Abstract
Quiescence, a hallmark of adult neural stem cells (NSCs), is required for maintaining the NSC pool to support life-long continuous neurogenesis in the adult dentate gyrus (DG). Whether long-lasting epigenetic modifications maintain NSC quiescence over the long term in the adult DG is not well-understood. Here we show that mice with haploinsufficiency of Setd1a, a schizophrenia risk gene encoding a histone H3K4 methyltransferase, develop an enlarged DG with more dentate granule cells after young adulthood. Deletion of Setd1a specifically in quiescent NSCs in the adult DG promotes their activation and neurogenesis, which is countered by inhibition of the histone demethylase LSD1. Mechanistically, RNA-sequencing and CUT & RUN analyses of cultured quiescent adult NSCs reveal Setd1a deletion-induced transcriptional changes and many Setd1a targets, among which down-regulation of Bhlhe40 promotes quiescent NSC activation in the adult DG in vivo. Together, our study reveals a Setd1a-dependent epigenetic mechanism that sustains NSC quiescence in the adult DG.
Similar content being viewed by others
Introduction
Adult neurogenesis occurs throughout life in the dentate gyrus (DG) of the hippocampus in almost all mammals examined and contributes to specific brain functions, whereas its dysregulation has been implicated in a number of brain disorders1,2,3,4,5,6,7,8,9. In the adult mouse DG, radial glia-like cells (RGLs) are bona fide multipotent adult neural stem cells (NSCs)10,11 and an overwhelming majority of them reside in a state of reversible cell cycle arrest, named quiescence12. Once activated, RGLs give rise to new dentate granule cells and astrocytes and are often depleted10,13,14,15. Consequently, there is an age-dependent decline in the size of the adult NSC pool accompanied by reduced adult hippocampal neurogenesis16,17,18,19. Similarly, a number of genetic manipulations that promote precocious activation of quiescent RGLs in mice, such as deletion of FoxOs20,21 or Mfge822, leads to premature depletion of the NSC pool during early postnatal stages, resulting in decreased adult hippocampal neurogenesis later in life. Given that quiescence is required to maintain life-long continuous neurogenesis12,23, a better understanding of the molecular mechanisms regulating adult NSC quiescence is of significant importance.
A growing body of evidence indicates that the quiescent state of RGLs is not a dormant state, but is actively regulated in the adult brain12,24,25,26,27,28. Extracellular signals from the local niche29,30,31,32 converge in RGLs, and downstream intrinsic factors, including metabolites33,34,35,36, signaling pathways37, cell cycle regulators38, and transcription factors39,40,41,42, mediate the effects of niche signals to balance RGL quiescence versus activation12,23. For example, a number of neuronal activity-dependent niche factors have been shown to dynamically regulate RGL quiescence32, such as GABA30 and sFRP329,43. On the other hand, as a hallmark of adult NSCs, quiescence can be maintained over months and years before RGL reactivation44, and even upon deletion of factors such as PTEN10, Mfge822, and GABARA30 that promote quiescent RGL activation, the majority of RGLs remain quiescent. Therefore, beyond mechanisms that dynamically regulate quiescent RGL activation upon demand, there could exist a fundamental mechanism to ensure sustained adult NSC quiescence over the long term. Epigenetic mechanisms, including long-lasting DNA and histone modifications, are ideal candidates45. While epigenetic mechanisms have been shown to affect the identity and differentiation of adult NSCs and their progeny46,47,48,49,50,51,52, whether and how epigenetic mechanisms may regulate adult NSC quiescence in the DG is not well-understood53,54,55.
Setd1a encodes a histone methyltransferase that deposits mono-/di-/trimethylation epigenetic marks on the lysine 4 residue in histone 3 tails (H3K4me1/2/3)56. In humans, rare heterozygous loss-of-function variants in the Setd1a gene increase the risk of schizophrenia (SCZ) about 35-fold57,58,59. Several Setd1a+/− mouse models have been generated to investigate how this SCZ-associated chromatin regulator affects brain functions and behaviors60,61,62. Some common findings include reduced levels of H3K4me3, disrupted expression of synaptic genes, defective synaptic transmission, aberrant morphology in neurons60,61,62, and behavioral abnormalities, such as deficits in spatial working memory and recognition memory60,61. Many of these deficits can be rescued with pharmacological inhibition of LSD1, a histone demethylase that counters the effect of Setd1a60,63, implying an epigenetic mechanism of Setd1a in these biological processes. To date, all published mouse studies have been focused on the role of Setd1a in the prefrontal cortex and striatum60,61,62, whereas its role in the hippocampus and neurogenesis has not been investigated. Here, we show that adult mice with Setd1a haploinsufficiency develop an enlarged DG. We further investigate the specific role of Setd1a in regulating adult hippocampal neurogenesis and identify an epigenetic mechanism that sustains NSC quiescence in the adult mouse DG.
Results
Increased number of dentate granule cells in adult mice with Setd1a haploinsufficiency
We first crossed Setd1afl/fl mice64 with Nestin-Cre mice65 to delete Setd1a in the embryonic mouse brain. The efficacy of Setd1a deletion was confirmed by Western blotting with E17.5 brains (Supplementary Fig. 1a, b). Homozygous conditional knockout mice died by postnatal day 2 (P2) and heterozygous conditional knockout mice (hereafter, referred to as cHet) survived into adulthood. Interestingly, the DG was substantially enlarged in 3-month-old cHet mice compared to wildtype (Setd1afl/+, WT) littermates (Fig. 1a, b). We measured the size of neuronal layers in the hippocampus in every 6th section across the entire dorsal-ventral axis and found specific enlargement of the dentate granule cell layer, but not CA1 or CA3 neuronal layers in cHet compared to WT mice (Fig. 1a, b). Specifically, cHet mice exhibited increased layer thickness, but not density of dentate granule cells, compared to WT littermates (Fig. 1c, d). Further time course analysis showed no significant difference between WT and cHet mice in the size of the dentate granule cell layer at P14 and P28, but an increased size in cHet mice at 3 and 9 months (Fig. 1b and Supplementary Fig. 1c–h), indicating an adult-onset phenotype.
a, b Sample images (a; scale bars: 200 μm) and quantification of areas of the dentate granule cell layer (GCL), CA1 and CA3 neuronal layers of every 6th brain section along the dorsal-ventral axis of the hippocampus in 3-month-old WT and cHet mice (b). Values represent mean ± SEM (DG: n = 4/WT, 6/cHet; CA1: n = 4/WT, 3/cHet; CA3: n = 4/WT, 3/cHet). c, d Sample confocal images of DAPI staining of the suprapyramidal blade of the DG in 3-month-old WT and cHet mice (c; scale bar: 20 μm) and quantification of the thickness of GCL and densities of dentate granule cells (d). Each dot represents data from one section (left panel) or one mouse (right panel). Values represent mean ± SEM (n = 3/WT, 3/cHet). e, f Sample confocal images of immunostaining of various markers and DAPI (e; scale bars: 20 μm) and quantification of the densities of Nestin+GFAP+ radial glial-like neural stem cells (RGLs), Tbr2+ intermediate progenitor cells (IPCs) and DCX+ immature neurons (INs) in the DG of WT and cHet mice at P28, 3 months and 9 months (f). In f, each dot represents data from one mouse. Values represent mean ± SEM (n = 3/WT, 3/cHet for each condition). g, h Sample confocal images of immunostaining of RGL markers Nestin, GFAP and Sox2 and proliferation marker MCM2 RGLs in the DG of 3-month-old WT and cHet mice (g; scale bars: 20 μm) and quantification of percentages of MCM2+Nestin+GFAP+Sox2+ active RGLs among Nestin+GFAP+Sox2+ total RGLs in the DG of P28, 3-month-old and 9-month-old WT and cHet mice (h). Arrowheads mark MCM2+Nestin+Sox2+GFAP+ active RGLs and arrows mark MCM2-Nestin+Sox2+GFAP+ quiescent RGLs (g). In h, each dot represents data from one mouse. Values represent mean ± SEM (n = 5/WT, 5/cHet for 3 months and n = 3/WT, 3/cHet for P28 and 9 months). Two-tailed Student’s t-test is performed for statistical analysis and P values are listed. Source data are provided as a Source Data file.
We next examined the cellular mechanism underlying the increased number of dentate granule cells in the adult cHet mice. The adult DG harbors a population of quiescent RGLs, which can be activated to give rise to new dentate granule cells throughout life1,8,66, and thus potentially contribute to the increased number of dentate granule cells. To examine adult hippocampal neurogenesis, we quantified densities of Nestin+GFAP+ RGLs, Tbr2+ intermediate progenitor cells (IPCs), and DCX+ immature neurons (INs) in the DG at P28, 3 months and 9 months. Compared to WT littermates, densities of IPCs and INs increased at P28 and 3 months, but decreased at 9 months in cHet mice, whereas the density of RGLs in cHet mice was lower at all time points examined with increasing differences over time (Fig. 1e, f). Next, we directly examined quiescent RGL activation and found an increased percentage of MCM2+Nestin+GFAP+Sox2+ active RGLs among Nestin+GFAP+Sox2+ total RGLs in the DG of cHet mice at P28, 3 months and 9 months compared to WT littermates (Fig. 1g, h).
Taken together, these results show that heterozygous Setd1a deletion in the embryonic nervous system results in the enlargement of DG in adulthood. Mechanistically, Setd1a haploinsufficiency leads to elevated activation of quiescent RGLs in the adult DG, resulting in an increased number of dentate granule cells and accelerated depletion of RGLs over time. Given the strong phenotype in heterozygous deletion animals (Fig. 1f–h), these findings suggest an indispensable role of Setd1a for the long term maintenance of RGL quiescence and the NSC pool in the adult DG.
Elevated activation of quiescent RGLs in the postnatal DG upon induced Setd1a deletion
In our Nestin-cre-based mice, Setd1a was deleted in all cells in the embryonic nervous system and there could be both non-cell autonomous effects and indirect effects of epigenetic inheritance from development on RGLs in the adult DG. To further support our model and examine the cell autonomous direct role of Setd1a specifically in quiescent RGLs, we generated sets of Setd1a inducible heterozygous and homozygous conditional deletion mice with matched controls, including Hopx-CreERT2::Setd1afl/+::H2B-GFP (iHet), Hopx-CreERT2::Setd1afl/fl::H2B-GFP (iKO), and Hopx-CreERT2::Setd1+/+::H2B-GFP (WT), by crossing mice with the Setd1afl/fl conditional allele with the Hopx-CreERT2 driver that specifically targets quiescent RGLs in the adult DG44 and has an inducible nuclear localized H2B-GFP reporter67. We injected 6-week-old mice with tamoxifen once daily for 3 days and analyzed the efficacy of Setd1a deletion in RGLs at 3 and 6 days after the last injection (dpi) by immunohistology (Supplementary Fig. 2a). Quantification of Setd1a expression showed reduced levels in RGLs of iHet mice at 3 and 6 dpi with further decreased levels in iKO mice, especially at 6 dpi (Supplementary Fig. 2b–e). We next examined RGL activation in these animals at 7 dpi (Fig. 2a). Quantification revealed a significant increase in the percentage of MCM2+Sox2+Nestin+GFP+ active RGLs among Sox2+Nestin+GFP+ total RGLs in the DG of both iHet and iKO mice compared to WT littermates (Fig. 2b, c). We then examined the consequence of elevated quiescent RGL activation at 14 and 30 dpi (Fig. 2d). At 14 dpi, quantification showed significantly increased percentages of Tbr2+ IPCs and DCX+ INs and decreased percentages of Nestin+GFAP+GFP+ RGLs among all GFP+ cells in the iHet and iKO mice compared to WT mice (Fig. 2e, f). At 30 dpi, there were still increased percentages of INs and decreased percentages of RGLs, but no difference in percentages of IPCs, among all GFP+ cells in the iHet and iKO mice compared to WT mice (Fig. 2g, h).
a–c Deletion of Setd1a specifically in RGLs promotes their activation in the adult DG. Shown in (a) is a schematic diagram of the experimental design. Shown in (b) are sample confocal images of immunostaining for various markers. Scale bars, 20 μm (inset, 5 μm). Arrowheads mark MCM2+Nestin+Sox2+GFP+ active RGLs and arrows mark MCM2-Nestin+Sox2+GFP+ quiescent RGLs. Shown in (c) is the quantification of the percentage of MCM2+Nestin+Sox2+GFP+ active RGLs among Nestin+Sox2+GFP+ total RGLs in the DG. Each dot represents data from one mouse. Values represent mean ± SEM (n = 3/WT, 4/iHet, 4/iKO; *P < 0.05). d–h Deletion of Setd1a in RGLs led to increased neurogenesis in the adult DG. Shown in (d) is a schematic diagram of the experimental design. Also shown are sample confocal images of immunostaining for various markers and DAPI at 14 (e) and 30 (g) days after the last tamoxifen injection (scale bars, 50 μm) and quantifications of percentages of Nestin+GFAP+GFP+ RGLs, Tbr2+GFP+ IPCs and DCX+GFP+ INs among total GFP+ cells in the DG of adult WT, iHet and iKO mice at 14 (f) and 30 (h) days after the last tamoxifen injection. In (f–h), each dot represents data from one mouse. Values represent mean ± SEM (14 days: RGLs: n = 3/WT, 4/iHet, 3/iKO; IPCs: n = 4/WT, 5/iHet, 3/iKO; INs: n = 4/WT, 5/iHet, 3/iKO; 30 days: RGLs: n = 3/WT, 4/iHet, 3/iKO; IPCs: n = 3/WT, 5/iHet, 3/iKO; INs: n = 4/WT, 5/iHet, 3/iKO). Two-tailed Student’s t-test is performed for statistical analysis and P values are listed. Source data are provided as a Source Data file.
We also examined whether the role of Setd1a is specific to quiescent RGLs by analyzing NSCs in the develo** DG. Our previous study has shown that Hopx-CreERT2 targets the population of NSCs in the develo** DG that gradually transitions into quiescent RGLs from P7 to P1428,44, therefore we used the same set of animals. We injected tamoxifen once at P1 and examined the proliferation of Sox2+Nestin+GFP+ NSCs at P3, P7 and P14 (Supplementary Fig. 3a). We did not detect any differences in the percentage of MCM2+Sox2+Nestin+GFP+ NSCs among Sox2+Nestin+GFP+ total NSCs in the DG of WT, iHet and iKO mice at P3 and P7, when almost all NSCs were proliferating (Supplementary Fig. 3b–e). In contrast, at P14 when about 70% of NSCs were quiescent in the WT DG, the percentages of MCM2+Sox2+Nestin+GFP+ active NSCs among Sox2+Nestin+GFP+ total NSCs were increased in iHet and iKO mice (Supplementary Fig. 3f, g).
Together, results from this cell type-specific inducible system, particularly the significant effect in iHet mice, suggest a critical and highly sensitive cell autonomous role of Setd1a in maintaining RGL quiescence in the DG.
Suppression of Setd1a-deficient quiescent RGL activation in the adult DG by LSD1 inhibition
To investigate the molecular mechanism underlying Setd1a-dependent maintenance of RGL quiescence in the adult DG, we first asked whether Setd1a functions through an epigenetic mechanism, given that Setd1a is known to exert functions independent of its H3K4 methyltransferase activity, such as mediating DNA damage responses68. LSD1, a histone demethylase for H3K4me1/2, is known to counteract the H3K4 methyltransferase function of Setd1a60,63. For example, pharmacological LSD1 inhibitors TCP69 or ORY-100170,71 rescued deficits in axon branching and abnormal behaviors of Setd1a+/- mice60. TCP is a prototype LSD1 inhibitor that also inhibits two major isoforms of monoamine oxidase MAO-A and MAO-B69. ORY-1001 is a TCP derivative that exhibits most potent and selective LSD1 inhibition reported to date; ORY-1001 has an over 1000-fold selectivity for LSD1 compared to MAOs60,70,71. We injected tamoxifen once daily for 3 days into 6-week-old WT, iHet, and iKO mice followed by TCP or ORY-1001 injection once daily for 7 days, and then performed analysis one day after the last injection of LSD1 inhibitors (Fig. 3a). Immunohistological analysis confirmed that these two LSD inhibitors indeed elevated levels of H3K4me1/2 as well as H3K4me3 in RGLs in WT animals (Supplementary Fig. 4). Quantification showed that either TCP or ORY-1001 treatment decreased percentages of MCM2+Sox2+Nestin+GFP+ active RGLs among Sox2+Nestin+GFP+ total RGLs in both iHet and iKO mice (Fig. 3), supporting the notion that Setd1a maintains adult RGL quiescence via regulation of H3K4 methylation levels. Notably, LSD1 inhibition also decreased the level of quiescent adult RGL activation in WT mice (Fig. 3b, e). Collectively, these results suggest that Setd1a and LSD1, two epigenetic regulators, co-regulate the quiescent state of adult RGLs in the DG, but in the opposite manner.
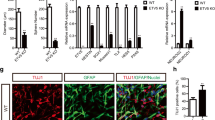
a A schematic diagram of the experimental design. b–d Sample confocal images of immunostaining for various markers. Scale bars: 20 μm. e Quantification of percentages of MCM2+Nestin+Sox2+GFP+ active RGLs among Nestin+Sox2+GFP+ total RGLs under the treatment of PBS (control) or LSD1 inhibitors (TCP or ORY-1001) in the DG of adult WT, iHet and iKO mice. Each dot represents data from one mouse. Values represent mean ± SEM (WT: n = 4/PBS, 3/TCP, 3/ORY-1001; iHet: n = 3/PBS, 3/TCP, 4/ORY-1001; iKO: n = 3/PBS, 3/TCP, 3/ORY-1001). Two-tailed Student’s t-test is performed for statistical analysis and P values are listed. Source data are provided as a Source Data file.
Epigenetically regulated transcriptomic changes in Setd1a-deficient adult NSCs
To further investigate the molecular mechanism underlying Setd1a-dependent maintenance of NSC quiescence, we turned to a culture model that mimics in vivo NSC quiescence by exposing adult NSC cultures72 to BMP4/FGF236,73,74. We derived adult NSCs from the DG of 7 ~ 8-week-old Setd1afl/fl::Hopx-CreERT2::H2B-GFP mice in the presence of EGF and FGF2 (Supplementary Fig. 5a). At 2 days after exposure to BMP4/FGF2 without EGF, almost all Sox2+ NSCs became Ki67-, indicative of quiescence (Supplementary Fig. 5a, b). Importantly, these quiescent adult NSCs were able to re-enter the cell cycle after replating in the presence of EGF/FGF2 without BMP4 (Supplementary Fig. 5a, b). We added lentivirus expressing Cre-GFP or GFP alone (control) to the quiescent adult NSC culture at 2 days after BMP4/FGF2 treatment (Supplementary Fig. 5c). About 80% of NSCs were infected by lentivirus as indicated by GFP expression and Setd1a levels were diminished at day 5 (3 days after viral infection) with Cre-GFP expression (Supplementary Fig. 5d–f).
We next performed time course analyzes of these NSC cultures at days 4, 5, 6, 7 and 9 for immunostaining and RNA-seq (Supplementary Fig. 5c). We first examined whether our culture model could recapitulate the elevated activation of Setd1a-deficient quiescent RGLs in the adult brain in vivo. Indeed, deletion of Setd1a increased the percentage of Ki67+GFP+ proliferating NSCs among total GFP+ NSCs at days 7 and 9, but not at day 5 (Fig. 4a, b). We next examined differentially expressed genes (DEGs) between Setd1a-deficient (expressing Cre-GFP) and WT (expressing GFP) NSCs with RNA-seq from days 4 to 9 (Supplementary Dataset 1). GO analysis showed that the downregulated DEGs from Setd1a-deficient NSCs were related to cell cycle and DNA repair pathways between days 5 and 7, whereas cell cycle genes were upregulated in Setd1a-deficient NSCs between days 6 and 9 (Fig. 4c), consistent with the observation of elevated activation of Setd1a-deficient quiescent NSCs at days 7 and 9 (Fig. 4a, b). Our previous single-cell RNA-seq analysis revealed a shift in the energy source and metabolism when adult quiescent RGLs transit into active states in the DG in vivo24. Specifically, lysosomal and metabolic pathways enriched in quiescent adult RGLs, such as fatty acid, sphingolipid and glutathione metabolism, were downregulated upon adult RGL activation24. Similarly, KEGG pathway analysis of downregulated DEGs from Setd1a-deficient NSCs showed downregulated lysosomal and metabolism pathways, including fatty acid, glutathione and sphingolipid metabolism (Fig. 4d). These results suggest that Setd1a deficiency results in transcriptomic alterations in cultured quiescent adult NSCs that mimic quiescent adult RGL activation in the DG in vivo.
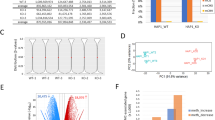
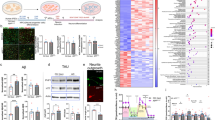
a, b Sample confocal images of immunostaining of Ki67, GFP and DAPI (a; scale bars: 20 μm) and quantification of percentages of Ki67+GFP+ active NSCs among total GFP+ cells at days 5, 7 and 9 after lentivirus infection to express GFP or Cre-GFP in Setd1a-deficient NSCs at day 2. In b, each dot represents data from one image. Values represent mean ± SEM (n = 3 independent cultures for each condition). c, d Pathway analysis of differentially expressed genes upon Setd1a deletion in adult NSCs. Shown in (c) are heatmaps of GO analysis (biological process) of downregulated and upregulated genes in NSCs from days 4 to 9 with GFP or Cre-GFP expression, indicating dysregulated expression of cell cycle genes and DNA repair genes. Shown in (d) is a heatmap of KEGG pathway analysis of downregulated genes from days 4 to 9, revealing downregulated metabolic pathways. e, f Sample confocal images of immunostaining of Ki67, GFP, and DAPI (e; scale bars: 20 μm) and quantification of percentages of Ki67+GFP+ proliferating NSCs among total GFP+ cells under the treatment of LSD1 inhibitor ORY-1001 or DMSO as control at days 5, 7 and 9 with lentivirus to express GFP or Cre-GFP at day 2 (f). Values represent mean ± SEM (n = 3 independent cultures for each condition). g-j RNA-seq analysis of WT (GFP) and Setd1a-difficient (Cre-GFP) NSCs under the treatment of DMSO (control), or LSD1 inhibitor ORY-1001. Shown in (g) is a PCA plot of biological replicates. Shown in (h) are Volcano plots of differentially expressed genes under different conditions. Shown in (i and j) are Venn diagrams of genes that were dysregulated by Setd1a deletion and were rescued by LSD1 inhibitor treatment (left) and GO analysis (biological process) of these genes (right). Two-tailed Student’s t-test is performed for statistical analysis and P values are listed. Source data are provided as a Source Data file.
To assess whether Setd1a also functions through an epigenetic mechanism to maintain quiescence of cultured adult NSCs as in vivo (Fig. 3), we treated cultured Setd1a-deficient quiescent NSCs with the LSD1 inhibitor ORY-1001 from day 2 (Supplementary Fig. 5c). Indeed, ORY-1001 treatment suppressed the elevated activation of Setd1a-deficient quiescent NSCs in vitro (Fig. 4e, f). To obtain mechanistic insights into suppression effects of ORY-1001, we performed RNA-seq and found that ORY-1001 treatment dramatically reduced the number of DEGs between Setd1a-deficient and WT NSCs at day 7, from 4,333 DEGs between KO + DMSO and WT + DMSO conditions to only 120 DEGs between KO + ORY and WT + ORY conditions (Fig. 4g, h and Supplementary Dataset 1). As a result, most of the Setd1a deficiency-induced upregulated and downregulated DEGs were rescued by the ORY-1001 treatment (Fig. 4i, j). GO analysis revealed that these rescued DEGs are related to cell division, cell cycle and metabolism pathways (Fig. 4i, j).
Collectively, our in vitro results corroborated our in vivo findings and suggested that Setd1a deficiency-induced and epigenetically regulated transcriptomic changes in quiescent adult NSCs in vitro mimic in vivo adult RGL activation. This in vitro adult NSC model is therefore well suited for identification of Setd1a epigenetic targets that may mediate its effect on maintaining adult RGL quiescence in vivo.
Bhlhe40, a Setd1a epigenetic target, maintains RGL quiescence in the adult DG
To identify Setd1a-target genes, we performed a Setd1a CUT & RUN assay of cultured WT quiescent adult NSCs at day 5 and identified 6969 Setd1a binding sites (Fig. 5a, b and Supplementary Dataset 2). About 66% of the Setd1a peaks were localized at promoter regions, which is consistent with the known enrichment of H3K4me3 epigenetic marks at active promoters75 (Fig. 5b). To identify potential quiescence-related genes directly regulated by Setd1a, we compared genes harboring Setd1a peaks at promoters with DEGs in Setd1a-deficit NSCs from our RNA-seq data at day 5, a time point just before the elevated activation of Setd1a-deficient quiescent NSCs (Fig. 4a, b). We found 330 overlapped genes with downregulated DEGs and 276 overlapped genes with upregulated DEGs (Fig. 5c and Supplementary Dataset 2). Given that Setd1a is associated with active promoters and transcription activation75, we focused on 330 overlapped genes among downregulated DEGs and our further analysis identified 27 genes whose downregulation may potentially regulate somatic stem cell quiescence based on published studies (Fig. 5d). For example, deletion of either Bhlhe40 or Atr has been reported to activate quiescent muscle stem cells in adult mice76,77 (Fig. 5d). A number of genes, such as Dazap2, Fbxl5 and Pgk1, have been reported to be involved in key signaling pathways regulating quiescence of somatic stem cells, such as Wnt, mTOR and PTEN pathways78,79,80 (Fig. 5d). In particular, our previous single-cell RNA-seq analysis revealed downregulation of Bhlhe41 during RGL transition from quiescence to activation in the adult DG24. Bhlhe40 is a paralogous gene of Bhlhe41, and Bhlhe40/41 form a homodimer or a heterodimer to function as transcriptional repressors81. In cultured quiescent adult NSCs, the expression level of Bhlhe40 was much higher than that of Bhlhe41 (Supplementary Fig. 6a). While no studies so far have reported the role of Bhlhe40/41 in the maintenance of adult NSC quiescence, depletion of Bhlhe40 has been shown to activate quiescent muscle stem cells in adult mice76 and we found that downregulation of Bhlhe40 in Setd1a-deficient NSCs was rescued by LSD1 inhibition (Fig. 5f, g). All together, these analyzes raised the possibility that Bhlhe40 is one of the Setd1a targets that may regulate adult NSC quiescence.
a Correlation of two independent Setd1a CUT & RUN analysis. b Distribution of CUT & RUN Setd1a peaks among genomic features in cultured quiescent NSCs at day 5. c Venn diagram showing the overlap between downregulated (left panel) and upregulated (right panel) genes in Setd1a-deficient compared to WT quiescent NSCs at day 5 from bulk RNA-seq data and genes with Setd1a peaks at the promoter regions in WT quiescent NSCs at day 5 from Setd1a CUT & RUN analysis. d Heatmaps showing Setd1a-target genes related to somatic stem cell quiescence. These representative genes were selected from 330 shared genes in the Venn diagram in (c), including genes reported to be important for maintaining quiescence of somatic stem cells in the adult mice and genes involved in the signaling pathways regulating quiescence of NSCs. HSCs, hematopoietic stem cells; HpSCs, hepatocyte stem cells; MuSCs, muscle stem cells; NSCs, neural stem cells. e Sequencing tracks of example Setd1a targets: Bhlhe40, Bhlhe41, Atr, Dazap2 and Fbxl5 from Setd1a CUT & RUN (left) and RNA-seq (right). f, g Heatmaps showing downregulation of Bhlhe40 and Bhlhe41 expression in Setd1a-deficient quiescent NSCs at indicated time points (f) and RNA-seq track of Bhlhe40 loci under different conditions (g). Note that downregulation of Bhlhe40 expression in Setd1a-deficient quiescent NSCs was rescued by ORY-1001 treatment at day 7.
To directly test this possibility, we expressed Bhlhe40 shRNA with an mCherry reporter in cultured quiescent adult NSCs with lentivirus and qPCR analysis showed that Bhlhe40 shRNA1 and shRNA2 decreased levels of Bhlhe40 transcripts by approximately 30% and 70%, respectively (Supplementary Fig. 6b). Quantification further showed that Bhlhe40 knockdown by either shRNA1 or shRNA2 led to increased percentages of Ki67+mCherry+ cells among all mCherry+ cells with shRNA2 exhibiting an earlier and larger effect than shRNA1, indicating that Bhlhe40 maintains adult NSC quiescence in a dose-dependent manner (Fig. 6a, b). To corroborate these in vitro findings, we injected lentivirus into the DG of 2-month-old WT mice to knockdown Bhlhe40 in adult RGLs. Quantification showed an increase in the percentage of MCM2+Sox2+Nestin+mCherry+ active RGLs among Sox2+Nestin+mCherry+ total infected RGLs for both shRNA1 and shRNA2 with a much stronger effect for shRNA2 (Fig. 6c, d).
a, b Sample confocal images of immunostaining for mCherry, Ki67 and DAPI (a; scale bars: 20 μm) and quantification of percentages of Ki67+mCherry+ proliferating NSCs among total mCherry+ NSCs at days 5 and 7 in culture (b). In b, each dot represents data from one culture. Values represent mean ± SEM (n = 3 independent cultures for each condition). c, d Sample confocal images of immunostaining of various markers (c; scale bars: 20 μm) and quantification of percentages of mCherry+MCM2+Sox2+Nestin+ active RGLs among mCherry+Sox2+Nestin+ total infected RGLs in the DG of 2-month-old WT mice 7 days after lentiviral injection (d). Arrowheads mark infected MCM2+ active RGLs and arrows mark infected MCM2- quiescent RGLs (c). In d, each dot represents data from one mouse. Values represent mean ± SEM (n = 5 mice for each condition). Two-tailed Student’s t-test is performed for statistical analysis and P values are listed. Source data are provided as a Source Data file.
Together, these results identified Bhlhe40 as one major epigenetic target of Setd1a that maintains adult RGL quiescence in the DG.
Discussion
Our comprehensive in vivo and in vitro studies with cellular, molecular and epigenetic analyzes identified a Setd1a-dependent epigenetic mechanism that sustains the quiescence of adult RGLs in the mouse DG. Beyond previous findings12,23 that provided insights into transcriptional and post-transcriptional control of adult NSC quiescence by genetic manipulations of niche factors29,30,31, cell cycle regulators38, transcription factors39,40,41,42, or metabolic enzymes12, our findings add an additional layer of epigenetic regulation to the quiescence control of adult NSCs and highlight the complexity of NSC quiescence regulation during life-long adult hippocampal neurogenesis. Importantly, the identified epigenetic mechanism potentially provides a foundation to ensure long term quiescence of adult NSCs, from which dynamic regulators can fine tune levels of adult NSC activation and neurogenesis upon demand. The significant effect of removing even one copy of Seta1a highlights the sensitive and indispensable role of such an epigenetic mechanism in sustaining NSC quiescence in the adult brain (Figs. 1f, h, 2c, and 3e,). Given that somatic stem cells in many adult tissue compartments are quiescent, our findings may have broad implications82,83,84.
Due to the long-lasting feature of epigenetic modifications, epigenetic mechanisms are ideal for sustained regulation, such as maintaining cell type identity throughout life85,86. Indeed, epigenetic mechanisms are known to regulate adult neurogenesis in the context of establishing cell type identity and regulating differentiation46,47,48. Here we identified an epigenetic mechanism via Setd1a-dependent H3K4 methylation for long term maintenance of NSC quiescence in the adult DG. A previous study showed that inhibition of histone deacetylases (HDACs) by valproic acid suppresses seizure-induced cell proliferation in the epileptic adult DG54. A further study showed that HDAC3 controls G2/M phase cell cycle progression of adult neural progenitors, therefore not quiescence maintenance87. Using a cell type-specific inducible system, we provided evidence for a cell autonomous and direct role of Sedt1a in maintaining adult NSC quiescence and further identified Bhlhe40 as one of Setd1a targets that maintains adult NSC quiescence both in vitro and in vivo. Notably, the impact of Bhlhe40-downregulation on quiescent adult NSC activation appeared to be much stronger than Setd1a downregulation both in vitro and in vivo (Figs. 2c, 4b, 6b and 6d), indicating that Bhlhe40 is a key factor maintaining adult RGL quiescence in the DG, whereas Setd1a is an upstream epigenetic regulator. Interestingly, Notch signaling is elevated in Bhlhe40-deficient muscle stem cells76. Notch signaling is essential for both proliferative and quiescent states of NSCs88, and promotes proliferation and quiescence through dynamic expression of different Notch effector proteins89. The downstream mechanism by which Bhlhe40 regulates quiescence of adult RGLs remains to be determined. In the current study, we identified 330 Setd1a targets and a list of 27 potential targets that may regulate somatic stem cell quiescence (Fig. 5), but only validated the function of Bhlhe40. It is possible that Setd1a deficiency-induced activation of quiescent adult NSCs is a convergent effect of dysregulating multiple quiescence regulators. Our study provides a list of candidates for future studies to test their roles in regulating adult NSC quiescence in vitro and in vivo. A previous study has shown that depletion of Bhlhe40 activates quiescent muscle stem cells in adult mice76. In addition, specific deletion of Setd1a in adult long term hematopoietic stem cells (LT-HSCs) results in G0-to-G1 transition and progression through cell cycle as well as increased numbers of Ki67+ LT-HSCs90. Whether the expression of Bhlhe40 is downregulated in Setd1a-deficient LT-HSCs, which contributes to the activation of Setd1a-deficient LT-HSCs, remains to be examined. Together with these findings, our study points to a common mechanism that can maintain quiescence of adult somatic stem cells in different tissues.
Setd1a is one of six major H3K4 methyltransferases expressed in mammalian cells91. Whether other H3K4 methyltransferases, such as Setd1b and Mll1-4, also regulate quiescence of adult RGLs in the DG remains to be determined. However, activation of quiescent RGLs by heterozygous Setd1a deletion indicates that loss of Setd1a was not compensated by other H3K4 methyltransferases, thus the function of Setd1a on maintaining quiescence of adult RGLs is not redundant. Our results from pharmacological inhibition of LSD1 with TCP and ORY-1001 suggest that Setd1a-mediated maintenance of RGL quiescence in the adult DG is dependent on its H3K4 methyltransferase activity. A previous study reported that inhibition of LSD1 with polyamine analogs not only increased global levels of H3K4 methylation, but also H3K9ac, an epigenetic mark at active promoters92,93, at the promoters of SFRP1, SFRP5, SFRP5 and GATA5 in human colon carcinoma cells94. Although TCP specifically inhibits H3K4 demethylation without affecting the deacetylase activity69, we cannot rule out the possibility that TCP or ORY-1001 treatment increases H3K9ac levels at promoters of certain genes in quiescent RGLs.
Setd1a is an established risk gene for SCZ57,58. Different from previous studies that explored the effects of Setd1a deficiency on neuronal functions with mouse models or human induced pluripotent stem cell-derived neuronal models60,61,62,95, our study uncovered an unexpected role of Setd1a in the long-term maintenance of NSC quiescence during adult hippocampal neurogenesis and showed that Setd1a haploinsufficiency, which has been used to model SCZ60,61,62, leads to reduced adult hippocampal neurogenesis later in life in mice (Fig. 1f). SCZ patients exhibit learning and memory deficits and studies have suggested deficits of adult hippocampal neurogenesis in SCZ96,97,98. It is interesting to speculate that deficits in Setd1a-mediated adult NSC maintenance and decreased adult hippocampal neurogenesis may occur later in life in SCZ patients with Setd1a mutations and contribute to their cognitive deficits.
Methods
Animals
All experimental procedures with mice used in this study were performed in accordance with protocols approved by the Institutional Animal Care and Use Committee of University of Pennsylvania. All mice were kept in cages with bedding materials and housed in a 14-hour light/10-hour dark cycle with regular food and water changes, an ambient temperature of 21–23 °C and humidity levels between 40% and 60% (protocol numbers: 806245, 806246). Setd1afl/fl mice were previously generated in Dr. Suming Huang’s lab64. Nestin-Cre+/Tg mice (Jackson Laboratory; 003771) were used to generate conditional knockout mice. These mice are referred to as Setd1afl/+ (WT) and Nestin-Cre::Setd1afl/+ (cHet). To generate Setd1a cell type-specific inducible knockout animals, we crossed Setd1afl/fl mice with Hopx-CreERT2::H2B-GFP mice44. Hopx-CreERT2::H2B-GFP mice were generated by crossing Hopx-CreERT2 knock-in mice (Jackson Laboratory; 017606) that harbored a tamoxifen-inducible CreERT2 fusion gene with a Cre-reporter mouse line Rosa26flox-stop-flox-H2B-GFP mice (from lab of Dr. Z. Josh Huang) harboring loxP sites on either side of a STOP sequence and upstream of a fusion H2B-GFP protein cassette67. These mice are referred to as Hopx-CreERT2::Setd1a+/+:: H2B-GFP (WT), Hopx-CreERT2::Setd1afl/+::H2B-GFP (iHet), and Hopx-CreERT2::Setd1afl/fl::H2B-GFP (iKO). Both male and female mice were used for all experiments, and no obvious sex phenotype was observed in any of the experiments and data were combined.
Primary mouse adult neural stem cells
Adult NSCs were derived from the DG of 7 ~ 8-week-old Hopx-CreERT2::Setd1afl/fl::H2B-GFP mice and cultured in proliferation medium containing Neurobasal medium (GIBCO), 20 ng/ml FGF2 (Peprotech), 20 ng/ml EGF (Peprotech), 2% B27 (v/v, GIBCO), 1% Glutamax (GIBCO), and 1% Penicillin/Streptomycin, on culture dishes precoated with poly-D-lysine (Sigma) and laminin (Sigma) as previously described72. To induce quiescence, cells were first plated into the proliferation medium at a density of 35,000–65,000 cells/cm2, and after 16 h fresh medium was added without EGF and with 20 ng/ml BMP4 (EMSCO/FISHER) and 20 ng/ml FGF2 (quiescent medium). For reactivation, after 3 days in the quiescence medium, cells were passaged with Accutase (Thermo Fisher) and plated into a proliferation medium at a density of 35,000–65,000 cells/cm2.
Antibodies
Anti-Nestin (Aves labs, Cat#NES, 1:500), Anti-GFAP (Dako, Cat# Z0334, 1:500), Anti-GFAP (Sigma, Cat# MAB360, 1:500), Anti-TBR2 (Abcam, Cat#ab183991, 1:200), Anti-DCX (EMD Millipore, Cat#AB2253, 1:200), Anti-DCX (Santa Cruz, Cat# sc-8066, 1:200), Anti-Mcm2 (BD, Cat#610701, 1:150), Anti-Sox2 (Invitrogen, Cat#14-9811-82, 1:150), Anti-GFP (Rockland, Cat#600101215, 1:500), Anti-mCherry (Biorbyt, Cat#orb11618, 1:500), Anti-Ki67 (BD Biosciences, Cat#550609, 1:150), Anti-Setd1a (Abcam, Cat#ab70378, 1:150), Anti-Setd1a (Santa Cruz, Cat#sc-515590, 1:150), Anti-α-Tubulin (Cell Signaling, Cat#3873 S, 1:5000), Anti-H3K4me1 (Abcam, Cat#ab8895, 1:150), Anti-H3K4me2 (Abcam, Cat#ab7766, 1:150), Anti-H3K4me3 (Abcam, Cat#ab8580, 1:150)
Western blot analysis
Brain tissues were lysed in ice-cold RIPA buffer (Thermofisher) containing protease inhibitor cocktail (Sigma) and phosphatase inhibitors (Cell Signaling). The lysates were sonicated and subjected to SDS-PAGE (BIO-RAD). The proteins on the gel were transferred to a nitrocellulose membrane (BIO-RAD), which was then blocked with 5% milk/TBST for 1 h at room temperature. The blot was incubated with primary antibodies in 5% milk/TBST overnight at 4 °C. After 3 washes in TBST, the blot was incubated with HRP-conjugated secondary antibodies in 5% milk/TBST for 1 h at room temperature. After 3 washes in TBST, ECL (Thermofisher) was used to detect immunoreactive bands on the blot using an Amersham Imager 600. Primary antibodies are listed in the Supplementary Dataset 3. Quantification of bands was performed using ImageJ software.
Tissue processing and immunohistology
For immunostaining of brain sections, animals were transcardially perfused with ice-cold PBS, followed by fresh ice-cold 4% paraformaldehyde in PBS, and brains were post-fixed overnight in the same fixative at 4 °C as previously described99. Samples were cryoprotected in 30% sucrose in PBS overnight at 4 °C, embedded in tissue freezing medium and sectioned coronally (35 μm thickness) on a Leica CM3050S cryostat. Brain sections were washed with PBS, and then blocked and permeabilized with the blocking solution (2% donkey serum, 3% Bovine serum albumin, and 0.3% Triton X-100 in PBS) for 1 h at room temperature, followed by incubation with primary antibodies diluted in the blocking solution at 4 °C overnight. After washing with PBS, secondary antibodies diluted in blocking solution were applied to the sections for 1 h at room temperature. Nuclei were visualized by incubating for 10 min with DAPI (BD Biosciences) in PBS. Brain sections were mounted with Aqua-Mount Mounting Medium (EMSCO/FISHER) and analyzed. Brain sections immunostained for Nestin and MCM2 underwent antigen retrieval where sections were incubated in 1× Target Retrieval Solution (Agilent Dako) at 95 °C for 20 min in a steamer, then room temperature for 20 min before primary antibody incubation. If GFP immunostaining was performed in conjunction with Nestin or MCM, then GFP primary and secondary antibody steps were completed prior to antigen retrieval. For immunostaining of cultured cells, cells were fixed with fresh 4% paraformaldehyde in PBS for 10 min. The following procedure was the same as the immunostaining for brain sections. All the antibodies used are listed in the Supplementary Dataset 3.
Confocal microscopy and image processing
Images were taken as z stacks using a Zeiss LSM 780 confocal microscope (Carl Zeiss). 20×, 40× or 63× objective was used for imaging. All confocal images were blindly acquired between experimental and control groups under the same laser power and gain. Images were analyzed using ImageJ software.
A confirmed RGL in Figs. 1e, 2e, g needed to satisfy the following criteria: (1) is located in the SGZ of the DG; (2) has a distinct Nestin+GFAP+ radial process; (3) has a DAPI+ nucleus that is largely surrounded by GFAP immunostaining signal in the same focal plane; and (for Fig. 2, g) (4) has a GFP+ nucleus. A confirmed RGL in Figs. 2b, 3b–d and Supplementary Fig. 4a–c needed to satisfy the following criteria: (1) is located in the SGZ of the DG; (2) has a distinct Nestin+ radial process; (3) has a Sox2 +GFP+ nucleus in the same focal plane. A confirmed RGL in Fig. 1g needed to satisfy the following criteria, (1) located in the SGZ of the DG; (2) has a distinct Nestin+GFAP+ radial process; (3) has a Sox2 + nucleus in the same focal plane. A confirmed RGL in Fig. 6c needed to satisfy the following criteria: (1) is located in the SGZ of the DG; (2) has a distinct Nestin+ radial process; (3) has a Sox2 + nucleus in the same focal plane; (4) has an mCherry+ cell body. A confirmed RGL in Supplementary Fig. 2b,d needed to satisfy the following criteria: (1) is located in the SGZ of the DG; (2) has a distinct Nestin+ radial process; (3) has a DAPI+GFP+ nucleus in the same focal plane.
Tamoxifen injection
A stock solution of 66.67 mg/mL tamoxifen (Thomas Scientific) was prepared in a 5:1 solution of corn oil:ethanol at 37 °C with occasional vortexing until dissolved44. Tamoxifen was injected intraperitoneally with one injection daily for 3 days for 6-week-old mice (Fig. 2a, d and Supplementary Fig. 2a) and once for P1 mice (Supplementary Fig. 3a).
TCP preparation and injection
Tranylcypromine (TCP, Sigma) was dissolved in PBS at a stock concentration of 0.75 mg/ml by vortexing. Mice at 17–20 g body weight were injected 2.7 mg/kg/day; mice at 21–28 g body weight were injected 3.0 mg/kg/day as previously described60. 6-week-old animals were intraperitoneally injected for 7 consecutive days following tamoxifen injection (Fig. 3a).
ORY-1001 preparation and injection
ORY-1001 (Cayman Chemical) was dissolved in PBS at a stock concentration of 1 mg/ml by vortexing. Mice at 17–20 g body weight were injected 9.0 μg/kg/day; mice at 21–28 g body weight were injected 10.0 μg/kg/day as previously descrived60. 6-week-old animals were intraperitoneally injected for 7 consecutive days following tamoxifen injection (Fig. 3a).
RNA-seq library preparation, sequencing and analyzes
RNA-seq libraries were prepared based on the SMART-seq2 method as previously described with minor modifications100,101. Briefly, for RT, 3.2 μl RNA (100 ng/μl) was combined with 0.25 μl RNase inhibitor (NEB) and 1 μl CDS primer (10 mM, 5ʹ-AAGCAGTGG TATCAACGCAGAGTACT30VN-3ʹ) in an 8-well PCR tube strip and incubated at 70 °C for 2 min and immediately placed on ice. Then, 2 μl of 5× SMARTScribe RT buffer (TaKaRa), 0.5 μl of DTT (100 mM). 0.3 μl of MgCl2 (200 mM), 1 μl of dNTPs (10 mM), 1 μl of TSO primer (10 μM, 5ʹ-AAGCAGTGGTATCAACGCAGAGTACATrGrGrG-3ʹ) 0.25 μl of RNase inhibitor (NEB), and 0.5 μl SMARTScribe reverse transcriptase (TaKaRa) was added to the reaction. RT was performed under the following conditions: 42 °C for 90 min, 10 cycles of 50 °C for 2 min and 42 °C for 2 min, 70 °C for 15 min and 4 °C indefinitely. For cDNA amplification, 2 μl of the RT reaction was combined with 2.5 μl of 10× Advantage 2 buffer (Takara), 2.5 μl of 2.5 mM dNTPs (Takara), 0.25 μl of 10 μM IS PCR primer (5ʹ-AAGCAGTGGTATCAACGCAGAGT-3ʹ), 17.25 μl nuclease-free water, and 0.5 μl Advantage 2 DNA polymerase (Takara). Thermocycling conditions were as follows: 94 °C for 3 min, 9 cycle of 94 °C for 15 s, 65 °C for 30 s, and 68 °C for 6 min, 72 °C for 10 min, and 4 °C indefinitely. Amplified cDNA was then purified using 0.8× AMPure XP beads (Beckman Coulter), eluted in nuclease-free water and quantified following the instructions of Qubit dsDNA HS assay kit (Thermo Fisher). For tagmentation of cDNA, 50 pg cDNA in 1μl nuclease-free water, 2.5 μl 2×TD buffer (20 mM Tris/PH 8.0, 10 mM MgCl2, and 16% PEG 8000) and 0.5 μl adapter-loaded Tn5 transposase (Lucigen) were mixed and incubated at 55 °C for 15 min and the reaction was terminated by adding 1.25 μl 0.2% SDS and incubated at room temperature for 10 min. Fragments were amplified by adding 16.75 μl nuclease-free water, 1 μl of Nextera i7 primer (10 mM), 1 μl of Nextera i5 primer (10 mM), and 25 μl KAPA HiFi hotstart readymix (EMSCO/FISHER). Thermocycling conditions were as follows: 72 °C for 5 min, 95 °C for 1 min, 14 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, 72 °C for 1 min, and 4 °C indefinitely. PCR products were then purified twice with 0.8× AMPure XP beads and eluted in nuclease-free water. Samples were quantified by qPCR (KAPA) and pooled at equal molar amounts. The average fragment size of the final library fragment was quantified by bioanalyzer (Agilent), and the final concentration of the library was determined by qPCR (KAPA). About 2.7 pmol DNA was loaded on NextSeq high output kit (75 cycles, Illumina) and subjected to a NextSeq 550 sequencer (Illumina). Raw sequencing data from RNA-seq were demultiplexed with bcl2fastq2 v2.17.1.14 (Illumina), and adapters were trimmed using Trimmomatic v0.32102. Alignments were made using STAR v2.5.2a103 to GENCODE mouse reference genome GRCm38. Only uniquely mapped reads were quantified at the gene level and summarized to gene counts using STAR-quantMode (Gene Counts), with multimap** and chimeric alignments discarded. Further analyzes were performed in R (v3.6.0). Differentially expressed genes were identified using edgeR104. Identified upregulated and downregulated gene lists were used for Gene Ontology (GO) enrichment and KEGG pathway analyzes using Metascape Database105. Fold changes of target differentially expressed genes were visualized in heatmaps by “ComplexHeatmap” package106 in R.
Administration of LSD1 inhibitor into primary adult NSC cultures
LSD1 inhibitor, ORY-1001 (Cayman Chemical), was dissolved at 30 μM in 100% DMSO. For inhibiting LSD1 enzymatic activity in cultured quiescent NSCs, ORY-1001 was diluted to 15 nM in fresh quiescence medium. Quiescence medium without ORY-1001 was first added to primary NSC cultures to induce quiescence. After 48 h, medium was replaced with fresh quiescence medium containing 15 nM ORY-1001 as well as lentivirus expressing either GFP or GFP-Cre. Quiescence medium with 15 nM ORY-1001 was replaced every 2 days until cultures were harvested for immunostaining or RNA-seq experiments.
CUT & RUN assay and bioinformatic analyzes
The Setd1a CUT & RUN samples were prepared with CUT & RUN kit (EpiCypher). Briefly, cultured quiescent NSCs were collected in a 1.5 ml tube containing wash buffer. Cells were captured with ConA beads and incubated with Setd1a antibody (Abcam: ab70378) overnight at 4 °C in antibody incubation buffer. After unbound antibodies were washed away, pAG-MNase was added and incubated for 10 min at RT. After washing, CaCl2 was added and incubated for 2 h at 4 °C, and the reaction was stopped by adding stop buffer. The protein-DNA complexes were released by incubating at 37 °C for 10 min. The supernatant was transferred to a new Lo-bound tube. DNA was then purified using a DNA cleanup kit. Sequencing libraries were prepared using the NEBNext Ultra II DNA library preparation kit for Illumina (New England Biolabs) according to the manufacturer’s instructions. Briefly, end repair was conducted at 20 °C for 30 min, followed by dA-tailing at 65 °C for 30 min. After adapter ligation, the DNA fragments were purified by 0.9× volume of SPRIselect beads (EMSCO/FISHER) followed by 14 cycles PCR amplification with NEBNext Ultra II Q5 Master Mix (New England Biolabs). The PCR products were cleaned up with 0.9× volume of SPRIselect beads. The CUT & RUN libraries were quantified using qPCR (KAPA) and pooled at equal molar amounts, and quality control was performed with the Bioanalyzer High Sensitivity DNA Analysis (Agilent). The libraries were sequenced on the NextSeq high output kit (75 cycles, Illumina). For the analysis of the CUT & RUN data, all the reads were trimmed, and then uniquely mapped to the mm10 reference genome using Bowtie2107. Low map** quality reads were filtered by SAMtools108. Peak calling was performed using MACS2 with default parameters109. The intersected peaks defined in both replicates were considered high confidence peaks. The BAM files were converted to Bigwig format using deepTools110. The normalized read densities of merged replicates were visualized using the Integrative Genomics Viewer (IGV). Peak annotation was performed with ChIPseeker111.
Plasmid constructs
For knockdown experiments for mouse Bhlhe40, the short hairpin RNA sequence was cloned into the lentiviral vector expressing mCherry under the control of the hPGK promoter and the specific shRNA under the control of U6 promoter (Addgene plasmid #128073). All shRNA sequences are listed in the Supplementary Dataset 3. The efficacy of each Bhlhe40 shRNA was confirmed in cultured quiescent NSCs.
Lentivirus preparation and stereotaxic injection in the adult DG
When HEK293T cells cultured on the 15 cm dish reached 80% confluence, pLV-eGFP112 (Addgene) or pLV-EGFP-Cre113 vector (Addgene) was co-transfected with pCMV-VSV-G (Addgene) and HIV-1 packaging plasmid Δ8.9114 into HEK293T cells with LipoD293 in vitro transfection reagent (SignaGen). For lentivirus expressing specific shRNA against mouse Bhlhe40, mCherry and shRNA were co-expressed under the control of the hPGK and U6 promoters, respectively, using the pLK0.1 mCherry plasmid (Addgene). After co-transfection, the culture medium was collected once every 24 h for three times. The lentivirus in the collected medium was concentrated by ultracentrifugation at 106,800 g (25,000 rpm) for 1.5 h at 4 °C, with the supernatant discarded and pellet reconstituted with PBS. The titer of virus was measured by counting the number of infected cell clusters in a serial dilution experiment. Two-month-old mice were anesthetized and placed in a David Kopf Instruments stereotaxic frame (model 1900) equipped with a digital manipulator and a UMP3-1 Ultra pump. Mice were kept deeply anesthetized as assessed by monitoring pinch withdrawal and respiration rate. Viral injections were targeted to the dentate gyrus (A/P: −2.0 mm; M/L: 1.5 mm; D/V: −2.6 mm). The injections were performed at a rate of 0.2 μl/min. The needle was left in place for 10 min after each injection to minimize upward flow of viral solution after raising the needle.
RNA extraction and qPCR
Cell cultures were dissolved in 1 ml Trizol (Thermo Fisher) for 15 min and 200 μl chloroform was then added. After centrifugation 12000 g for 15 min at 4 °C, about 400 μl supernatant was carefully taken out and mixed with the same volume of ethanol, and the mixture was transferred to Zymo-SpinTM IC Column in a collection tube from the kit of Zymo RNA clean & concentrator kit (Zymo Research). After centrifugation, 400 μl RNA Prep buffer, 700 μl RNA wash buffer and 400 μl RNA wash buffer was sequentially added to the column with each procedure followed by centrifugation. Finally, about 15 μl nuclease-free water was added to elute the column to collect the RNA sample. RNA concentration and quality were assessed using a Nanodrop 2000. Subsequently, about 1 μg RNA was used for reverse transcription (RT) to synthesize cDNA with the SuperScript III First-Strand Synthesis System (Thermo Fisher). Quantitative RT-PCR was then performed using SYBR Green (Thermo Fisher) and the StepOnePlus Real-Time PCR system (Applied biosystems). Quantitative levels for all genes were normalized to the house-kee** gene GAPDH and expressed relative to the relevant control samples. All primer sequences are listed in the Supplementary Dataset 3.
Quantification and Statistical analyzes
Data in figure panels reflect several independent experiments performed on different days. Unpaired two-tailed Student’s t-tests were performed with GraphPad Prism. Quantitative results are expressed as the means ± SEM. A P value < 0.05 was considered significant. Statistical significance level was set as follows: *if P < 0.05, **if P < 0.01, ***if P < 0.001, n.s: P > 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq and CUT & RUN data generated in this study have been deposited in the NCBI’s Gene Expression Omnibus database under accession code GSE250278. Source data are provided with this paper. No custom software algorithms were generated. Any additional information required to reanalyze the data reported in this paper is available from the corresponding authors upon request. Source data are provided with this paper.
References
Ming, G. L. & Song, H. Adult neurogenesis in the mammalian brain: significant answers and significant questions. Neuron 70, 687–702 (2011).
Sahay, A., Wilson, D. A. & Hen, R. Pattern separation: a common function for new neurons in hippocampus and olfactory bulb. Neuron 70, 582–588 (2011).
Denoth-Lippuner, A. & Jessberger, S. Formation and integration of new neurons in the adult hippocampus. Nat Rev Neurosci 22, 223–236 (2021).
Toda, T., Parylak, S. L., Linker, S. B. & Gage, F. H. The role of adult hippocampal neurogenesis in brain health and disease. Mol Psychiatry 24, 67–87 (2019).
Kempermann, G. et al. Human adult neurogenesis: evidence and remaining questions. Cell Stem Cell 23, 25–30 (2018).
Terreros-Roncal, J. et al. Impact of neurodegenerative diseases on human adult hippocampal neurogenesis. Science 374, 1106–1113 (2021).
Zhou, Y. et al. Molecular landscapes of human hippocampal immature neurons across lifespan. Nature 607, 527–533 (2022).
Ming, G. L. & Song, H. Adult neurogenesis in the mammalian central nervous system. Annu Rev Neurosci 28, 223–250 (2005).
Christian, K. M., Song, H. & Ming, G. L. Functions and dysfunctions of adult hippocampal neurogenesis. Annu Rev Neurosci 37, 243–262 (2014).
Bonaguidi, M. A. et al. In vivo clonal analysis reveals self-renewing and multipotent adult neural stem cell characteristics. Cell 145, 1142–1155 (2011).
Sun, G. J. et al. Latent tri-lineage potential of adult hippocampal neural stem cells revealed by Nf1 inactivation. Nat Neurosci 18, 1722–1724 (2015).
Urban, N., Blomfield, I. M. & Guillemot, F. Quiescence of adult mammalian neural stem cells: a highly regulated rest. Neuron 104, 834–848 (2019).
Encinas, J. M. et al. Division-coupled astrocytic differentiation and age-related depletion of neural stem cells in the adult hippocampus. Cell Stem Cell 8, 566–579 (2011).
Pilz, G. A. et al. Live imaging of neurogenesis in the adult mouse hippocampus. Science 359, 658–662 (2018).
Bottes, S. et al. Long-term self-renewing stem cells in the adult mouse hippocampus identified by intravital imaging. Nat Neurosci 24, 225–233 (2021).
Kuhn, H. G., Dickinson-Anson, H. & Gage, F. H. Neurogenesis in the dentate gyrus of the adult rat: age-related decrease of neuronal progenitor proliferation. J Neurosci 16, 2027–2033 (1996).
Ibrayeva, A. et al. Early stem cell aging in the mature brain. Cell Stem Cell 28, 955–966.e957 (2021).
Audesse, A. J. & Webb, A. E. Mechanisms of enhanced quiescence in neural stem cell aging. Mech Ageing Dev 191, 111323 (2020).
Wu, Y. et al. Chronic in vivo imaging defines age-dependent alterations of neurogenesis in the mouse hippocampus. Nat Aging 3, 380–390 (2023).
Paik, J. H. et al. FoxOs cooperatively regulate diverse pathways governing neural stem cell homeostasis. Cell Stem Cell 5, 540–553 (2009).
Renault, V. M. et al. FoxO3 regulates neural stem cell homeostasis. Cell Stem Cell 5, 527–539 (2009).
Zhou, Y. et al. Autocrine Mfge8 signaling prevents developmental exhaustion of the adult neural stem cell pool. Cell Stem Cell 23, 444–452.e444 (2018).
Urban, N. Could a different view of quiescence help us understand how neurogenesis is regulated? Front Neurosci 16, 878875 (2022).
Shin, J. et al. Single-cell RNA-seq with waterfall reveals molecular cascades underlying adult neurogenesis. Cell Stem Cell 17, 360–372 (2015).
Ding, W. Y., Huang, J. & Wang, H. Waking up quiescent neural stem cells: molecular mechanisms and implications in neurodevelopmental disorders. PLoS Genet 16, e1008653 (2020).
Blasco-Chamarro, L. & Farinas, I. Fine-tuned rest: unveiling the regulatory landscape of adult quiescent neural stem cells. Neuroscience 525, 26–37 (2023).
Lampada, A. & Taylor, V. Notch signaling as a master regulator of adult neurogenesis. Front Neurosci 17, 1179011 (2023).
Zhang, F., Yoon, K., Kim, N. S., Ming, G. L. & Song, H. Cell-autonomous and non-cell-autonomous roles of NKCC1 in regulating neural stem cell quiescence in the hippocampal dentate gyrus. Stem Cell Reports 18, 1468–1481 (2023).
Jang, M. H. et al. Secreted frizzled-related protein 3 regulates activity-dependent adult hippocampal neurogenesis. Cell Stem Cell 12, 215–223 (2013).
Song, J. et al. Neuronal circuitry mechanism regulating adult quiescent neural stem-cell fate decision. Nature 489, 150–154 (2012).
Mira, H. et al. Signaling through BMPR-IA regulates quiescence and long-term activity of neural stem cells in the adult hippocampus. Cell Stem Cell 7, 78–89 (2010).
Song, J., Sun, J., Olsen, R. H. J., Ming, G.-l & Song, H. Neuronal circuitry mechanisms regulating adult mammalian neurogenesis. Cold Spring Harbor Perspective in Biology 8, a018937 (2016).
Knobloch, M. et al. Metabolic control of adult neural stem cell activity by Fasn-dependent lipogenesis. Nature 493, 226–230 (2013).
Knobloch, M. et al. A fatty acid oxidation-dependent metabolic shift regulates adult neural stem cell activity. Cell Rep 20, 2144–2155 (2017).
Petrelli, F. et al. Mitochondrial pyruvate metabolism regulates the activation of quiescent adult neural stem cells. Sci Adv 9, eadd5220 (2023).
Kobayashi, T. et al. Enhanced lysosomal degradation maintains the quiescent state of neural stem cells. Nat Commun 10, 5446 (2019).
Gengatharan, A. et al. Adult neural stem cell activation in mice is regulated by the day/night cycle and intracellular calcium dynamics. Cell 184, 709–722 e713 (2021).
Fong, B. C. et al. The Rb/E2F axis is a key regulator of the molecular signatures instructing the quiescent and activated adult neural stem cell state. Cell Rep 41, 111578 (2022).
Andersen, J. et al. A transcriptional mechanism integrating inputs from extracellular signals to activate hippocampal stem cells. Neuron 83, 1085–1097 (2014).
Fan, W. et al. The transcriptional co-activator Yap1 promotes adult hippocampal neural stem cell activation. EMBO J 42, e110384 (2023).
Urban, N. et al. Return to quiescence of mouse neural stem cells by degradation of a proactivation protein. Science 353, 292–295 (2016).
Blomfield, I. M. et al. Id4 promotes the elimination of the pro-activation factor Ascl1 to maintain quiescence of adult hippocampal stem cells. Elife 8, e48561 (2019).
Sun, J. et al. A septo-temporal molecular gradient of sfrp3 in the dentate gyrus differentially regulates quiescent adult hippocampal neural stem cell activation. Mol Brain 8, 52 (2015).
Berg, D. A. et al. A common embryonic origin of stem cells drives developmental and adult neurogenesis. Cell 177, 654–668.e615 (2019).
Yao, B. et al. Epigenetic mechanisms in neurogenesis. Nat Rev Neurosci 17, 537–549 (2016).
Lim, D. A. et al. Chromatin remodelling factor Mll1 is essential for neurogenesis from postnatal neural stem cells. Nature 458, 529–533 (2009).
Delgado, R. N. et al. Maintenance of neural stem cell positional identity by mixed-lineage leukemia 1. Science 368, 48–53 (2020).
Toda, T. et al. Nup153 interacts with Sox2 to enable bimodal gene regulation and maintenance of neural progenitor cells. Cell Stem Cell 21, 618–634.e617 (2017).
Li, X. et al. Ten-eleven translocation 2 interacts with forkhead box O3 and regulates adult neurogenesis. Nat Commun 8, 15903 (2017).
Guerra, M. V. et al. H3K9 methyltransferases Suv39h1 and Suv39h2 control the differentiation of neural progenitor cells in the adult hippocampus. Front Cell Dev Biol 9, 778345 (2021).
Zocher, S. et al. De novo DNA methylation controls neuronal maturation during adult hippocampal neurogenesis. EMBO J 40, e107100 (2021).
Jobe, E. M. & Zhao, X. DNA methylation and adult neurogenesis. Brain Plast 3, 5–26 (2017).
Zocher, S. & Toda, T. Epigenetic aging in adult neurogenesis. Hippocampus 33, 347–359 (2023).
Jessberger, S. et al. Epigenetic modulation of seizure-induced neurogenesis and cognitive decline. J Neurosci 27, 5967–5975 (2007).
Ma, D. K. et al. Neuronal activity-induced Gadd45b promotes epigenetic DNA demethylation and adult neurogenesis. Science 323, 1074–1077 (2009).
Wang, S. et al. SETD1A Mediated H3K4 Methylation and Its Role in Neurodevelopmental and Neuropsychiatric Disorders. Front Mol Neurosci 14, 772000 (2021).
Singh, T. et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature 604, 509–516 (2022).
Singh, T. et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat Neurosci 19, 571–577 (2016).
Takata, A. et al. Loss-of-function variants in schizophrenia risk and SETD1A as a candidate susceptibility gene. Neuron 82, 773–780 (2014).
Mukai, J. et al. Recapitulation and reversal of schizophrenia-related phenotypes in Setd1a-deficient mice. Neuron 104, 471–487.e412 (2019).
Nagahama, K. et al. Setd1a insufficiency in mice attenuates excitatory synaptic function and recapitulates schizophrenia-related behavioral abnormalities. Cell Rep 32, 108126 (2020).
Chen, R. et al. Cell type-specific mechanism of Setd1a heterozygosity in schizophrenia pathogenesis. Sci Adv 8, eabm1077 (2022).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Tusi, B. K. et al. Setd1a regulates progenitor B-cell-to-precursor B-cell development through histone H3 lysine 4 trimethylation and Ig heavy-chain rearrangement. FASEB J 29, 1505–1515 (2015).
Tronche, F. et al. Disruption of the glucocorticoid receptor gene in the nervous system results in reduced anxiety. Nat Genet 23, 99–103 (1999).
Bond, A. M., Ming, G. L. & Song, H. Adult mammalian neural stem cells and neurogenesis: five decades later. Cell Stem Cell 17, 385–395 (2015).
Wong, S. Z. H. et al. In vivo clonal analysis reveals spatiotemporal regulation of thalamic nucleogenesis. PLoS Biol 16, e2005211 (2018).
Hoshii, T. et al. A non-catalytic function of SETD1A regulates cyclin K and the dna damage response. Cell 172, 1007–1021.e1017 (2018).
Lee, M. G., Wynder, C., Schmidt, D. M., McCafferty, D. G. & Shiekhattar, R. Histone H3 lysine 4 demethylation is a target of nonselective antidepressive medications. Chem Biol 13, 563–567 (2006).
Maes, T., Carceller, E., Salas, J., Ortega, A. & Buesa, C. Advances in the development of histone lysine demethylase inhibitors. Curr Opin Pharmacol 23, 52–60 (2015).
Maes, T. et al. ORY-1001, a Potent and Selective Covalent KDM1A Inhibitor, for the Treatment of Acute Leukemia. Cancer Cell 33, 495–511.e412 (2018).
Ma, D. K., Chiang, C. H., Ponnusamy, K., Ming, G. L. & Song, H. G9a and Jhdm2a regulate embryonic stem cell fusion-induced reprogramming of adult neural stem cells. Stem Cells 26, 2131–2141 (2008).
Martynoga, B. et al. Epigenomic enhancer annotation reveals a key role for NFIX in neural stem cell quiescence. Genes Dev 27, 1769–1786 (2013).
Marques-Torrejon, M. A. et al. LRIG1 is a gatekeeper to exit from quiescence in adult neural stem cells. Nat Commun 12, 2594 (2021).
Santos-Rosa, H. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002).
Sun, H. et al. Stra13 regulates satellite cell activation by antagonizing Notch signaling. J Cell Biol 177, 647–657 (2007).
Salvi, J. S. et al. ATR activity controls stem cell quiescence via the cyclin F-SCF complex. Proc Natl Acad Sci USA 119, e2115638119 (2022).
Lukas, J. et al. Dazap2 modulates transcription driven by the Wnt effector TCF-4. Nucleic Acids Res 37, 3007–3020 (2009).
Qian, X. et al. PTEN suppresses glycolysis by dephosphorylating and inhibiting autophosphorylated PGK1. Mol Cell 76, 516–527.e517 (2019).
Yamauchi, T., Nishiyama, M., Moroishi, T., Kawamura, A. & Nakayama, K. I. FBXL5 inactivation in mouse brain induces aberrant proliferation of neural stem progenitor cells. Mol Cell Biol 37, e00470–16 (2017).
Asanoma, K. et al. Regulation of the mechanism of TWIST1 transcription by BHLHE40 and BHLHE41 in cancer cells. Mol Cell Biol 35, 4096–4109 (2015).
de Morree, A. & Rando, T. A. Regulation of adult stem cell quiescence and its functions in the maintenance of tissue integrity. Nat Rev. Mol. Cell Biol. 24, 334–354 (2023).
Radak, M. & Fallahi, H. The epigenetic regulation of quiescent in stem cells. Glob Med Genet 10, 339–344 (2023).
Cho, I. J. et al. Mechanisms, hallmarks, and implications of stem cell quiescence. Stem Cell Reports 12, 1190–1200 (2019).
Nicetto, D. & Zaret, K. S. Role of H3K9me3 heterochromatin in cell identity establishment and maintenance. Curr Opin Genet Dev 55, 1–10 (2019).
Kim, M. & Costello, J. DNA methylation: an epigenetic mark of cellular memory. Exp Mol Med 49, e322 (2017).
Jiang, Y. & Hsieh, J. HDAC3 controls gap 2/mitosis progression in adult neural stem/progenitor cells by regulating CDK1 levels. Proc Natl Acad Sci USA 111, 13541–13546 (2014).
Imayoshi, I., Sakamoto, M., Yamaguchi, M., Mori, K. & Kageyama, R. Essential roles of notch signaling in maintenance of neural stem cells in develo** and adult brains. J Neurosci 30, 3489–3498 (2010).
Sueda, R., Imayoshi, I., Harima, Y. & Kageyama, R. High Hes1 expression and resultant Ascl1 suppression regulate quiescent vs. active neural stem cells in the adult mouse brain. Genes Dev 33, 511–523 (2019).
Arndt, K. et al. SETD1A protects HSCs from activation-induced functional decline in vivo. Blood 131, 1311–1324 (2018).
Hyun, K., Jeon, J., Park, K. & Kim, J. Writing, erasing and reading histone lysine methylations. Exp Mol Med 49, e324 (2017).
Gates, L. A. et al. Acetylation on histone H3 lysine 9 mediates a switch from transcription initiation to elongation. J Biol Chem 292, 14456–14472 (2017).
Karmodiya, K., Krebs, A. R., Oulad-Abdelghani, M., Kimura, H. & Tora, L. H3K9 and H3K14 acetylation co-occur at many gene regulatory elements, while H3K14ac marks a subset of inactive inducible promoters in mouse embryonic stem cells. BMC Genomics 13, 424 (2012).
Huang, Y. et al. Inhibition of lysine-specific demethylase 1 by polyamine analogues results in reexpression of aberrantly silenced genes. Proc Natl Acad Sci USA 104, 8023–8028 (2007).
Wang, S. et al. Loss-of-function variants in the schizophrenia risk gene SETD1A alter neuronal network activity in human neurons through the cAMP/PKA pathway. Cell Rep 39, 110790 (2022).
Kang, E., Wen, Z., Song, H., Christian, K. M. & Ming, G. L. Adult neurogenesis and psychiatric disorders. Cold Spring Harb Perspect Biol 8, a019026 (2016).
Allen, K. M., Fung, S. J. & Weickert, C. S. Cell proliferation is reduced in the hippocampus in schizophrenia. Aust N Z J Psychiatry 50, 473–480 (2016).
Reif, A. et al. Neural stem cell proliferation is decreased in schizophrenia, but not in depression. Mol Psychiatry 11, 514–522 (2006).
Duan, X. et al. Disrupted-in-schizophrenia 1 regulates integration of newly generated neurons in the adult brain. Cell 130, 1146–1158 (2007).
Weng, Y. L. et al. An intrinsic epigenetic barrier for functional axon regeneration. Neuron 94, 337–346.e336 (2017).
Zhang, F. et al. Epitranscriptomic regulation of cortical neurogenesis via Mettl8-dependent mitochondrial tRNA m(3)C modification. Cell Stem Cell 30, 300–311.e311 (2023).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10, 1523 (2019).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137 (2008).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res 44, W160–W165 (2016).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Enomoto, M., Bunge, M. B. & Tsoulfas, P. A multifunctional neurotrophin with reduced affinity to p75NTR enhances transplanted Schwann cell survival and axon growth after spinal cord injury. Exp Neurol 248, 170–182 (2013).
Truitt J. M., et al. Inhibition of IKKbeta Reduces Ethanol Consumption in C57BL/6J Mice. eNeuro 3, (2016).
Kim, Y. S. et al. Characterizing the conductance underlying depolarization-induced slow current in cerebellar Purkinje cells. J Neurophysiol 109, 1174–1181 (2013).
Acknowledgements
We thank members of Ming and Song laboratories for comments and suggestions, and B. Temsamrit, E. LaNoce, A. Angelucci, and G. Alepa for laboratory support. This work was supported by grants from National Institutes of Health (R35NS097370 and R01MH125528 to G-l.M., R01CA264932 and R01CA 260729 to S.H., and R35NS116843 and RF1AG079557 to H.S.), and from Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (to G-l.M.).
Author information
Authors and Affiliations
Contributions
T.Z. and Y.H performed all the analyzes. B.Y. and S.H. provided Setd1af/f mice. T.Z., Y.H., G.M. and H.S. conceived the project and wrote the manuscript with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, T., Hong, Y., Yan, B. et al. Epigenetic maintenance of adult neural stem cell quiescence in the mouse hippocampus via Setd1a. Nat Commun 15, 5674 (2024). https://doi.org/10.1038/s41467-024-50010-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-50010-y
- Springer Nature Limited