 reveals that the 51YXXY54 motif indeed folds into a 310-helix structure that was predicted to be key for toxSAS neutralization, scaffolded by β-strands β1 and β3 (Fig. 1a and Supplementary Fig. 1b). This suggests that, even in the absence of the toxin, ATfaRel2 is primed for efficient FaRel2 neutralization. Guided by the structural similarities between ATfaRel2 and pZFDCapRel, we probed the role of the individual residues of the 51YXXY54 motif in the neutralization of FaRel2 toxicity in vivo (Fig. 1e). While Y54A substitution fully ablates the neutralizing activity of ATfaRel2, I45A and Y51A result in modest but clearly detectable defects in FaRel2 neutralization. Our isothermal titration calorimetry (ITC) assays lend further support to the in vivo neutralization results. To overcome the challenges of producing an otherwise highly toxic FaRel2, we followed a well-established substitution strategy for RSH enzymes10,12,13 and we used a catalytically impaired Y128F-substituted variant, FaRel2-Y128F. The substitution interferes with the accommodation of the acceptor nucleotide in the active site but is located far from the predicted pZFDCapRel–toxSYNTH interface. We characterized complex formation between FaRel2-Y128F and wild-type (WT) ATfaRel2 or variants carrying substitutions in the 51YXXY54 motif (Fig. 1f,g and Supplementary Table 3). V43A and A50M substitutions had a negligible effect on complex stability. These mutant antitoxins bound FaRel2-Y128F with KD values of 78.8 nM and 38.0 nM, respectively, compared with a KD value of 35.2 nM in the case of WT ATfaRel2. The destabilizing effect was more pronounced in the case of I45A and Y51A variants. While these substitutions resulted in 25-fold and 43-fold reductions in affinity, respectively, the remaining affinity was still sufficient for partial neutralization in vivo (Fig. 1e). Lastly, the Y54A variant, which was unable to neutralize FaRel2 in vivo, had a 250-fold lower affinity to the toxin than WT ATfaRel2.
FaRel2 binds ATP in a partially folded state
Next, we determined the structure of the catalytically impaired FaRel2-Y128F bound to the non-hydrolyzable ATP analog adenosine-5′-[(α,β)-methyleno]triphosphate (APCPP) at 2.6-Å resolution (Fig. 2a). The toxSYNTH domain of FaRel2 shares the overall topology of other nucleotide pyrophophotransferases, retaining a presumably ancestral fold composed of a central five-stranded β-sheet framed by two α-helices, α3 and α6 (refs. 13,14) (Fig. 2b). The core toxSYNTH domain is very similar to that of the (pp)pApp alarmone synthetase Tas1 (Protein Data Bank (PDB) 6OX6) and the tRNA-pyrophosphokinase CapRelSJ46 (PDB 7ZTB), superimposing with r.m.s.d. values of 1.0 Å and 0.8 Å, respectively (Supplementary Fig. 3a,b).
a, The structure of Coprobacillus sp. D7 FaRel2 in complex with APCPP bound to the pyrophosphate donor site. The adenine base is coordinated by conserved residues R64, R95 and Q147, as well as residue G93, while E145 provides the specificity for adenosine nucleotides. R64 provides additional van der Waals contacts with the ribose moiety. The 5′ triphosphate moiety is stabilized by R64, K66, S70, K74 and R77, while the Mg2+ ion is held in place by the catalytic D90. b, Topology diagram of FaRel2, highlighting the N-terminal region involved in tRNA recognition (yellow), the toxSYNTH domain (dark blue, with the catalytic region outlined by a dashed line) and the disordered C-terminal region (dark magenta). The acceptor site is highlighted by the G-loop (β3–β4 loop) in red and the donor site involving β1, β2 and α3 is shaded in pink. c, Superposition of the FaRel2:APCPP complex on the structure of S. aureus RelP in complex with APCPP (PDB 6EWZ; turquoise). The ordered N terminus of FaRel2 that is lacking in RelP is highlighted in yellow. d, Superposition of the complexes of FaRel2:APCPP (dark blue), RelP:APCPP (turquoise) and Thermus thermophilus RelNTD:AMP (dark orange; PDB 6S2U) illustrates the conserved mode of pyrophosphage donor coordination in RSH pyrophosphotransferases.
The catalytic core of RSH enzymes is typically decorated by regulatory elements that exert allosteric control on the enzymatic domains12,15. The FaRel2 toxSYNTH domain has a well-resolved N-terminal α-helical extension comprising helices α1 and α2, with residues K28 and R29 from the α2–α3 loop being crucial for tRNA binding7 (Supplementary Fig. 1c). While the N terminus is unresolved (Fig. 2c) in the non-toxic (p)ppGpp alarmone synthetases RelP15,16 and RelQ17, the N terminus folds back toward the toxSYNTH core in FaRel2 and intercalates α1 and α2 between α3 and α5, providing an anchor point for tRNAs near the active site G-loop (Fig. 2a,c). The C-terminal region of SYNTH and toxSYNTH domains of pyrophosphotransferases have varied architectures composed predominantly of small α-helical domains6,10,13,14,18,19; the C-terminal bundle of four α-helices of SASs RelQ and RelP, which synthesizes (p)ppGpp, acts as an oligomerization interface15,17, toxSYNTH of the monomeric Tas1 is followed by a small α-helical domain6 and the SYNTH domain of long RHSs is followed by a flexible core domain with a high α-helical propensity18. In the case of FaRel2 complexed with APCPP, the C terminus is disordered and not visible in the electron density (Fig. 2d).
Despite these differences, the FaRel2-bound APCPP superimposes remarkably well with the APCPP bound in the donor site of other (p)ppGpp-synthesizing RSHs SAS RelP and RelQ, as well as the long RSH Rel13,15,17 (Fig. 2d). As in the other alarmone synthetases, the adenosine group stacks with the conserved R64 and R95 from β1 and β2, with β5 E145 hydrogen bonding the adenosine NH2 group (Supplementary Fig. 1c). The R64 residue has a crucial role in providing van der Waals contacts to accommodate the ribose while directly coordinating the 5′ α and β phosphates together with K66. The strongly basic α4 that follows β1 further stabilizes the triphosphates through S70, K74 and R77 (Fig. 2a). At the pyrophosphate acceptor side of the active site, the conformation of the G-loop and the orientation of the base-coordinating F128 (Y128 in the WT FaRel2) deviate from what was observed in precatalytic and postcatalytic complexes of (p)ppGpp synthetases15,17 (Supplementary Fig. 3c,d). It is tempting to speculate that, while the ground state of FaRel2 is primed to bind ATP, efficient tRNA binding would likely involve further conformational rearrangements, such as folding of the C terminus and alignment of the active site.
ATfaRel2 binds FaRel2 by β-sheet extension
To uncover the mechanism of toxin neutralization, we determined the structure of the ATfaRel2:FaRel2 complex. The structure reveals a ATfaRel22:FaRel22 heterotetrametric arrangement (Fig. 3a–c). The 2:2 stoichiometry was confirmed in solution by SEC (62 kDa versus the theoretical 68 kDa; Fig. 3d) and was consistent with the ITC measurements (Fig. 1f).
a, Structure of the heterotetrametic ATfaRel22:FaRel22 complex. Antitoxin units are colored light blue and pink, while the toxin units are colored dark blue and gold. The C-terminal region of FaRel2, disordered in the FaRel2:APCPP structure, in the TA complex folds into a small α-helical subdomain. b, Stabilization of the ATfaRel2:FaRel2 heterodimer by the primary complex interface. The ATP-binding catalytic sites have key structural roles in the interface. c, The secondary interface ATfaRel2:FaRel2 is formed at the two-fold symmetry axis of the complex. The interface comprises (1) cross-coordination of R47 from each antitoxin unit; (2) formation of an alternative hydrophobic interface between ATfaRel2 and FaRel2 (that is, different from the YXXY motif); and (3) interactions between the C-terminal α-helical regions of the two FaRel2 units. d, Analytical SEC of ATfaRel22:FaRel2-Y128F2 (dark-blue trace), ATfaRel2-R47A2:FaRel2-Y128F2 (lilac), FaRel2-Y128F (yellow) and ATfaRel2 (red). e, Probing of the secondary FaRel2:ATfaRel2 interface through toxicity neutralization assays. Serial dilutions of E. coli strains expressing FaRel2 alone or coexpressed with ATfaRel2 (WT or R47A, Y57A and Y59A variants) were plated on solid LB medium and scored after 16 h at 37 °C. f,g, Binding of R47A-substituted (f) and F59A-substituted (g) ATfaRel2 to Y128F-substituted FaRel2, monitored by ITC.
The C-terminal region of FaRel2, disordered in the FaRel2:APCPP complex, folds into an α-helical subdomain upon complex formation and provides a dimerization interface mediated only by toxin–toxin interactions. By contrast, in the bound state, the conformation of ATfaRel2 matches that of the free antitoxin, with both structures superimposing with an r.m.s.d. of 0.4 Å (Supplementary Fig. 3e). The primary interface between ATfaRel2 and FaRel2 has an area of 1,190.0 Å2, with ATfaRel2 sterically blocking access of the ATP substrate to the pyrophosphate donor binding site of the toxSYNTH domain (Fig. 3b). Through this large interface, ATfaRel2 contacts several key functional regions of FaRel2: (1) the long N-terminal α-helix α3 (a structural element that is often involved in allosteric crosstalk in many RSH enzymes and interacts with α1 from ATfaRel2); (2) the basic α-helix α4 (involved in the stabilization of the ATP triphosphate group, which is coordinated by ATfaRel2 through β3 and the β3–α1 loop); (3) the central β-sheet (through β1, β2 and β5, which harbor the catalytic center and adenine coordinating residues); and (4) the C-terminal cap of the α-helix α6 (which is disordered in the FaRel2:APCPP complex and folds into an α-helical dimerization region when bound to ATfaRel2). The only notable exception was the predicted tRNA recognition site that remains solvent accessible (Supplementary Fig. 3f).
The center of the neutralization interface is formed by the antiparallel β-strand interaction between FaRel2 β1 and ATfaRel2 β3 that connects the β-sheets of both proteins, extending the core of the complex. Residues I42–T46 of β3 form multiple van der Waals contacts with FaRel2 β1, further stabilizing the complex. Residues V43 and I45 serve as a scaffold, orienting the YXXY 310-helix that anchors ATfaRel2 (Fig. 3b). As predicted for pZFDCapRel (ref. 10), this hydrophobic tether projects Y51 and Y54 into the ATP-binding site through a π-stacking arrangement with R64 and R95, which precludes adenine coordination to the pyrophosphate donor site of FaRel2. These results suggest that, while this mechanism of neutralization is likely the same at the structural level to that proposed for CapRelSJ46, the energetics of neutralization are certainly different, with the dynamic association of pZFDCapRel regulating toxSYNTHCapRel in cis, contrasting with the stable in trans neutralization in the ATfaRel2:FaRel2 complex. These differences could have important implications for triggering these systems.
Dimerization enhances toxin neutralization by ATfaRel2
Compared with pZFDCapRel, ATfaRel2 is considerably more tolerant to substitutions in the toxin-binding interface (compare Fig. 1e with Extended Data Fig. 3j in ref. 10). The structure of the ATfaRel22:FaRel22 complex reveals that ATfaRel2 engages the neighboring FaRel2 in the heterotetramer through a secondary interface that is half the size of the primary one (~550.0 Å2 versus 1,190.0 Å2), thus effectively crosslinking the complex (Supplementary Fig. 4a–c). We hypothesized that the stable oligomeric nature of ATfaRel22:FaRel22 compensates for the lack of TA colocalization enforced in the monomeric CapRelSJ46 through fusion of the toxin and antitoxin domains into one polypeptide. On this basis, substitutions disrupting the ATfaRel22:FaRel22 oligomerization interface (and not affecting the primary neutralization interface at the active site) would compromise the TA recognition.
To test this hypothesis, we subjected the secondary interface to single-residue substitutions and assessed the complex stability in vivo through toxicity neutralization assays (Fig. 3e). The three targeted residues R47A, Y57A and F59A of ATfaRel2 are all distant from the main contact interface that blocks access of ATP to the active site (Fig. 3c). R47 is located on the two-fold symmetry axis of the complex and the side chains of R47 from each ATfaRel2 interlock through π–π interactions. Y57 and F59 are part of a small hydrophobic core that defines the secondary oligomerization interface. The R47A substitution resulted in a modest defect, whereas Y57A and F59A compromised the neutralization severely (Fig. 3d).
The direct interrogation of these interactions by ITC was in good agreement with the in vivo data. The R47A substitution efficiently perturbs the secondary interface and decouples the highly cooperative tetramer formation observed in the WT protein (Fig. 3f and Supplementary Table 3). The first high-affinity binding event (KD = 35 nM with a stoichiometry of 0.4) followed a lower-affinity recognition event (KD = 750 nM with a stoichiometry of 0.8). This likely represents the initial neutralization of FaRel2 (with a 2:1 TA ratio) followed by the formation of a less stable 2:2 tetramer. The impact of F59A on the affinity was even stronger, with a 60-fold decrease in affinity and a confirmed 1:1 binding molar ratio, indicating an interaction mediated by only the primary interface (Fig. 3g and Supplementary Table 3). These results were consistent with SEC experiments that revealed a decrease in size of the TA complex from the estimated 70.8 kDa of the WT (consistent with a 2:2 A:T stoichiometry) to 51.3 kDa suggestive of a ATfaRel2-R47A:FaRel2-Y128F2 complex (1:2 A:T stoichiometry) (Fig. 3d and Supplementary Table 2). The observation of a stable ATfaRel2-R47A:FaRel2-Y128F2 complex in SEC matched the high-affinity interaction observed by ITC with ATfaRel2-R47A (Supplementary Table 2). It is, thus, likely that the C-terminal region of FaRel2 that folds upon TA complex formation and provides a large FaRel2:FaRel2 interface in the complex is still capable of partially stabilizing the oligomer against the effect of mild substitutions such as R47A but not against F59A, which had a major effect on complex formation.
FaRel2 C terminus stabilizes the heterotetrameric complex
Upon formation of the ATfaRel22:FaRel22 complex, the C-terminal regions of the two individual FaRel2 toxin polypeptides fold into a dimerization region with four α-helices that contributes 670 Å2 to the TA interface (Supplementary Fig. 4d). This folding upon binding interaction likely has an important role in the overall stability of the heterotetramer. Guided by the structure of FaRel2 bound to APCPP, we constructed a truncated version of FaRel2 (FaRel2-Δ166–206) lacking the C-terminal disordered part. FaRel2-Δ166–206 interacted with ATfaRel2 with a KD of 3.3 μM, as measured by ITC (Supplementary Fig. 4e). This ~90-fold drop in affinity of FaRel2-Δ166–206 for ATfaRel2 underscores the strong contribution of oligomerization to the overall energetics of complex formation. In the case of the WT TA complex, the binding was both entropically and enthalpically driven. The entropic penalty from the folding of the FaRel2 C terminus was likely compensated for by the configurational entropy associated with the large hydrophobic surface buried upon binding (Supplementary Table 3). Together with the strong enthalpic component that accompanied the oligomerization through the C-terminal α-helical region, this resulted in very stable heterotetramerization. Because the FaRel2-Δ166–206 truncation removes the enthalpic contribution of the C terminus, binding to ATfaRel2 was, as expected, predominantly entropically driven, resulting in a less stable complex (Supplementary Table 3). Collectively, these results suggest that, while the main TA interface drives toxin neutralization, the oligomerization further stabilizes the interaction between ATfaRel2 and the toxSYNTH domain of FaRel2. Additional contacts of the main interface YXXY motif with the folded C-terminal α-helical FaRel2:FaRel2 interface link the oligomerization with toxin neutralization, hinting at a potential allosteric path for activation and toxin release.
tRNA-pyrophosphorylating toxins specifically bind tRNA
Given the low concentrations toxSAS toxins are typically found in the cell (below the detection levels of current techniques)20,21, tRNA-phosphorylating activity would likely depend on a strong and specific association with tRNAs. We used ITC to examine the tRNA binding capacity of a representative toxSAS and housekee** SASs: tRNA-phosphorylating Coprobacillus sp. D7 FaRel2 and Mycobacterium phage Phrann PhRel, (pp)pApp-synthesizing C. marina FaRel and (p)ppGpp-synthesizing Staphylococcus aureus SAS RelQ. FaRel2 and PhRel bound deacylated initiator tRNAifMet with similar affinities (KD values of 483 nM and 825 nM, respectively; Fig. 4a,b and Supplementary Table 3). (pp)pApp synthetase FaRel had a 37-fold lower affinity to tRNAifMet (KD value of 17.8 µM). No tRNAifMet binding was observed for (p)ppGpp-producing Enterococcus faecalis SAS RelQ (Fig. 4c,d and Supplementary Table 3). Lastly, E. faecalis RelQ was shown to interact with a short single-stranded model mRNA(MF) coding for MF dipeptide22. Our ITC experiments demonstrated that S. aureus RelQ similarly bound mRNA(MF) with submicromolar affinity (KD value of 922 nM) (Fig. 4e and Supplementary Table 3).
a–d, Binding of FaRel2-Y128F (a), PhRel2-Y143F (b), FaRel-Y175F (c) and WT RelQSa (d) to deacylated initiator tRNAifMet, monitored by ITC. e, Binding of mRNA(MF) to WT RelQSa, monitored by ITC. f,g, Binding of APCPP to FaRel2-Y128F (f) and ATfaRel22:FaRel2-Y128F2 complex (g), monitored by ITC. h, Binding of tRNAifMet to the ATfaRel22:FaRel2-Y128F2 complex, monitored by ITC. i, Structure of ATfaRel22:FaRel2-Y128F2 bound to APCPP (green). The unbiased mFo-DFc electron density map corresponding to the bound APCPP is shown in gray. j, Details of the coordination of APCPP (green) in the acceptor site when bound to ATfaRel22:FaRel2-Y128F2. The adenosine base is coordinated by the G-loop F128 and the 5′ β and γ phosphates extend to the basic patch of α4. There, they bind in a reverse orientation compared with APCPP in the FaRel2:APCPP complex, underscoring that the nucleotide is bound in a state incompatible with phosphate transfer. For comparison, the APCPP in the orientation observed in the complex with FaRel2 is shown in light yellow. Active site residues of FaRel2 are labeled in black and the residues from the YXXY motif of ATfaRel2 are labeled in light blue. k, FaRel2-Y128F:APCPP complex with APCPP placed in the donor site in a catalytically compatible orientation, presented in the same pose as i. In the absence of ATfaRel2, α7 and α8 are not visible in the electron density.
ATfaRel2 interferes with APCPP binding but not tRNA recognition
While association with ATfaRel2 decreased the affinity (KD) to APCPP ~20-fold, from 2.1 to 41.3 μM (Fig. 4f,g and Supplementary Table 3), the affinity to deacylated tRNAifMet was virtually the same for the free toxin and inactive TA complex (KD value of 483 nM versus 460 nM) (Fig. 4h and Supplementary Table 3). In the cases of both free monomeric FaRel2 and the heterotetrametric ATfaRel2:FaRel2 complex, the tRNA binding had 1:1 stoichiometry with respect to FaRel2.
To further investigate the effect of AtfaRel2 binding on the interaction of FaRel2 with ATP, we determined the structure of the ATfaRel2:FaRel2 complex bound to APCPP (Fig. 4i). The high protein concentration intrinsic of the crystal lattice combined with high nucleotide concentration used for soaking facilitated the binding of APCPP to a partially blocked active site. As predicted from the structure of ATfaRel2:FaRel2, the coordination sites for the adenine, ribose and α phosphate groups at the pyrophosphate donor site are blocked by ATfaRel2. Thus, APCPP is bound in the pyrophosphate acceptor site in a conformation incompatible with pyrophosphate transfer (Fig. 4j). The adenine base is coordinated by Y128 and R95, resembling the expected coordination of the terminal adenine of the CCA tRNA moiety. The β and γ phosphates further anchor the nucleotide; however, they are observed in a reversed orientation compared with the FaRel2:APCPP complex (Fig. 4j–k).
It is instructive to compare the local charge distributions in the active sites of tRNA-modifying FaRel2 to those of (pp)pGpp and (p)ppApp alarmone synthetases Rel13, RelP15 and Tas1 (ref. 6). All alarmone synthetases have a large positive patch that accommodates diphosphate and triphosphate nucleotide substrates, located on the acceptor site close to the conserved Y residue that interacts with the acceptor base (Supplementary Fig. 4f–h). This positive patch is considerably smaller in FaRel2 (Supplementary Fig. 4i), which explains the misorientation of the β and γ phosphates of APCPP bound in the acceptor site of FaRel2 and the lack of alarmone synthetase activity of the tRNA-targeting toxSAS.
Collectively, our results demonstrate that FaRel2 is neutralized by the ATfaRel2 antitoxin by compromising the accommodation of ATP in the toxSYNTH active site without affecting the interaction with uncharged tRNAs. This suggests that the ATfaRel22:FaRel22:tRNA2 ternary complex could be preformed in the cell, with the toxSAS neutralized in the complex until activation is triggered.
toxSAS neutralization is defined by catalytic activity
Prompted by the conceptual differences between Tis1-mediated neutralization of Tas1 and ATfaRel2-mediated neutralization of FaRel2, we next used AlphaFold223 to explore the general principles underlying the mechanisms of toxin neutralization across known toxSAS functional diversity (Fig. 5 and Supplementary Fig. 5a–l). On the toxSAS toxin side, AlphaFold2 predicts a strong conservation of a core toxSYNTH fold decorated with a variety of insertions at the N and C termini (Fig. 5a–e). On the antitoxin side, the pZFD fold is found as either a standalone neutralizing domain or part of multidomain antitoxins combined with either Tis1 or Tis1-like (Fig. 5f–h) or PanA domains11,24 (Supplementary Fig. 5a–j).
a–f, AlphaFold2-generated structural models of Coprobacillus sp. D7 FaRel2 (a), M. tuberculosis CapRel (unfused) (b), M. tuberculosis PhRel (c), bacteriophage Lily PhRel2 (d) and C. marina FaRel (e) and the AT2faRel:FaRel complex (f). The Tis1-like NTD of AT2faRel is colored dark green and the pZFD/ATfaRel2-like CTD is colored light cyan. AlphaFold2 predicts that FaRel is neutralized by the Tis1-like domain through the pyrophosphate acceptor site. g,h, Structural superposition of the Tis1-like NTD of AT2faRel (dark green) on Tis1 (PDB 6OX6), colored brown (g) and the CTD (light cyan) on pZFDCapRel (PDB 7ZTB), colored light teal (h). i, Probing of the secondary FaRel2:ATfaRel2 interface through toxicity neutralization assays. Serial dilutions of E. coli strains expressing either AT2faRelNTD or AT2faRelCTD alone or together with FaRel were plated on solid LB medium and scored after 16 h at 37 °C.
Structural predictions of the different neutralized complexes uncovered a general trend. Translation-targeting tRNA-pyrophosphorylating toxSASs such as fused and split CapRel, FaRel2, PhRel and PhRel2 are inhibited through the pyrophosphate donor site (Supplementary Fig. 5a–j). Conversely, metabolism-targeting (pp)pApp-producing toxSASs such as FaRel and Tas1 are neutralized through the pyrophosphate acceptor site (Fig. 5f and Supplementary Fig. 5k,l). The generality of this observation holds even in cases of multidomain antitoxins. In the case of the translation-targeting PhRel2 of bacteriophage Lily and Bacillus subtilis Ia1a. These toxins are neutralized by multidomain antitoxins that contain a pZFD fold subtype PAD1 (panacea-associated domain 1)11. However, ATphRel2 antitoxins neutralize the toxins analogously to the pZFD-mediated neutralization of CapRel and FaRel2 (Supplementary Fig. 5f,g)24. By contrast, the (pp)pApp-producing FaRel is inhibited by the Tis1-like domain of AT2faRel in a manner analogous to Tis1-mediated inhibition of Tas1 (Fig. 5f and Supplementary Fig. 5k,l).
We validated the structural predictions of Alphafold2 through mutagenesis and toxicity neutralization assays. In the case of PhRel2, as we showed previously24, the isolated PAD1 domain of B. subtilis Ia1a ATphRel2 is sufficient to neutralize the toxin. In the case of FaRel:AT2faRel, the Tis1-like N-terminal domain (NTD) of AT2faRel (Fig. 5g) is sufficient to neutralize the (pp)pApp synthetase FaRel, while the pZFD C-terminal domain (CTD; Fig. 5h) has no neutralizing activity (Fig. 5i). Interestingly, this loss of neutralizing activity by the CTD of AT2faRel is accompanied by the loss of the YXXY recognition motif of the pZFD (Fig. 5f). Collectively, our results suggest a coupling between the substrate specificity of toxSASs and the mechanism of neutralization.
Discussion
Our current mechanistic understanding of toxSYNTH inhibition by immunity proteins and antitoxins is largely based on two landmark studies. The first study involves the structure of the neutralized complex between a monomeric (p)ppApp-producing toxSAS Tas1 and its immunity protein Tis1 (ref. 6). Ahmad and colleagues proposed that the enzymatic activity of Tas1 was suppressed by Tis1 through distortion of the acceptor nucleotide-binding site of toxSYNTH. The second study involves the structure of the tRNA-pyrophosphorylating toxSAS CapRelSJ46 (ref. 10), a fused monocistronic TA, with the antitoxin part comprising two anchor regions and a pZFD. In this structure, CapRelSJ46 appears in a catalytically competent state (that is not autoinhibited by the pZFD), as the antitoxin domain was in a conformation compatible with the enzymatic activity of toxSYNTH. Further exploration of the structural dynamics of CapRelSJ46 with AlphaFold2 (ref. 23) predicted a possible mechanism of autoinhibition mediated by the pZFD blocking the donor nucleotide-binding site of toxSYNTH (Fig. 6a,b).
Metabolism-targeting and translation-targeting toxSAS are inhibited by different strategies. a, Enzymatic activity of metabolism-targeting toxSASs is suppressed by inhibition of the recruitment of the pyrophosphate acceptor substrate. b, Translation-targeting toxSASs are neutralized by inhibition of the recruitment of the pyrophosphate donor, while binding of the pyrophosphate acceptor substrate, tRNA, is permitted. This strategy is analogous to that used for the regulation of long (p)ppGpp-producing RSHs Rel, RelA and SpoT; while the affinity to the GTP or GDP substrate nucleotide is constitutive, binding of the ATP nucleotide to the pyrophosphate donor site is under strict allosteric control.
Our study provides detailed mechanistic understanding of toxSAS regulation and allows generalizing the observations previously made with CapRelSJ46 and Tas1:Tis1 to the whole toxSAS superfamily. We put forward a model that couples toxSAS substrate specificity to the inhibition strategies used by antitoxins and immunity proteins (Fig. 6a,b). Type II TA modules are notoriously evolutionarily promiscuous, with members of the same toxin family being neutralized by unrelated antitoxins through different mechanisms24,25,26. toxSAS TAs display a strikingly clear-cut dichotomy of neutralization mechanisms. Metabolism-targeting (pp)pApp synthetases such as Tas1 and FaRel are inhibited by Tis1-like antitoxins through the occlusion of the pyrophosphate acceptor nucleotide-binding site6 (Fig. 6a). Conversely, translation-targeting tRNA pyrophosphotransferases such as FaRel2 and CapRel are inhibited by pZFD interfering with the binding site for pyrophosphate donor ATP (Fig. 6b), which echoes the regulatory strategy used in the autoinhibition of multidomain ‘long’ (p)ppGpp-synthesizing housekee** RSHs12,27. While the antitoxins and immunity proteins mediating the two neutralization strategies have diverged in sequence, the neutralizing elements display strong structural conservation.
Type II TA antitoxins often rely on unstructured elements and disordered domains that fold upon binding for toxin neutralization25,28,29,30,31,32,33,34. In many cases, the structural plasticity of the disordered region of antitoxins allows them to couple toxin neutralization with transcriptional autoregulation29,35,36 to balance the cellular T:A ratio. Interestingly, this feature of TA regulation seems so prevalent that, in the exception provided by the GraTA system (from the RelBE superfamily), which contains a well-folded and globular antitoxin, the disordered region leaps to the toxin GraT retaining a role as a crucial regulatory element37. The structurally defined lock-and-key neutralization specificity of toxSASs is, thus, uncommon and may underlie their ‘switch’ nature. In this sense, it is instructive to compare the structural–energetic interplay of the fused CapRelSJ46 TA with the bipartite ATfaRel2:FaRel2 TA. The forced colocalization of the toxin and antitoxin elements of CapRelSJ46 is compatible with a conformationally dynamic enzyme and offsets the entropic penalty associated with the antitoxin assuming a compact and structured neutralizing state (Supplementary Fig. 6a). These intrinsic dynamics facilitate the formation of the CapRelSJ46:Gp57 complex that triggers the enzyme. By contrast, in the bipartite ATfaRel2:FaRel2 TA system, the free antitoxin naturally assumes the optimal conformation for neutralization. The entropic penalty associated with FaRel2 folding is compensated by the formation of the heterotetramer and a tight TA complex. Thus, the loss of colocalization is compensated by the stabilizing effect of oligomerization. Our findings suggest that offsetting this oligomeric structure could be the key to triggering these bipartite systems (Supplementary Fig. 6b).
ZFDs perform various molecular recognition functions38 facilitated by a marked structural plasticity39. Structurally related pZFDs mediate both phage recognition and toxin autoinhibition in fused tRNA-pyrophosphorylating TA CapRelSJ46 (ref. 10). PAD1 has the same fold as pZFD and directly mediates the neutralization of B. subtilis Ia1a and Clostridium hylemonae the tRNA-pyrophosphorylating PhRel2 toxSAS11,24. Conversely, the pZFD-like CTD of AT2faRel is not essential for the inhibition of the (pp)pApp synthetase FaRel, which is neutralized by its Tis1-like NTD (Fig. 5c,b). This suggests that the pZFD-like domain of AT2faRel performs a different sensory function. This evolutionary dynamic is reminiscent of PanA11 and HigA37,40 antitoxins. While, in the case of single-domain antitoxins, PanA directly mediates toxin neutralization, in multidomain antitoxins, PanA domains are not the direct neutralization element; instead, these domains were hypothesized to sense the TA-activating triggers24. Establishing the putative sensory functions of pZFDs in translation-targeting and metabolism-targeting toxSASs is one of the remaining challenges in the field.
Methods
Plasmid construction
Fragments of WT aTfaRel2 and its substituted variants (V43A, I45A, A50M, Y51A and Y54A) were PCR-amplified with primers VTK198 and VTK199 and templates VHp278 (WT), VHp1225 (V43A), VHp1226 (I45A), VHp1227 (A50M), VHp1236 (Y51A) or VHp1228 (Y54A). Using Gibson assembly, the resulting linear DNA fragment was inserted into linearized pMG25 using pMG HiFi For and pMG HiFi Rev primers.
Sequence analysis
Representative FaRel2 and cognate ATFaRel2 sequences were retrieved using webFlaGs41, implementing the protein basic local alignment search tool (BLASTp) in the National Center for Biotechnology Information (NCBI) RefSeq Select database and otherwise default settings. Sequences were aligned using MAFFT version 7.490 with the L-INS-i strategy42.
Toxicity neutralization assays
The experiments were performed as described previously7. The assays were performed on Luria–Bertani (LB) medium plates (BD). We used the Escherichia coli BW25113 strain cotransformed with two different plasmid systems for controllable expression of toxins and antitoxins. We used a pair of compatible plasmids: pMG25 for antitoxin expression (high copy number, ColE1 origin of replication (pUC), AmpR, antitoxin expressed under the control of IPTG-inducible PA1/04/03 promoter43] and pBAD33 for toxin expression (medium copy number, p15A origin of replication, CmlR, toxins expressed under the control of arabinose-inducible PBAD promoter44). The cells were grown in liquid LB medium (BD) supplemented with 0.2% glucose (repression conditions), 100 µg ml−1 ampicillin (AppliChem) and 20 µg ml−1 chloramphenicol (AppliChem). Serial dilutions were spotted on solid LB plates supplemented with 0.2% arabinose, as well as 100 µg ml−1 ampicillin (AppliChem) and 20 µg ml−1 chloramphenicol (AppliChem), and bacterial growth was scored after 16-h incubation at 37 °C.
Protein purification
ATfaRel2, Farel2-Y128F and the different variants of the proteins were expressed in E. coli BL21DE3. The proteins were produced with a His6 tag at the N terminus, followed by a tobacco etch virus (TEV) protease cleavage site for ATfaRel2 and the different ATfaRel2 variants and a SUMO (small ubiquitin-like modifier) tag for FaRel2-Y128F. Cultures were grown in LB medium supplemented with kanamycin (50 μg ml−1) at 37 °C with aeration. Expression was induced with 0.5 mM IPTG when the cells carrying the plasmid reached an optical density at 600 nm (OD600 nm) of ~0.5–0.8. After induction, the cells were harvested 16 h later by centrifugation and resuspended with buffer (25 mM HEPES pH 7.6, 1 M NaCl, 5 mM MgCl2 and 1 mM TCEP) supplied with cOmplete protease inhibitor cocktail (Roche). The resuspended cells were flash-frozen in liquid nitrogen and stored at −80 °C.
The cell extracts were lysed using an Emulsiflex cell disruptor and the lysate was centrifuged to remove cell debris for 45 min at 25,000g. In both cases, the supernatant was loaded onto a 1-ml HiTrap Ni-NTA column (Cytiva) coupled to a fast protein liquid chromatography (FPLC) system (ÄKTA Explorer) equilibrated with buffer A (25 mM HEPES pH 7.6, 1 M NaCl, 5 mM MgCl2, 1 mM TCEP and 20 mM imidazole). The column was washed with a linear gradient of buffer B (25 mM HEPES pH 7.6, 1 M NaCl, 5 mM MgCl2, 1 mM TCEP and 500 mM imidazole). After tag removal, all individual proteins were further purified by SEC in a Superdex 75 Increase 10/30 (Cytiva) column using 25 mM HEPES pH 7.6, 300 mM NaCl, 2 mM MgCl2 and 1 mM TCEP. Sample purity was confirmed by SDS–PAGE. The ATfaRel2:FaRel2-Y128F complex was obtained by mixing both proteins in a 1:1.2 molar ratio with an excess of antitoxin that was separated by SEC.
Analytical SEC
For the analytical SEC, 150 μl of each protein at a concentration of 1 mg ml−1 was loaded on a Superdex 300 Increase 1030 column (Cytiva) previously equilibrated in SEC buffer (25 mM HEPES pH 7.6, 200 mM NaCl, 2 mM MgCl2 and 1 mM TCEP). The progress of the chromatography was monitored by the OD280 value.
Crystallization
Before crystallization, FaRel2-Y128F, ATfaRel2 and the ATfaRel2 variants were purified from the TEV cleavage reaction in the SEC buffer and concentrated to 8–10 mg ml−1. Screening of the crystallization conditions was carried out using the sitting-drop vapor diffusion method. The drops were set up in Swiss (MRC) 96-well two-drop UVP sitting-drop plates using the Mosquito HTS system (TTP Labtech). Then, 0.1-μl drops of protein and precipitant solution were equilibrated to 80 μl of precipitant solution in the reservoir. Commercially available screens LMB and SG1 (Molecular Dimensions) were used to test the crystallization conditions. The conditions resulting in diffracting crystals are listed in Supplementary Table 1. The crystals used for data collection the 0.2-μl drops used in the screens were scaled to 2-μl drops.
Structure determination
All the data were processed with the XDS suite45 and scaled with Aimless46. In all cases, the unit cell content was estimated with the program MATTHEW COEF from the CCP4 program suite47. The crystals of ATfaRel2 diffracted on average to ~1.3 Å. The analysis of the crystal anisotropy by the STARANISO server (http://staraniso.globalphasing.org/) proposed a resolution of 1.24 Å (with 1.24 Å in a*, 1.33 Å in b* and 1.43 Å in c*). We used Arcimboldo_Lite48 to perform ab initio phasing and solved the structure of ATfaRel2 in combination with Phaser49 and SHELXE50,51. The solution contained 75 of the 99 residues; the final structure was completed by manual building using Coot52 and refined with Buster/TNT53 (R/Rfree = 18.6/20.8).
In the case of the crystals of the FaRel2:APCPP complex, the analysis of the diffraction data suggested a resolution of 2.62 Å (with diffraction limits of 2.53 Å in a*, 2.67 Å in b* and 2.97 Å in c*) based on which we selected 2.62 Å as the resolution cut-ff. We used the coordinates of CapRelSJ46 (PDB 7ZTB) as the search model for the toxSYNTH domain of FaRel2 in complex with APCPP. The molecular replacement (MR) solution from Phaser49 was used in combination with Rosetta as implemented in the MR-Rosetta suite from the Phenix package54. After several iterations of manual building with Coot52 and maximum likelihood refinement as implemented in Buster/TNT53, the model was extended to cover all the residues (R/Rfree = 19.4/25.5).
The crystals of the ATfaRel2:FaRel2 complex were obtained in the P212121 space group. The anisotropic analysis of the diffraction data suggested a resolution of 2.14 Å (with diffraction limits of 2.25 Å in a*, 3.00 Å in b* and 2.13 Å in c*). We used the refined coordinates of FaRel2 (PDB 8PU4, this work) and ATfaRel2 (PDB 8PU2, this work) as the search model for phasing and estimated the unit cell content with MATTHEW COEF from the CCP4 program suite47. The MR solution from Phaser49 was completed by manual building with Coot52 and refined with Buster/TNT53 (R/Rfree = 18.4/23.8). To obtain the structure of the ATfaRel2:FaRel2:APCPP complex, the P212121 space group did not tolerate soaking; therefore, we grew crystals in a different condition (F4132 space group). The coordinates of FaRel2 (PDB 8PU4, this work) and ATfaRel2 (PDB 8PU2, this work) were used for MR with Phaser49 and modeling was completed by manual building with Coot52 and refinement with Buster/TNT53 (R/Rfree = 21.8/23.2). Supplementary Table 1 details all the X-ray data collection and refinement statistics.
ITC
All titrations were performed with an Affinity ITC (TA instruments) at 25 °C. For antitoxin versus toxin titrations, ATfaRel2 and its substituted variants were loaded in the instrument syringe at 200–150 μM and FaRel2-Y128F was used in the cell at 15–20 μM. In the case of the titrations of tRNA and mRNA versus toxSASs, 150 μM tRNA or mRNA was titrated into 15 μM FaRel2-Y128F, PhRel-Y143F, FaRel-Y175F, RelQ or the ATfaRel2:FaRel2-Y128F complex. For the titrations with nucleotides, APCPP was loaded in the instrument syringe at 180 μM and FaRel2-Y128F or the ATfaRel2:FaRel2-Y128F complex was used in the cell at 15 μM. All titrations were performed in 25 mM HEPES pH 7.6, 300 mM NaCl, 2 mM MgCl2 and 1 mM TCEP. Final concentrations were verified by the OD280 value using a Nanodrop One (Thermo Fisher Scientific). All ITC measurements were performed by titrating a constant volume of 2 μl into the ITC cell using a constant stirring rate of 75 rpm. All data were processed, buffer-corrected and analyzed using the NanoAnalyse and Origin software packages. Supplementary Table 3 details all the thermodynamic parameters derived from the ITC titrations.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data generated in this study are provided in the Supplementary Information provided with this paper. All coordinates were deposited in the PDB under accession numbers 8PU1, 8PU2, 8PU3 and 8PU4. Data are also available from the corresponding authors upon request.
References
Atkinson, G. C., Tenson, T. & Hauryliuk, V. The RelA/SpoT homolog (RSH) superfamily: distribution and functional evolution of ppGpp synthetases and hydrolases across the tree of life. PLoS ONE 6, e23479 (2011).
Lemos, J. A., Lin, V. K., Nascimento, M. M., Abranches, J. & Burne, R. A. Three gene products govern (p)ppGpp production by Streptococcus mutans. Mol. Microbiol. 65, 1568–1581 (2007).
Nanamiya, H. et al. Identification and functional analysis of novel (p)ppGpp synthetase genes in Bacillus subtilis. Mol. Microbiol. 67, 291–304 (2008).
Srivatsan, A. et al. High-precision, whole-genome sequencing of laboratory strains facilitates genetic studies. PLoS Genet. 4, e1000139 (2008).
Jimmy, S. et al. A widespread toxin–antitoxin system exploiting growth control via alarmone signaling. Proc. Natl Acad. Sci. USA 117, 10500–10510 (2020).
Ahmad, S. et al. An interbacterial toxin inhibits target cell growth by synthesizing (p)ppApp. Nature 575, 674–678 (2019).
Kurata, T. et al. RelA–SpoT homolog toxins pyrophosphorylate the CCA end of tRNA to inhibit protein synthesis. Mol. Cell 81, 3160–3170 (2021).
Geiger, T., Kastle, B., Gratani, F. L., Goerke, C. & Wolz, C. Two small (p)ppGpp synthases in Staphylococcus aureus mediate tolerance against cell envelope stress conditions. J. Bacteriol. 196, 894–902 (2014).
Takada, H. et al. Ribosome association primes the stringent factor Rel for tRNA-dependent locking in the A-site and activation of (p)ppGpp synthesis. Nucleic Acids Res. 49, 444–457 (2021).
Zhang, T. et al. Direct activation of a bacterial innate immune system by a viral capsid protein. Nature 612, 132–140 (2022).
Kurata, T. et al. A hyperpromiscuous antitoxin protein domain for the neutralization of diverse toxin domains. Proc. Natl Acad. Sci. USA 119, e2102212119 (2022).
Roghanian, M. et al. (p)ppGpp controls stringent factors by exploiting antagonistic allosteric coupling between catalytic domains. Mol. Cell 81, 3310–3322 (2021).
Tamman, H. et al. A nucleotide-switch mechanism mediates opposing catalytic activities of Rel enzymes. Nat. Chem. Biol. 16, 834–840 (2020).
Hogg, T., Mechold, U., Malke, H., Cashel, M. & Hilgenfeld, R. Conformational antagonism between opposing active sites in a bifunctional RelA/SpoT homolog modulates (p)ppGpp metabolism during the stringent response [corrected]. Cell 117, 57–68 (2004).
Manav, M. C. et al. Structural basis for (p)ppGpp synthesis by the Staphylococcus aureus small alarmone synthetase RelP. J. Biol. Chem. 293, 3254–3264 (2018).
Steinchen, W. et al. Structural and mechanistic divergence of the small (p)ppGpp synthetases RelP and RelQ. Sci. Rep. 8, 2195 (2018).
Steinchen, W. et al. Catalytic mechanism and allosteric regulation of an oligomeric (p)ppGpp synthetase by an alarmone. Proc. Natl Acad. Sci. USA 112, 13348–13353 (2015).
Tamman, H. et al. Structure of SpoT reveals evolutionary tuning of catalysis via conformational constraint. Nat. Chem. Biol. 19, 334–345 (2022).
Pausch, P. et al. Structural basis for regulation of the opposing (p)ppGpp synthetase and hydrolase within the stringent response orchestrator Rel. Cell Rep. 32, 108157 (2020).
LeRoux, M., Culviner, P. H., Liu, Y. J., Littlehale, M. L. & Laub, M. T. Stress can induce transcription of toxin–antitoxin systems without activating toxin. Mol. Cell 79, 280–292 (2020).
Overgaard, M., Borch, J., Jorgensen, M. G. & Gerdes, K. Messenger RNA interferase RelE controls relBE transcription by conditional cooperativity. Mol. Microbiol. 69, 841–857 (2008).
Beljantseva, J. et al. Negative allosteric regulation of Enterococcus faecalis small alarmone synthetase RelQ by single-stranded RNA. Proc. Natl Acad. Sci. USA 114, 3726–3731 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
& Ernits, K. et al. The structural basis of hyperpromiscuity in a core combinatorial network of type II toxin–antitoxin and related phage defense systems. Proc. Natl Acad. Sci. USA 120, e2305393120 (2023).
Garcia-Pino, A., Zenkin, N. & Loris, R. The many faces of Fic: structural and functional aspects of Fic enzymes. Trends Biochem. Sci. 39, 121–129 (2014).
Loris, R. & Garcia-Pino, A. Disorder- and dynamics-based regulatory mechanisms in toxin–antitoxin modules. Chem. Rev. 114, 6933–6947 (2014).
Ainelo, A. et al. The structure of DarB in complex with Rel(NTD) reveals nonribosomal activation of Rel stringent factors. Sci. Adv. 9, eade4077 (2023).
Dalton, K. M. & Crosson, S. A conserved mode of protein recognition and binding in a ParD–ParE toxin–antitoxin complex. Biochemistry 49, 2205–2215 (2010).
De Jonge, N. et al. Rejuvenation of CcdB-poisoned gyrase by an intrinsically disordered protein domain. Mol. Cell 35, 154–163 (2009).
Engel, P. et al. Adenylylation control by intra- or intermolecular active-site obstruction in Fic proteins. Nature 482, 107–110 (2012).
Garcia-Pino, A. et al. Allostery and intrinsic disorder mediate transcription regulation by conditional cooperativity. Cell 142, 101–111 (2010).
Garcia-Pino, A. et al. Doc of prophage P1 is inhibited by its antitoxin partner Phd through fold complementation. J. Biol. Chem. 283, 30821–30827 (2008).
Kamada, K., Hanaoka, F. & Burley, S. K. Crystal structure of the MazE/MazF complex: molecular bases of antidote–toxin recognition. Mol. Cell 11, 875–884 (2003).
Sterckx, Y. G. et al. A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2–ParE2 antitoxin–toxin complex. J. Mol. Biol. 428, 1589–1603 (2016).
Garcia-Pino, A. et al. An intrinsically disordered entropic switch determines allostery in Phd–Doc regulation. Nat. Chem. Biol. 12, 490–496 (2016).
Jurenas, D., Van Melderen, L. & Garcia-Pino, A. Mechanism of regulation and neutralization of the AtaR–AtaT toxin–antitoxin system. Nat. Chem. Biol. 15, 285–294 (2019).
Talavera, A. et al. A dual role in regulation and toxicity for the disordered N-terminus of the toxin GraT. Nat. Commun. 10, 972 (2019).
Laity, J. H., Lee, B. M. & Wright, P. E. Zinc finger proteins: new insights into structural and functional diversity. Curr. Opin. Struct. Biol. 11, 39–46 (2001).
Azarkan, M., Martinez-Rodriguez, S., Buts, L., Baeyens-Volant, D. & Garcia-Pino, A. The plasticity of the β-trefoil fold constitutes an evolutionary platform for protease inhibition. J. Biol. Chem. 286, 43726–43734 (2011).
Hadzi, S. et al. Ribosome-dependent Vibrio cholerae mRNAse HigB2 is regulated by a β-strand sliding mechanism. Nucleic Acids Res. 45, 4972–4983 (2017).
Saha, C. K., Sanches Pires, R., Brolin, H., Delannoy, M. & Atkinson, G. C. FlaGs and webFlaGs: discovering novel biology through the analysis of gene neighbourhood conservation. Bioinformatics 37, 1312–1314 (2021).
Katoh, K. & Standley, D. M. MAFFT: iterative refinement and additional methods. Methods Mol. Biol. 1079, 131–146 (2014).
Jaskólska, M. & Gerdes, K. CRP-dependent positive autoregulation and proteolytic degradation regulate competence activator Sxy of Escherichia coli. Mol. Microbiol. 95, 833–845 (2015).
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177, 4121–4130 (1995).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Collaborative Computational Project, Number 4.The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Sammito, M. et al. ARCIMBOLDO_LITE: single-workstation implementation and use. Acta Crystallogr. D Biol. Crystallogr. 71, 1921–1930 (2015).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Thorn, A. Experimental phasing: substructure solution and density modification as implemented in SHELX. Methods Mol. Biol. 1607, 357–376 (2017).
Uson, I. & Sheldrick, G. M. An introduction to experimental phasing of macromolecules illustrated by SHELX; new autotracing features. Acta Crystallogr. D Struct. Biol. 74, 106–116 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Smart, O. S. et al. Exploiting structure similarity in refinement: automated NCS and target-structure restraints in BUSTER. Acta Crystallogr. D Biol. Crystallogr. 68, 368–380 (2012).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Acknowledgements
This work was supported by the Fonds National de Recherche Scientifique (FNRS; CDR J.0068.19 and J.0065.23F, EQP UN.025.19 and PDR T.0066.18 and T.0090.22 to A.G.P.), European Research Council (ERC; CoG DiStRes, no. 864311 to A.G.P.), Joint Programming Initiative on Antimicrobial Resistance (JPIAMR; JPI-EC-AMR-R.8004.18 to A.G.P.), Fonds Jean Brachet and the Fondation Van Buuren (A.G.P.). This work was supported by the Swedish Research Council (Vetenskapsrådet; grants 2019-01085 and 2022-01603 to G.C.A. and 2021-01146 to V.H.), Crafoord Foundation (project grant no. 20220562 to V.H.), Estonian Research Council (PRG335 to V.H.) and Cancerfonden (20 0872 Pj to V.H.). G.C.A. and V.H. were also supported by a project grant from the Knut and Alice Wallenberg Foundation (2020-0037 to G.C.A.). A.C. was supported by the Fund for Research in Industry and Agronomy (FRIA; FC31211 to A.C.). We acknowledge the use of beamtimes PROXIMA 1 and 2A at the Soleil synchrotron (Gif-sur-Yvette, France) and I24 at the Diamond Light Source (United Kingdom).
Funding
Open access funding provided by Lund University.
Author information
Authors and Affiliations
Contributions
A.G.P. and V.H. drafted the paper with contributions from all authors and coordinated the study. V.H., A.G.P., G.C.A., A.T.P. and T.K. designed the experiments and analyzed the data. L.D.M., A.C., A.T.P. and T.K. performed the biochemical and microbiological experiments. D.E.B., S.Z. and A.C. performed the mutagenesis of FaRel2, ATfaRel2, FaRel and AT2faRel. L.D.M., A.C. and A.T.P. performed the biophysical measurements. L.D.M. crystallized the proteins. A.T.P. and A.G.P. determined the structures of FaRel2, ATfaRel2 and FaRel2:ATfaRel2. A.G.P. and G.C.A. performed the bioinformatic analyses. All authors approved the final revision of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Kevin Forsberg, Mahavir Singh and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Tables 1–4.
Supplementary Data 1
Strains.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dominguez-Molina, L., Kurata, T., Cepauskas, A. et al. Mechanisms of neutralization of toxSAS from toxin–antitoxin modules. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-024-01630-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-024-01630-4
- Springer Nature America, Inc.